New R-Based Methodology to Optimize the Identification of Root Endophytes against Heterobasidion parviporum
Abstract
1. Introduction
2. Materials and Methods
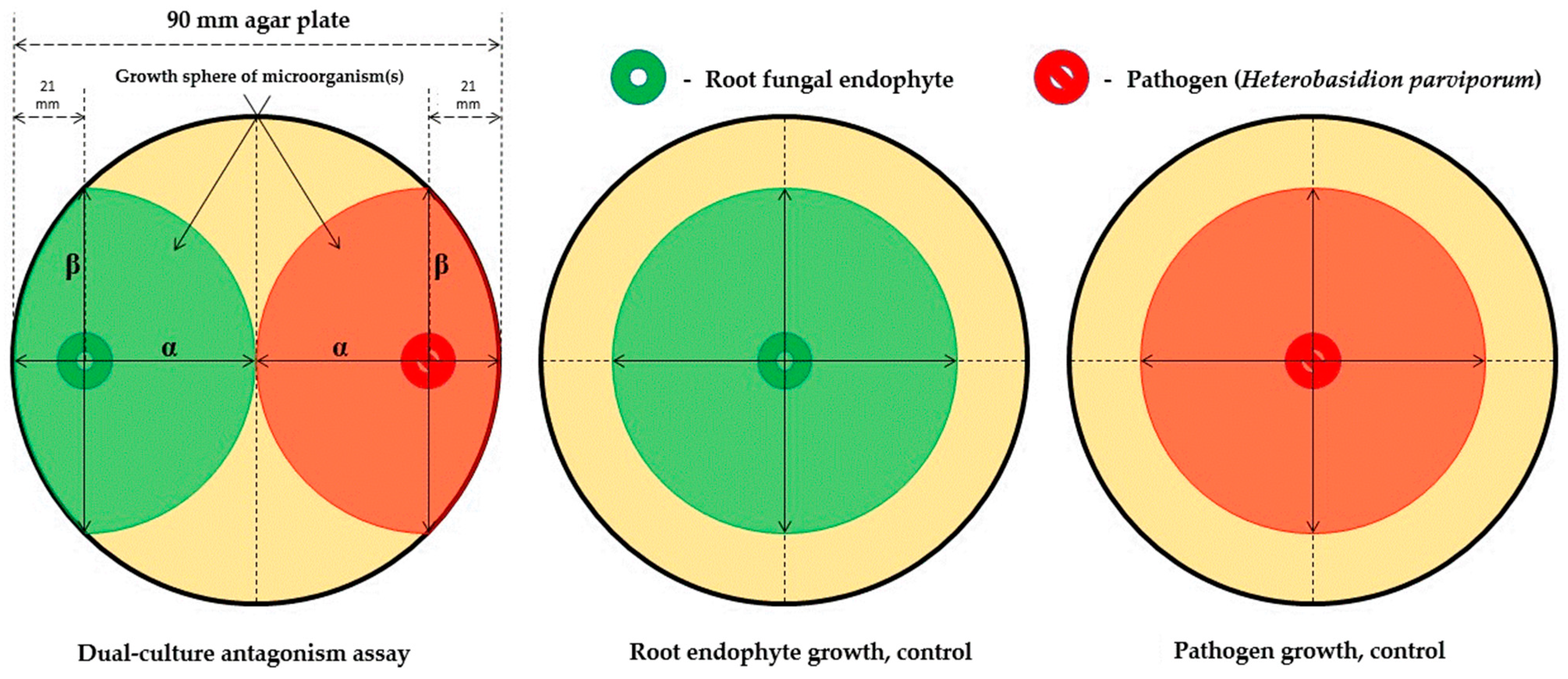
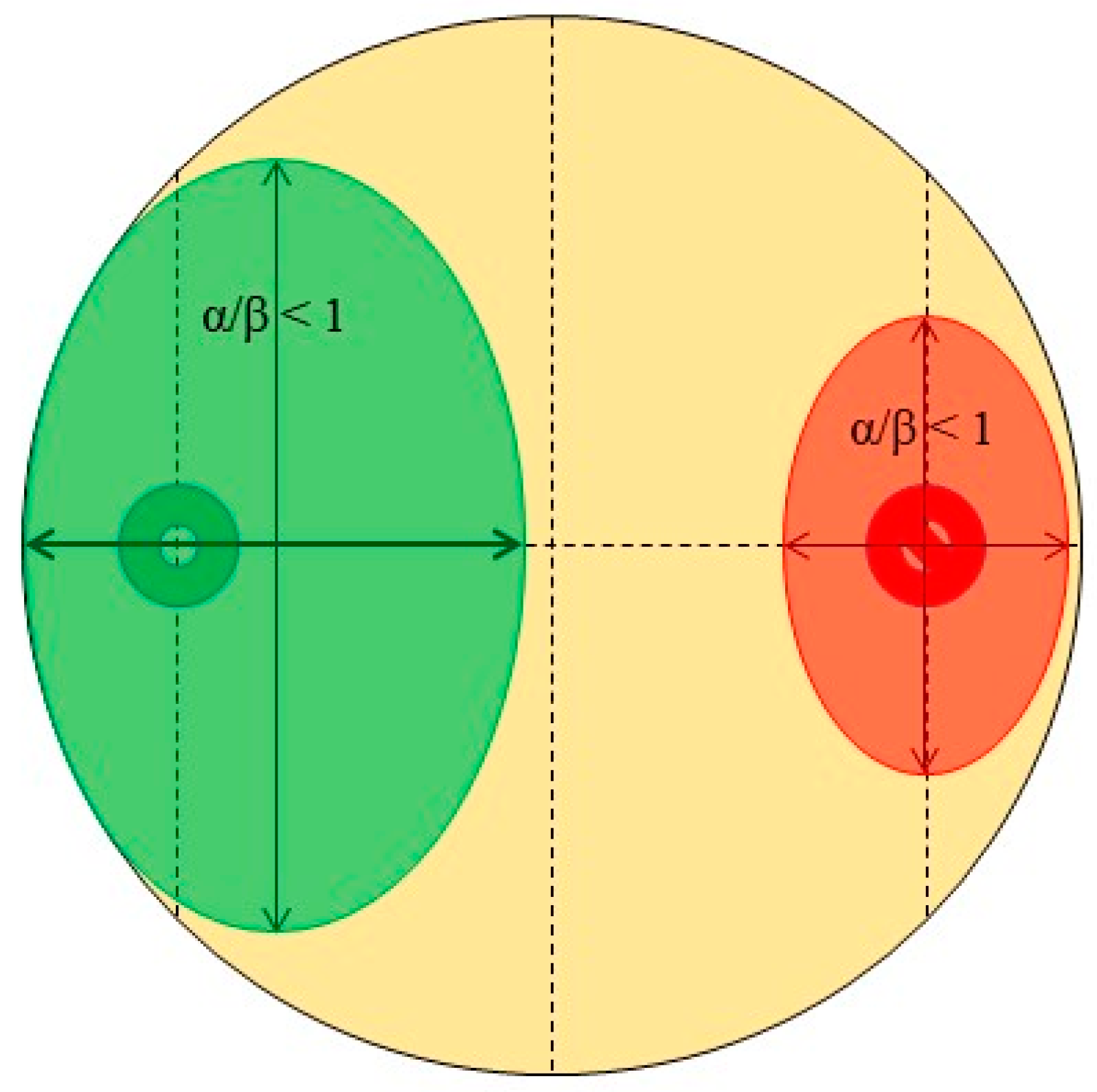
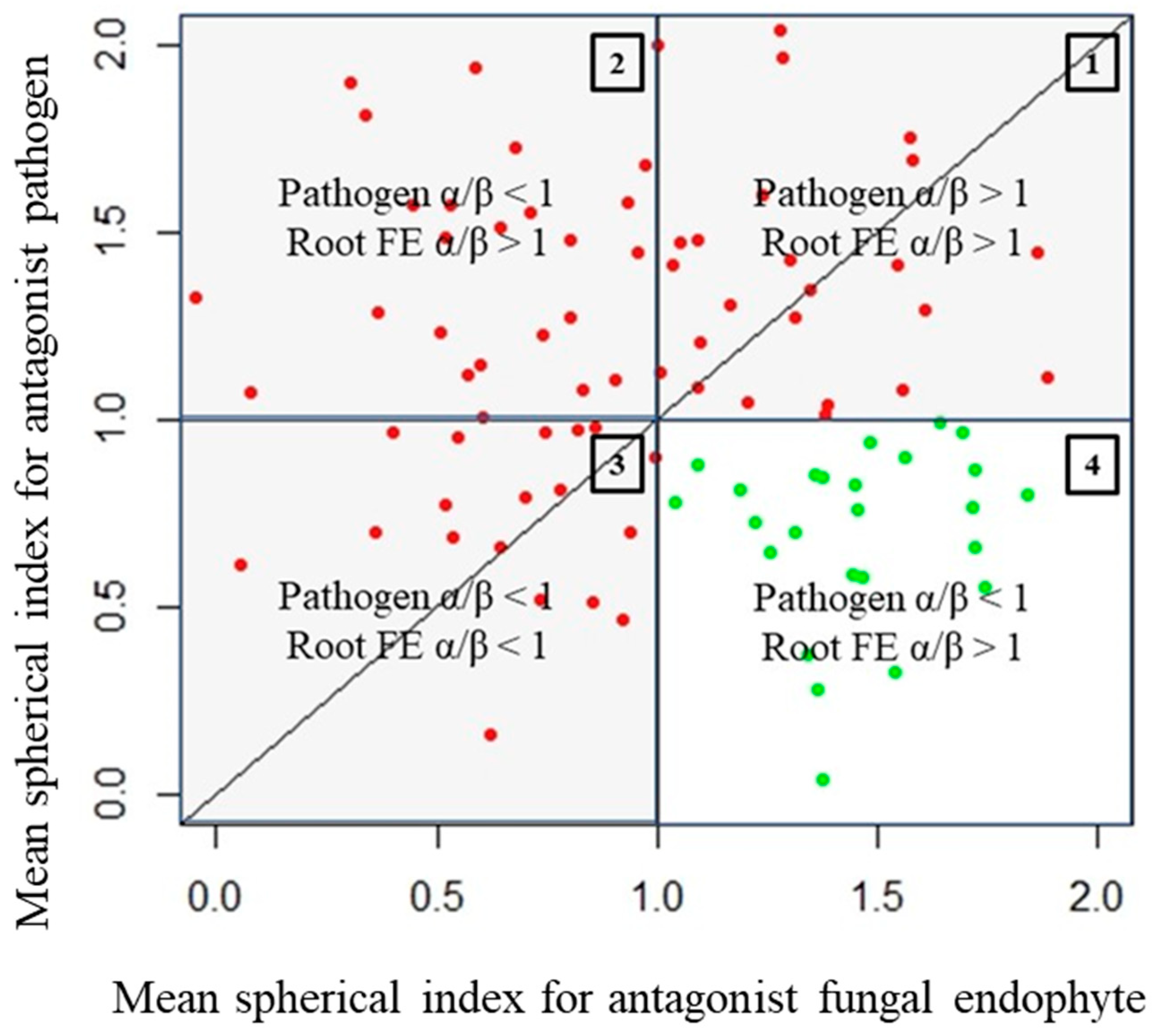
- Quadrant described by pathogen, α/β > 1, and endophyte, α/β > 1 (quadrant 1): There is clearly no antagonism occurring in these samples as the growth of neither organism along the colinear axis appears to be suppressed or prohibited.
- Quadrant described by pathogen, α/β > 1, and endophyte, α/β < 1 (quadrant 2): The root endophytes described by their datapoints in this quadrant likely have no effect on the growth of the pathogen (or may conversely even be getting suppressed by the pathogen themselves) as its progression along the colinear axis is higher in comparison to its orthogonal spread and also higher in comparison to the progression of the antagonist along the colinear axis itself.
- Quadrant described by pathogen, α/β < 1, and endophyte, α/β < 1 (quadrant 3): There is likely no interaction at all between the root endophytes and the pathogen in the cases described by this quadrant as both spherical indices are less than 1; this likely indicates purely natural growth with no antagonistic effects taking place.
3. Results
3.1. Identification and Diversity of Root Endophytes
3.2. Antagonisms Assay
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carroll, G. Fungal Endophytes in Stems and Leaves: From Latent Pathogen to Mutualistic Symbiont. Ecology 1988, 69, 2–9. [Google Scholar] [CrossRef]
- Saikkonen, K.; Faeth, S.H.; Helander, M.; Sullivan, T.J. Fungal endophytes. A continuum of interactions with host plants. Annu. Rev. Ecol. Syst. 1998, 29, 319–343. [Google Scholar]
- Clay, K. Fungi and the food of the gods. Nature 2004, 427, 401–402. [Google Scholar] [CrossRef]
- Petrini, O. Fungal Endophytes of Tree Leaves. In Microbial Ecology of Leaves; Andrews, J.H., Hirano, S.S., Eds.; Springer: New York, NY, USA, 1991; pp. 179–197. [Google Scholar]
- Terhonen, E.; Blumenstein, K.; Kovalchuk, A.; Asiegbu, F.O. Forest Tree Microbiomes and associated Fungal Endophytes: Functional Roles and Impact on Forest Health. Forests 2019, 10, 42. [Google Scholar] [CrossRef]
- Tellenbach, C.; Sieber, T.N. Do colonization by dark septate endophytes and elevated temperature affect pathogenicity of oomycetes? FEMS Microbiol. Ecol. 2012, 82, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Martín, J.A.; Witzell, J.; Blumenstein, K.; Rozpedowska, E.; Helander, M.; Sieber, T.N.; Gil, L. Resistance to Dutch Elm Disease Reduces Presence of Xylem Endophytic Fungi in Elms (Ulmus spp.). PLoS ONE 2013, 8, e56987. [Google Scholar]
- Terhonen, E.; Sipari, N.; Asiegbu, F.O. Inhibition of phytopathogens by fungal root endophytes of Norway spruce. Biol. Control 2016, 99, 53–63. [Google Scholar] [CrossRef]
- Jumpponen, A.; Trappe, J.M. Performance of Pinus contorta inoculated with two strains of root endophytic fungus, Phialocephala fortinii: Effects of synthesis system and glucose concentration. Can. J. Bot. 1998, 76, 1205–1213. [Google Scholar]
- Newsham, K.K. A meta-analysis of plant responses to dark septate root endophytes. New Phytol. 2011, 190, 783–793. [Google Scholar] [CrossRef]
- Vohník, M.; Lukančič, S.; Bahor, E.; Regvar, M.; Vosátka, M.; Vodnik, D. Inoculation of Rhododendron cv. Belle-Heller with two strains of Phialocephala fortinii in two different substrates. Folia Geobot. 2003, 38, 191. [Google Scholar] [CrossRef]
- Tellenbach, C.; Grünig, C.R.; Sieber, T.N. Negative effects on survival and performance of Norway spruce seedlings colonized by dark septate root endophytes are primarily isolate-dependent. Environ. Microbiol. 2011, 13, 2508–2517. [Google Scholar] [CrossRef]
- Fernando, A.A.; Currah, R.S. A comparative study of the effects of the root endophytes Leptodontidium orchidicola and Phialocephala fortinii (Fungi Imperfecti) on the growth of some subalpine plants in culture. Can. J. Bot. 1996, 74, 1071–1078. [Google Scholar] [CrossRef]
- Jumpponen, A.; Mattson, K.G.; Trappe, J.M. Mycorrhizal functioning of Phialocephala fortinii with Pinus contorta on glacier forefront soil: Interactions with soil nitrogen and organic matter. Mycorrhiza 1998, 7, 261–265. [Google Scholar] [CrossRef]
- Grünig, C.R.; Queloz, V.; Sieber, T.N.; Holdenrieder, O. Dark septate endophytes (DSE) of the Phialocephala fortinii s.l.—Acephala applanata species complex in tree roots: Classification, population biology, and ecology. Botany 2008, 86, 1355–1369. [Google Scholar] [CrossRef]
- Mandyam, K.; Jumpponen, A. Seeking the elusive function of the root-colonising dark septate endophytic fungi. Stud. Mycol. 2005, 53, 173–189. [Google Scholar] [CrossRef]
- Witzell, J.; Martín, J.A. Phenolic metabolites in the resistance of northern forest trees to pathogens—Past experiences and future prospects. Can. J. For. Res. 2008, 38, 2711–2727. [Google Scholar] [CrossRef]
- Schulz, B.J.E.; Boyle, C.J.C.; Sieber, T.N. Microbial Root Endophytes; Soil Biology; Springer: Berlin/Heidelberg, Germany, 2006; ISBN 978-3-540-33525-2. [Google Scholar]
- Terhonen, E.; Keriö, S.; Sun, H.; Asiegbu, F.O. Endophytic fungi of Norway spruce roots in boreal pristine mire, drained peatland and mineral soil and their inhibitory effect on Heterobasidion parviporum in vitro. Fungal Ecol. 2014, 9, 17–26. [Google Scholar] [CrossRef]
- Kovalchuk, A.; Mukrimin, M.; Zeng, Z.; Raffaello, T.; Liu, M.; Kasanen, R.; Sun, H.; Asiegbu, F.O. Mycobiome analysis of asymptomatic and symptomatic Norway spruce trees naturally infected by the conifer pathogens Heterobasidion spp. Environ. Microbiol. Rep. 2018, 10, 532–541. [Google Scholar] [CrossRef]
- Addy, H.D.; Piercey, M.M.; Currah, R.S. Microfungal endophytes in roots. Can. J. Bot. 2005, 83, 1–13. [Google Scholar] [CrossRef]
- Jumpponen, A.; Trappe, J.M. Dark septate endophytes: A review of facultative biotrophic root-colonizing fungi. New Phytol. 1998, 140, 295–310. [Google Scholar] [CrossRef]
- Queloz, V.; Sieber, T.N.; Holdenrieder, O.; McDonald, B.A.; Grünig, C.R. No biogeographical pattern for a root-associated fungal species complex. Glob. Ecol. Biogeogr. 2011, 20, 160–169. [Google Scholar] [CrossRef]
- Sietiö, O.-M.; Tuomivirta, T.; Santalahti, M.; Kiheri, H.; Timonen, S.; Sun, H.; Fritze, H.; Heinonsalo, J. Ericoid plant species and Pinus sylvestris shape fungal communities in their roots and surrounding soil. New Phytol. 2018, 218, 738–751. [Google Scholar] [CrossRef]
- Grünig, C.R.; Duò, A.; Sieber, T.N.; Holdenrieder, O. Assignment of species rank to six reproductively isolated cryptic species of the Phialocephala fortinii s.l.-Acephala applanata species complex. Mycologia 2008, 100, 47–67. [Google Scholar] [CrossRef]
- Stoyke, G.; Currah, R.S. Endophytic fungi from the mycorrhizae of alpine ericoid plants. Can. J. Bot. 1991, 69, 347–352. [Google Scholar] [CrossRef]
- Tejesvi, M.V.; Sauvola, T.; Pirttilä, A.M.; Ruotsalainen, A.L. Neighboring Deschampsia flexuosa and Trientalis europaea harbor contrasting root fungal endophytic communities. Mycorrhiza 2013, 23, 1–10. [Google Scholar] [CrossRef]
- Grünig, C.R.; Sieber, T.N.; Rogers, S.O.; Holdenrieder, O. Spatial distribution of dark septate endophytes in a confined forest plot. Mycol. Res. 2002, 106, 832–840. [Google Scholar] [CrossRef]
- Grünig, C.R.; McDonald, B.A.; Sieber, T.N.; Rogers, S.O.; Holdenrieder, O. Evidence for subdivision of the root-endophyte Phialocephala fortinii into cryptic species and recombination within species. Fungal Genet. Biol. 2004, 41, 676–687. [Google Scholar] [CrossRef]
- Queloz, V.; Duò, A.; Grünig, C.R. Isolation and characterization of microsatellite markers for the tree-root endophytes Phialocephala subalpina and Phialocephala fortinii s.s. Mol. Ecol. Resour. 2008, 8, 1322–1325. [Google Scholar] [CrossRef]
- Queloz, V.; Grunig, C.R.; Sieber, T.N.; Holdenrieder, O. Monitoring the spatial and temporal dynamics of a community of the tree-root endophyte Phialocephala fortinii s.l. New Phytol. 2005, 168, 651–660. [Google Scholar] [CrossRef]
- Queloz, V.; Duò, A.; Sieber, T.N.; Grünig, C.R. Microsatellite size homoplasies and null alleles do not affect species diagnosis and population genetic analysis in a fungal species complex. Mol. Ecol. Resour. 2010, 10, 348–367. [Google Scholar] [CrossRef]
- Sieber, T.N. Endophytic fungi in forest trees: Are they mutualists? Fungal Biol. Rev. 2007, 21, 75–89. [Google Scholar] [CrossRef]
- Saikkonen, K. Forest structure and fungal endophytes. Fungal Biol. Rev. 2007, 21, 67–74. [Google Scholar] [CrossRef]
- Garbelotto, M.; Gonthier, P. Biology, epidemiology, and control of Heterobasidion species worldwide. Annu. Rev. Phytopathol. 2013, 51, 39–59. [Google Scholar] [CrossRef]
- Gonthier, P.; Garbelotto, M.M.; Nicolotti, G. Seasonal Patterns of Spore Deposition of Heterobasidion Species in Four Forests of the Western Alps. Phytopathology 2005, 95, 759–767. [Google Scholar] [CrossRef]
- Lind, M.; Stenlid, J.; Olson, Å. Chapter Twelve—Heterobasidion annosum s.l. Genomics. In Advances in Botanical Research; Martin, F.M., Ed.; Academic Press: London, UK, 2014; Volume 70, pp. 371–396. [Google Scholar]
- Asiegbu, F.O.; Adomas, A.; Stenlid, J. Conifer root and butt rot caused by Heterobasidion annosum (Fr.) Bref. s.l. Mol. Plant Pathol. 2005, 6, 395–409. [Google Scholar] [CrossRef]
- Korhonen, K. Intersterility groups of Heterobasidion annosum. Commun. Inst. For. Fenn. 1978, 94, 1–25. [Google Scholar]
- Capretti, P.; Korhonen, K.; Mugnai, L.; Romagnoli, C. An intersterility group of Heterobasidion annosum specialized to Abies alba. Eur. J. For. Pathol. 1990, 20, 231–240. [Google Scholar] [CrossRef]
- Rishbeth, J. Dispersal of Fomes annosus Fr. and Peniophora gigantea (Fr.) massee. Trans. Br. Mycol. Soc. 1959, 42, 243–260. [Google Scholar] [CrossRef]
- Isomäki, A.; Kallio, T. Consequences of injury caused by timber harvesting machines on the growth and decay of spruce (Picea abies (L.) Karst.). Acta For. Fenn. 1974, 136, 1–34. [Google Scholar] [CrossRef]
- Redfern, D.B.; Stenlid, J. Spore dispersal and infection. In Heterobasidion Annosum: Biology, Ecology, Impact and Control; Woodward, S., Stenlid, J., Karjalainen, R., Hüttermann, A., Eds.; CAB International: Wallingford, UK, 1998; pp. 105–124. [Google Scholar]
- Piri, T. The spreading of the S type of Heterobasidion annosum from Norway spruce stumps to the subsequent tree stand. Eur. J. For. Pathol. 1996, 26, 193–204. [Google Scholar] [CrossRef]
- Oliva, J.; Zhao, A.; Zarei, S.; Sedlák, P.; Stenlid, J. Effect of temperature on the interaction between Phlebiopsis gigantea and the root-rot forest pathogen Heterobasidion spp. For. Ecol. Manag. 2015, 340, 22–30. [Google Scholar] [CrossRef]
- Piri, T.; Hamberg, L. Persistence and infectivity of Heterobasidion parviporum in Norway spruce root residuals following stump harvesting. For. Ecol. Manag. 2015, 353, 49–58. [Google Scholar] [CrossRef]
- Marx, D.H. The influence of ectotrophic mycorrhizal fungi on the resistance of pine roots to pathogenic infections. I. Antagonism of mycorrhizal fungo to root pathogenic fungi and soil bacteria. Phytopathology 1969, 59, 153–163. [Google Scholar]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basidiomycetes—Application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Ryberg, M.; Hartmann, M.; Branco, S.; Wang, Z.; Godhe, A.; Wit, P.D.; Sánchez-García, M.; Ebersberger, I.; de Sousa, F.; et al. Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol. Evol. 2013, 4, 914–919. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Stöver, B.C.; Müller, K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010, 11, 7. [Google Scholar] [CrossRef] [PubMed]
- Blumenstein, K. Endophytic Fungi in Elms: Implications for the Integrated Management of Dutch Elm Disease. Ph.D. Thesis, Swedish University of Agricultural Sciences, Alnarp, Sweden, 2016. [Google Scholar]
- Martín, J.A.; Macaya-Sanz, D.; Witzell, J.; Blumenstein, K.; Gil, L. Strong in vitro antagonism by elm xylem endophytes is not accompanied by temporally stable in planta protection against a vascular pathogen under field conditions. Eur. J. Plant Pathol. 2015, 142, 185–196. [Google Scholar] [CrossRef]
- Sun, X.; Guo, L.-D. Endophytic fungal diversity: Review of traditional and molecular techniques. Mycology 2012, 3, 65–76. [Google Scholar]
- Siddique, A.B.; Khokon, A.M.; Unterseher, M. What do we learn from cultures in the omics age? High-throughput sequencing and cultivation of leaf-inhabiting endophytes from beech (Fagus sylvatica L.) revealed complementary community composition but similar correlations with local habitat conditions. MycoKeys 2017, 20, 1–16. [Google Scholar] [CrossRef]
- Lazarević, J.; Menkis, A. Fungi inhabiting fine roots of Pinus heldreichii in the Montenegrin montane forests. Symbiosis 2018, 74, 189–197. [Google Scholar] [CrossRef]
- Ahlich, K.; Sieber, T.N. The profusion of dark septate endophytic fungi in non-ectomycorrhizal fine roots of forest trees and shrubs. New Phytol. 1996, 132, 259–270. [Google Scholar] [CrossRef]
- Sieber, T.N.; Grünig, C.R. Fungal root endophytes. In Plant Roots: The Hidden Half, 4th ed.; Eshel, A., Beeckman, T., Eds.; CRC Press LLC: Boca Raton, FL, USA, 2013; pp. 38.1–38.49. [Google Scholar]
- Grünig, C.R.; Duò, A.; Sieber, T.N. Population genetic analysis of Phialocephala fortinii s.l. and Acephala applanata in two undisturbed forests in Switzerland and evidence for new cryptic species. Fungal Genet. Biol. 2006, 43, 410–421. [Google Scholar] [CrossRef]
- Stroheker, S.; Dubach, V.; Queloz, V.; Sieber, T.N. Resilience of Phialocephala fortinii s.l.—Acephala applanata communities—Effects of disturbance and strain introduction. Fungal Ecol. 2018, 31, 19–28. [Google Scholar] [CrossRef]
- Bensch, K.; Braun, U.; Groenewald, J.Z.; Crous, P.W. The genus Cladosporium. Stud. Mycol. 2012, 72, 1–401. [Google Scholar] [CrossRef]
- Paul, N.C.; Yu, S.H. Two species of endophytic Cladosporium in Pine trees in Korea. Mycobiology 2008, 36, 211–216. [Google Scholar] [CrossRef]
- Yuan, Z.; Chen, Y.; Yang, Y. Diverse non-mycorrhizal fungal endophytes inhabiting an epiphytic, medicinal orchid (Dendrobium nobile): Estimation and characterization. World J. Microbiol. Biotechnol. 2009, 25, 295. [Google Scholar] [CrossRef]
- Sutton, J.C.; Liu, W.; Ma, J.; Brown, W.G.; Stewart, J.F.; Walker, G.D. Evaluation of the fungal endophyte Clonostachys rosea as an inoculant to enhance growth, fitness and productivity of crop plants. Acta Hortic. 2008, 782, 279–286. [Google Scholar] [CrossRef]
- Manici, L.M.; Kelderer, M.; Caputo, F.; Saccà, M.L.; Nicoletti, F.; Topp, A.R.; Mazzola, M. Involvement of Dactylonectria and Ilyonectria spp. in tree decline affecting multi-generation apple orchards. Plant Soil 2018, 425, 217–230. [Google Scholar] [CrossRef]
- Weber, R.W.S.; Entrop, A.-P. Dactylonectria torresensis as the Main Component of the Black Root Rot Complex of Strawberries and Raspberries in Northern Germany. Erwerbs-Obstbau 2017, 59, 157–169. [Google Scholar] [CrossRef]
- Nigro, F.; Antelmi, I.; Sion, V.; Parente, P.; Pacifico, A. First Report of Dactylonectria torresensis Causing Foot and Root Rot of Olive Trees. Plant Dis. 2018. [Google Scholar] [CrossRef]
- Jankowiak, R.; Stępniewska, H.; Szwagrzyk, J.; Bilański, P.; Gratzer, G. Characterization of Cylindrocarpon-like species associated with litter in the old-growth beech forests of Central Europe. For. Pathol. 2016, 46, 582–594. [Google Scholar] [CrossRef]
- Demers, J.E.; Gugino, B.K.; Jiménez-Gasco, M.d.M. Highly Diverse Endophytic and Soil Fusarium oxysporum Populations Associated with Field-Grown Tomato Plants. Appl. Environ. Microbiol. 2015, 81, 81–90. [Google Scholar] [CrossRef]
- González-Teuber, M.; Vilo, C.; Bascuñán-Godoy, L. Molecular characterization of endophytic fungi associated with the roots of Chenopodium quinoa inhabiting the Atacama Desert, Chile. Genom. Data 2017, 11, 109–112. [Google Scholar] [CrossRef]
- Rodriguez, R.J.; White, J.F., Jr.; Arnold, A.E.; Redman, R.S. Fungal endophytes: Diversity and functional roles: Tansley review. New Phytol. 2009, 182, 314–330. [Google Scholar] [CrossRef]
- Castañares, E.; Stenglein, S.A.; Dinolfo, M.I.; Moreno, M.V. Fusarium tricinctum Associated with Head Blight on Wheat in Argentina. Plant Dis. 2010, 95, 496. [Google Scholar] [CrossRef]
- Görke, C. Mykozönosen von Wurzel und Stamm von Jungbäumen unterschiedlicher Bestandsbegründungen. Available online: https://www.schweizerbart.de/publications/detail/isbn/9783443590758 (accessed on 28 February 2019).
- Sieber, T.N. Endophytische Pilze von Winterweizen (Triticum vulgare Vill.): Ein Vergleich Zwischen Weizen aus Gebeiztem und Solchem aus Ungebeiztem Saatgut; ETH Zürich: Zürich, Switzerland, 1985. [Google Scholar]
- Stark, C.; Babik, W.; Durka, W. Fungi from the roots of the common terrestrial orchid Gymnadenia conopsea. Mycol. Res. 2009, 113, 952–959. [Google Scholar] [CrossRef] [PubMed]
- Petrini, O.; Fisher, P.; Petrini, L. Fungal endophytes of bracken (Pteridium aquilinum), with some reflections on their use in biological control. Sydowia 1992, 44, 282–293. [Google Scholar]
- Cabral, A.; Rego, C.; Nascimento, T.; Oliveira, H.; Groenewald, J.Z.; Crous, P.W. Multi-gene analysis and morphology reveal novel Ilyonectria species associated with black foot disease of grapevines. Fungal Biol. 2012, 116, 62–80. [Google Scholar] [CrossRef] [PubMed]
- Blumenstein, K.; Albrectsen, B.R.; Martín, J.A.; Hultberg, M.; Sieber, T.N.; Helander, M.; Witzell, J. Nutritional niche overlap potentiates the use of endophytes in biocontrol of a tree disease. BioControl 2015, 60, 655–667. [Google Scholar] [CrossRef]
- Sánchez Márquez, S.; Bills, G.F.; Domínguez Acuña, L.; Zabalgogeazcoa, I. Endophytic mycobiota of leaves and roots of the grass Holcus lanatus. Fungal Divers. 2010, 41, 115–123. [Google Scholar] [CrossRef]
- Nicoletti, R.; Fiorentino, A.; Scognamiglio, M. Endophytism of Penicillium _Species in Woody Plants. TOMYCJ 2014, 8, 1–26. [Google Scholar] [CrossRef]
- Courtois, H. The mycoflora in the crown region and rhizosphere of Norway spruce (Picea abies), and its significance for interpreting forest decline symptoms. Angew. Bot. 1990, 64, 381–392. [Google Scholar]
- Bezerra, J.D.P.; Santos, M.G.S.; Svedese, V.M.; Lima, D.M.M.; Fernandes, M.J.S.; Paiva, L.M.; Souza-Motta, C.M. Richness of endophytic fungi isolated from Opuntia ficus-indica Mill. (Cactaceae) and preliminary screening for enzyme production. World J. Microbiol. Biotechnol. 2012, 28, 1989–1995. [Google Scholar] [CrossRef] [PubMed]
- Allen, T.R.; Millar, T.; Berch, S.M.; Berbee, M.L. Culturing and direct DNA extraction find different fungi from the same ericoid mycorrhizal roots. New Phytol. 2003, 160, 255–272. [Google Scholar] [CrossRef]
- Menkis, A.; Vasaitis, R. Fungi in Roots of Nursery Grown Pinus sylvestris: Ectomycorrhizal Colonisation, Genetic Diversity and Spatial Distribution. Microb. Ecol. 2011, 61, 52–63. [Google Scholar] [CrossRef]
- Summerbell, R.C. Microfungi associated with the mycorrhizal mantle and adjacent microhabitats within the rhizosphere of black spruce. Can. J. Bot. 1989, 67, 1085–1095. [Google Scholar] [CrossRef]
- An, H.; Liu, Y.; Zhao, X.; Huang, Q.; Yuan, S.; Yang, X.; Dong, J. Characterization of cadmium-resistant endophytic fungi from Salix variegata Franch. in Three Gorges Reservoir Region, China. Microbiol. Res. 2015, 176, 29–37. [Google Scholar] [CrossRef]
- Schlegel, M.; Dubach, V.; von Buol, L.; Sieber, T.N. Effects of endophytic fungi on the ash dieback pathogen. FEMS Microbiol. Ecol. 2016, 92. [Google Scholar] [CrossRef]
- Kwaśna, H.; Mazur, A.; Łabędzki, A.; Kuźmiński, R.; Łakomy, P. Communities of fungi in decomposed wood of oak and pine. For. Res. Pap. 2016, 77, 261–275. [Google Scholar] [CrossRef]
- Vaz, A.B.M.; Mota, R.C.; Bomfim, M.R.Q.; Vieira, M.L.A.; Zani, C.L.; Rosa, C.A.; Rosa, L.H. Antimicrobial activity of endophytic fungi associated with Orchidaceae in Brazil. Can. J. Microbiol. 2009, 55, 1381–1391. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Lie, T.K.; Qian, X.; Zheng, Z.; Huang, Y.; Shen, Y. Species Diversity, Distribution, and Genetic Structure of Endophytic and Epiphytic Trichoderma Associated with Banana Roots. Microb. Ecol. 2011, 61, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Werner, C.; Petrini, O.; Hesse, M. Degradation of the polyamine alkaloid aphelandrine by endophytic fungi isolated from Aphelandra tetragona. FEMS Microbiol. Lett. 1997, 155, 147–153. [Google Scholar] [CrossRef][Green Version]
- Kattner, D.; Schönhar, S. Occurrence of microscopic fungi in fine roots of apparently healthy Norway spruce on various sites. Mitteilungen Verein Für Forstliche Standortskunde Forstpflanzenzüchtung 1990, 35, 39–43. [Google Scholar]
- Gan, H.; Churchill, A.C.L.; Wickings, K. Invisible but consequential: Root endophytic fungi have variable effects on belowground plant–insect interactions. Ecosphere 2017, 8, e01710. [Google Scholar] [CrossRef]
- Schlegel, M.; Münsterkötter, M.; Güldener, U.; Bruggmann, R.; Duò, A.; Hainaut, M.; Henrissat, B.; Sieber, C.M.K.; Hoffmeister, D.; Grünig, C.R. Globally distributed root endophyte Phialocephala subalpina links pathogenic and saprophytic lifestyles. BMC Genom. 2016, 17, 1015. [Google Scholar] [CrossRef]
- Bußkamp, J. Schadenserhebung, Kartierung und Charakterisierung des “Diplodia-Triebsterbens” der Kiefer, Insbesondere des Endophytischen Vorkommens in den Klimasensiblen Räumen und Identifikation von den in Kiefer (Pinus sylvestris) Vorkommenden Endophyten. Ph.D. Thesis, University of Kassel, Kassel, Germany, 2018. [Google Scholar]
| Host Species | Common Name | Native to | Abbreviation |
|---|---|---|---|
| Picea abies | Norway spruce | Northern, Central, and Eastern Europe | Abie |
| Picea glauca | White/Canadian spruce | Alaska through central Canada to Newfoundland | Glau |
| Picea omorika | Serbian spruce | Endemic to Drina river valley, Serbia | Omori |
| Picea pungens | Blue/Colorado spruce | Rocky Mountains, USA | Pung |
| Picea sitchensis | Sitka spruce | West coast Canada, down to California | Sitch |
| Pinus jeffreyi | Jeffrey pine | Oregon-California, USA | Jeff |
| Pinus peuce | Macedonian/Balkan pine | Mountains of Balkan region | Peuc |
| Pinus sylvestris | Scots pine | Eurasia | Sylv |
| Identification | Best Match Accession no. | Order | Class | Picea | Pinus | Sum | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| abies | glauca | omorika | sitchensis | jeffreyi | peuce | sylvestris | |||||
| Pseudogymnoascus sp. | MH857254 | Incertae sedis | Ascomycetes | 1 | 1 | ||||||
| Cladosporium sp. | MH863129 | Capnodiales | Dothideomycetes | 2 | 2 | ||||||
| Pyrenochaeta sp. | KT269928 | Pleosporales | Dothideomycetes | 2 | 4 | 6 | |||||
| Pyrenochaeta sp. | LT821390 | Pleosporales | Dothideomycetes | 1 | 7 | 8 | |||||
| Penicillium sp. | MG821367 | Eurotiales | Eurotiomycetes | 1 | 1 | ||||||
| Penicillium sp. | MH865458 | Eurotiales | Eurotiomycetes | 1 | 1 | ||||||
| PAC b | AY606286 | Helotiales | Leotiomycetes | 1 | 1 | ||||||
| Fungal sp. | KY322665 | NA | NA | 2 | 2 | ||||||
| Pezizales sp. | MH859398 | Pezizales | Pezizomycetes | 1 | 1 | ||||||
| Clonostachys sp. | KY977560 | Hypocreales | Sordariomycetes | 3 | 3 | ||||||
| Dactylonectria sp. | MH865183 | Hypocreales | Sordariomycetes | 8 | 1 | 9 | |||||
| Fusarium sp. | MG252283 | Hypocreales | Sordariomycetes | 2 | 6 | 8 | |||||
| Fusarium sp. | MG704912 | Hypocreales | Sordariomycetes | 2 | 2 | ||||||
| Ilyonectria sp. | MH865177 | Hypocreales | Sordariomycetes | 1 | 2 | 3 | |||||
| Trichoderma sp. | MH930456 | Hypocreales | Sordariomycetes | 5 | 5 | ||||||
| Unknown | NA | NA | 6 | 6 | |||||||
| Unknown | NA | NA | 2 | 2 | |||||||
| Unknown | NA | NA | 1 | 1 | |||||||
| Unknown | NA | NA | 3 | 3 | |||||||
| TOTAL | 11 | 9 | 9 | 10 | 18 | 2 | 6 | 65 | |||
| Sample ID | Host | Best Match | Accession | Mi/Qc a | Region | Our Definition |
|---|---|---|---|---|---|---|
| Abie23 | Picea abies | Clonostachys candelabrum | KY977560 | 100/100 | Croatia | Clonostachys sp. |
| Abie26 | Picea abies | Fusarium tricinctum | MG704912 | 100/099 | S. Korea | Fusarium sp. |
| Abie27 | Picea abies | Pyrenochaeta sp. | KT269928 | 099/100 | Greece | Pyrenochaeta sp. |
| Glau13 | Picea glauca | Dactylonectria torresensis | MH865183 | 098/100 | Portugal | Dactylonectria sp. |
| Glau17 | Picea glauca | Dactylonectria torresensis | MH865183 | 099/100 | Portugal | Dactylonectria sp. |
| Glau18 | Picea glauca | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Glau21 | Picea glauca | Dactylonectria torresensis | MH865183 | 100/100 | Portugal | Dactylonectria sp. |
| Jeff49 | Pinus jeffreyi | Fusarium solani | MG252283 | 099/100 | China | Fusarium sp. |
| Jeff51 | Pinus jeffreyi | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Jeff56 | Pinus jeffreyi | Cladosporium subinflatum | MH863129 | 099/100 | Slovenia | Cladosporium sp. |
| Jeff59 | Pinus jeffreyi | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Jeff60 | Pinus jeffreyi | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Jeff61 | Pinus jeffreyi | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Jeff65 | Pinus jeffreyi | Pyrenochaeta inflorescentiae | LT821390 | 099/100 | Germany | Pyrenochaeta sp. |
| Omori01 | Picea omorika | Penicillium glandicola | MG821367 | 099/100 | Italy | Penicillium sp. |
| Omori02 | Picea omorika | Penicillium granulatum | MH865458 | 099/100 | USA | Penicillium sp. |
| Omori03 | Picea omorika | Fungal sp. KK15 | KY322665 | 099/100 | Montenegro | NA |
| Omori07 | Picea omorika | Phialocephala fortinii | AY606286 | 099/100 | Sweden | PAC b |
| Omori08 | Picea omorika | Ilyonectria lusitanica | MH865177 | 099/100 | Portugal | Ilyonectria sp. |
| Peuc68 | Pinus peuce | Dactylonectria torresensis | MH865183 | 100/100 | Portugal | Dactylonectria sp. |
| Peuc69 | Pinus peuce | Pseudogymnoascus pannorum | MH857254 | 100/100 | Germany | Pseudogymnoascus sp. |
| Sylv36 | Pinus sylvestris | Trichoderma koningii | MH930456 | 099/100 | Spain | Trichoderma sp. |
| Sylv37 | Pinus sylvestris | Trichoderma koningii | MH930456 | 099/100 | Spain | Trichoderma sp. |
| Sylv38 | Pinus sylvestris | Ascobolus lineolatus | MH859398 | 097/099 | Tanzania | Pezizales sp. |
| Sample ID | Host | Identification | Sampling Time (days) a | ||
|---|---|---|---|---|---|
| 3 | 7 | 10 | |||
| Abie25 | Picea abies | Fusarium sp. | N | Y | N |
| Abie29 | Picea abies | Unknown | N | N | Y |
| Glauc15 | Picea glauca | Dactylonectria sp. | N | Y | Y |
| Glauc19 | Picea glauca | Dactylonectria sp. | N | N | Y |
| Glauc13 | Picea glauca | Dactylonectria sp. | N | Y | N |
| Glauc18 | Picea glauca | Pyrenochaeta sp. | N | Y | Y |
| Omori01 | Picea omorika | Penicillium sp. | N | Y | N |
| Omori03 | Picea omorika | Fungal sp. | N | Y | Y |
| Omori07 | Picea omorika | PAC b | N | N | Y |
| Omori08 | Picea omorika | Ilyonectria sp. | N | Y | N |
| Omori06 | Picea omorika | Fungal sp. | N | Y | N |
| Jeff50 | Pinus jeffreyi | Fusarium sp. | N | Y | Y |
| Jeff55 | Pinus jeffreyi | Fusarium sp. | N | N | Y |
| Jeff58 | Pinus jeffreyi | Fusarium sp. | N | N | Y |
| Jeff62 | Pinus jeffreyi | Pyrenochaeta sp. | N | N | Y |
| Jeff63 | Pinus jeffreyi | Pyrenochaeta sp. | N | N | Y |
| Jeff64 | Pinus jeffreyi | Unknown | N | N | Y |
| Jeff51 | Pinus jeffreyi | Pyrenochaeta sp. | N | Y | N |
| Jeff56 | Pinus jeffreyi | Cladosporium sp. | N | Y | N |
| Jeff65 | Pinus jeffreyi | Pyrenochaeta sp. | N | N | Y |
| Sitch40 | Pinus sitchensis | Pyrenochaeta sp. | N | N | Y |
| Sitch41 | Pinus sitchensis | Pyrenochaeta sp. | N | N | Y |
| Sitch42 | Pinus sitchensis | Unknown | N | N | Y |
| Sitch46 | Pinus sitchensis | Unknown | N | Y | Y |
| Sitch47 | Pinus sitchensis | Pyrenochaeta sp. | N | N | Y |
| Sitch48 | Pinus sitchensis | Pyrenochaeta sp. | N | Y | Y |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rigerte, L.; Blumenstein, K.; Terhonen, E. New R-Based Methodology to Optimize the Identification of Root Endophytes against Heterobasidion parviporum. Microorganisms 2019, 7, 102. https://doi.org/10.3390/microorganisms7040102
Rigerte L, Blumenstein K, Terhonen E. New R-Based Methodology to Optimize the Identification of Root Endophytes against Heterobasidion parviporum. Microorganisms. 2019; 7(4):102. https://doi.org/10.3390/microorganisms7040102
Chicago/Turabian StyleRigerte, Linda, Kathrin Blumenstein, and Eeva Terhonen. 2019. "New R-Based Methodology to Optimize the Identification of Root Endophytes against Heterobasidion parviporum" Microorganisms 7, no. 4: 102. https://doi.org/10.3390/microorganisms7040102
APA StyleRigerte, L., Blumenstein, K., & Terhonen, E. (2019). New R-Based Methodology to Optimize the Identification of Root Endophytes against Heterobasidion parviporum. Microorganisms, 7(4), 102. https://doi.org/10.3390/microorganisms7040102





