Targeted High-Throughput Sequencing Identifies Predominantly Fungal Pathogens in Patients with Clinically Infectious, Culture-Negative Endophthalmitis in South India
Abstract
1. Introduction
2. Results
2.1. Microbiology Culture
2.2. Taxonomic Analysis by NGS
2.3. Agreement between Culture and HTS Techniques for Culture-Positive Samples
2.4. HTS Analysis for Culture-Negative Endophthalmitis
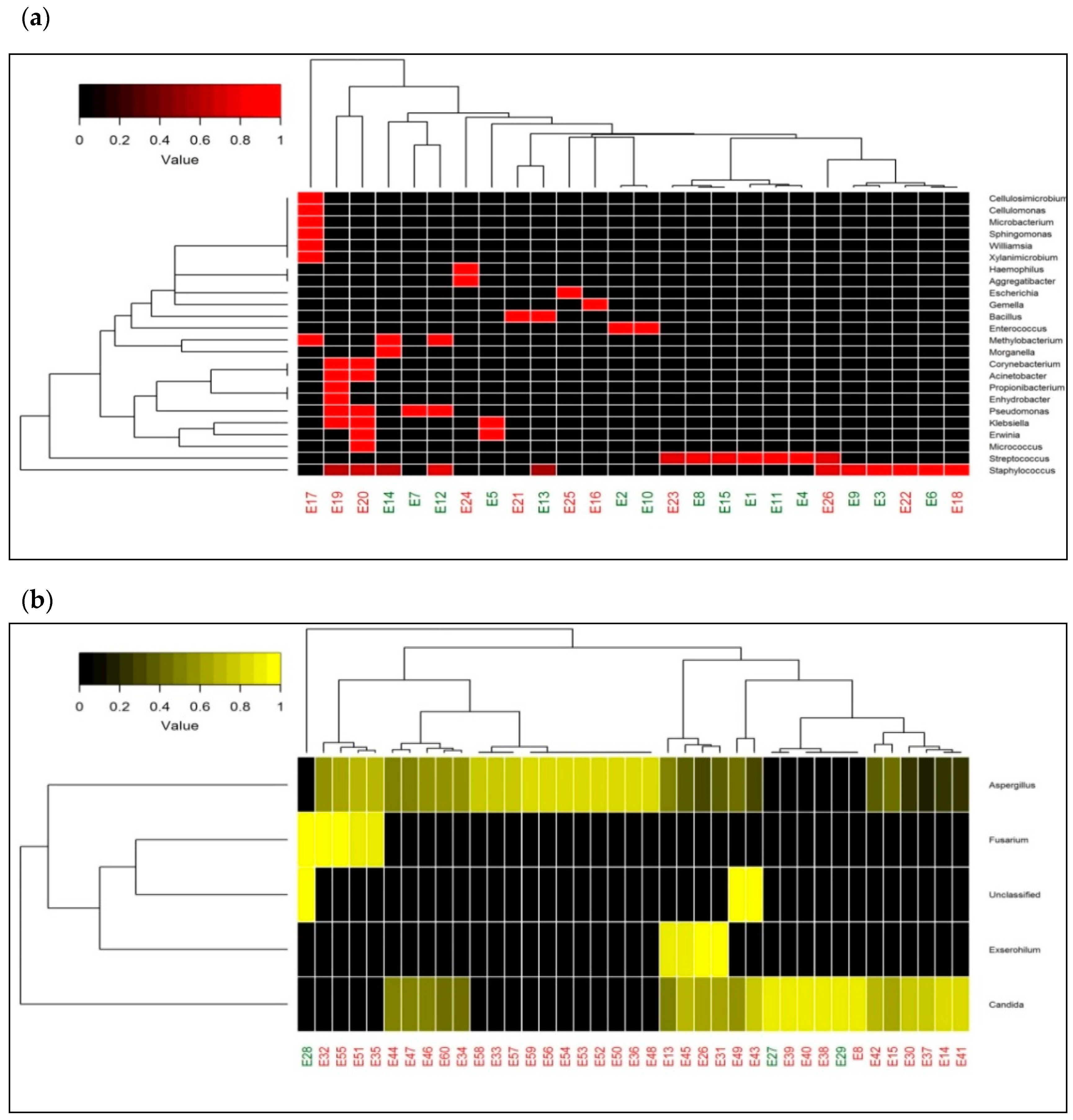
2.5. Heatmap with Bacterial and Fungal Genera
3. Discussion
4. Materials and Methods
4.1. Ethics
4.2. Clinical Samples
4.3. PCR Amplification, Illumina HiSeq Sequencing, and NGS Library Preparation
4.4. Taxonomic Analysis by NGS
4.5. Identification of Contaminant Microbial Reads
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Durand, M.L. Endophthalmitis. Clin. Microbiol. Infect. 2013, 19, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Chiquet, C.; Lina, G.; Benito, Y.; Cornut, P.L.; Etienne, J.; Romanet, J.P.; Denis, P.; Vandenesch, F. Polymerase chain reaction identification in aqueous humor of patients with postoperative endophthalmitis. J. Cataract Refract. Surg. 2007, 33, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Shukla, P.K. Use of PCR Targeting of Internal Transcribed Spacer Regions and Single-Stranded Conformation Polymorphism Analysis of Sequence Variation in Different Regions of rRNA Genes in Fungi for Rapid Diagnosis of Mycotic Keratitis. J. Clin. Microbiol. 2005, 43, 662. [Google Scholar] [CrossRef] [PubMed]
- Hong, B.K.; Lee, C.S.; Van Gelder, R.N.; Garg, S.J. Emerging techniques for pathogen discovery in endophthalmitis. Curr. Opin. Ophthalmol. 2015, 26, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Doan, T.; Wilson, M.R.; Crawford, E.D.; Chow, E.D.; Khan, L.M.; Knopp, K.A.; O’Donovan, B.D.; Xia, D.; Hacker, J.K.; Stewart, J.M.; et al. Illuminating uveitis: Metagenomic deep sequencing identifies common and rare pathogens. Genome Med. 2016, 8, 90. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.Y.; Akileswaran, L.; Tibbetts, M.D.; Garg, S.J.; Van Gelder, R.N. Identification of torque teno virus in culture-negative endophthalmitis by representational deep DNA sequencing. Ophthalmology 2015, 122, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Anis, E.; Hawkins, I.K.; Ilha, M.R.S.; Woldemeskel, M.W.; Saliki, J.T.; Wilkes, R.P. Evaluation of targeted next-generation sequencing for detection of bovine pathogens in clinical samples. J. Clin. Microbiol. 2018, 56, e00399-18. [Google Scholar] [CrossRef] [PubMed]
- Deshmukh, D.; Joseph, J.; Chakrabarti, M.; Sharma, S.; Jayasudha, R.; Sama, K.C.; Sontam, B.; Tyagi, M.; Narayanan, R.; Shivaji, S. New insights into culture negative endophthalmitis by unbiased next generation sequencing. Sci. Rep. 2019, 9, 844. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Miller, S.; Chiu, C.Y. Clinical metagenomic next-generation sequencing for pathogen detection. Annu. Rev. Pathol. 2018, 14, 319–338. [Google Scholar] [CrossRef] [PubMed]
- Dave, V.P.; Sharma, S.; Dave, P.J.; Joseph, J.; Pappuru, R.R. Clinical presentations, diagnostic dilemma and management outcomes of chronic postoperative endophthalmitis caused by Stephanoascus ciferrii. Retin. Cases Brief Rep. 2018. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]

| Culture Positive (18) | Culture Negative (42) | |
|---|---|---|
| Demographic Characteristics | ||
| Age in years (mean:range) | 37.7:2–74 | 38.7:2–81 |
| Sex (male:female) | 15:3 | 28:14 |
| Diagnosis | ||
| Traumatic | 8 | 24 |
| Post-operative | 6 | 15 |
| Endogenous | 4 | 3 |
| Initial Visual Acuity | ||
| Eviseration/Phthisis | 0 | 0 |
| <(20/200) | 17 | 34 |
| >(20/20)–(20/200)< | 1 | 8 |
| =20/20 | 0 | 0 |
| Microbiology | Streptococcus pneumonia (4) Streptococcus mitis (1) | |
| Bacteria | Staphylococcus epidermidis (4) Staphylococcus haemolyticus (1) Enterococcus casseliflavus (1) Enterococcus faecalis (1) Klebsiella pneumoniae (1) Corneybacterium pseudodiphtherticum (1) Bacillus licheniformis (1) | |
| Fungi | Candida trophicalis (1) Fusarium solani (1) Candida albicans (1) | |
| S. No. | Sample ID | Culture Report | HTS Identification Taxonomic Lineage with Abundance |
|---|---|---|---|
| 1 | E1 | Streptococcus pneumoniae | Streptococcus = 99% |
| 2 | E2 | Enterococcus casseliflavus | Enterococcus = 85% Unclassified = 10% |
| 3 | E3 | Staphylococcus epidermidis | Staphylococcus = 89% Unclassified = 10% |
| 4 | E4 | Streptococcus pneumoniae | Streptococcus = 97% |
| 5 | E5 | Klebsiella pneumoniae | Klebsiella = 90% Erwinia = 5% |
| 6 | E6 | Staphylococcus epidermidis | Staphylococcus = 92% Planococcaceae = 6% |
| 7 | E7 | Corneybacterium pseudodiphtherticum | Pseudomonas = 94% Pseudomonadaceae = 5% |
| 8 | E8 | Streptococcus pneumoniae | Streptococcus = 94% |
| 9 | E9 | Staphylococcus epidermidis | Staphylococcus = 86% Planococcaceae = 7% Unclassified = 3% |
| 10 | E10 | Enterococcus faecalis | Enterococcus = 85% Unclassified = 10% Enterococcaceae = 4% |
| 11 | E11 | Streptococcus mitis | Streptococcus = 95% |
| 12 | E12 | Staphylococcus epidermidis | Pseudomonas = 65% Methylobacterium = 14% Staphylococcus = 13% |
| 13 | E13 | Bacillus licheniformis | Bacillus = 86% Staphylococcus = 3% |
| 14 | E14 | Staphylococcus haemolyticus | Methylobacterium = 84% Staphylococcus = 4% Morganllea = 4% |
| 15 | E15 | Streptococcus pneumoniae | Streptococcus = 94% |
| S. No. | Sample ID | Culture Report | HTS Identification Taxonomic Lineage with Abundance |
|---|---|---|---|
| 1 | E27 | Candida trophicalis | Candida = 99% |
| 2 | E28 | Fusarium solani | Fusarium = 59% Unclassified = 41% |
| 3 | E29 | (1) Candida albicans (2) Staphylococcus aureus | Candida = 100% |
| S. No. | Sample ID | HTS Identification Taxonomic Lineage with Abundance |
|---|---|---|
| 1 | E17C | Enterobacteriaceae = 53%, Paracoccus = 13%, Acinetobacter = 5%, Staphylococcus = 4% Enhydrobacter = 3%, Corynebacterium = 3% Rhodobacteraceae = 3% |
| 2 | E26C | Streptococcus = 78%, Propionibacterium = 7% Abiotrophia = 3% |
| 3 | E53C | Enhydrobacter = 32%, Sphingobium = 21% Flavobacterium = 11%, Pseudomonas = 9% Caulobacteraceae = 3% |
| 4 | E54C | Enterobacteriaceae = 52%, Corynebacterium = 9% Citrobacter = 8%, Lactobacillus = 5% Finegoldia = 4%, Erwinia = 3% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gandhi, J.; Jayasudha, R.; Naik, P.; Sharma, S.; Dave, V.P.; Joseph, J. Targeted High-Throughput Sequencing Identifies Predominantly Fungal Pathogens in Patients with Clinically Infectious, Culture-Negative Endophthalmitis in South India. Microorganisms 2019, 7, 411. https://doi.org/10.3390/microorganisms7100411
Gandhi J, Jayasudha R, Naik P, Sharma S, Dave VP, Joseph J. Targeted High-Throughput Sequencing Identifies Predominantly Fungal Pathogens in Patients with Clinically Infectious, Culture-Negative Endophthalmitis in South India. Microorganisms. 2019; 7(10):411. https://doi.org/10.3390/microorganisms7100411
Chicago/Turabian StyleGandhi, Jaishree, Rajagopalaboopathi Jayasudha, Poonam Naik, Savitri Sharma, Vivek Pravin Dave, and Joveeta Joseph. 2019. "Targeted High-Throughput Sequencing Identifies Predominantly Fungal Pathogens in Patients with Clinically Infectious, Culture-Negative Endophthalmitis in South India" Microorganisms 7, no. 10: 411. https://doi.org/10.3390/microorganisms7100411
APA StyleGandhi, J., Jayasudha, R., Naik, P., Sharma, S., Dave, V. P., & Joseph, J. (2019). Targeted High-Throughput Sequencing Identifies Predominantly Fungal Pathogens in Patients with Clinically Infectious, Culture-Negative Endophthalmitis in South India. Microorganisms, 7(10), 411. https://doi.org/10.3390/microorganisms7100411





