1. Introduction
Despite Australia’s vast landmass, an estimated 85% of the total population live within the coastal zone [
1]. Ongoing rapid urban expansion, strong industrialization, and modern agriculture have caused intense degradation of coastal riparian vegetation [
2], which in turn has led to the progressive deterioration of natural ecosystems [
3], and riverine and estuarine water quality [
4]. Furthermore, increased human activities have significantly changed the nutrient flux into these aquatic ecosystems [
5]. Excessive levels of inorganic nutrients such as nitrogen and phosphorus accelerate the eutrophication of waterways and estuaries [
6]. Eutrophication is one of the biggest threats to coasts and estuaries globally [
5], causing severe degradation of water quality, loss of biodiversity and richness, ecological shifts, and ecosystem malfunction [
7]. Such changes are also likely to adversely affect coastal communities in terms of employment and tourism opportunities and recreational activities. Compounding the issue further, in areas of extreme rainfall, unpredictable flood events drastically alter the quality of run-off and, therefore, delivery of catchment-derived sediments and nutrients [
8]. A substantial number of other pollutants such as trace metals, pesticides, and hydrocarbons have also been introduced into coastal waters as a result of human activities, increased urbanization, and as a consequence of greater impervious surfaces constructed within catchments [
9].
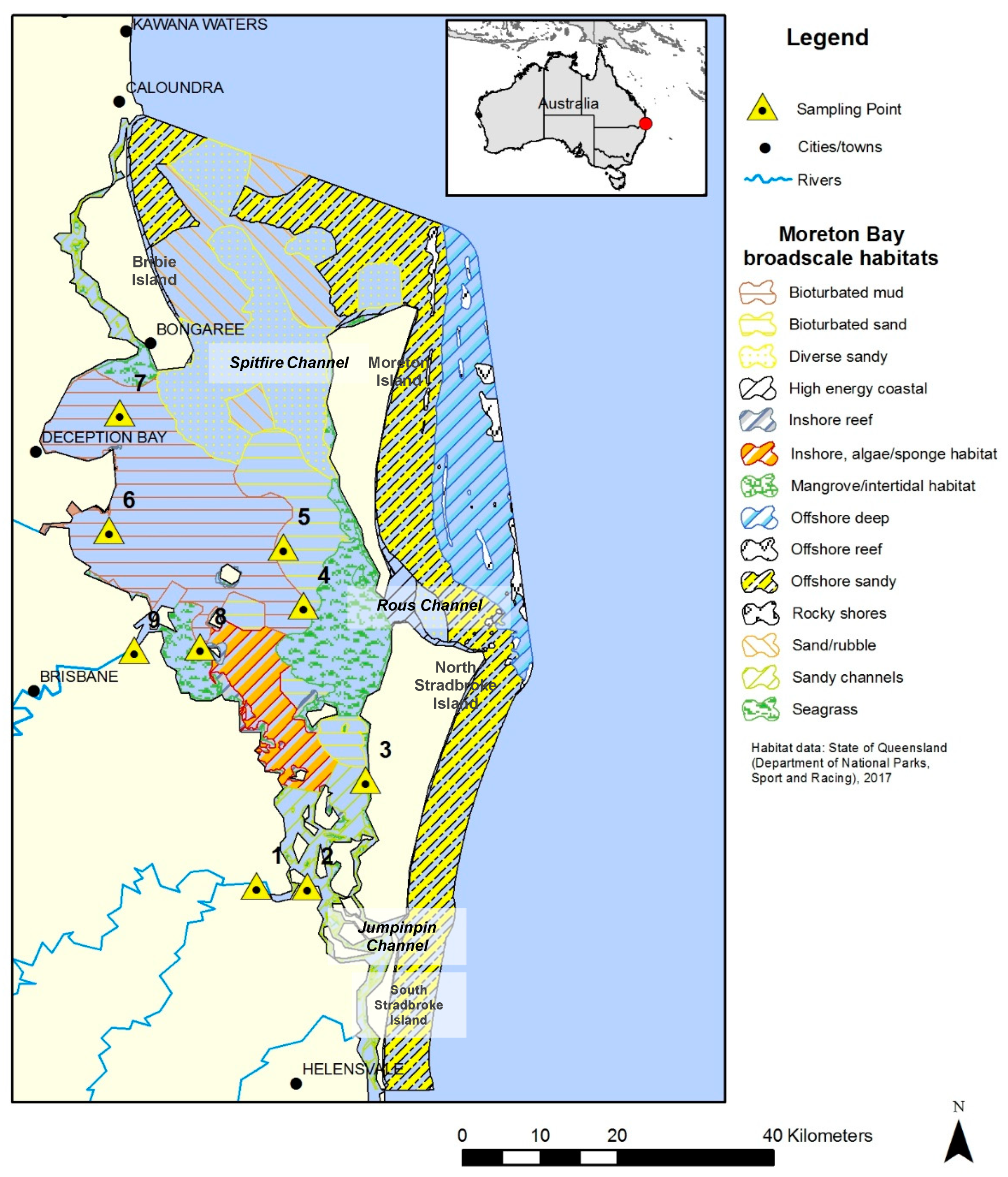
One catchment/ecosystem of interest is Moreton Bay in South East Queensland, Australia (see
Figure 1). Moreton Bay is a major sub-tropical shallow embayment. The bay is large (~1500 km
2) and 125 km long, with a broad (35 km) opening at its widest west–east axis. This lagoon-like bay is surrounded by four large continental sand islands—Bribie, Moreton, North Stradbroke, and South Stradbroke [
10]. The Bay has three openings to the Pacific Ocean. The northern passage between the Bribie and Moreton Islands is Spitfire Channel, which is the largest opening with the greatest oceanic exchange. Rous Channel, a smaller southern passage between the Moreton and North Stradbroke Islands, has a more restricted flow. Lastly, Jumpinpin Channel in the southern Bay between North and South Stradbroke Islands has the lowest degree of flushing [
11]. The greater Moreton Bay catchment comprises 21,200 km
2 of land, with four main rivers that drain into Moreton Bay: the Caboolture, Pine, Brisbane, Logan-Albert, and Pimpama Rivers. The river outflow includes effluent from riparian agriculture, treated sewage, and residential and industrial discharges, in addition to urban litter and pollution [
10].
The Bay provides aquaculture, industrial, residential, and shipping services, making it vulnerable to environmental stresses. Being subtropical, Moreton Bay supports a diverse community of marine megafauna such as fish, migratory whales, corals, and subtropical species including sea turtles, dugongs, and Australian humpback dolphins [
10]. The Bay also provides a variety of habitats for wildlife including extensive wetlands, shallow sandbanks, mudflats, mangrove forests, coral reefs, seagrass meadows, and myriad creeks, brooks, and rivulets [
10]. Moreton Bay has been recently declared a marine park with recognition as a heritage site [
12]. Historically, the Bay has been exploited for coral mining, sand mining, intense trawling, recreational fishing, and ecotourism [
8]. In addition to increased urbanization in the adjacent catchment, such human activities adversely affect the ecology of this embayment. This subtropical region is predisposed to seasonal and mildly monsoonal rainfall during the wet season when freshwater runoff into the bay increases markedly, with occasional coastal flooding events [
10]. Sediments, nutrients, and pollutants are mobilized and rapidly delivered to coastal waters [
13]. The eastern and northern regions of the Bay are relatively well-flushed by oceanic exchange, whereas residence times in the middle, southern, and western Bay are generally longer (>40 days) [
11]. As a result, riverine input has a greater influence on the southern and western Bay, where sediments are characterized by higher silt, organic carbon, and nutrient content [
14]). Nutrients and chlorophyll in the western Bay have been reported to be 10-fold to 100-fold higher than in the eastern Bay [
15]. Despite these large gradients, planktonic production remains the primary carbon input throughout the Bay, supplemented primarily by seagrass and mangrove production depending on local habitat type [
16]. The Bay is thought to be nitrogen-limited, with nitrogen fixation accounting for 60% of the nitrogen input. Point source inputs, e.g., from wastewater treatment facilities and the Port of Brisbane, account for 85% of the phosphorus input and 25% of the nitrogen input [
17]. It should be noted, however, that point source inputs have significantly reduced over the last 25 years with the move to tertiary wastewater treatment; as a result, agricultural runoff is the key concern. Industrial activities at the Port and shipping channel in the western Bay have also been linked to habitat degradation from toxic biocides derived from copper-based antifouling paints [
18].
In recent times, omics-based techniques and their integration have been used to advance our understanding of microbial physiology. Kimes et al. [
19] applied bacterial 16S rRNA gene sequencing and metabolite profiling techniques to characterize the microbial communities exposed to anthropogenic hydrocarbons and how their community composition and metabolic function impacted the bioremediation of oil spills. In a previous study, we used a similar approach to investigate surface water quality and characterize the bacterial population with respect to water contaminants in the Brisbane river [
20]. A multi-omics-based ecological analysis of coastal marine sediments was also used to assess the impact of anthropogenic factors arising from point and non-point pollution sources at a multi-commodity marine port [
21] and assess the impact of seasonal rainfall and urban pollution on marine sediment ecosystems [
8]. In this study, we extend the application of bacterial 16S rRNA gene sequencing and metabolomics to assess the pressures of urban expansion, industrialization, and other factors associated with increased human activities on the Moreton Bay ecosystem. Specific stressors on the sediment microbial communities that were studied at various sediment depths include increased organic carbon inputs, nutrients and pollutants. This was done to present a baseline set of observations for the Moreton Bay ecosystem during the dry season and characterize the sediment community composition and function at different depths. Utilizing the principles of untargeted metabolomics, this also has the potential to generate new hypotheses for follow-up research activities within the studied catchment relating to ecosystem function [
22]. Such an approach will provide insight into the influence of baseline residual contaminants within the sampled sediments, where the impact and influence of rainfall events and spike contamination events are removed. Together, this will enable a better understanding of the underlying envelope of conditions that define constructs of ecosystem health and resilience within Moreton Bay.
4. Discussion
The Australian and New Zealand Guidelines for Fresh and Marine Water Quality provide a framework to classify sampling sites based on their ecosystem condition [
26]. In this study, based on the Guidelines, Moreton Bay is considered a moderately disturbed ecosystem of high ecological value. As such, the physical and chemical characteristics are assessed against the sub-regional guidelines for Moreton Bay [
40]. Sites 1–3 and 6–9 are considered enclosed coastal waters or lower estuary sites, while sites 4 and 5 are considered open coastal estuary waters. Sites 1–5 and 8 are also noted to be of high ecological value marine/estuarine waters.
As previously stated, Moreton Bay receives water from a coastal and hinterland area of ~22,000 km
2. This includes rainwater runoff, effluent from riparian agriculture, treated sewage, residential and industrial outflow, and urban litter. Such modifications have led to deterioration of the bay water and sediment quality in terms of increased nutrient, heavy metals and contaminant fluxes. These sediments exposed to urban pollutants may have long-term human and environmental implications. Nutrients, such as nitrogen and phosphorus, are essential for plant and animal growth and nourishment. Over-abundance of these nutrients in water can result in over-stimulation of growth of nuisance aquatic plants and algae. The total nitrogen concentration was above the trigger value of 150 μg N L
-1 for all samples. The total organic content (TOC), in addition to sediment grain size, is known to influence the adsorption capacity of sediments and also the binding of organic pollutants to the sediments [
41]. The combined high percentage (58–100%) of coarse and fine sand found in the sediments with a high TOC concentration of (2.07–6.24 mg TOC g
-1 dry weight) may increase the adsorption capacity (larger surface area) of the sediments and result in considerably higher binding of organic pollutants (hydrophobicity of TOC). No trends, however, were observed between samples. The TOC data were used to normalize the organic pollutant data as per standard practice. Heavy metal pollution of marine sediments is typically associated with human activities such as mining or discharges from manufacturing industries [
41]. High concentrations of metals in sediments can potentially have toxic effects on resident aquatic ecosystem and can make fish, crustaceans, and bivalves unsuitable for human consumption [
42]. Coates-Marnane et al. [
12] reported trace metal (lead, zinc and copper) deposits in sediment core samples collected in the Moreton Bay region. Heavy metals, albeit below trigger values, were found in our previous studies at all sites sampled in this region [
8]. All organic pollutants except pindolol were found in most of the samples from various sampling sites. A few samples sites reported organic pollutants above the threshold for chemical stressors at the 25% probability of biological effects (80–100 μg g
-1 OC dry weight) [
43]. Organic pollutants reported above the threshold were penicillin V and DL-atenolol for site 1; penicillin V, diphenhydramine, promethazine, carbamazepine, N,N-diethyl-meta-toluamide, and DL-atenolol for site 3; promethazine and DL-atenolol for site 4; promethazine, N,N-diethyl-meta-toluamide, DL-atenolol for sites 5 and 6; and penicillin V, promethazine, and carbamazepine for site 9. No organic pollutants were reported above the threshold for sites 2, 7, and 8.
Large populations of sulfate and sulfur metabolizing bacterial communities (
Deltaproteobacteria, 30%;
Gammaproteobacteria, 12%) and organohalide respiring bacteria (
Dehalococcoidetes, 8.7%) were detected in collected sediment samples. Members of the families
Desulfobacteraceae (11.4%) and
Syntrophobacteraceae (11%) were the most important taxonomic groups in terms of abundance and activity (in terms of metabolism). These results align well with the previous findings [
44]. Recent estimates suggest that 29% of organic matter deposited to the seafloor is remineralized by sulfate-reducing microorganisms (SRMs) such as
Deltaproteobacteria [
45]. SRMs cooperatively degrade organic matter and, as such, are conventionally regarded as the terminal components of anaerobic food webs [
46]. Overall, the SRMs were observed to work in commensal relationship with methanotrophs and methylotrophs, especially towards the lower end of bioturbated muds and overlapping zones of these muds and sandy channels. This was especially observed in the degradation pathways of aromatics and polychlorinated pollutants washed off from the riverine system. While the methylotrophs were active in the river sediments, especially at 20-cm depth, SRMs were more active in the bioturbated mud and sandy channels. The methylotrophs were observed to be involved in the degradation of PCBs and pyruvate metabolism; the products of these metabolic pathways were further processed in bioturbated muds (further elaborated below).
Methanotrophs such as
Methylococcaceae formed a considerable part of bioturbated mud and sandy channel communities. SRMs such as
Desulfovibrionaceae were observed to utilize the products of methanotrophs to generate CO
2 via sulfate acceptors via non-stoichiometric oxidation processes. Although the Bay is classified as one of the shallower water bodies, these demineralization activities in sandy channels and bioturbated muds showed similarities to studies of deeper water bodies [
47,
48]. Organohalide-respiring microorganisms (ORMs) such as
Dehalococcoidetes reduce chlorinated compounds into less toxic or harmless products through the anaerobic reductive dechlorination in the presence of an electron donor such as direct H
2 or fermentable organic substrates [
49].
Biogeochemical zonation in sediments was not measured in this study, as it was deemed out of scope for this research and relevant data from prior studies are limited. Moriarty et al. [
50] found that 90% of the sulfate reduction occurred in the top 5 cm of two cores from seagrass habitats in Moreton Bay. The authors also found that methanogenesis accounted for <2% of sediment carbon flow through the microbial population, which was consistent with the view that methanogenesis is low in Moreton Bay sediment. We expect that this short length scale of zonation reported in seagrass sediments by Moriarty et al. [
50] could vary between different habitat types, but there is a paucity of direct geochemical process measurements to validate this speculation. Alternatively, we examine the biogeochemical role of sediment bacterial populations based on metabolic phenotype (e.g.,
Figure 4).
Sediment accumulation in Moreton Bay is driven primarily by episodic delivery of riverine sediment loads during major floods, which form extensive deposits of bioturbated mud and sand in the eastern Bay (
Figure 1). Prior studies have reported high sediment accumulation rates in these depositional regions (>1 cm yr
-1, [
13,
51]) and shown that sediment at the depths sampled in this study (0–40 cm) can be attributed to anthropogenic activity in the catchment over the past 100 years. Depths of active bioturbation are unknown, but analysis of geochemical tracers in mud cores by Coates-Marnane et al. [
13] indicate recent mixing of sediments to depths >40 cm. Morelli and Gasparon [
12] found less mixing in cores from shallow and intertidal environments in Moreton Bay; cores near seagrass-dominated regions showed the lowest disturbance, with mixing confined to the top 10 cm.
Table 3 provides a summary of the bacterial 16S sequencing data based on unique features (out of top 25 features) per class, order, family and genus for each core depth and for each habitat. More unique features at various core depths were observed when examined at family and genus level. The taxonomic classification revealed that the sediment samples at all core depths and from different habitat horizons had ≤2 unique features based on class and order.
The unique features identified at the family level for sediment samples were Acidimicrobiaceae, Cryomorphaceae, Cystobacteraceae, Flammeovirgaceae and Rhodobacteraceae for core depth 10 cm; Ignavibacteriaceae for core depth 20 cm; Pelobacteraceae and Thermodesulfobacteraceae for core depth 30 cm; and Holophagaceae for core depth 40 cm. At the genus level, the unique features were Actibacter, Desulfococcus, Eudorea, and Methylonatrum for core depth 10 cm; Phaselicystis for core depth 20 cm; Desulfatiferula, Fervidicola, Isosphaera, Pelobacter, Thermacetogenium, and Thermococcoides for core depth 30 cm; Smithelia for core depth 40 cm.
At the family level, Acidimicrobiaceae, Cytophagaceae and Halothiobacillaceae were identified as the unique features in river sediments. The unique features in sandy channel sediments were Pelobacteraceae and Thermotogaceae; in bioturbated mud they were Nitrospinaceae and Rhodobacteraceae and in seagrass they were Cystobacteraceae and Saprospiraceae. At the genus level, no unique features were identified in seagrass. The unique features were Actibacter and Planctomyces for river sediments; Desulfobacca, Ignavibacterium, Pelobacter, Smithelia, and Spirochaeta for sandy channel, and Desulfatiferula and Nitrospina for bioturbated mud.
The bacterial 16S rRNA gene sequencing datasets were collated and analyzed using multivariate statistics in order to assess Moreton Bay sites from the perspective of depth-profiling. As such, the sites were pooled and core depth was used as a factor for investigation. The main points of difference in bacterial families identified between sampled depths using multivariate statistics are detailed in
Table 4. It should be noted that members of family
Desulfobulbaceae that belong to SRMs and the family
Caldilineaceae that belong to ORMs were identified as significant bacterial features distinguishing between sediments samples from different core depths. These observations align well with the predicted metabolic functions based on the bacterial 16S rRNA gene sequencing data. Dehalogenation and sulfate reduction were found to be two most significant metabolic functions performed by the bacterial community.
Table 5 presents a summary of predicted phenotypic classification using multivariate statistics. The metabolomics data for depth-profiling was assessed using multivariate statistics. The absence of any significant metabolites indicates that the microbial metabolome is largely similar within the sediments sampled at various core depths, that is, the abundance of metabolites either did not meet the fold change criteria or the
p-value threshold. Microbial metabolomes from deeper sediments suggested differences between microbial communities based on their metabolic functions.
Aerobic organisms were found to be significantly more abundant within top 20 cm sediments. Microbes in this region are known to source their energy from methane and other inorganic compounds containing sulfur, nitrogen or iron. Microbes at depths of 30 cm and 40 cm were anaerobic in nature and sourced their energy from both organic and inorganic compounds. Aerobic bacteria such as Rhizobiaceae, Conexibacteraceae and Flavobacteriaceae were dominant in the top 10 cm of sediment. Anaerobic bacteria such as Methylococcaceae and Desulfuromonadaceae were also found to dominate. Methanotroph bacteria (Methylococcaceae) were the most abundant species at depths of 10 cm and 20 cm. Nitrogen fixing, diazotrophic, aerobic bacteria such as Rhizobiaceae were also identified within the top layer. Anaerobic bacteria such as Thermoactinomycetaceae were dominant at depths of 30 cm and 40 cm. SRMs such as Ectothiorhodospiraceae, Granulosicoccaceae and Chromatiaceae often found in marine environments were also identified in the sediments from Moreton Bay. The halotolerant Oceanospirillaceae, often found in marine environments, were also detected in sediments.
Methylcoccaceae were some of the differential communities in the marine ecosystem. The metabolomics output suggested that certain family members utilized the high nitrogenous substrates present in sediments (See
Table 1) to feed it back to glycolysis via the pentose phosphate pathway to pyruvate metabolism, with additional feeding from starch metabolism by other communities. Recent studies have indicated that the salt tolerant members of
Methylcoccaceae such as
Methylococcus capsulatus have the ability to process nitrogenous substrates via the Benson-Calvin cycle in the presence of methane. This can also be inferred from the considerable upregulation of palmitic acid, which is known to be a major metabolite of this family and other methanotrophs [
52]. Similarly,
Geobacteraceae activity suggested their role in both the production of isoleucine and valine by recruiting 2-hydroxy butanoate from propanoate metabolism, with pyruvic acid as a precursor, as demonstrated in an earlier study [
53]. The metabolic activity also suggested a conversion by Fe (III) mobilization in the overlapping regions between bioturbated mud and sandy channels. Similarly, in the bioturbated mud, these bacteria appeared to use fumarate as an electron acceptor to degrade the benzoic acid (the product of PCB degradation in the riverine ecosystem) via muconic acid intermediate to pyruvate metabolism. This ability has been suggested by Röling [
54], where these bacteria work in commensalism with sulfate-reducing bacteria to degrade organic pollutants such as PCBs and similar polychlorinated compounds resulting from human activities.
A high β-alanine content in the riverine sediments indicated the presence of high degradation of organic matter in sediments, especially at shallow depths. Generally, in marine environments, β-alanine is formed as a degradation product of aspartic acid and a high Asp/ β-alanine ratio indicates the presence of fresh organic matter [
55,
56]. However, in the current study, it appears that a high perturbation in riverine and bioturbated sediments, combined with high nitrogen content, resulted in considerably high biosynthesis of β-alanine through other pathways as well. Similarly, a high β-alanine content (very low Asp/ β-alanine ratio) in this study also indicates a considerable decrease in the amount of fresh water (with respect to saline water) among the sediment samples. These outputs also indicated a high upregulation of β-alanine-pyruvate transaminase amongst the microbial communities in the river and bioturbated mud ecosystems. This output is in line with the previously studied geo-biochemical pathways occurring due to the polluting activities in the estuarine sediments [
56,
57].
The bacterial 16S rRNA gene sequencing datasets were also assessed from the perspective of habitat horizons. The main points of difference in terms of taxonomy (bacterial families) and phenotype identified between sampled horizons using multivariate statistics are detailed in
Table 6 and
Table 7, respectively.
Bacteria such as Sphingomonadaceae that degrade aromatic organics were abundant in the river habitat. Bacteria with diverse characteristics were found to dominate in the river habitat such as thermophilic Hydrogenophilaceae, diazotrophic Oxalobacteraceae and the halotolerant mesophilic chemolithotrophic bacteria Halothiobacillaceae. It is worth noting that bacteria from the family Neisseriaceae were also identified in the river habitat. Members of this family are known to cause gonorrhea and meningitis. Streptomycin-producing bacteria such as Streptomycetaceae were also found in the river habitat.
Organisms in the river habitat were more likely to be photoheterotrophs which use light energy but rely on carbohydrates, fatty acids and alcohols as a carbon source. Most microbes in the river habitat reduce nitrates to release nitrogen into the atmosphere and oxidize sulfur. Nitrogen-fixing bacteria were dominant in the sandy channel habitat. Lignin-degrading bacteria were most likely to be abundant in the bioturbated mud.
The comparison of bacterial 16S rRNA gene sequencing and metabolomics datasets suggest increased anoxygenic conditions in the sediments. This was most marked in the bioturbated mud site sediments. These results indicate a decrease in faunal activity in bioturbated mud sediments (
Figure 2) and may result in decreased oxygen supply. Similar results for sediments from Moreton Bay were reported by our group during the dry season [
8]. This possibly resulted in an increase in the population of SRM such as
Desulfobulbaceae,
Desulfovibrionaceae and other similar species. The increased population of SRM may have changed the metabolite constitution of those sediments. This is indicated by an increased fatty acid metabolism. A putative indicator of increased fatty acid metabolism was palmitic acid (
Table 2). The role of these bacteria in β-oxidation has been reported earlier [
8,
58,
59] and our results are in alignment with these observations. An increased population of SRM also has an impact on sugar and sucrose metabolism [
8]. This was particularly evident in river sediment samples by the presence of trehalose and maltose (
Table 2).
The metabolomics dataset was subjected to pathway impact analysis (
Figure 2) and pathway analysis (
Figure 3). Based on the statistically significant metabolites identified using Fisher’s LSD analysis (
Table 2), sandy channel and river habitat sites were found to be very active.
Supplementary Table S7 presents the predicted metabolic pathways based on significant metabolites identified from sediment samples collected from various habitat sites as determined by the Fisher’s LSD analysis. Glycerolipid metabolism was the only common pathway observed in all habitat sediment sites. The presence of metabolites such as inosinic acid and deoxycytidine in bioturbated mud and river sediment samples, respectively, suggests that a considerable amount of biotransformation related to purine and pyrimidine metabolism occurred at those sites. The bacterial population present at the mouths of the rivers, most likely, were able to divert the pyrimidine metabolism via pantothenate and/or β-alanine intermediate pathways.
Bioturbated mud sediment samples exhibited fructose and mannose metabolism indicated by the presence of mannitol. We observed succinic acid in river and sandy channel sediments, which was indicative of propanoate metabolism, butanoate metabolism, citrate cycle, and glyoxylate and dicarboxylate metabolism. Detection of glycolic acid in river sediment samples also indicated glyoxylate and dicarboxylate metabolism. The detection of valine and α-ketoisovaleric acid, especially in the sandy channel habitat samples, could be attributed to metabolism of complex amino acids such as valine, leucine and isoleucine possibly via propanoate metabolism, butanoate metabolism, pyruvate metabolism, citrate cycle, or β-alanine pathways. Interestingly, 2-pyrocatechuic acid was observed in river sediment samples. This is indicative of a siderophore biosynthesis pathway. Siderophores are ferric iron chelators secreted by microorganisms for survival in low-iron environments, which was supported by the observed low concentrations of iron in the sampled sediments. Other notable metabolic pathways found to be unique in river habitat were D-alanine metabolism, pyruvate metabolism and selenoamino acid metabolism.