The Purple Sea Urchin Strongylocentrotus purpuratus Demonstrates a Compartmentalization of Gut Bacterial Microbiota, Predictive Functional Attributes, and Taxonomic Co-Occurrence
Abstract
1. Introduction
2. Materials and Methods
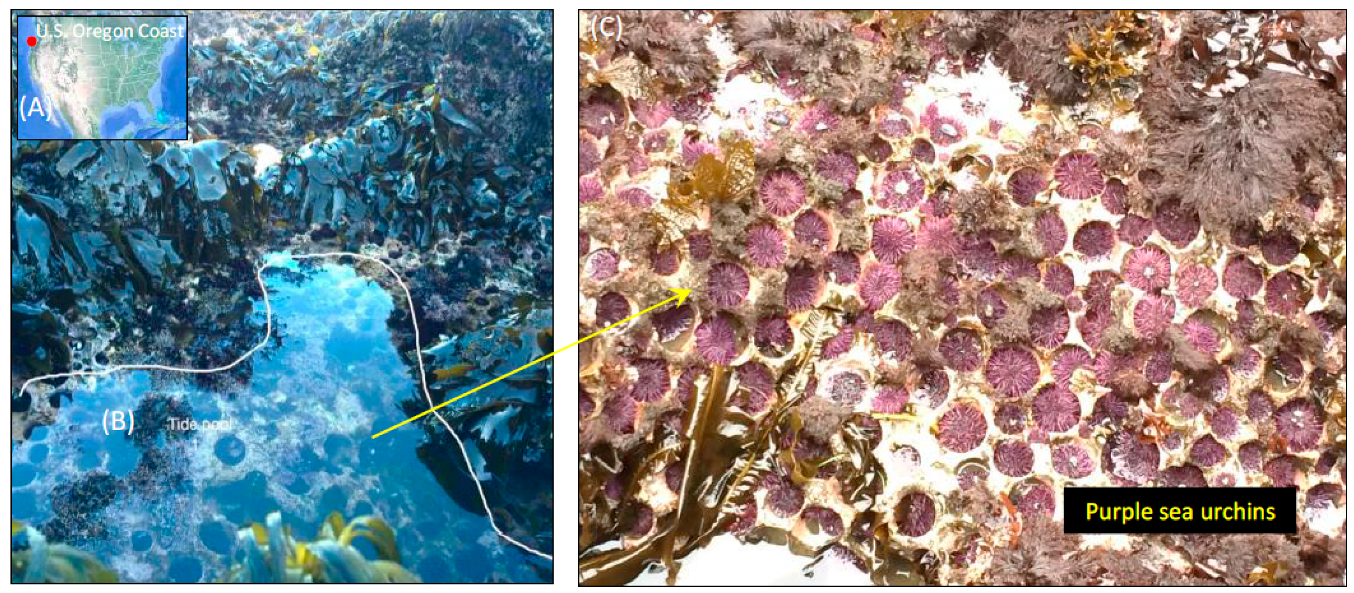
2.1. Collection and Sample Preparation of S. Purpuratus
2.2. Community DNA Extraction, Illumina MiSeq Sample Preparation, and High-Throughput Sequencing
2.3. Quality Assessment and Filtering
2.4. Taxonomic Distribution
2.5. Alpha Diversity
2.6. Beta Diversity
2.7. Predicted Functional Analysis
2.8. Co-Occurrence Analysis of Microbial Taxa
3. Results
3.1. Environmental Conditions and Sea Urchin Measurements
3.2. Quality Assessment and Sample Statistics
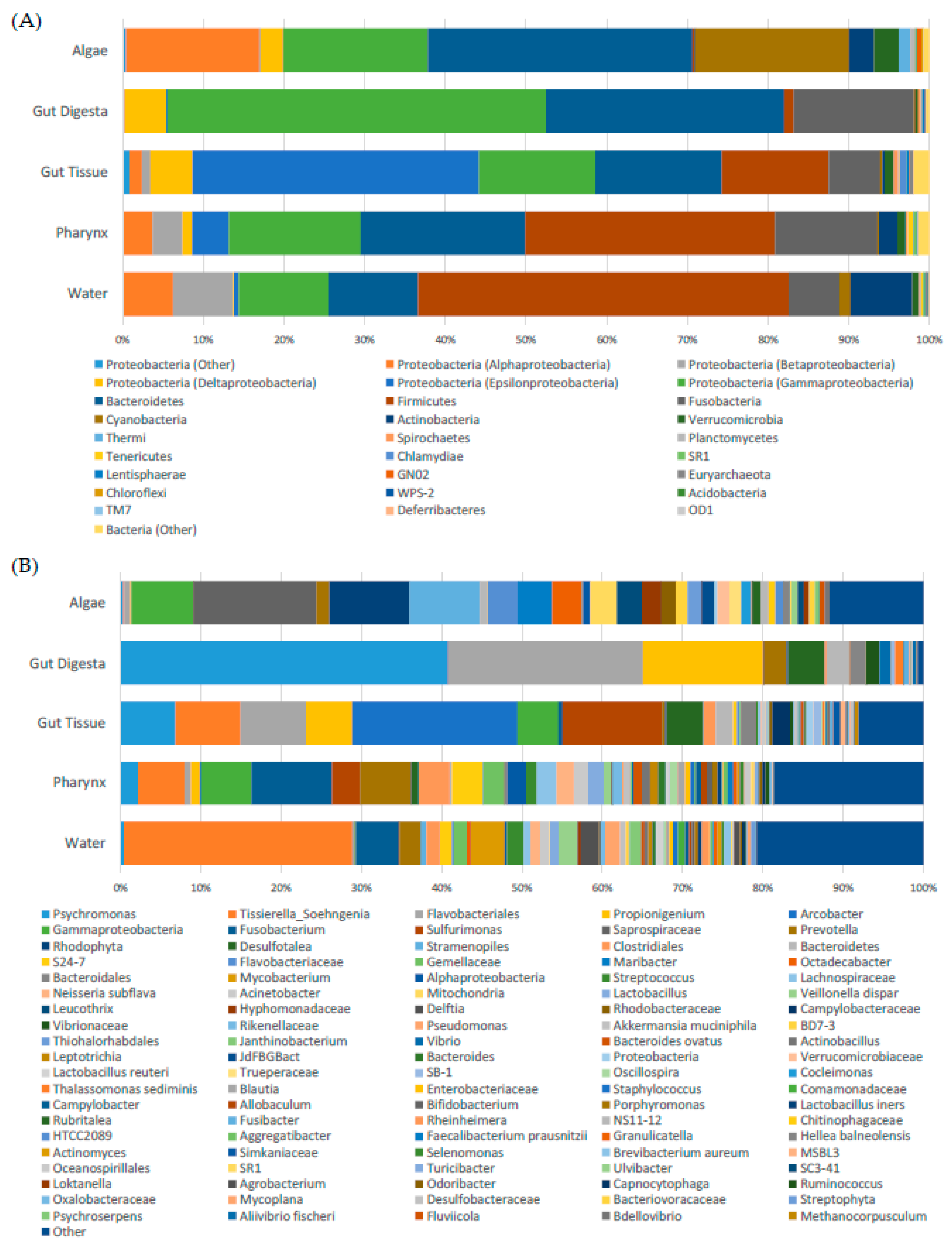
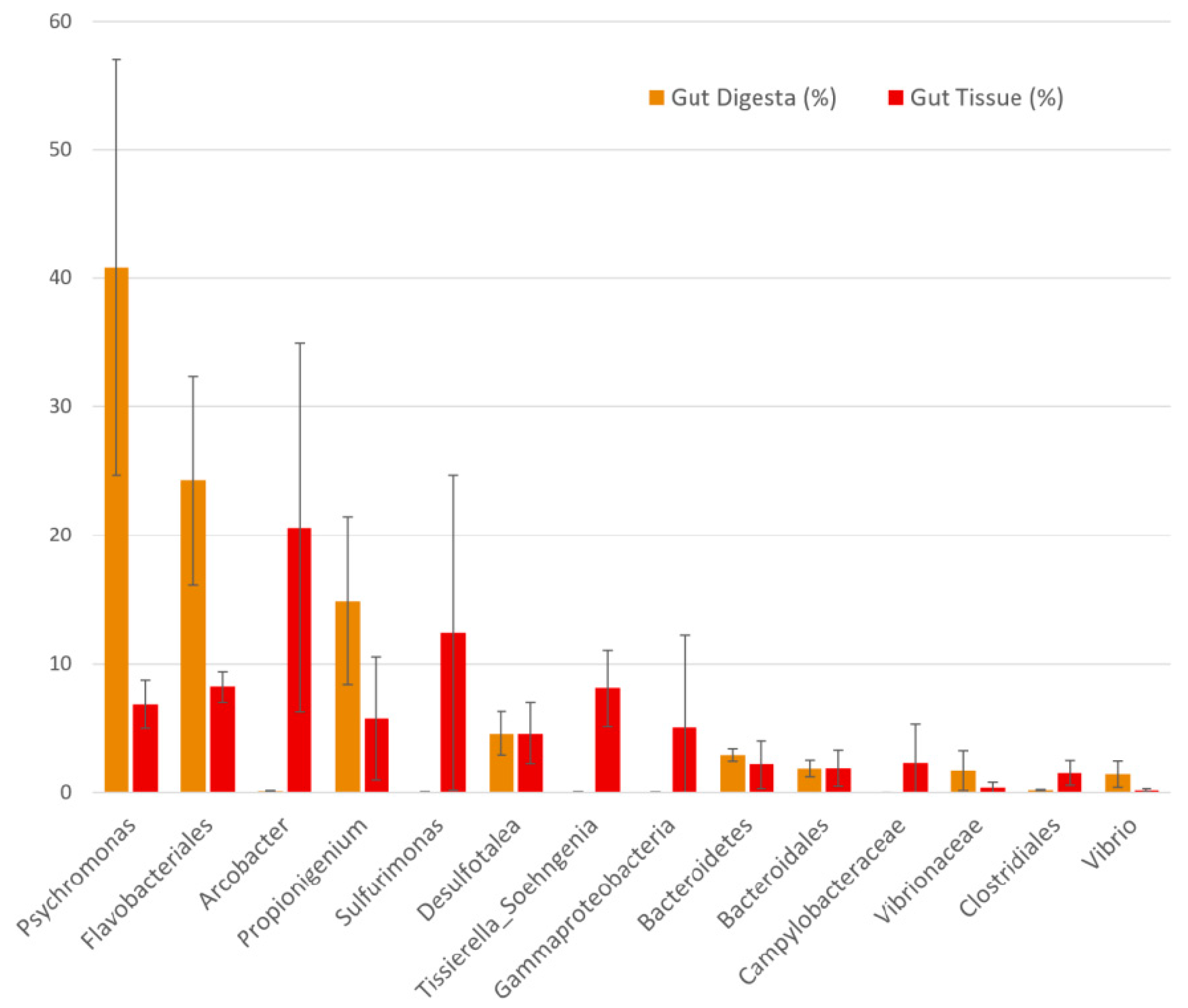
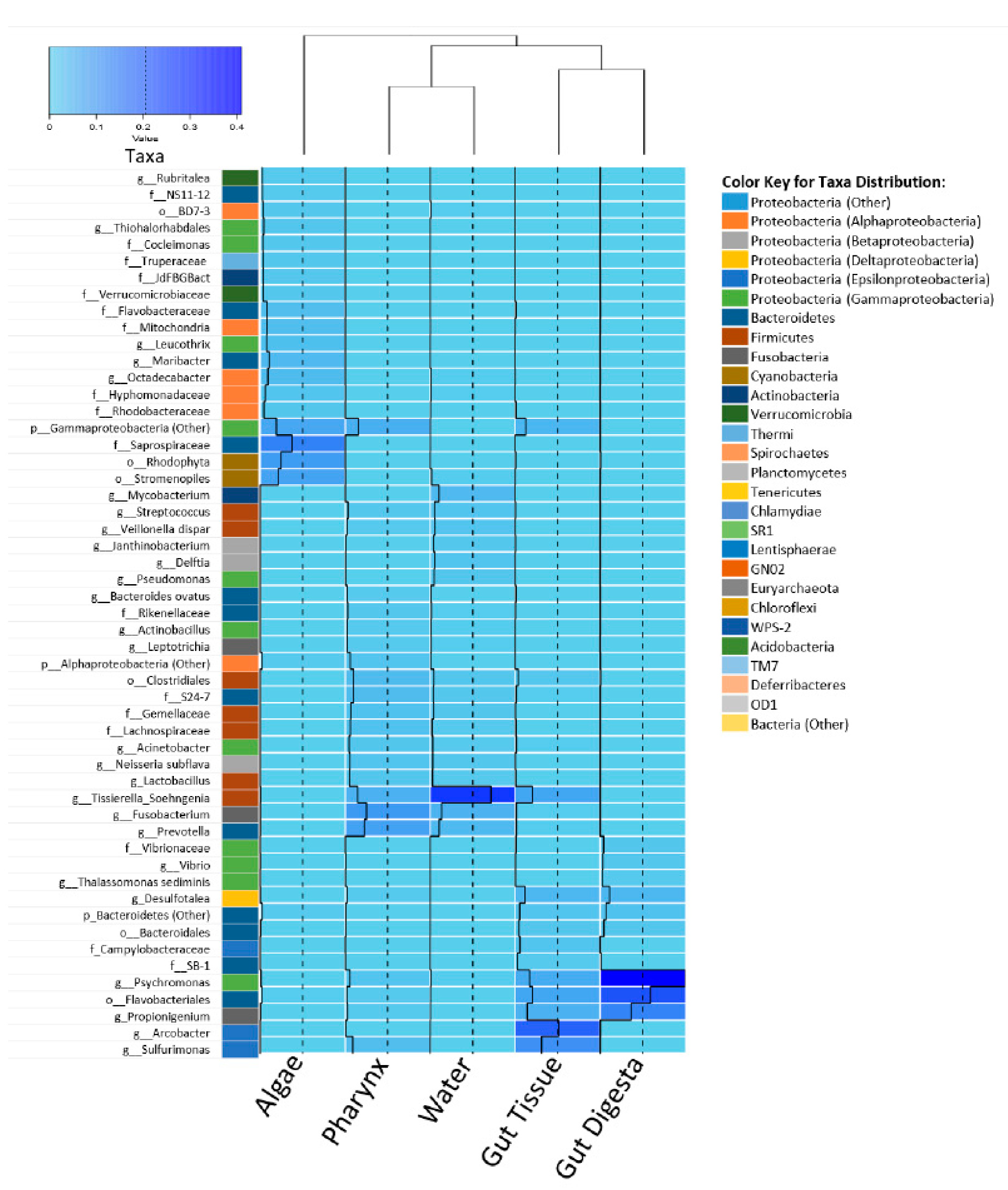
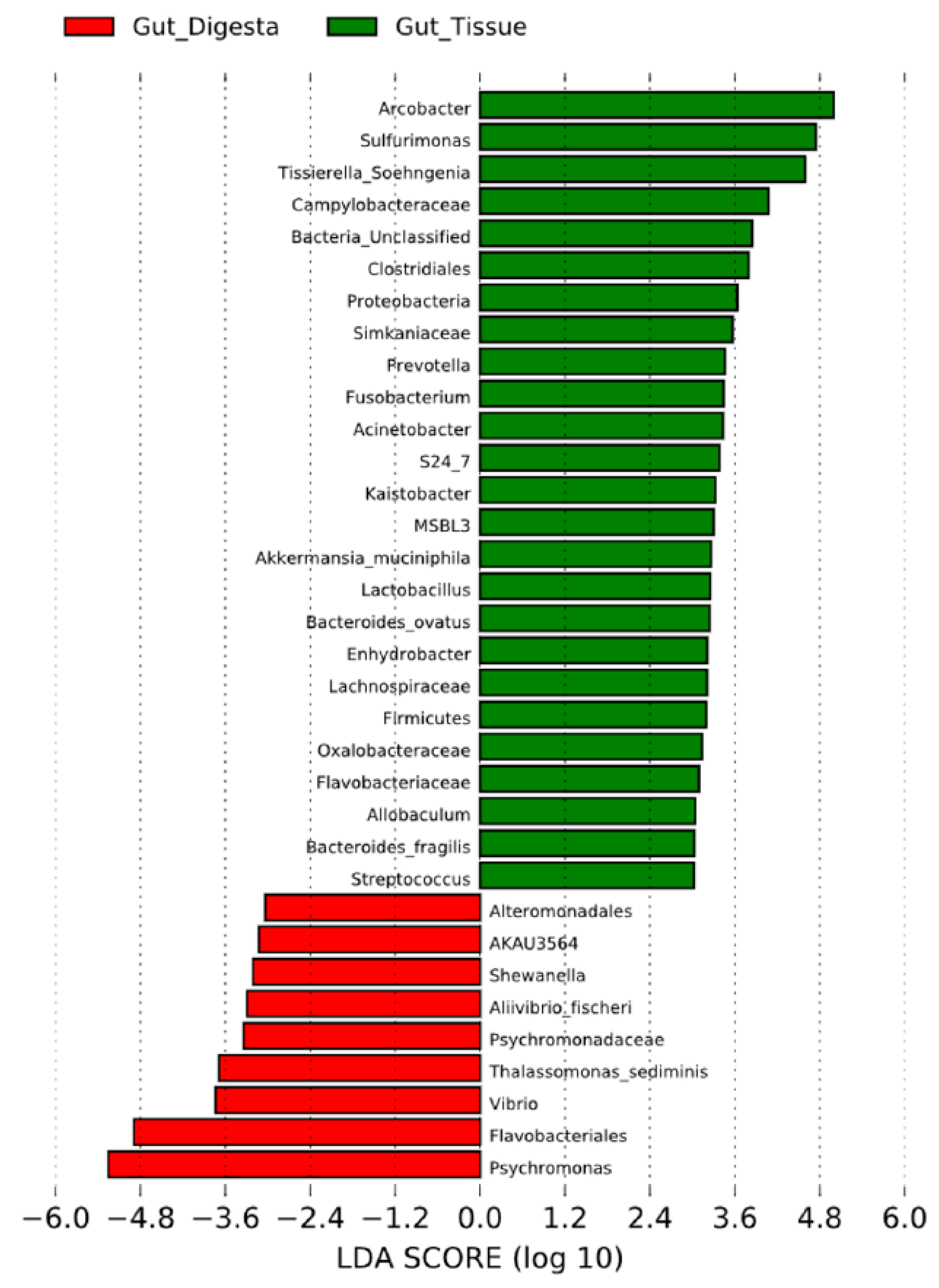
3.3. Taxonomic Distribution across Samples
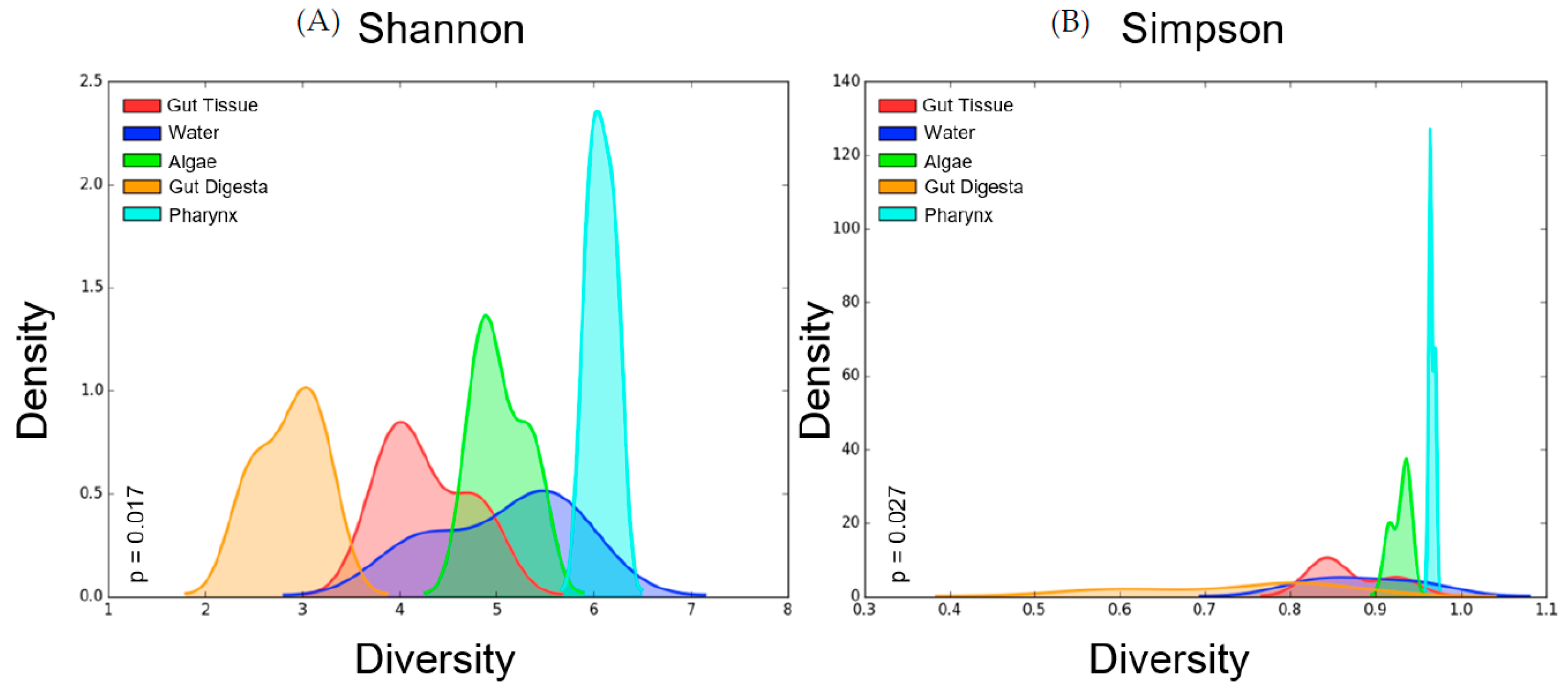
3.4. Alpha Diversity
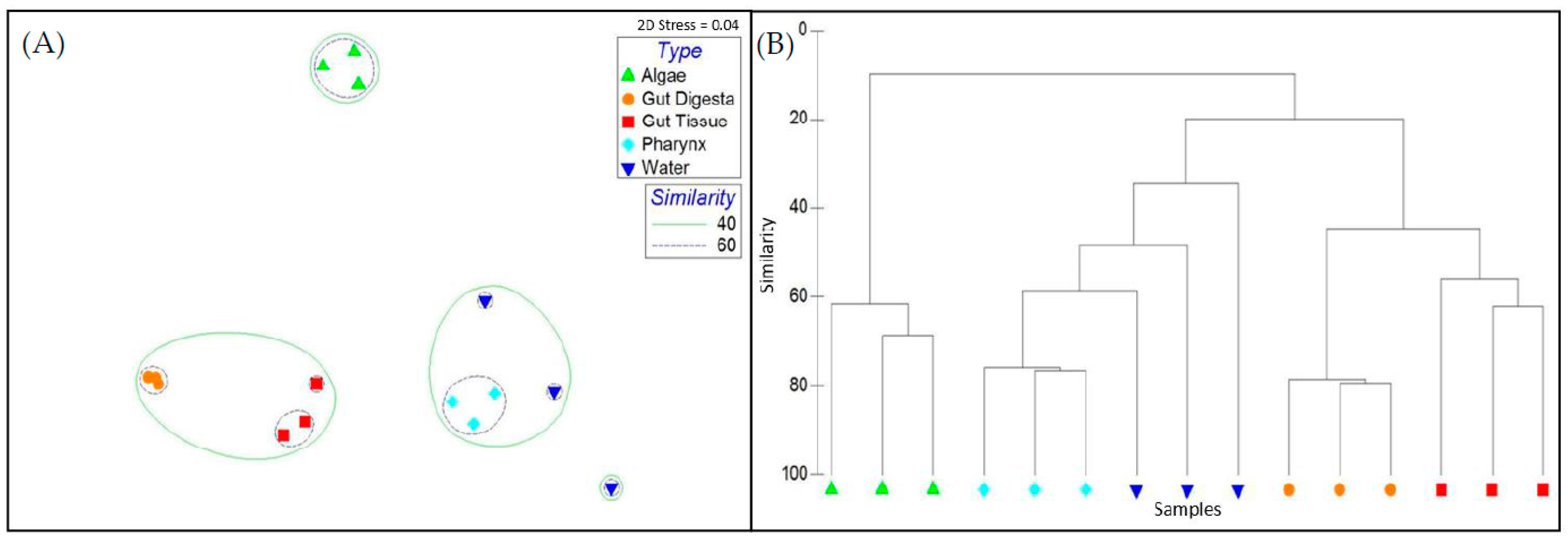
3.5. Beta Diversity
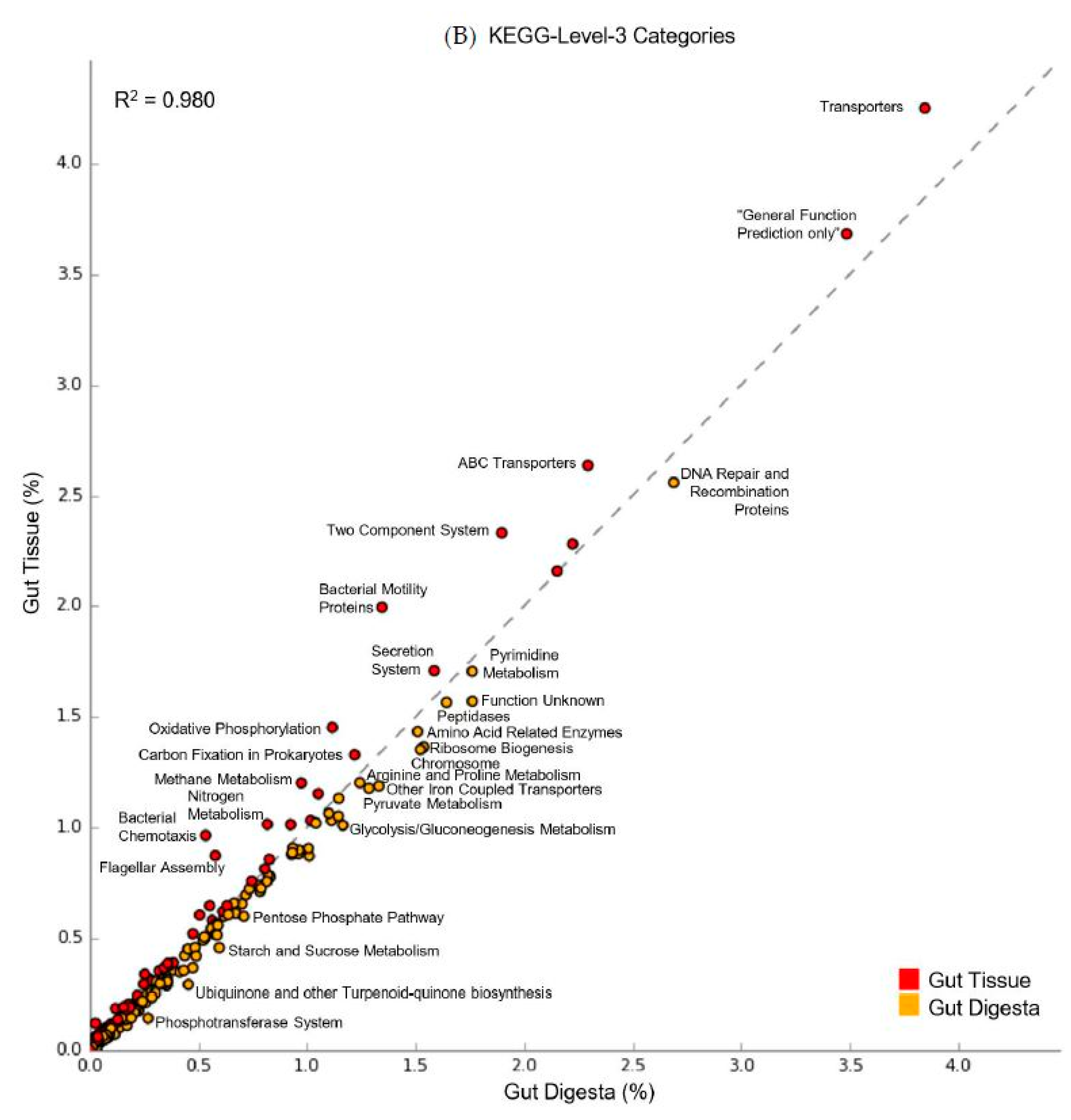
3.6. Predicted Functional Capacity
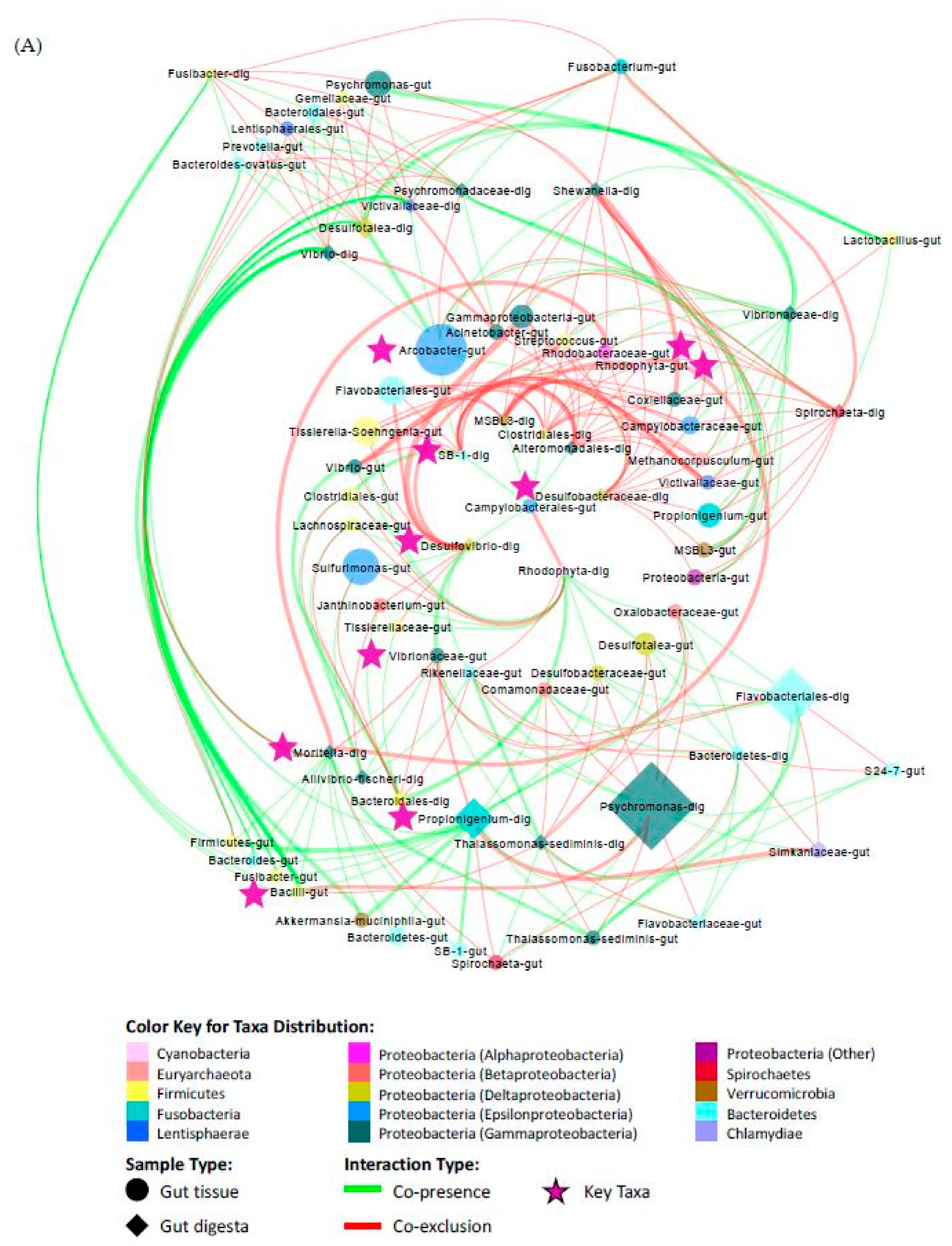
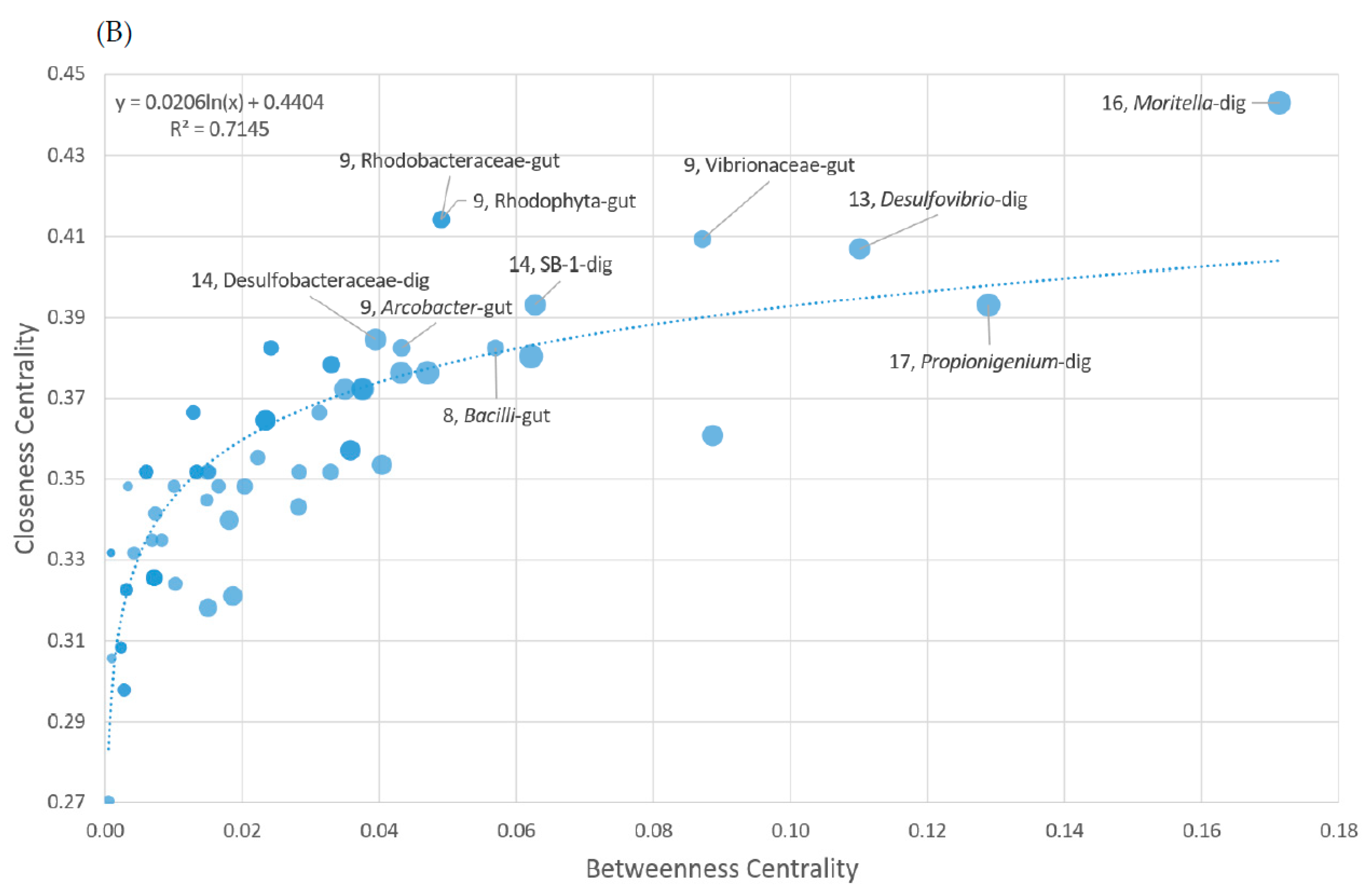
3.7. Co-Presence, Co-Exclusion, and Key Taxa in the Gut Environment
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ebert, T.A.; Schroeter, S.C.; Dixon, J.D.; Kalvass, P. Settlement patterns of red and purple sea urchins (Strongylocentrotus franciscanus and S. purpuratus) in California, USA. Mar. Ecol. Prog. Ser. 1994, 111, 41–52. [Google Scholar] [CrossRef]
- Dethier, M.N. Disturbance and recovery in intertidal pools: Maintenance of mosaic patterns. Ecol. Monogr. 1984, 54, 99–118. [Google Scholar] [CrossRef]
- Lafferty, K.D.; Kushner, D. Population regulation of the purple sea urchin (Strongylocentrotus purpuratus) at the California Channel Islands. In Proceedings of the 5th California Islands Symposium; Browne, D.R., Mitchell, K.L., Chaney, H.W., Eds.; Publication 99-0038; Minerals Management Service: Santa Barbara, CA, USA, 2000; pp. 379–381. [Google Scholar]
- Watanabe, J.M.; Harrold, C. Destructive grazing by sea urchins Strongylocentrotus spp. in a central California kelp forest: Potential roles of recruitment, depth, and predation. Mar. Ecol. Prog. Ser. 1991, 125–141. [Google Scholar] [CrossRef]
- Metaxas, A.; Scheibling, R.E. Community structure and organization of tidepools. Mar. Ecol. Prog. Ser. 1993, 187–198. [Google Scholar] [CrossRef]
- Tegner, M.J.; Dayton, P.K. Ecosystem effects of fishing in kelp forest communities. ICES J. Mar. Sci. 2000, 57, 579–589. [Google Scholar] [CrossRef]
- Davidson, T.M.; Grupe, B.M. Habitat modification in tidepools by bioeroding sea urchins and implications for fine-scale community structure. Mar. Ecol. 2015, 36, 185–194. [Google Scholar] [CrossRef]
- Sea Urchin Genome Sequencing Consortium. The genome of the sea urchin Strongylocentrotus purpuratus. Science 2006, 314, 941–952. [Google Scholar] [CrossRef]
- Lasker, R.; Giese, A.C. Nutrition of the sea urchin, Strongylocentrotus purpuratus. Biol. Bull. 1954, 106, 328–340. [Google Scholar] [CrossRef]
- Holland, N.D.; Ghiselin, M.T. A comparative study of gut mucous cells in thirty-seven species of the class Echinoidea (Echinodermata). Biol. Bull. 1970, 138, 286–305. [Google Scholar] [CrossRef]
- De Ridder, C.; Jangoux, M. Digestive system: Echinoidea. In Echinoderm Nutrition; Jangoux, M., Lawrence, J.M., Eds.; A.A. Balkema Publ.: Rotterdam, The Netherlands, 1982; pp. 213–234. ISBN 90-6191-080-3. [Google Scholar]
- Ziegler, A.; Mooi, R.; Rolet, G.; De Ridder, C. Origin and evolutionary plasticity of the gastric caecum in sea urchins (Echinodermata: Echinoidea). BMC Evol. Biol. 2010, 10, 313. [Google Scholar] [CrossRef]
- Sawabe, T.; Oda, Y.; Shiomi, Y.; Ezura, Y. Alginate degradation by bacteria isolated from the gut of sea urchins and abalones. Microb. Ecol. 1995, 30, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Fong, W.; Mann, K.H. Role of gut flora in the transfer of amino acids through a marine food chain. Can. J. Fish. Aquat. Sci. 1980, 37, 88–96. [Google Scholar] [CrossRef]
- Hakim, J.A.; Koo, H.; Dennis, L.N.; Kumar, R.; Ptacek, T.; Morrow, C.D.; Lefkowitz, E.J.; Powell, M.L.; Bej, A.K.; Watts, S.A. An abundance of Epsilonproteobacteria revealed in the gut microbiome of the laboratory cultured sea urchin, Lytechinus variegatus. Front. Microbiol. 2015, 6, 1047. [Google Scholar] [CrossRef] [PubMed]
- Hakim, J.A.; Koo, H.; Kumar, R.; Lefkowitz, E.J.; Morrow, C.D.; Powell, M.L.; Watts, S.A.; Bej, A.K. The gut microbiome of the sea urchin, Lytechinus variegatus, from its natural habitat demonstrates selective attributes of microbial taxa and predictive metabolic profiles. FEMS Microbiol. Ecol. 2016, 92, fiw146. [Google Scholar] [CrossRef] [PubMed]
- Sauchyn, L.K.; Lauzon-Guay, J.S.; Scheibling, R.E. Sea urchin fecal production and accumulation in a rocky subtidal ecosystem. Aquat. Biol. 2011, 13, 215–223. [Google Scholar] [CrossRef]
- Sauchyn, L.K.; Scheibling, R.E. Degradation of sea urchin feces in a rocky subtidal ecosystem: Implications for nutrient cycling and energy flow. Aquat. Biol. 2009, 6, 99–108. [Google Scholar] [CrossRef]
- Sauchyn, L.K.; Scheibling, R.E. Fecal production by sea urchins in native and invaded algal beds. Mar. Ecol. Prog. Ser. 2009, 396, 35–48. [Google Scholar] [CrossRef]
- Schram, J.B.; Kobelt, J.N.; Dethier, M.N.; Galloway, A.W.E. Trophic transfer of macroalgal fatty acids in two urchin species: Digestion, egestion, and tissue building. Front. Ecol. Evol. 2018, 6, 83. [Google Scholar] [CrossRef]
- Troussellier, M.; Escalas, A.; Bouvier, T.; Mouillot, D. Sustaining rare marine microorganisms: Macroorganisms as repositories and dispersal agents of microbial diversity. Front. Microbiol. 2017, 8, 947. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Brothers, C.J.; Van Der Pol, W.J.; Morrow, C.D.; Hakim, J.A.; Koo, H.; McClintock, J.B. Ocean warming alters predicted microbiome functionality in a common sea urchin. Proc. R. Soc. B 2018, 285, 20180340. [Google Scholar] [CrossRef] [PubMed]
- Dabdoub, S.M.; Fellows, M.L.; Paropkari, A.D.; Mason, M.R.; Huja, S.S.; Tsigarida, A.A.; Kumar, P.S. PhyloToAST: Bioinformatics tools for species-level analysis and visualization of complex microbial datasets. Sci. Rep. 2016, 6, 29123. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Gonzalez Peña, A.; Goodrich, J.K.; Gordon, J.G.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Meth. 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Faust, K.; Sathirapongsasuti, J.F.; Izard, J.; Segata, N.; Gevers, D.; Raes, J.; Huttenhower, C. Microbial co-occurrence relationships in the human microbiome. PLOS Comput. Biol. 2012, 8, e1002606. [Google Scholar] [CrossRef] [PubMed]
- Faust, K.; Raes, J. Microbial interactions: From networks to models. Nat. Rev. Microbiol. 2012, 10, 538–550. [Google Scholar] [CrossRef] [PubMed]
- Faust, K.; Raes, J. CoNet app: Inference of biological association networks using Cytoscape. F1000Research 2016, 5, 1519. [Google Scholar] [CrossRef]
- Berry, D.; Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 2014, 5, 219. [Google Scholar] [CrossRef]
- Banerjee, S.; Schlaeppi, K.; Heijden, M.G. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 2018, 16, 567–576. [Google Scholar] [CrossRef]
- Song, S.J.; Amir, A.; Metcalf, J.L.; Amato, K.R.; Xu, Z.Z.; Humphrey, G.; Knight, R. Preservation methods differ in fecal microbiome stability, affecting suitability for field studies. mSystems 2016, 1, e00021-16. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4516–4522. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Eipers, P.; Little, R.B.; Crowley, M.; Crossman, D.K.; Lefkowitz, E.J.; Morrow, C.D. Getting started with microbiome analysis: Sample acquisition to bioinformatics. Curr. Protoc. Hum. Genet. 2014, 82, 18.8.1–18.8.29. [Google Scholar] [CrossRef] [PubMed]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Ausubel, F.M. Current Protocols in Molecular Biology; Ausubel, F.M., Brent, R., Kingston, R.F., Moore, D.D., Seidman, J.G., Smith, J.A., Struhl, K., Eds.; Publishing Associates and Wiley-Interscience: New York, NY, USA, 1987; ISBN 978-047-162-594-0. [Google Scholar]
- Cock, P.J.A.; Fields, C.J.; Goto, N.; Heuer, M.L.; Rice, P.M. The Sanger FASTQ File Format for Sequences with Quality Scores, and the Solexa/Illumina FASTQ Variants. Nucleic Acids Res. 2009, 38, 1767–1771. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S. FastQC: A quality Control Tool for High Throughput Sequence Data. 2010. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 12 January 2019).
- Gordon, A.; Hannon, G.J. “FASTX-Toolkit,” FASTQ/A Short-Reads pre-Processing Tools. 2012. Available online: http://hannonlab.cshl.edu/fastx_toolkit (accessed on 12 January 2019).
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. QIIME 2: Reproducible, interactive, scalable, and extensible microbiome data science. PeerJ 2018. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Holmes, S.P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 2017, 11, 2639–2643. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581. [Google Scholar] [CrossRef]
- Blaxter, M.; Mann, J.; Chapman, T.; Thomas, F.; Whitton, C.; Floyd, R.; Abebe, E. Defining operational taxonomic units using DNA barcode data. Philos. Trans. Royal Soc. B 2005, 1462, 1935–1943. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Desantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Anderson, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2011, 6, 610–618. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, J.I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57–59. [Google Scholar] [CrossRef] [PubMed]
- De Souza, R.S.C.; Okura, V.K.; Armanhi, J.S.L.; Jorrín, B.; Lozano, N.; Da Silva, M.J.; Imperial, J. Unlocking the bacterial and fungal communities assemblages of sugarcane microbiome. Sci. Rep. 2016, 6, 28774. [Google Scholar] [CrossRef] [PubMed]
- Pechal, J.L.; Schmidt, C.J.; Jordan, H.R.; Benbow, M.E. Frozen: Thawing and its effect on the postmortem microbiome in two pediatric cases. J. Forensic Sci. 2017, 62, 1399–1405. [Google Scholar] [CrossRef] [PubMed]
- Nearing, J.T.; Douglas, G.M.; Comeau, A.M.; Langille, M.G. Denoising the Denoisers: An independent evaluation of microbiome sequence error-correction approaches. PeerJ 2018, 6, e5364. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- De Cárcer, D.A.; Denman, S.E.; McSweeney, C.; Morrison, M. Evaluation of subsampling-based normalization strategies for tagged high-throughput sequencing datasets from gut microbiomes. Appl. Environ. Microbiol. 2011, 77, 8795–8798. [Google Scholar] [CrossRef]
- Pruesse, E.; Peplies, J.; Glöckner, F.O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 2012, 28, 1823–1829. [Google Scholar] [CrossRef]
- Cole, J.R.; Wang, Q.; Cardenas, E.; Fish, J.; Chai, B.; Farris, R.J.; Kulam-Syed-Mohideen, A.S.; McGarrell, D.M.; Marsh, T.; Garrity, G.M.; et al. The Ribosomal Database Project: Improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 2008, 37 (Suppl. 1), D141–D145. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Glöckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef] [PubMed]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef] [PubMed]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Hill, T.C.; Walsh, K.A.; Harris, J.A.; Moffett, B.F. Using ecological diversity measures with bacterial communities. FEMS Microbiol. Ecol. 2003, 43, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Marcon, E.; Scotti, I.; Hérault, B.; Rossi, V.; Lang, G. Generalization of the partitioning of Shannon diversity. PLoS ONE 2014, 9, e90289. [Google Scholar] [CrossRef] [PubMed]
- Simpson, E.H. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Kruskal, W.H.; Wallis, W.A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- Bray, J.R.; Curtis, J.T. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monograph. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Anderson, M.J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001, 26, 32–46. [Google Scholar] [CrossRef]
- Clarke, K.R. Non-parametric multivariate analyses of changes in community structure. Austral Ecol. 1993, 18, 117–143. [Google Scholar] [CrossRef]
- Oksanen, J.; Kindt, R.; Legendre, P.; O’Hara, B.; Stevens, M.H.H.; Oksanen, M.J.; Suggests, M. The vegan package. Community Ecol. Package 2007, 10, 631–637. [Google Scholar]
- Clarke, K.R.; Gorley, R.N. PRIMER v6: User Manual/Tutorial (Plymouth Routines in Multivariate Ecological Research); PRIMER-E Ltd.: Plymouth, UK, 2006. [Google Scholar]
- Lozupone, C.; Lladser, M.E.; Knights, D.; Stombaugh, J.; Knight, R. UniFrac: An effective distance metric for microbial community comparison. ISME J. 2011, 5, 169–172. [Google Scholar] [CrossRef] [PubMed]
- gplots: Various R Programming Tools for Plotting Data. Available online: cran.r-project.org/web/packages/gplots (accessed on 22 October 2018).
- RColorBrewer: ColorBrewer Palettes. Available online: cran.r-project.org/web/packages/RColorBrewer (accessed on 22 October 2018).
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.A. The use of multiple measurements in taxonomic problems. Ann. Hum. Genet. 1936, 7, 179–188. [Google Scholar] [CrossRef]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Thurber, R.L.V.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Choma, M.; Bárta, J.; Šantrůčková, H.; Urich, T. Low abundance of Archaeorhizomycetes among fungi in soil metatranscriptomes. Sci. Rep. 2016, 6, 38455. [Google Scholar] [CrossRef]
- Galton, F. Regression towards mediocrity in hereditary stature. J. R. Anthropol. Inst. 1886, 15, 246–263. [Google Scholar] [CrossRef]
- Pearson, K. Note on regression and inheritance in the case of two parents. Proc. R. Soc. Lond. 1895, 58, 240–242. [Google Scholar] [CrossRef]
- Spearman, C. The proof and measurement of association between two things. Am. J. Psychol. 1904, 15, 72–101. [Google Scholar] [CrossRef]
- Kullback, S.; Leibler, R.A. On information and sufficiency. Ann. Math. Stat. 1951, 22, 79–86. [Google Scholar] [CrossRef]
- Cover, T.M.; Thomas, J.A. Information Theory and Statistics. In Elements of Information Theory; John Wiley and Sons, Inc.: New York, NY, USA, 1991; Volume 1, pp. 279–335. ISBN 0-471-06259-6. [Google Scholar]
- Brown, M.B. 400: A method for combining non-independent, one-sided tests of significance. Biometrics 1975, 31, 987–992. [Google Scholar] [CrossRef]
- Wiese, R.; Eiglsperger, M.; Kaufmann, M. yFiles—Visualization and Automatic Layout of Graphs. In Graph Drawing Software (Mathematics and Visualization); Jünger, M., Mutzel, P., Eds.; Springer: Berlin/Heidelberg, Germany, 2004; pp. 173–191. ISBN 978-3-642-18638-7. [Google Scholar]
- Assenov, Y.; Ramírez, F.; Schelhorn, S.E.; Lengauer, T.; Albrecht, M. Computing topological parameters of biological networks. Bioinformatics 2007, 24, 282–284. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Wang, H.; Dsouza, M.; Lou, J.; He, Y.; Dai, Z.; Brookes, P.C.; Xu, J.; Gilbert, J.A. Geographic patterns of co-occurrence network topological features for soil microbiota at continental scale in eastern China. ISME J. 2016, 10, 1891. [Google Scholar] [CrossRef]
- Wang, Z.; Bafadhel, M.; Haldar, K.; Spivak, A.; Mayhew, D.; Miller, B.E.; Tal-Singer, R.; Johnston, S.L.; Ramsheh, M.Y.; Barer, M.R. Lung microbiome dynamics in chronic obstructive pulmonary disease exacerbations. Eur. Respir. J. 2016, 47, 1082–1092. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Sheng, H.F.; He, Y.; Wu, J.Y.; Jiang, Y.X.; Tam, N.F.Y.; Zhou, H.W. Comparison of the levels of bacterial diversity in freshwater, intertidal wetland, and marine sediments by using millions of Illumina tags. Appl. Environ. Microbiol. 2012, 78, 8264–8271. [Google Scholar] [CrossRef]
- Wirsen, C.O.; Sievert, S.M.; Cavanaugh, C.M.; Molyneaux, S.J.; Ahmad, A.; Taylor, L.; Delong, E.F.; Taylor, C.D. Characterization of an autotrophic sulfide-oxidizing marine Arcobacter sp. that produces filamentous sulfur. Appl. Environ. Microbiol. 2002, 68, 316–325. [Google Scholar] [CrossRef]
- Sievert, S.M.; Hügler, M.; Taylor, C.D.; Wirsen, C.O. Sulfur Oxidation at Deep-Sea Hydrothermal Vents. In Microbial Sulfur Metabolism; Dahl, C., Friedrich, C.G., Eds.; Springer: Berlin/Heidelberg, Germany, 2008; pp. 238–258. ISBN 978-3-540-72682-1. [Google Scholar]
- Inagaki, F.; Takai, K.; Kobayashi, H.; Nealson, K.H.; Horikoshi, K. Sulfurimonas autotrophica gen. nov.; sp. nov.; a novel sulfur-oxidizing ε-proteobacterium isolated from hydrothermal sediments in the Mid-Okinawa Trough. Int. J. Syst. Evol. Microbiol. 2003, 53, 1801–1805. [Google Scholar] [CrossRef]
- Campbell, B.J.; Engel, A.S.; Porter, M.L.; Takai, K. The versatile ε-proteobacteria: Key players in sulphidic habitats. Nat. Rev. Microbiol. 2006, 4, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Durand, L.; Zbinden, M.; Cueff-Gauchard, V.; Duperron, S.; Roussel, E.G.; Shillito, B.; Cambon-Bonavita, M.A. Microbial diversity associated with the hydrothermal shrimp Rimicaris exoculata gut and occurrence of a resident microbial community. FEMS Microbiol. Ecol. 2009, 71, 291–303. [Google Scholar] [CrossRef] [PubMed]
- Romero, J.; Garcia-Varela, M.; Laclette, J.; Espejo, R. Bacterial 16S rRNA gene analysis revealed that bacteria related to Arcobacter spp. constitute an abundant and common component of the oyster microbiota (Tiostrea chilensis). Microb. Ecol. 2002, 44, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.W.; Hwang, J.S.; Cheung, M.K.; Kwan, H.S.; Wong, C.K. Dietary analysis on the shallow-water hydrothermal vent crab Xenograpsus testudinatus using Illumina sequencing. Mar. Biol. 2015, 162, 1787–1798. [Google Scholar] [CrossRef]
- Urakawa, H.; Dubilier, N.; Fujiwara, Y.; Cunningham, D.E.; Kojima, S.; Stahl, D.A. Hydrothermal vent gastropods from the same family (Provannidae) harbour ε-and γ-proteobacterial endosymbionts. Environ. Microbiol. 2005, 7, 750–754. [Google Scholar] [CrossRef] [PubMed]
- Beinart, R.A.; Sanders, J.G.; Faure, B.; Sylva, S.P.; Lee, R.W.; Becker, E.L.; Gartman, A.; Luther, G.W.; Seewald, J.S.; Fisher, C.R. Evidence for the role of endosymbionts in regional-scale habitat partitioning by hydrothermal vent symbioses. Proc. Natl. Acad. Sci. USA 2012, 109, E3241–E3250. [Google Scholar] [CrossRef] [PubMed]
- Beinart, R.; Gartman, A.; Sanders, J.; Luther, G.; Girguis, P. The uptake and excretion of partially oxidized sulfur expands the repertoire of energy resources metabolized by hydrothermal vent symbioses. Proc. R. Soc. B 2015, 282, 20142811. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.S. Prokaryotes and the input of polyunsaturated fatty acids to the marine food web. FEMS Microbiol. Lett. 2003, 219, 1–7. [Google Scholar] [CrossRef]
- Kawasaki, K.; Nogi, Y.; Hishinuma, M.; Nodasaka, Y.; Matsuyama, H.; Yumoto, I. Psychromonas marina sp. nov.; a novel halophilic, facultatively psychrophilic bacterium isolated from the coast of the Okhotsk Sea. Int. J. Syst. Evol. Microbiol. 2002, 52, 1455–1459. [Google Scholar] [CrossRef]
- Groudieva, T.; Grote, R.; Antranikian, G. Psychromonas arctica sp. nov.; a novel psychrotolerant, biofilm-forming bacterium isolated from Spitzbergen. Int. J. Syst. Evol. Microbiol. 2003, 53, 539–545. [Google Scholar] [CrossRef]
- Breezee, J.; Cady, N.; Staley, J. Subfreezing growth of the sea ice bacterium “Psychromonas ingrahamii”. Microb. Ecol. 2004, 47, 300–304. [Google Scholar] [CrossRef] [PubMed]
- Hosoya, S.; Jang, J.-H.; Yasumoto-Hirose, M.; Matsuda, S.; Kasai, H. Psychromonas agarivorans sp. nov.; a novel agarolytic bacterium. Int. J. Syst. Evol. Microbiol. 2009, 59, 1262–1266. [Google Scholar] [CrossRef] [PubMed]
- Schink, B. The genus Propionigenium. In The Prokaryotes; Springer: New York, NY, USA, 2006; pp. 955–959. ISBN 978-0387-25494-4. [Google Scholar]
- Smith, P.; Willemsen, D.; Popkes, M.; Metge, F.; Gandiwa, E.; Reichard, M.; Valenzano, D.R. Regulation of life span by the gut microbiota in the short-lived African turquoise killifish. eLife 2017, 66, e27014. [Google Scholar] [CrossRef] [PubMed]
- Cardona, E.; Gueguen, Y.; Magré, K.; Lorgeoux, B.; Piquemal, D.; Pierrat, F.; Noguier, F.; Saulnier, D. Bacterial community characterization of water and intestine of the shrimp Litopenaeus stylirostris in a biofloc system. BMC Microbiol. 2016, 16, 157. [Google Scholar] [CrossRef] [PubMed]
- Thomas, F.; Barbeyron, T.; Tonon, T.; Génicot, S.; Czjzek, M.; Michel, G. Characterization of the first alginolytic operons in a marine bacterium: From their emergence in marine Flavobacteriia to their independent transfers to marine Proteobacteria and human gut Bacteroides. Environ. Microbiol. 2012, 14, 2379–2394. [Google Scholar] [CrossRef] [PubMed]
- Rabus, R.; Ruepp, A.; Frickey, T.; Rattei, T.; Fartmann, B.; Stark, M.; Bauer, M.; Zibat, A.; Lombardot, T.; Becker, I.; Amann, J.; et al. The genome of Desulfotalea psychrophila, a sulfate-reducing bacterium from permanently cold Arctic sediments. Environ. Microbiol. 2004, 6, 887–902. [Google Scholar] [CrossRef]
- Lockhart, C.; Fraga, D. Characterization of the arginine kinase from Desulfotalea psychrophila LSv54: The effects of environmental conditions and catalytic domain sequence variations on enzymatic turnover. FASEB J. 2007, 21, A299. [Google Scholar]
- Frøslev, T.G.; Kjøller, R.; Bruun, H.H.; Ejrnæs, R.; Brunbjerg, A.K.; Pietroni, C.; Hansen, A.J. Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates. Nat. Commun. 2017, 8, 1188. [Google Scholar] [CrossRef]
- Konstantinidis, K.T.; Tiedje, J.M. Genomic insights that advance the species definition for prokaryotes. Proc. Natl. Acad. Sci. USA 2005, 102, 2567–2572. [Google Scholar] [CrossRef]
- Stackebrandt, E.; Goebel, B.M. Taxonomic note: A place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Evol. Microbiol. 1994, 44, 846–849. [Google Scholar] [CrossRef]
- Tamar, E.; Koler, M.; Vaknin, A. The role of motility and chemotaxis in the bacterial colonization of protected surfaces. Sci. Rep. 2016, 6, 19616. [Google Scholar] [CrossRef] [PubMed]
- Thorsen, M.S. Microbial activity, oxygen status and fermentation in the gut of the irregular sea urchin Echinocardium cordatum (Spatangoida: Echinodermata). Mar. Biol. 1998, 132, 423–433. [Google Scholar] [CrossRef]
- Meziti, A.; Kormas, K.A.; Pancucci-Papadopoulou, M.A.; Thessalou-Legaki, M. Bacterial phylotypes associated with the digestive tract of the sea urchin Paracentrotus lividus and the ascidian Microcosmus sp. Russ. J. Mar. Biol. 2007, 33, 84–91. [Google Scholar] [CrossRef]
- Guerinot, M.L.; Fong, W.; Patriquin, D.G. Nitrogen fixation (acetylene reduction) associated with sea urchins (Strongylocentrotus droebachiensis) feeding on seaweeds and eelgrass. J. Fish. Res. Board Can. 1977, 34, 416–420. [Google Scholar] [CrossRef]
- Guerinot, M.L.; Patriquin, D.G. N2-fixing vibrios isolated from the gastrointestinal tract of sea urchins. Can. J. Microbiol. 1981, 27, 311–317. [Google Scholar] [CrossRef] [PubMed]
- Achá, D.; Hintelmann, H.; Yee, J. Importance of sulfate reducing bacteria in mercury methylation and demethylation in periphyton from Bolivian Amazon region. Chemosphere 2011, 82, 911–916. [Google Scholar] [CrossRef] [PubMed]
- Roalkvam, I.; Drønen, K.; Stokke, R.; Daae, F.L.; Dahle, H.; Steen, I.H. Physiological and genomic characterization of Arcobacter anaerophilus IR-1 reveals new metabolic features in Epsilonproteobacteria. Front. Microbiol. 2015, 6, 987. [Google Scholar] [CrossRef]
- Becker, P.T.; Samadi, S.; Zbinden, M.; Hoyoux, C.; Compère, P.; De Ridder, C. First insights into the gut microflora associated with an echinoid from wood falls environments. Cah. Biol. Mar. 2009, 50, 343–352. [Google Scholar]



| Sample | Raw Reads | Trimmed Reads | Unfilt. OTUs | Filt. OTUs | Cond. OTUs | Cond. OTUs Med. | Cond. OTUs Min. | Shannon Diversity | Simpson Diversity |
|---|---|---|---|---|---|---|---|---|---|
| Algae 1 | 154,561 | 121,543 | 4985 | 1345 | 268 | 259 | 243 | 5.3590 | 0.9376 |
| Algae 2 | 137,323 | 103,116 | 2973 | 855 | 226 | 220 | 204 | 4.7960 | 0.9340 |
| Algae 3 | 160,926 | 115,779 | 4745 | 1383 | 329 | 325 | 302 | 4.9626 | 0.9156 |
| Gut Digesta 1 | 76,329 | 60,871 | 2133 | 580 | 155 | 155 | 151 | 3.1936 | 0.8260 |
| Gut Digesta 2 | 74,249 | 61,267 | 2144 | 599 | 119 | 119 | 113 | 2.4760 | 0.5991 |
| Gut Digesta 3 | 123,640 | 99,546 | 3056 | 611 | 126 | 121 | 106 | 2.9289 | 0.7891 |
| Gut Tissue 1 | 128,539 | 90,412 | 4384 | 1094 | 328 | 322 | 301 | 4.1236 | 0.8440 |
| Gut Tissue 2 | 68,644 | 51,895 | 2311 | 898 | 273 | 273 | 273 | 3.8787 | 0.8436 |
| Gut Tissue 3 | 123,735 | 89,547 | 4765 | 1418 | 403 | 397 | 368 | 4.8038 | 0.9250 |
| Pharynx Tissue 1 | 107,276 | 72,954 | 5663 | 1558 | 430 | 430 | 418 | 6.0719 | 0.9642 |
| Pharynx Tissue 2 | 86,515 | 58,281 | 4417 | 1488 | 431 | 431 | 430 | 5.9389 | 0.9632 |
| Pharynx Tissue 3 | 92,987 | 63,405 | 4918 | 1489 | 402 | 402 | 397 | 6.2246 | 0.9697 |
| Water 1 | 123,154 | 81,885 | 4725 | 1679 | 504 | 504 | 487 | 5.2838 | 0.8797 |
| Water 2 | 137,044 | 93,713 | 6127 | 1087 | 403 | 400 | 386 | 5.7289 | 0.9546 |
| Water 3 | 119,824 | 85,613 | 5608 | 1406 | 400 | 400 | 377 | 4.2286 | 0.8211 |
| Summary | total = 1,714,746 | total = 1,249,827 | total = 44,664 | total = 4290 | total = 776 | total = 776 | total = 776 | avg. = 4.6666 | avg. = 0.8778 |
| Diversity Measure | Unfilt. OTUs | Filt. OTUs | Cond. OTUs | Cond. OTUs Med. | Cond. OTUs Min |
|---|---|---|---|---|---|
| ANOSIM | R = 0.93185 | R = 0.93185 | R = 0.94074 | R = 0.94074 | R = 0.94222 |
| Adonis | R2 = 0.69518 | R2 = 0.71145 | R2 = 0.74894 | R2 = 0.74951 | R2 = 0.75688 |
| Sample Type | Phylum | Taxon | Av. Ab. | Clos. Cent. | Bet. Cent. | Deg. | Pos. Deg. | Neg. Deg. |
|---|---|---|---|---|---|---|---|---|
| Gut Digesta | Fusobacteria | Propionigenium | 14.89% | 0.39 | 0.13 | 17 | 14 | 4 |
| Gut Digesta | Proteobacteria (Gammaproteobacteria) | Moritella | 0.15% | 0.44 | 0.17 | 16 | 10 | 6 |
| Gut Digesta | Bacteroidetes | SB-1 | 0.38% | 0.39 | 0.06 | 14 | 3 | 11 |
| Gut Digesta | Proteobacteria (Deltaproteobacteria) | Desulfobacteraceae | 0.24% | 0.38 | 0.04 | 14 | 2 | 12 |
| Gut Digesta | Proteobacteria (Deltaproteobacteria) | Desulfovibrio | 0.24% | 0.41 | 0.11 | 13 | 6 | 7 |
| Gut Tissue | Proteobacteria (Alphaproteobacteria) | Rhodobacteraceae | 0.34% | 0.41 | 0.05 | 9 | 1 | 8 |
| Gut Tissue | Cyanobacteria | Rhodophyta | 0.22% | 0.41 | 0.05 | 9 | 1 | 8 |
| Gut Tissue | Proteobacteria (Gammaproteobacteria) | Vibrionaceae | 0.36% | 0.41 | 0.09 | 9 | 5 | 5 |
| Gut Tissue | Proteobacteria (Epsilonproteobacteria) | Arcobacter | 20.59% | 0.38 | 0.04 | 9 | 7 | 2 |
| Gut Tissue | Firmicutes | Bacilli | 0.16% | 0.38 | 0.06 | 8 | 6 | 2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hakim, J.A.; Schram, J.B.; Galloway, A.W.E.; Morrow, C.D.; Crowley, M.R.; Watts, S.A.; Bej, A.K. The Purple Sea Urchin Strongylocentrotus purpuratus Demonstrates a Compartmentalization of Gut Bacterial Microbiota, Predictive Functional Attributes, and Taxonomic Co-Occurrence. Microorganisms 2019, 7, 35. https://doi.org/10.3390/microorganisms7020035
Hakim JA, Schram JB, Galloway AWE, Morrow CD, Crowley MR, Watts SA, Bej AK. The Purple Sea Urchin Strongylocentrotus purpuratus Demonstrates a Compartmentalization of Gut Bacterial Microbiota, Predictive Functional Attributes, and Taxonomic Co-Occurrence. Microorganisms. 2019; 7(2):35. https://doi.org/10.3390/microorganisms7020035
Chicago/Turabian StyleHakim, Joseph A., Julie B. Schram, Aaron W. E. Galloway, Casey D. Morrow, Michael R. Crowley, Stephen A. Watts, and Asim K. Bej. 2019. "The Purple Sea Urchin Strongylocentrotus purpuratus Demonstrates a Compartmentalization of Gut Bacterial Microbiota, Predictive Functional Attributes, and Taxonomic Co-Occurrence" Microorganisms 7, no. 2: 35. https://doi.org/10.3390/microorganisms7020035
APA StyleHakim, J. A., Schram, J. B., Galloway, A. W. E., Morrow, C. D., Crowley, M. R., Watts, S. A., & Bej, A. K. (2019). The Purple Sea Urchin Strongylocentrotus purpuratus Demonstrates a Compartmentalization of Gut Bacterial Microbiota, Predictive Functional Attributes, and Taxonomic Co-Occurrence. Microorganisms, 7(2), 35. https://doi.org/10.3390/microorganisms7020035





