Cryptosporidium Prevalence in Calves and Geese Co-Grazing on Four Livestock Farms Surrounding Two Reservoirs Supplying Public Water to Mainland Orkney, Scotland
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Sample Processing and Analysis
2.2.1. Processing Faecal Samples
- Calf faecal samples: 250 μg of sample was added to 200 µL lysis buffer (T1 buffer, Macherey-Nagel, Duren, Germany. NZ740952250).
- Goose faecal samples: Salt flotation, using approximately 3 g of faecal sample, was performed [2], following which the final pellet was re-suspended in 200 µL lysis buffer. The extra salt flotation step was performed on goose samples due to the higher fibre content of these samples, which requires a further processing step prior to DNA extraction.
2.2.2. DNA Extraction
2.2.3. PCR Sequencing and Analysis
2.2.4. Subtyping C. parvum-Positive Samples
2.2.5. Processing and Analysis of Water Samples
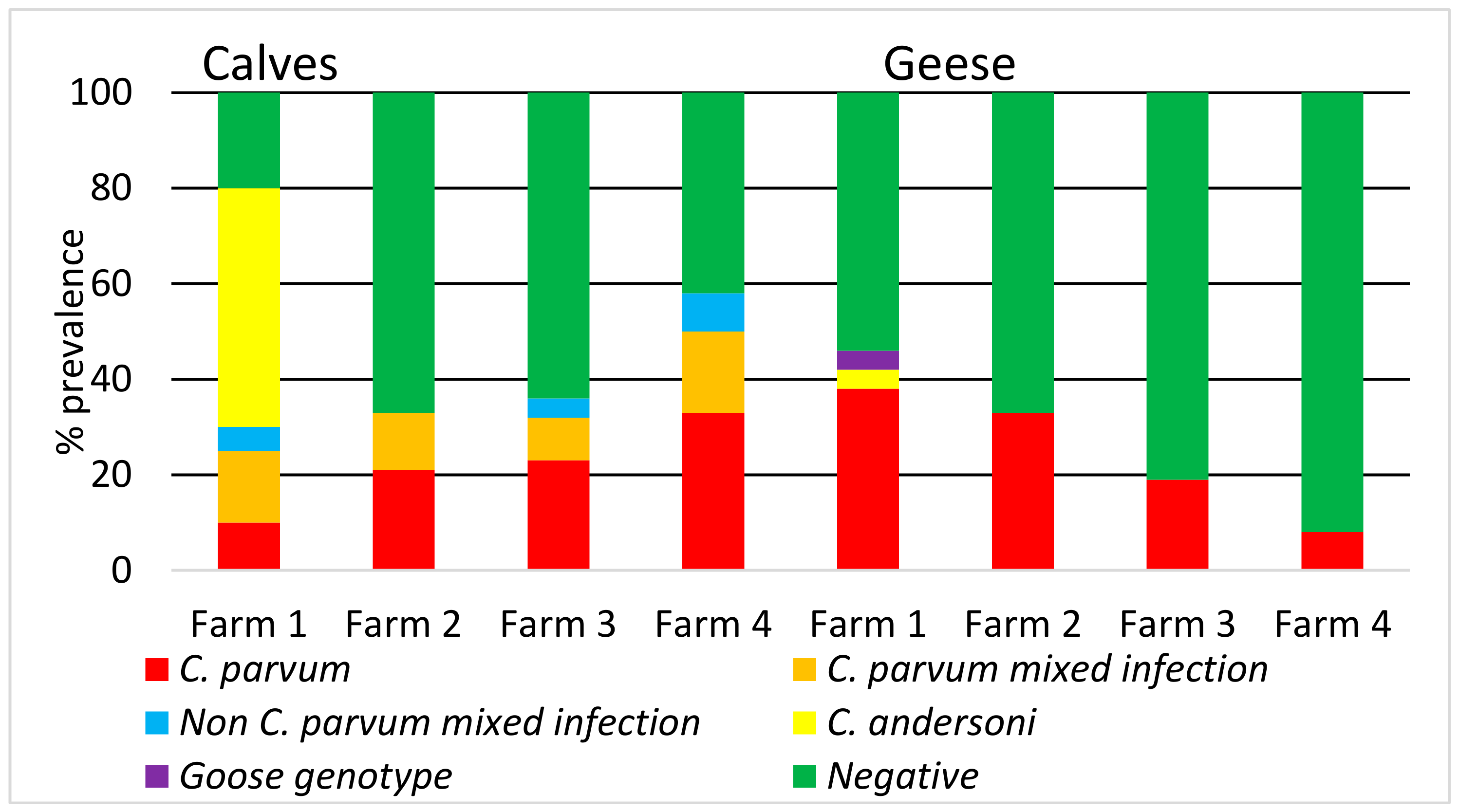
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Santin, M. Clinical and subclinical infections with Cryptosporidium in animals. N Z Vet. J. 2013, 61, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Chalmers, R.M.; Elwin, K.; Thomas, A.L.; Guy, E.C.; Mason, B. Long-term Cryptosporidium typing reveals the aetiology and species-specific epidemiology of human cryptosporidiosis in England and Wales, 2000 to 2003. Euro. Surveillance: Bulletin Europeen Sur Les Maladies Transmissibles = European Communicable Disease Bulletin 2009, 14. [Google Scholar] [CrossRef] [PubMed]
- Ryan, U.; Fayer, R.; Xiao, L. Cryptosporidium species in humans and animals: Current understanding and research needs. Parasitology 2014, 141, 1667–1685. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.P.; Clifton-Hadley, F.A.; Cheney, T.; Giles, M. Prevalence and molecular typing of Cryptosporidium in dairy cattle in England and Wales and examination of potential on-farm transmission routes. Vet. Parasitol. 2014, 204, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Atwill, E.; Johnson, E.; Klingborg, D.; Veserat, G.; Markegard, G.; Jensen, W.; Pratt, D.; Delmas, R.; George, H.; Forero, L.; et al. Age, geographic and temporal distribution of fecal shedding of Cryptosporidium parvum oocysts in cow-calf herds. Am. J. Vet. Res. 1999, 60, 420–425. [Google Scholar] [PubMed]
- De Waele, V.; Berzano, M.; Speybroeck, N.; Berkvens, D.; Mulcahy, G.M.; Murphy, T.M. Peri-parturient rise of Cryptosporidium oocysts in cows: New insights provided by duplex quantitative real-time PCR. Vet. Parasitol. 2012, 189, 366–368. [Google Scholar] [CrossRef] [PubMed]
- Sturdee, A.P.; Bodley-Tickell, A.T.; Archer, A.; Chalmers, R.M. Long-term study of Cryptosporidium prevalence on a lowland farm in the United Kingdom. Vet. Parasitol. 2003, 116, 97–113. [Google Scholar] [CrossRef]
- Zahedi, A.; Paparini, A.; Jian, F.; Robertson, I.; Ryan, U. Public health significance of zoonotic Cryptosporidium species in wildlife: Critical insights into better drinking water management. Int. J. Parasitol. Parasites. Wildl. 2016, 5, 88–109. [Google Scholar] [CrossRef]
- Zahedi, A.; Monis, P.; Gofton, A.W.; Oskam, C.L.; Ball, A.; Bath, A.; Bartkow, M.; Robertson, I.; Ryan, U. Cryptosporidium species and subtypes in animals inhabiting drinking water catchments in three states across Australia. Water Res. 2018, 134, 327–340. [Google Scholar] [CrossRef]
- Wells, B.; Shaw, H.; Hotchkiss, E.; Gilray, J.; Ayton, R.; Green, J.; Katzer, F.; Wells, A.; Innes, E. Prevalence, species identification and genotyping Cryptosporidium from livestock and deer in a catchment in the Cairngorms with a history of a contaminated public water supply. Parasit. Vectors. 2015, 8, 66. [Google Scholar] [CrossRef]
- Thomson, S.; Hamilton, C.A.; Hope, J.C.; Katzer, F.; Mabbott, N.A.; Morrison, L.J.; Innes, E.A. Bovine cryptosporidiosis: Impact, host-parasite interaction and control strategies. Vet. Res. 2017, 48, 42. [Google Scholar] [CrossRef] [PubMed]
- Meinhardt, P.L.; Casemore, D.P.; Miller, K.B. Epidemiologic aspects of human cryptosporidiosis and the role of waterborne transmission. Epidemiol. Rev. 1996, 18, 118–136. [Google Scholar] [CrossRef] [PubMed]
- Robinson, G.; Chalmers, R.M.; Stapleton, C.; Palmer, S.R.; Watkins, J.; Francis, C.; Kay, D. A whole water catchment approach to investigating the origin and distribution of Cryptosporidium species. J. Appl. Microbiol. 2011, 111, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Government, S. The Cryptosporidium (Scottish Water) Directions; Scottish Government: Edinburgh, UK, 2003.
- Jellison, K.L.; Lynch, A.E.; Ziemann, J.M. Source tracking identifies deer and geese as vectors of human-infectious Cryptosporidium genotypes in an urban/suburban watershed. Environ. Sci. Technol. 2009, 43, 4267–4272. [Google Scholar] [CrossRef] [PubMed]
- Kassa, H.; Harrington, B.J.; Bisesi, M.S. Cryptosporidiosis: A brief literature review and update regarding Cryptosporidium in feces of Canada geese (Branta canadensis). J. Environ. Health 2004, 66, 34–40. [Google Scholar]
- Gorham, T.J.; Lee, J. Pathogen loading from canada geese faeces in freshwater: potential risks to human health through recreational water exposure. Zoonoses Public Health 2016, 63, 177–190. [Google Scholar] [CrossRef]
- Elmberg, J.; Berg, C.; Lerner, H.; Waldenstrom, J.; Hessel, R. Potential disease transmission from wild geese and swans to livestock, poultry and humans: A review of the scientific literature from a One Health perspective. Infect Ecol. Epidemiol. 2017, 7, 1300450. [Google Scholar] [CrossRef]
- Thomson, S.; Jonsson, N.; Innes, E.A.; Katzer, F. A multiplex PCR test to identify four common cattle adapted Cryptosporidium species. Parasitol. Open 2016. In Press. [Google Scholar] [CrossRef]
- Hall, T. BioEdit: A user-friendly biological sequence alignment editor and analysis programme for Windows 95/98/NT. Nucl. Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Brook, E.J.; Anthony Hart, C.; French, N.P.; Christley, R.M. Molecular epidemiology of Cryptosporidium subtypes in cattle in England. Vet. J. 2009, 179, 378–382. [Google Scholar] [CrossRef]
- Sulaiman, I.M.; Hira, P.R.; Zhou, L.; Al-Ali, F.M.; Al-Shelahi, F.A.; Shweiki, H.M.; Iqbal, J.; Khalid, N.; Xiao, L. Unique endemicity of cryptosporidiosis in children in Kuwait. J. Clin. Microbiol. 2005, 43, 2805–2809. [Google Scholar] [CrossRef] [PubMed]
- Environment Agency. The Microbiology of Drinking Water (2010)—Part 14—Methods for the Isolation, Identification and Enumeration of Cryptosporidium Oocysts and Giardia Cysts; Environment Agency: Bristol, UK, 2010. [Google Scholar]
- Mosier, D.A.; Oberst, R.D. Cryptosporidiosis. A global challenge. Ann. N Y Acad. Sci. 2000, 916, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Blanchard, P.C. Diagnostics of Dairy and Beef Cattle Diarrhea. Vet. Clin. N. Am. Food A 2012, 28, 443–464. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.I.; Yoon, K.J. An overview of calf diarrhea – infectious etiology, diagnosis, and intervention. J. Vet. Sci. 2014, 15, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Plutzer, J.; Tomor, B. The role of aquatic birds in the environmental dissemination of human pathogenic Giardia duodenalis cysts and Cryptosporidium oocysts in Hungary. Parasitol. Int. 2009, 58, 227–231. [Google Scholar] [CrossRef]
- Grimason, A.M.; Smith, H.V.; Parker, J.F.; Jackson, M.H.; Smith, P.G.; Girdwood, R.W. Occurrence of Giardia sp. cysts and Cryptosporidium sp. oocysts in faeces from public parks in the west of Scotland. Epidemiol. Infect 1993, 110, 641–645. [Google Scholar]
- Smith, H.V.; Brown, J.; Coulson, J.C.; Morris, G.P.; Girdwood, R.W. Occurrence of oocysts of Cryptosporidium sp. in Larus spp. gulls. Epidemiol. Infect. 1993, 110, 135–143. [Google Scholar] [CrossRef]
- Xiao, L.; Feng, Y. Zoonotic cryptosporidiosis. FEMS Immunol. Med. Microbiol. 2008, 52, 309–323. [Google Scholar] [CrossRef]
- Feng, Y.; Alderisio, K.A.; Yang, W.; Blancero, L.A.; Kuhne, W.G.; Nadareski, C.A.; Reid, M.; Xiao, L. Cryptosporidium genotypes in wildlife from a new york watershed. Appl. Environ. Microbiol. 2007, 73, 6475–6483. [Google Scholar] [CrossRef]
- Zhou, L.; Kassa, H.; Tischler, M.L.; Xiao, L. Host-adapted Cryptosporidium spp. in Canada geese (Branta canadensis). Appl. Environ. Microbiol. 2004, 70, 4211–4215. [Google Scholar] [CrossRef]
- Graczyk, T.K.; Cranfield, M.R.; Fayer, R.; Trout, J.; Goodale, H.J. Infectivity of Cryptosporidium parvum oocysts is retained upon intestinal passage through a migratory water-fowl species (Canada goose, Branta canadensis). Trop. Med. Int. Health 1997, 2, 341–347. [Google Scholar] [CrossRef] [PubMed]
- McDonald, S.; Berzano, M.; Ziegler, P.; Murphy, T.M.; Holden, N.M. Qualitative risk assessment of surface water contamination with Cryptosporidium Sp Oocysts: A case study of three agricultural catchments. Hum Ecol. Risk Assess. 2011, 17, 813–825. [Google Scholar] [CrossRef]


| Location | Area (ha) | Number—August Count | Mean Number over 5 Years | ||||
|---|---|---|---|---|---|---|---|
| 2012 | 2013 | 2014 | 2015 | 2016 | |||
| North Ronaldsay | 690 | 389 | 132 | 355 | 546 | 401 | 365 |
| Sanday | 5043 | 2591 | 1780 | 2083 | 2613 | 2578 | 2329 |
| Westray | 4713 | 840 | 1223 | 983 | 962 | 1082 | 967 |
| Papa Westray | 933 | 343 | 157 | 501 | 61 | 263 | |
| Eday | 2745 | 1138 | 1221 | 708 | 566 | 814 | 889 |
| Small Holms (Faray, Muckle Green Holm) | 265 | 92 | NC | NC | NC | NC | N/A |
| Stronsay | 3430 | 951 | 1895 | 1978 | 1732 | 3477 | 2007 |
| Shapinsay | 2948 | 1765 | 1423 | 1282 | 1563 | 1915 | 1590 |
| Rousay/Eynhallow | 4935 | 399 | 113 | 576 | 447 | 568 | 419 |
| Egilsay | 650 | 0 | 36 | 146 | 176 | 20 | 76 |
| Wyre | 311 | 0 | 0 | 0 | 27 | 10 | 7 |
| Gairsay | 240 | 55 | 80 | 160 | 47 | 20 | 72 |
| Auskerry | 85 | 20 | 30 | NC | 25 | 12 | 22 |
| East Mainland | 52,325 | 2216 | 2233 | 1862 | 1952 | 2931 | 2239 |
| West Mainland | 8409 | 7660 | 9759 | 7012 | 7835 | 8135 | |
| Copinsay | 73 | 0 | 0 | NC | 0 | 0 | N/A |
| Burray | 1098 | 731 | 750 | 571 | 466 | 352 | 574 |
| South Ronaldsay | 4980 | 1234 | 1370 | 2233 | 2113 | 2021 | 1794 |
| Hoy & South Walls | 14,558 | 107 | 271 | 0 | 495 | 0 | 175 |
| Flotta / Fara / Switha | 1212 | 87 | 25 | 58 | 6 | 142 | 64 |
| Graemsay | 409 | 0 | 0 | 0 | 105 | 11 | 23 |
| Swona | 92 | NC | 0 | NC | NC | NC | N/A |
| TOTAL | 101,735 | 21,367 | 20,242 | 22,911 | 21,354 | 24,250 | Mean 22,025 |
| Year | Total Orkney |
|---|---|
| 2018 | 63,534 |
| 2017 | 63,045 |
| 2016 | 46,678 |
| 2015 | 56,151 |
| 2014 | 65,067 |
| 2013 | 63,665 |
| 2012 | 74,913 |
| MEAN | 61,865 |
| Farm | Numbers of Calf Samples | Numbers of Goose Samples |
|---|---|---|
| 1 | 20 | 24 |
| 2 | 24 | 24 |
| 3 | 22 | 26 |
| 4 | 12 | 26 |
| Total | 78 | 100 |
| Cryptosporidium Species | Identified Cryptosporidium Species Calves Stool Samples N (%) | Geese Stool Samples N (%) | *** Water Samples N (%) |
|---|---|---|---|
| C. parvum | 19 (15/78) | 24 (24/100) | 33 (15/46) |
| * C. parvum mixed infection | 13 (10/78) | 0 | 0 |
| ** Non C. parvum mixed infection | 6 (5/78) | 0 | 0 |
| C. andersoni | 9 (7/78) | 1 (1/100) | 39 18/46) |
| C. bovis | 1 (1/78) | 0 | 0 |
| C. ubiquitum | 0 | 0 | 2 (1/46) |
| Goose genotype | 0 | 1 (1/100) | 0 |
| Total No. | 78 | 100 | 46 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wells, B.; Paton, C.; Bacchetti, R.; Shaw, H.; Stewart, W.; Plowman, J.; Katzer, F.; Innes, E.A. Cryptosporidium Prevalence in Calves and Geese Co-Grazing on Four Livestock Farms Surrounding Two Reservoirs Supplying Public Water to Mainland Orkney, Scotland. Microorganisms 2019, 7, 513. https://doi.org/10.3390/microorganisms7110513
Wells B, Paton C, Bacchetti R, Shaw H, Stewart W, Plowman J, Katzer F, Innes EA. Cryptosporidium Prevalence in Calves and Geese Co-Grazing on Four Livestock Farms Surrounding Two Reservoirs Supplying Public Water to Mainland Orkney, Scotland. Microorganisms. 2019; 7(11):513. https://doi.org/10.3390/microorganisms7110513
Chicago/Turabian StyleWells, Beth, Claire Paton, Ross Bacchetti, Hannah Shaw, William Stewart, James Plowman, Frank Katzer, and Elisabeth A Innes. 2019. "Cryptosporidium Prevalence in Calves and Geese Co-Grazing on Four Livestock Farms Surrounding Two Reservoirs Supplying Public Water to Mainland Orkney, Scotland" Microorganisms 7, no. 11: 513. https://doi.org/10.3390/microorganisms7110513
APA StyleWells, B., Paton, C., Bacchetti, R., Shaw, H., Stewart, W., Plowman, J., Katzer, F., & Innes, E. A. (2019). Cryptosporidium Prevalence in Calves and Geese Co-Grazing on Four Livestock Farms Surrounding Two Reservoirs Supplying Public Water to Mainland Orkney, Scotland. Microorganisms, 7(11), 513. https://doi.org/10.3390/microorganisms7110513






