Bacterial Endophyte Colonization and Distribution within Plants
Abstract
1. Introduction
2. Recruitment of Bacterial Endophytes by Host Plants
3. Attachment of Bacterial Endophytes to the Host Plant Surface
4. Entry of Bacterial Endophytes into the Host Plant
5. Bacterial Niches inside the Host Plant
6. Bacterial Genes Involved in Plant Colonization
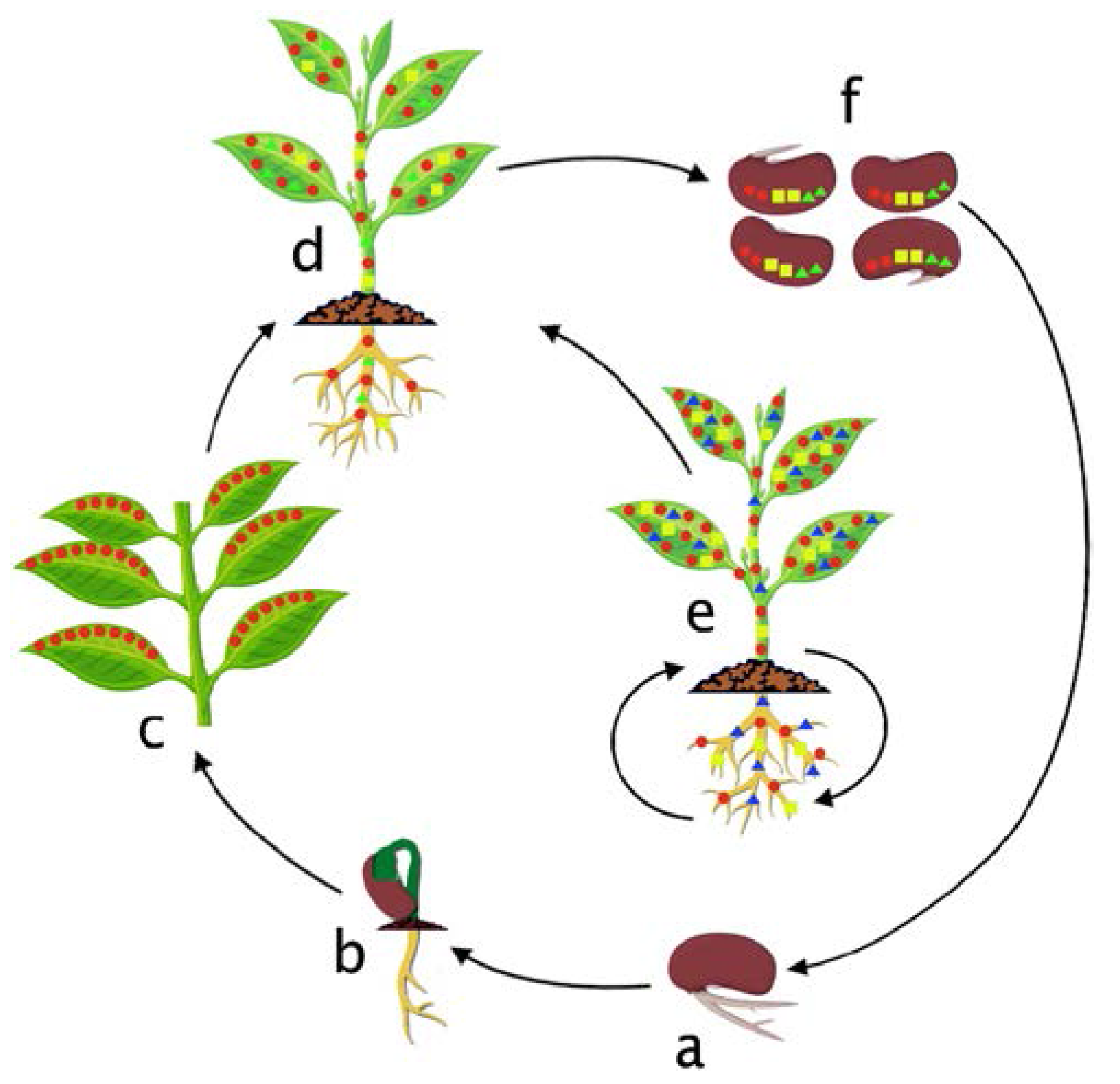
7. Colonization Cycle of Bacterial Endophytes in the Host Plant
8. Methods Used in Colonization Studies
8.1. Cultivation Based Studies
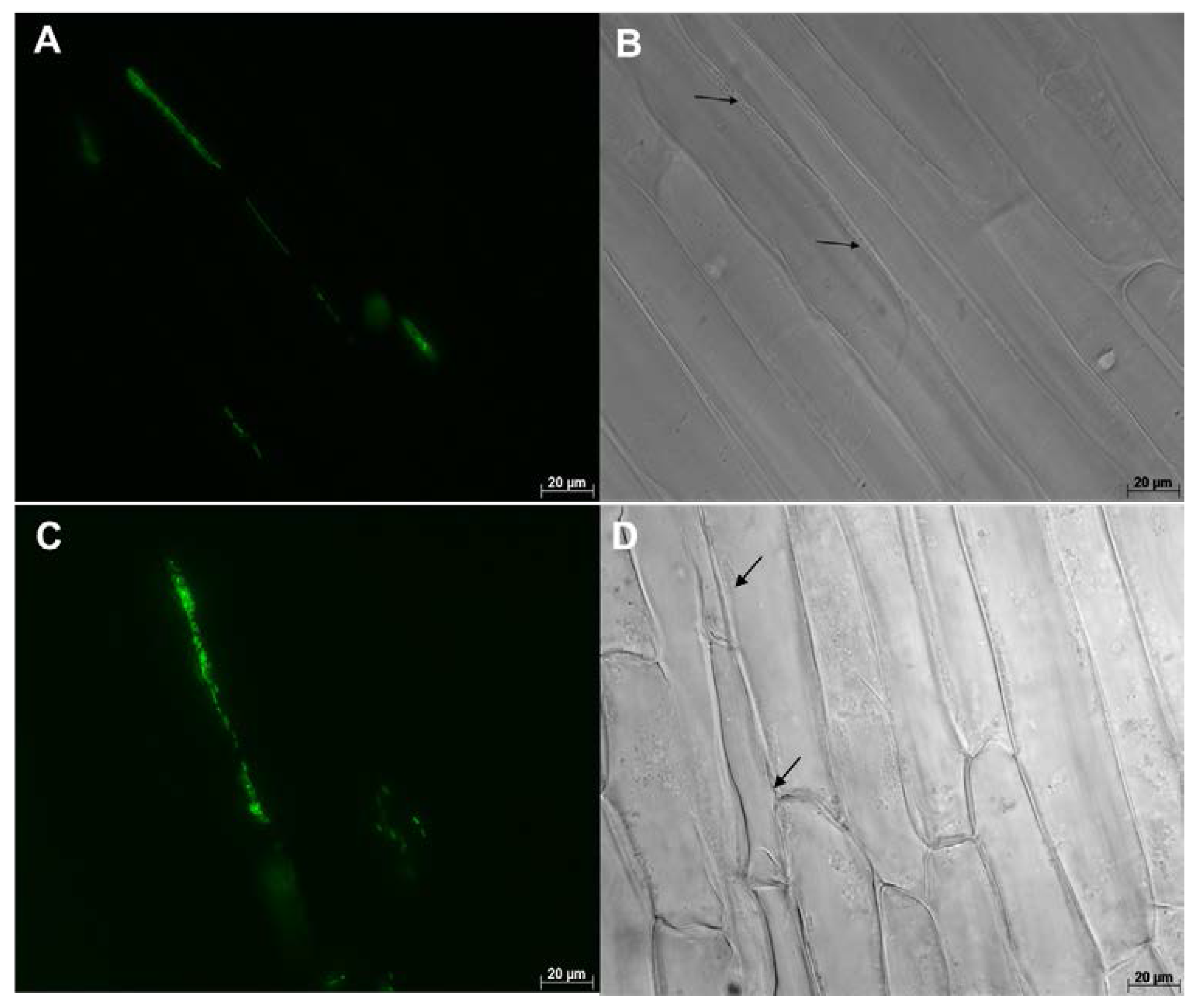
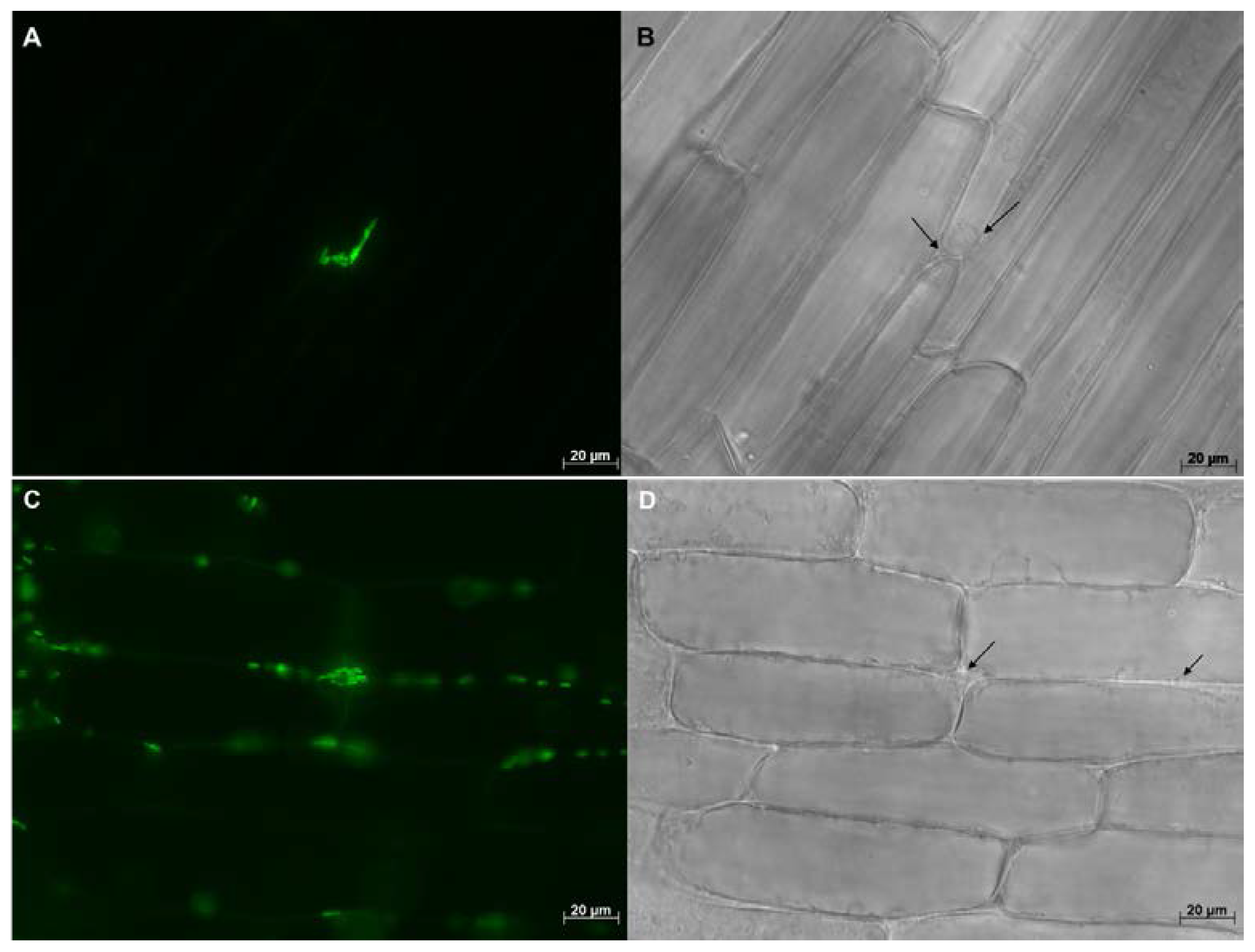
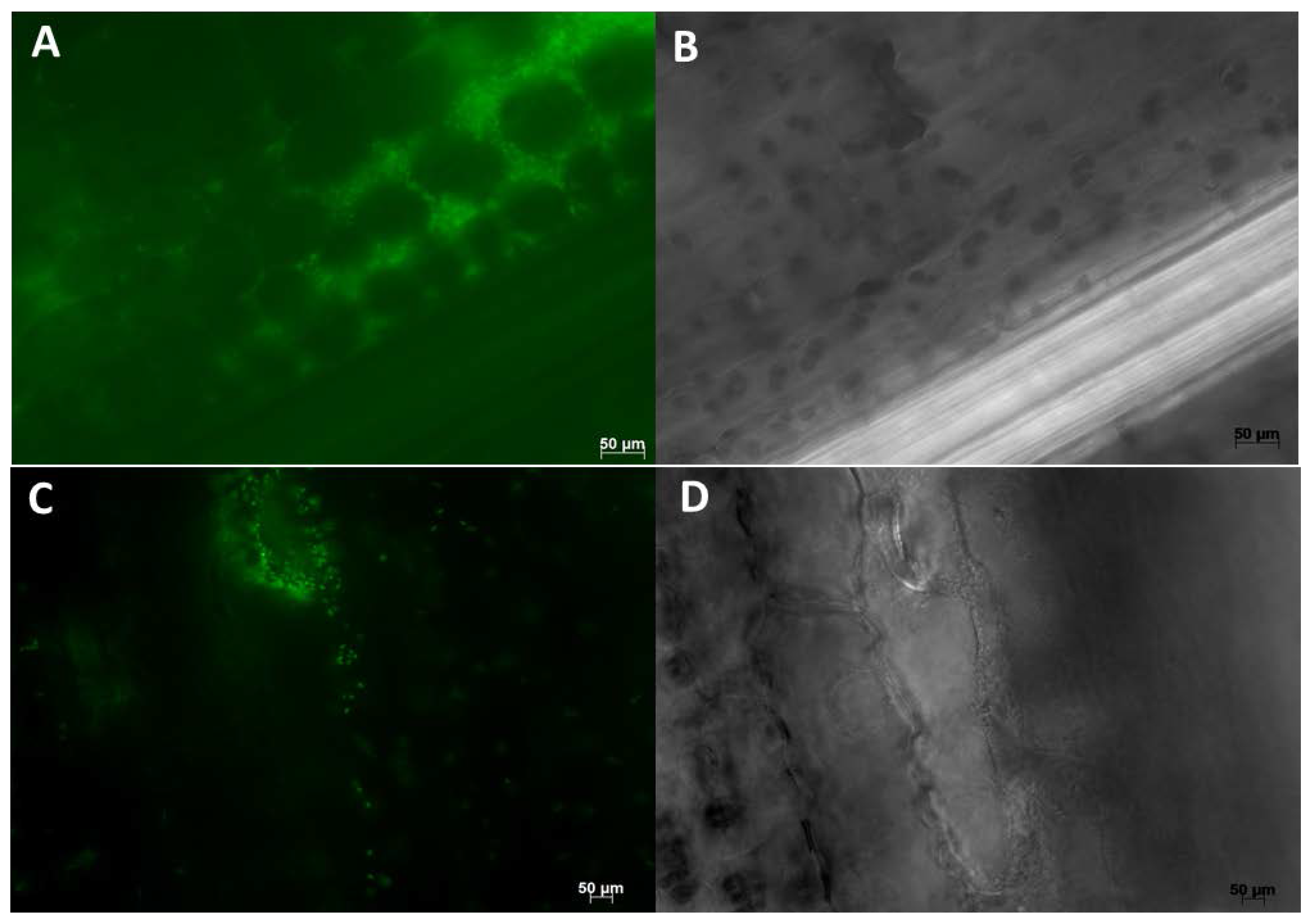
8.2. Microscopy Based Studies
8.3. Genomics Based Studies
9. Poplar Endophytes and Their Colonization Efficiency in Crop Plants
10. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wilson, D. Endophyte: The evolution of a term, and clarification of its use and definition. Oikos 1995, 73, 274–276. [Google Scholar] [CrossRef]
- Dobereiner, J. History and new perspectives of diazotrophs in association with non-leguminous plants. Symbiosis 1992, 13, 1–13. [Google Scholar]
- Schulz, B.; Boyle, C. What are endophytes? In Microbial Root Endophytes; Schulz, B., Boyle, C., Sieber, T., Eds.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 1–14. [Google Scholar]
- Hardoim, P.R.; van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The hidden world within Plants: Ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef] [PubMed]
- Sessitsch, A.; Hardoim, P.; Döring, J.; Weilharter, A.; Krause, A.; Woyke, T.; Mitter, B.; Hauberg-Lotte, L.; Friedrich, F.; Rahalkar, M.; et al. Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol. Plant Microbe Interact. 2012, 25, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Taghavi, S.; Garafola, C.; Monchy, S.; Newman, L.; Hoffman, A.; Weyens, N.; Barac, T.; Vangronsveld, J.; Van Der Lelie, D. Genome survey and characterization of endophytic bacteria exhibiting a beneficial effect on growth and development of poplar trees. Appl. Environ. Microbiol. 2009, 75, 748–757. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, S.; Okubo, T.; Anda, M.; Nakashita, H.; Yasuda, M.; Sato, S.; Kaneko, T.; Tabata, S.; Eda, S.; Momiyama, A.; et al. Community-and genome-based views of plant-associated bacteria: Plant-bacterial interactions in soybean and rice. Plant Cell Physiol. 2010, 51, 1398–1410. [Google Scholar] [CrossRef] [PubMed]
- Turner, T.R.; James, E.K.; Poole, P.S. The plant microbiome. Genome Biol. 2013, 14, 209. [Google Scholar] [CrossRef] [PubMed]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; Ver Loren van Themaat, E.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 2012, 488, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L.; Oakley, B.; Xin, G.; Kang, J.W.; Singleton, G.; Khan, Z.; Vajzovic, A.; Staley, J.T. Diazotrophic endophytes of native black cottonwood and willow. Symbiosis 2009, 47, 23–33. [Google Scholar] [CrossRef]
- Wemheuer, F.; Kaiser, K.; Karlovsky, P.; Daniel, R.; Vidal, S.; Wemheuer, B. Bacterial endophyte communities of three agricultural important grass species differ in their response towards management regimes. Sci. Rep. 2017, 7, 40914. [Google Scholar] [CrossRef] [PubMed]
- Elbeltagy, A.; Nishioka, K.; Sato, T.; Suzuki, H.; Ye, B.; Hamada, T.; Isawa, T.; Mitsui, H. Endophytic colonization and in planta nitrogen fixation by a Herbaspirillum sp. isolated from wild rice species. Appl. Environ. Microbiol. 2001, 67, 5285–5293. [Google Scholar] [CrossRef] [PubMed]
- Hurek, T.; Handley, L.L.; Reinhold-Hurek, B.; Piché, Y.; De, C.; Pavillon, C.; Laval, U.; Gk-p, C. Azoarcus grass endophytes contribute fixed nitrogen to the plant in an unculturable state. Mol. Plant Microbe Interact. 2002, 15, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Iniguez, A.L.; Dong, Y.; Triplett, E.W. Nitrogen fixation in wheat provided by Klebsiella pneumoniae 342. Mol. Plant Microbe Interact. 2004, 17, 1078–1085. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Shen, D.; Song, W. Rice endophyte Pantoea agglomerans YS19 promotes host plant growth and affects allocations of host photosynthates. J. Appl. Microbiol. 2006, 100, 938–945. [Google Scholar] [CrossRef] [PubMed]
- Momose, A.; Ohtake, N.; Sueyoshi, K.; Sato, T.; Nakanishi, Y.; Akao, S.; Ohyama, T. Nitrogen Fixation and Translocation in Young Sugarcane (Saccharum officinarum L.) Plants Associated with Endophytic Nitrogen-Fixing Bacteria. Microbes Environ. 2009, 24, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Botta, A.L.; Santacecilia, A.; Ercole, C.; Cacchio, P.; Del Gallo, M. In vitro and in vivo inoculation of four endophytic bacteria on Lycopersicon esculentum. N. Biotechnol. 2013, 30, 666–674. [Google Scholar] [CrossRef] [PubMed]
- Kandel, S.L.; Herschberger, N.; Kim, S.H.; Doty, S.L. Diazotrophic endophytes of poplar and willow for growth promotion of rice plants in nitrogen-limited conditions. Crop Sci. 2015, 55, 1765–1772. [Google Scholar] [CrossRef]
- Knoth, J.L.; Kim, S.-H.; Ettl, G.J.; Doty, S.L. Effects of cross host species inoculation of nitrogen-fixing endophytes on growth and leaf physiology of maize. GCB Bioenergy 2012, 5, 408–418. [Google Scholar] [CrossRef]
- Gaiero, J.R.; McCall, C.A.; Thompson, K.A.; Day, N.J.; Best, A.S.; Dunfield, K.E. Inside the root microbiome: Bacterial root endophytes and plant growth promotion. Am. J. Bot. 2013, 100, 1738–1750. [Google Scholar] [CrossRef] [PubMed]
- Lebeis, S.L. The potential for give and take in plant-microbiome relationships. Front. Plant Sci. 2014, 5, 287. [Google Scholar] [CrossRef] [PubMed]
- Rosenblueth, M.; Martínez-Romero, E. Bacterial endophytes and their interactions with hosts. Mol. Plant Microbe Interact. 2006, 19, 827–837. [Google Scholar] [CrossRef] [PubMed]
- Bastian, F.; Cohen, A.; Piccoli, P.; Luna, V.; Baraldi, R.; Bottini, R. Production of indole-3-acetic acid and gibberellins A1 and A3 by Acetobacter diazotrophicus and Herbaspirillum seropedicae in chemically-defined culture media. Plant Growth Regul. 1998, 24, 7–11. [Google Scholar] [CrossRef]
- Bhattacharjee, R.B.; Singh, A.; Mukhopadhyay, S.N. Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: Prospects and challenges. Appl. Microbiol. Biotechnol. 2008, 80, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Santi, C.; Bogusz, D.; Franche, C. Biological nitrogen fixation in non-legume plants. Ann. Bot. 2013, 111, 743–767. [Google Scholar] [CrossRef] [PubMed]
- Gillis, M.; Kersters, K.; Hoste, B.; Janssens, D.; Kroppenstedt, R.M.; Stephan, M.P. Acetobacter diazotrophicus sp. nov., a Nitrogen-Fixing Acetic Acid Bacterium Associated with Sugarcane. Int. J. Syst. Bacteriol. 1989, 39, 361–364. [Google Scholar] [CrossRef]
- Dong, Z.; Canny, M.J.; McCully, M.E.; Roboredo, M.R.; Cabadilla, C.F.; Ortega, E.; Rodes, R. A Nitrogen-Fixing Endophyte of Sugarcane Stems (A New Role for the Apoplast). Plant Physiol. 1994, 105, 1139–1147. [Google Scholar] [CrossRef] [PubMed]
- Reinhold-Hurek, B.; Hurek, T.; Gillis, M.; Hoste, B.; Vancanneyt, M.; Kersters, K.; De Ley, J. Azoarcus gen. nov., Nitrogen-Fixing Proteobacteria Associated with Roots of Kallar Grass (Leptochloa fusca (L.) Kunth), and Description of Two Species, Azoarcus indigens sp. nov. and Azoarcus communis sp. nov. Int. J. Syst. Bacteriol. 1993, 43, 574–584. [Google Scholar] [CrossRef]
- Riggs, P.J.; Chelius, M.K.; Iniguez, A.L.; Kaeppler, S.M.; Triplett, E.W. Enhanced maize productivity by inoculation with diazotrophic bacteria. Aust. J. Plant Physiol. 2001, 28, 829–836. [Google Scholar]
- Olivares, J.; Bedmar, E.J.; Sanjuán, J. Biological nitrogen fixation in the context of global change. Mol. Plant Microbe Interact. 2013, 26, 486–494. [Google Scholar] [CrossRef] [PubMed]
- Reinhold-Hurek, B.; Hurek, T. Life in grasses: Diazotrophic endophytes. Trends Microbiol. 1998, 6, 139–144. [Google Scholar] [CrossRef]
- Doty, S.L.; Sher, A.W.; Fleck, N.D.; Khorasani, M.; Bumgarner, R.E.; Khan, Z.; Ko, A.W.K.; Kim, S.-H.; DeLuca, T.H. Variable Nitrogen Fixation in Wild Populus. PLoS ONE 2016, 11, e0155979. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L. Endophytic N-Fixation: Controversy and a Path Forward. In Functional Importance of the Plant Microbiome; Doty, S.L., Ed.; Springer International Publishing: New York, NY, USA, 2017; pp. 17–58. [Google Scholar]
- Erisman, J.W.; Sutton, M.A.; Galloway, J.; Klimont, Z.; Winiwarter, W. How a century of ammonia synthesis changed the world. Nat. Geosci. 2008, 1, 636–639. [Google Scholar] [CrossRef]
- Howarth, R.W. Coastal nitrogen pollution: A review of sources and trends globally and regionally. Harmful Algae 2008, 8, 14–20. [Google Scholar] [CrossRef]
- Reay, D.S.; Davidson, E.A.; Smith, K.A.; Smith, P.; Melillo, J.M.; Dentener, F.; Crutzen, P.J. Global agriculture and nitrous oxide emissions. Nat. Clim. Chang. 2012, 2, 410–416. [Google Scholar] [CrossRef]
- Robarge, W.P.; Walker, J.T.; McCulloch, R.B.; Murray, G. Atmospheric concentrations of ammonia and ammonium at an agricultural site in the southeast United States. Atmos. Environ. 2002, 36, 1661–1674. [Google Scholar] [CrossRef]
- Driscoll, C.T.; Whitall, D.; Aber, J.; Boyer, E.; Castro, M.; Cronan, C.; Goodale, C.L.; Groffman, P.; Hopkinson, C.; Lambert, K.; et al. Nitrogen Pollution in the Northeastern United States: Sources, Effects, and Management Options. Bioscience 2003, 53, 357. [Google Scholar] [CrossRef]
- Santoyo, G.; Moreno-Hagelsieb, G.; del Carmen Orozco-Mosqueda, M.; Glick, B.R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 2016, 183, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Spaepen, S.; Vanderleyden, J. Auxin and plant-microbe interactions. Cold Spring Harb. Perspect. Biol. 2011, 3, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Gamalero, E.; Glick, B.R. Mechanisms used by plant growth-promoting bacteria. In Bacteria in Agrobiology: Plant Nutrient Management; Maheshwari, D.K.K., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 17–47. ISBN 978-3-642-21060-0. [Google Scholar]
- Shi, Y.; Lou, K.; Li, C. Promotion of plant growth by phytohormone-producing endophytic microbes of sugar beet. Biol. Fertil. Soils 2009, 45, 645–653. [Google Scholar] [CrossRef]
- Xin, G.; Zhang, G.; Kang, J.W.; Staley, J.T.; Doty, S.L. A diazotrophic, indole-3-acetic acid-producing endophyte from wild cottonwood. Biol. Fertil. Soils 2009, 45, 669–674. [Google Scholar] [CrossRef]
- Barra, P.J.; Inostroza, N.G.; Acuña, J.J.; Mora, M.L.; Crowley, D.E.; Jorquera, M.A. Formulation of bacterial consortia from avocado (Persea americana Mill.) and their effect on growth, biomass and superoxide dismutase activity of wheat seedlings under salt stress. Appl. Soil Ecol. 2016, 102, 80–91. [Google Scholar] [CrossRef]
- Khan, Z.; Rho, H.; Firrincieli, A.; Hung, S.H.; Luna, V.; Masciarelli, O.; Kim, S.H.; Doty, S.L. Growth enhancement and drought tolerance of hybrid poplar upon inoculation with endophyte consortia. Curr. Plant Biol. 2016, 6, 38–47. [Google Scholar] [CrossRef]
- Ahmed, E.; Holmström, S.J.M. Siderophores in environmental research: Roles and applications. Microb. Biotechnol. 2014, 7, 196–208. [Google Scholar] [CrossRef] [PubMed]
- Dias, A.C.F.; Costa, F.E.C.; Andreote, F.D.; Lacava, P.T.; Teixeira, M.A.; Assumpção, L.C.; Araújo, W.L.; Azevedo, J.L.; et al. Isolation of micropropagated strawberry endophytic bacteria and assessment of their potential for plant growth promotion. World J. Microbiol. Biotechnol. 2009, 25, 189–195. [Google Scholar] [CrossRef]
- Oteino, N.; Lally, R.D.; Kiwanuka, S.; Lloyd, A.; Ryan, D.; Germaine, K.J.; Dowling, D.N. Plant growth promotion induced by phosphate solubilizing endophytic Pseudomonas isolates. Front. Microbiol. 2015, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Passari, A.K.; Mishra, V.K.; Gupta, V.K.; Yadav, M.K.; Saikia, R.; Singh, B.P. In Vitro and In Vivo Plant Growth Promoting Activities and DNA Fingerprinting of Antagonistic Endophytic Actinomycetes Associates with Medicinal Plants. PLoS ONE 2015, 10, e0139468. [Google Scholar] [CrossRef] [PubMed]
- Joe, M.M.; Devaraj, S.; Benson, A.; Sa, T. Isolation of phosphate solubilizing endophytic bacteria from Phyllanthus amarus Schum & Thonn: Evaluation of plant growth promotion and antioxidant activity under salt stress. J. Appl. Res. Med. Aromat. Plants 2016. [Google Scholar] [CrossRef]
- Khan, M.S.; Zaidi, A.; Ahmad, E. Mechanism of phosphate solubilization and physiological functions of phosphate-solubilizing microorganisms. In Phosphate Solubilizing Microorganisms: Principles and Applications of Microphos Technology; Khan, M.S., Zaidi, A., Musarrat, J., Eds.; Springer International Publishing: Cham, Switzerland, 2014; Volume 108, ISBN 978-3-319-08215-8. [Google Scholar]
- Friesen, M.L.; Porter, S.S.; Stark, S.C.; Von Wettberg, E.J.; Sachs, J.L. Microbially Mediated Plant Functional Traits-2011.pdf. Annu. Rev. Ecol. Evol. Syst. 2011, 42, 23–46. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; Lugtenberg, B. Biotechnological Applications of Bacterial Endophytes. Curr. Biotechnol. 2014, 3, 60–75. [Google Scholar] [CrossRef]
- Glick, B.R. Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol. Res. 2014, 169, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Esmaeel, Q.; Pupin, M.; Kieu, N.P.; Chataigné, G.; Béchet, M.; Deravel, J.; Krier, F.; Höfte, M.; Jacques, P.; Leclère, V. Burkholderia genome mining for nonribosomal peptide synthetases reveals a great potential for novel siderophores and lipopeptides synthesis. Microbiologyopen 2016, 1–15. [Google Scholar]
- Larran, S.; Simon, M.R.; Moreno, M.V.; Siurana, M.P.S.; Perell, A. Endophytes from wheat as biocontrol agents against tan spot disease. Biol. Control 2016, 92, 17–23. [Google Scholar] [CrossRef]
- Kandel, S.L.; Firrincieli, A.; Joubert, P.M.; Okubara, P.A.; Leston, N.D.; McGeorge, K.M.; Mugnozza, G.S.; Harfouche, A.; Kim, S.H.; Doty, S.L. An in vitro study of bio-control and plant growth promotion potential of salicaceae endophytes. Front. Microbiol. 2017, 8, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Tapias, D.; Moreno-Galvan, A.; Pardo-Diaz, S.; Obando, M.; Rivera, D.; Bonilla, R. Effect of inoculation with plant growth-promoting bacteria (PGPB) on amelioration of saline stress in maize (Zea mays). Appl. Soil Ecol. 2012, 61, 264–272. [Google Scholar] [CrossRef]
- Ali, S.; Charles, T.C.; Glick, B.R. Amelioration of high salinity stress damage by plant growth-promoting bacterial endophytes that contain ACC deaminase. Plant Physiol. Biochem. 2014, 80, 160–167. [Google Scholar] [CrossRef] [PubMed]
- Naveed, M.; Mitter, B.; Reichenauer, T.G.; Wieczorek, K.; Sessitsch, A. Increased drought stress resilience of maize through endophytic colonization by Burkholderia phytofirmans PsJN and Enterobacter sp. FD17. Environ. Exp. Bot. 2014, 97, 30–39. [Google Scholar] [CrossRef]
- Qin, S.; Zhang, Y.J.; Yuan, B.; Xu, P.Y.; Xing, K.; Wang, J.; Jiang, J.H. Isolation of ACC deaminase-producing habitat-adapted symbiotic bacteria associated with halophyte Limonium sinense (Girard) Kuntze and evaluating their plant growth-promoting activity under salt stress. Plant Soil 2014, 374, 753–766. [Google Scholar] [CrossRef]
- Yaish, M.W.; Antony, I.; Glick, B.R. Isolation and characterization of endophytic plant growth-promoting bacteria from date palm tree (Phoenix dactylifera L.) and their potential role in salinity tolerance. Antonie Van Leeuwenhoek 2015, 107, 1519–1532. [Google Scholar] [CrossRef] [PubMed]
- Senga, R.-A.; Alegria Terrazas, S.; Balbirnie, K.; Blank, M.; Janiak, A.; Szarejko, I.; Chmielewska, B.; Karcz, J.; Morris, J.; Hedley, P.E.; et al. Root Hair Mutations Displace the Barley Rhizosphere Microbiota. Front. Plant Sci. 2017, 8, 1–15. [Google Scholar]
- Kawasaki, A.; Donn, S.; Ryan, P.R.; Mathesius, U.; Devilla, R.; Jones, A.; Watt, M. Microbiome and exudates of the root and rhizosphere of brachypodium distachyon, a model for wheat. PLoS ONE 2016, 11, e0164533. [Google Scholar] [CrossRef] [PubMed]
- Pétriacq, P.; Williams, A.; Cotton, A.; McFarlane, A.E.; Rolfe, S.A.; Ton, J. Metabolite profiling of non-sterile rhizosphere soil. Plant J. 2017, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Kost, T.; Stopnisek, N.; Agnoli, K.; Eberl, L.; Weisskopf, L. Oxalotrophy, a widespread trait of plant-associated Burkholderia species, is involved in successful root colonization of lupin and maize by Burkholderia phytofirmans. Front. Microbiol. 2014, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zúñiga, A.; Poupin, M.J.; Donoso, R.; Ledger, T.; Guiliani, N.; Gutiérrez, R.A.; González, B. Quorum sensing and indole-3-acetic acid degradation play a role in colonization and plant growth promotion of Arabidopsis thaliana by Burkholderia phytofirmans PsJN. Mol. Plant Microbe Interact. 2013, 26, 546–553. [Google Scholar] [CrossRef] [PubMed]
- Schikora, A.; Schenk, S.T.; Hartmann, A. Beneficial effects of bacteria-plant communication based on quorum sensing molecules of the N-acyl homoserine lactone group. Plant Mol. Biol. 2016, 90, 605–612. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, A.L.; Lappala, C.R.; Morlen, R.P.; Pelletier, D.A.; Lu, T.Y.S.; Lankford, P.K.; Harwood, C.S.; Greenberg, E.P. LuxR- and luxI-type quorum-sensing circuits are prevalent in members of the populus deltoides microbiome. Appl. Environ. Microbiol. 2013, 79, 5745–5752. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Rothballer, M.; Hense, B.A.; SchrÃder, P. Bacterial quorum sensing compounds are important modulators of microbe-plant interactions. Front. Plant Sci. 2014, 5, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Wagner, M.R.; Lundberg, D.S.; del Rio, T.G.; Tringe, S.G.; Dangl, J.L.; Mitchell-Olds, T. Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat. Commun. 2016, 7, 12151. [Google Scholar] [CrossRef] [PubMed]
- Tan, Z.; Hurek, T.; Reinhold-Hurek, B. Effect of N-fertilization, plant genotype and environmental conditions on nifH gene pools in roots of rice. Environ. Microbiol. 2003, 5, 1009–1015. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Blanco, A.; Sicardi, M.; Frioni, L. Plant genotype and nitrogen fertilization effects on abundance and diversity of diazotrophic bacteria associated with maize (Zea mays L.). Biol. Fertil. Soils 2015, 51, 391–402. [Google Scholar] [CrossRef]
- Lundberg, D.S.; Lebeis, S.L.; Paredes, S.H.; Yourstone, S.; Gehring, J.; Malfatti, S.; Tremblay, J.; Engelbrektson, A.; Kunin, V.; del Rio, T.G.; et al. Defining the core Arabidopsis thaliana root microbiome. Nature 2012, 488, 86–90. [Google Scholar] [CrossRef] [PubMed]
- Meneses, C.H.S.G.; Rouws, L.F.M.; Simoes-Araujo, J.L.; Vidal, M.S.; Baldani, J.I. Exopolysaccharide production is required for biofilm formation and plant colonization by the nitrogen-fixing endophyte Gluconacetobacter diazotrophicus. Mol. Plant Microbe Interact. 2011, 24, 1448–1458. [Google Scholar] [CrossRef] [PubMed]
- Meneses, C.; Gonçalves, T.; Alquéres, S.; Rouws, L.; Serrato, R.; Vidal, M.; Baldani, J.I. Gluconacetobacter diazotrophicus exopolysaccharide protects bacterial cells against oxidative stress in vitro and during rice plant colonization. Plant Soil 2017, 416, 133–147. [Google Scholar] [CrossRef]
- Balsanelli, E.; De Baura, V.A.; De Oliveira Pedrosa, F.; De Souza, E.M.; Monteiro, R.A. Exopolysaccharide biosynthesis enables mature biofilm formation on abiotic surfaces by Herbaspirillum seropedicae. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wang, X.; Pu, H.; Liu, S.; Kan, J.; Jin, C. Recent advances in endophytic exopolysaccharides: Production, structural characterization, physiological role and biological activity. Carbohydr. Polym. 2017, 157, 1113–1124. [Google Scholar] [CrossRef] [PubMed]
- Balsanelli, E.; Serrato, R.V.; de Baura, V.A.; Sassaki, G.; Yates, M.G.; Rigo, L.U.; Pedrosa, F.O.; de Souza, E.M.; Monteiro, R.A. Herbaspirillum seropedicae rfbB and rfbC genes are required for maize colonization. Environ. Microbiol. 2010, 12, 2233–2244. [Google Scholar] [PubMed]
- Balsanelli, E.; Tuleski, T.R.; de Baura, V.A.; Yates, M.G.; Chubatsu, L.S.; de Oliveira Pedrosa, F.; de Souza, E.M.; Monteiro, R.A. Maize Root Lectins Mediate the Interaction with Herbaspirillum seropedicae via N-Acetyl Glucosamine Residues of Lipopolysaccharides. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Compant, S.; Reiter, B.; Nowak, J.; Sessitsch, A.; Clément, C.; Barka, E.A. Endophytic Colonization of Vitis vinifera L. by Plant Growth- Promoting Bacterium Burkholderia sp. Strain PsJN. Appl. Environ. Microbiol. 2005, 71, 1685–1693. [Google Scholar] [CrossRef] [PubMed]
- Vargas, L.; de Carvalho, T.L.G.; Ferreira, P.C.G.; Baldani, V.L.D.; Baldani, J.I.; Hemerly, A.S. Early responses of rice (Oryza sativa L.) seedlings to inoculation with beneficial diazotrophic bacteria are dependent on plant and bacterial genotypes. Plant Soil 2012, 356, 127–137. [Google Scholar] [CrossRef]
- Rangjaroen, C.; Sungthong, R.; Rerkasem, B.; Teaumroong, N.; Noisangiam, R.; Lumyong, S. Untapped Endophytic Colonization and Plant Growth-Promoting Potential of the Genus Novosphingobium to Optimize Rice Cultivation. Microbes Environ. 2017, 32, 84–87. [Google Scholar] [CrossRef] [PubMed]
- Romanstchuk, M. Plant pathogenic bacteria attach to surface. Annu. Rev. Phytopathol 1992, 30, 225–243. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Navarro, D.N.; Dardanelli, M.S.; Ruíz-Saínz, J.E. Attachment of bacteria to the roots of higher plants. FEMS Microbiol. Lett. 2007, 272, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Matthysse, A.G. Attachment of Agrobacterium to plant surfaces. Front. Plant Sci. 2014, 5, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Pankievicz, V.C.S.; Camilios-Neto, D.; Bonato, P.; Balsanelli, E.; Tadra-Sfeir, M.Z.; Faoro, H.; Chubatsu, L.S.; Donatti, L.; Wajnberg, G.; Passetti, F.; et al. RNA-seq transcriptional profiling of Herbaspirillum seropedicae colonizing wheat (Triticum aestivum) roots. Plant Mol. Biol. 2016, 90, 589–603. [Google Scholar] [CrossRef] [PubMed]
- Reinhold-Hurek, B.; Maes, T.; Gemmer, S.; Van Montagu, M.; Hurek, T. An endoglucanase is involved in infection of rice roots by the not-cellulose-metabolizing endophyte Azoarcus sp. strain BH72. Mol. Plant Microbe Interact. 2006, 19, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Naveed, M.; Mitter, B.; Yousaf, S.; Pastar, M.; Afzal, M.; Sessitsch, A. The endophyte Enterobacter sp. FD17: A maize growth enhancer selected based on rigorous testing of plant beneficial traits and colonization characteristics. Biol. Fertil. Soils 2014, 50, 249–262. [Google Scholar] [CrossRef]
- Prieto, P.; Schilirò, E.; Maldonado-González, M.M.; Valderrama, R.; Barroso-Albarracín, J.B.; Mercado-Blanco, J. Root hairs play a key role in the endophytic colonization of olive roots by Pseudomonas spp. with biocontrol activity. Microb. Ecol. 2011, 62, 435–445. [Google Scholar] [CrossRef] [PubMed]
- Rangel de Souza, A.L.S.; De Souza, S.A.; De Oliveira, M.V.V.; Ferraz, T.M.; Figueiredo, F.A.M.M.A.; Da Silva, N.D.; Rangel, P.L.; Panisset, C.R.S.; Olivares, F.L.; Campostrini, E.; et al. Endophytic colonization of Arabidopsis thaliana by Gluconacetobacter diazotrophicus and its effect on plant growth promotion, plant physiology, and activation of plant defense. Plant Soil 2016, 399, 257–270. [Google Scholar] [CrossRef]
- Germaine, K.; Keogh, E.; Garcia-Cabellos, G.; Borremans, B.; Lelie, D.; Barac, T.; Oeyen, L.; Vangronsveld, J.; Moore, F.P.; Moore, E.R.B.; et al. Colonisation of poplar trees by gfp expressing bacterial endophytes. FEMS Microbiol. Ecol. 2004, 48, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Mitter, B.; Pfaffenbichler, N.; Flavell, R.; Compant, S.; Antonielli, L.; Petric, A.; Berninger, T.; Naveed, M.; Sheibani-Tezerji, R.; von Maltzahn, G.; et al. A new approach to modify plant microbiomes and traits by introducing beneficial bacteria at flowering into progeny seeds. Front. Microbiol. 2017, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Glassner, H.; Zchori-Fein, E.; Yaron, S.; Sessitsch, A.; Sauer, U.; Compant, S. Bacterial niches inside seeds of Cucumis melo L. Plant Soil 2017. [Google Scholar] [CrossRef]
- Castanheira, N.L.; Dourado, A.C.; Pais, I.; Semedo, J.; Scotti-Campos, P.; Borges, N.; Carvalho, G.; Barreto Crespo, M.T.; Fareleira, P. Colonization and beneficial effects on annual ryegrass by mixed inoculation with plant growth promoting bacteria. Microbiol. Res. 2017, 198, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Fisher, P.J.; Petrini, O.; Lappin Scott, H.M. The distribution of some fungal and bacterial endophytes in maize (Zea mays L.). New Phytol. 1992, 122, 299–305. [Google Scholar] [CrossRef]
- Wei, C.-Y.; Lin, L.; Luo, L.-J.; Xing, Y.-X.; Hu, C.-J.; Yang, L.-T.; Li, Y.-R.; An, Q. Endophytic nitrogen-fixing Klebsiella variicola strain DX120E promotes sugarcane growth. Biol. Fertil. Soils 2013, 50, 657–666. [Google Scholar] [CrossRef]
- Gyaneshwar, P.; James, E.K.; Mathan, N.; Reddy, P.M.; Reinhold-hurek, B.; Jagdish, K. Endophytic Colonization of Rice by a Diazotrophic Strain of Serratia marcescens. J. Bacteriol. 2001, 183, 2634–2645. [Google Scholar] [CrossRef] [PubMed]
- James, E.K.; Gyaneshwar, P.; Mathan, N.; Barraquio, W.L.; Reddy, P.M.; Iannetta, P.P.M.; Olivares, F.L.; Ladha, J.K. Infection and colonization of rice seedlings by the plant growth-promoting bacterium Herbaspirillum seropedicae Z67. Mol. Plant Microbe Interact. 2002, 15, 894–906. [Google Scholar] [CrossRef] [PubMed]
- Lo Piccolo, S.; Ferraro, V.; Alfonzo, A.; Settanni, L.; Ercolini, D.; Burruano, S.; Moschetti, G. Presence of endophytic bacteria in Vitis vinifera leaves as detected by fluorescence in situ hybridization. Ann. Microbiol. 2010, 60, 161–167. [Google Scholar] [CrossRef]
- Frank, A.; Saldierna Guzmán, J.; Shay, J. Transmission of Bacterial Endophytes. Microorganisms 2017, 5, 70. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.; Reddy, K.M. Microscopic elucidation of abundant endophytic bacteria colonizing the cell wall-plasma membrane peri-space in the shoot-tip tissue of banana. AoB Plants 2013, 5, 1–12. [Google Scholar] [CrossRef]
- Pirttilä, A.M.; Laukkanen, H.; Pospiech, H. Detection of Intracellular Bacteria in the Buds of Scotch Pine (Pinus sylvestris L.) by In Situ Hybridization Detection of Intracellular Bacteria in the Buds of Scotch Pine (Pinus sylvestris L.) by In Situ Hybridization. Appl. Environ. Microbiol. 2000, 66, 3073–3077. [Google Scholar] [CrossRef] [PubMed]
- White, J.F.; Torres, M.S.; Somu, M.P.; Johnson, H.; Irizarry, I.; Chen, Q.; Zhang, N.; Walsh, E.; Tadych, M.; Bergen, M. Hydrogen peroxide staining to visualize intracellular bacterial infections of seedling root cells. Microsc. Res. Tech. 2014, 77, 566–573. [Google Scholar] [CrossRef] [PubMed]
- De Almeida, C.V.; Andreote, F.D.; Yara, R.; Tanaka, F.A.O.; Azevedo, J.L.; de Almeida, M. Bacteriosomes in axenic plants: Endophytes as stable endosymbionts. World J. Microbiol. Biotechnol. 2009, 25, 1757–1764. [Google Scholar] [CrossRef]
- Perrine-Walker, F.M.; Prayitno, J.; Rolfe, B.G.; Weinman, J.J.; Hocart, C.H. Infection process and the interaction of rice roots with rhizobia. J. Exp. Bot. 2007, 58, 3343–3350. [Google Scholar] [CrossRef] [PubMed]
- Paungfoo-Lonhienne, C.; Rentsch, D.; Robatzek, S.; Webb, R.I.; Sagulenko, E.; Näsholm, T.; Schmidt, S.; Lonhienne, T.G.A. Turning the table: Plants consume microbes as a source of nutrients. PLoS ONE 2010, 5, e11915. [Google Scholar] [CrossRef] [PubMed]
- Paungfoo-lonhienne, C.; Schmidt, S.; Webb, R.I.; Interactions, P.M. Rhizophagy—A new dimension of plant—Microbe interactions. In Molecular Microbial Ecology of the Rhizosphere; de Bruijn, F.J., Ed.; Wiley-Blackwell: Hoboken, NJ, USA, 2013; Volume 2, pp. 1201–1207. [Google Scholar]
- Choudhury, F.K.; Rivero, R.M.; Blumwald, E.; Mittler, R. Reactive Oxygen species, abiotic stress and stress combination. Plant J. 2017, 90, 856–867. [Google Scholar] [CrossRef] [PubMed]
- Baker, C.J.; Orlandi, E.W. Active oxygen in plant pathogenesis. Annu. Rev. Phytopathol. 1993, 33, 299–321. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Silva, H.; Klessig, D.F. Active Oxygen Species in the Induction of Plant Systemic Acquired Resistance by Salicylic Acid. Science 1993, 262, 1883–1886. [Google Scholar] [CrossRef] [PubMed]
- Mehdy, M.C. Active Oxygen Species in Plant Defense against Pathogens. Plant Physiol. 1994, 105, 467–472. [Google Scholar] [CrossRef] [PubMed]
- Alquéres, S.; Meneses, C.; Rouws, L.; Rothballer, M.; Baldani, I.; Schmid, M.; Hartmann, A. The bacterial superoxide dismutase and glutathione reductase are crucial for endophytic colonization of rice roots by Gluconacetobacter diazotrophicus PAL5. Mol. Plant Microbe Interact. 2013, 26, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Krause, A.; Ramakumar, A.; Bartels, D.; Battistoni, F.; Bekel, T.; Boch, J.; Böhm, M.; Friedrich, F.; Hurek, T.; Krause, L.; et al. Complete genome of the mutualistic, N2-fixing grass endophyte Azoarcus sp. strain BH72. Nat. Biotechnol. 2006, 24, 1385–1391. [Google Scholar] [CrossRef] [PubMed]
- Taghavi, S.; van der Lelie, D. Genome Sequence of the Plant Growth-Promoting Endophytic Bacterium Enterobacter sp. 638. Mol. Microb. Ecol. Rhizosph. 2013, 2, 899–908. [Google Scholar] [CrossRef] [PubMed]
- Mitter, B.; Petric, A.; Shin, M.W.; Chain, P.S.G.; Hauberg-Lotte, L.; Reinhold-Hurek, B.; Nowak, J.; Sessitsch, A. Comparative genome analysis of Burkholderia phytofirmans PsJN reveals a wide spectrum of endophytic lifestyles based on interaction strategies with host plants. Front. Plant Sci. 2013, 4, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Duan, J.; Charles, T.C.; Glick, B.R. A bioinformatics approach to the determination of genes involved in endophytic behavior in Burkholderia spp. J. Theor. Biol. 2014, 343, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Nelson, E.B. The seed microbiome: Origins, interactions, and impacts. Plant Soil 2017, 1–28. [Google Scholar] [CrossRef]
- Shade, A.; Jacques, M.A.; Barret, M. Ecological patterns of seed microbiome diversity, transmission, and assembly. Curr. Opin. Microbiol. 2017, 37, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Truyens, S.; Weyens, N.; Cuypers, A.; Vangronsveld, J. Bacterial seed endophytes: Genera, vertical transmission and interaction with plants. Environ. Microbiol. Rep. 2015, 7, 40–50. [Google Scholar] [CrossRef]
- Johnston-Monje, D.; Raizada, M.N. Conservation and diversity of seed associated endophytes in Zea across boundaries of evolution, ethnography and ecology. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L. Functional Importance of the Plant Endophytic Microbiome: Implications for Agriculture, Forestry, and Bioenergy. In Functional Importance of the Plant Microbiome; Doty, S.L., Ed.; Springer International Publishing: New York, NY, USA, 2017; pp. 1–5. [Google Scholar]
- Compant, S.; Clément, C.; Sessitsch, A. Plant growth-promoting bacteria in the rhizo- and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol. Biochem. 2010, 42, 669–678. [Google Scholar] [CrossRef]
- Martiny, J.B.H.; Jones, S.E.; Lennon, J.T.; Martiny, A.C. Microbiomes in light of traits: A phylogenetic perspective. Science 2015, 350, aac9323. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.K.; Archana, G. Diverse culturable diazotrophic endophytic bacteria from Poaceae plants show cross-colonization and plant growth promotion in wheat. Plant Soil 2017, 417, 99–116. [Google Scholar] [CrossRef]
- Hurek, T.; Reinhold-Hurek, B.; Van Montagu, M.; Kellenberger, E. Root colonization and systemic spreading of Azoarcus sp. strain BH72 in grasses. J. Bacteriol. 1994, 176, 1913–1923. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhao, H.; Chen, S. Colonization of maize and rice plants by strain Bacillus megaterium C4. Curr. Microbiol. 2006, 52, 186–190. [Google Scholar] [CrossRef] [PubMed]
- Bacilio-Jiménez, M.; Aguilar-Flores, S.; Del Valle, M.V.; Pérez, A.; Zepeda, A.; Zenteno, E. Endophytic bacteria in rice seeds inhibit early colonization of roots by Azospirillum brasilense. Soil Biol. Biochem. 2001, 33, 167–172. [Google Scholar] [CrossRef]
- Tian, B.; Zhang, C.; Ye, Y.; Wen, J.; Wu, Y.; Wang, H.; Li, H.; Cai, S.; Cai, W.; Cheng, Z.; et al. Beneficial traits of bacterial endophytes belonging to the core communities of the tomato root microbiome. Agric. Ecosyst. Environ. 2017, 247, 149–156. [Google Scholar] [CrossRef]
- Ji, X.; Lu, G.; Gai, Y.; Zheng, C.; Mu, Z. Biological control against bacterial wilt and colonization of mulberry by an endophytic Bacillus subtilis strain. FEMS Microbiol. Ecol. 2008, 65, 565–573. [Google Scholar] [CrossRef] [PubMed]
- Compant, S.; Kaplan, H.; Sessitsch, A.; Nowak, J.; Ait Barka, E.; Clement, C. Endophytic colonization of Vitis vinifera L. by Burkholderia phytofirmans strain PsJN: From the rhizosphere to inflorescence tissues. FEMS Microbiol. Ecol. 2008, 63, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Lowman, S.; Hou, G.; Nowak, J.; Flinn, B.; Mei, C. Growth promotion and colonization of switchgrass (Panicum virgatum) cv. Alamo by bacterial endophyte Burkholderia phytofirmans strain PsJN. Biotechnol. Biofuels 2012, 5, 37. [Google Scholar] [CrossRef] [PubMed]
- Govindarajan, M.; Balandreau, J.; Kwon, S.W.; Weon, H.Y.; Lakshminarasimhan, C. Effects of the inoculation of Burkholderia vietnamensis and related endophytic diazotrophic bacteria on grain yield of rice. Microb. Ecol. 2008, 55, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Govindarajan, M.; Balandreau, J.; Muthukumarasamy, R.; Revathi, G.; Lakshminarasimhan, C. Improved yield of micropropagated sugarcane following inoculation by endophytic Burkholderia vietnamiensis. Plant Soil 2006, 280, 239–252. [Google Scholar] [CrossRef]
- Doty, S.L.; Freeman, J.L.; Cohu, C.M.; Burken, J.G.; Firrincieli, A.; Simon, A.; Khan, Z.; Isebrands, J.G.; Lukas, J.; Blaylock, M.J. Enhanced Degradation of TCE on a Superfund Site Using Endophyte-Assisted Poplar Tree Phytoremediation. Environ. Sci. Technol. 2017, 51, 10050–10058. [Google Scholar] [CrossRef] [PubMed]
- Luna, M.F.; Galar, M.L.; Aprea, J.; Molinari, M.L.; Boiardi, J.L. Colonization of sorghum and wheat by seed inoculation with Gluconacetobacter diazotrophicus. Biotechnol. Lett. 2010, 32, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Rouws, L.F.M.; Meneses, C.H.S.G.; Guedes, H.V.; Vidal, M.S.; Baldani, J.I.; Schwab, S. Monitoring the colonization of sugarcane and rice plants by the endophytic diazotrophic bacterium Gluconacetobacter diazotrophicus marked with gfp and gusA reporter genes. Lett. Appl. Microbiol. 2010, 51, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Amaral, F.P.; Bueno, J.C.F.; Hermes, V.S.; Arisi, A.C.M. Gene expression analysis of maize seedlings (DKB240 variety) inoculated with plant growth promoting bacterium Herbaspirillum seropedicae. Symbiosis 2014, 62, 41–50. [Google Scholar] [CrossRef]
- Roncato-Maccari, L.D.B.; Ramos, H.J.O.; Pedrosa, F.O.; Alquini, Y.; Chubatsu, L.S.; Yates, M.G.; Rigo, L.U.; Steffens, M.B.R.; Souza, E.M. Endophytic Herbaspirillum seropedicae expresses nif genes in gramineous plants. FEMS Microbiol. Ecol. 2003, 45, 39–47. [Google Scholar] [CrossRef]
- Brusamarello-Santos, L.C.; Gilard, F.; Brulé, L.; Quilleré, I.; Gourion, B.; Ratet, P.; De Souza, E.M.; Lea, P.J.; Hirel, B. Metabolic profiling of two maize (Zea mays L.) inbred lines inoculated with the nitrogen fixing plant-interacting bacteria Herbaspirillum seropedicae and Azospirillum brasilense. PLoS ONE 2017, 12, e0174576. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Iniguez, A.L.; Ahmer, B.M.M.; Triplett, E.W. Kinetics and strain specificity of rhizosphere and endophytic colonization by enteric bacteria on seedlings of Medicago sativa and Medicago truncatula. Appl. Environ. Microbiol. 2003, 69, 1783–1790. [Google Scholar] [CrossRef] [PubMed]
- Sheng, X.F.; Xia, J.J.; Jiang, C.Y.; He, L.Y.; Qian, M. Characterization of heavy metal-resistant endophytic bacteria from rape (Brassica napus) roots and their potential in promoting the growth and lead accumulation of rape. Environ. Pollut. 2008, 156, 1164–1170. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.C.; Singh, A.; Chowdhury, S.P.; Tripathi, A.K. Endophytic colonization ability of two deep-water rice endophytes, Pantoea sp. and Ochrobactrum sp. using green fluorescent protein reporter. Biotechnol. Lett. 2004, 26, 425–429. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.C.; Ladha, J.K.; Tripathi, A.K. Evaluation of plant growth promoting and colonization ability of endophytic diazotrophs from deep water rice. J. Bacteriol. 2001, 91, 127–141. [Google Scholar] [CrossRef]
- Long, H.H.; Schmidt, D.D.; Baldwin, I.T. Native bacterial endophytes promote host growth in a species-specific manner; phytohormone manipulations do not result in common growth responses. PLoS ONE 2008, 3, e2702. [Google Scholar] [CrossRef] [PubMed]
- Duijff, B.J.; Gianinazzi-Pearson, V.; Lemanceau, P. Involvement of the outer membrane lipopolysaccharides in the endophytic colonization of tomato roots by biocontrol Pseudomonas fluorescens strain WCS417r. New Phytol. 1997, 135, 325–334. [Google Scholar] [CrossRef]
- Khan, Z.; Roman, D.; Kintz, T.; Delas Alas, M.; Yap, R.; Doty, S. Degradation, phytoprotection and phytoremediation of phenanthrene by endophyte Pseudomonas putida, PD1. Environ. Sci. Technol. 2014, 48, 12221–12228. [Google Scholar] [CrossRef] [PubMed]
- Andreote, F.D.; De Araújo, W.L.; De Azevedo, J.L.; Van Elsas, J.D.; Da Rocha, U.N.; Van Overbeek, L.S. Endophytic colonization of potato (Solanum tuberosum L.) by a novel competent bacterial endophyte, Pseudomonas putida strain P9, and its effect on associated bacterial communities. Appl. Environ. Microbiol. 2009, 75, 3396–3406. [Google Scholar] [CrossRef] [PubMed]
- Germaine, K.J.; Liu, X.; Cabellos, G.G.; Hogan, J.P.; Ryan, D.; Dowling, D.N. Bacterial endophyte-enhanced phytoremediation of the organochlorine herbicide 2,4-dichlorophenoxyacetic acid. FEMS Microbiol. Ecol. 2006, 57, 302–310. [Google Scholar] [CrossRef] [PubMed]
- Khan, Z.; Doty, S.L. Characterization of bacterial endophytes of sweet potato plants. Plant Soil 2009, 322, 197–207. [Google Scholar] [CrossRef]
- Oliveira, A.L.M.; Urquiaga, S.; Döbereiner, J.; Baldani, J.I. The effect of inoculating endophytic N2-fixing bacteria on micropropagated sugarcane plants. Plant Soil 2002, 242, 205–215. [Google Scholar] [CrossRef]
- Knoth, J.L.; Kim, S.; Ettl, G.J.; Doty, S.L. Biological nitrogen fixation and biomass accumulation within poplar clones as a result of inoculations with diazotrophic endophyte consortia. New Phytol. 2014, 201, 599–609. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.L.M.; Stoffels, M.; Schmid, M.; Reis, V.M.; Baldani, J.I.; Hartmann, A. Colonization of sugarcane plantlets by mixed inoculations with diazotrophic bacteria. Eur. J. Soil Biol. 2009, 45, 106–113. [Google Scholar] [CrossRef]
- Khan, Z.; Kandel, S.L.; Ramos, D.N.; Ettl, G.J.; Kim, S.; Doty, S.L. Increased Biomass of Nursery-Grown Douglas-Fir Seedlings upon Inoculation with Diazotrophic Endophytic Consortia. Forests 2015, 6, 3582–3593. [Google Scholar] [CrossRef]
- Chen, X.; Miche, L.; Sachs, S.; Wang, Q.; Buschart, A.; Yang, H.; Vera Cruz, C.M.; Hurek, T.; Reinhold-Hurek, B. Rice responds to endophytic colonization which is independent of the common symbiotic signaling pathway. New Phytol. 2015, 208, 531–543. [Google Scholar] [CrossRef] [PubMed]
- Rollins, J.A.; Wolpert, T.J.; Johnson, K.B.; Dickman, M.B.; Ciuffetti, L.M.; Lorang, J.M.; Tuori, R.P.; Martinez, J.P.; Sawyer, T.L.; Redman, R.S. Green Fluorescent Protein Is Lighting Up Fungal Biology. Appl. Environ. Microbiol. 2001, 67, 1987–1994. [Google Scholar]
- Reinhold-Hurek, B.; Hurek, T. Living inside plants: Bacterial endophytes. Curr. Opin. Plant Biol. 2011, 14, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Germaine, K.; Franks, A.; Ryan, D.J.; Dowling, D.N. Bacterial endophytes: Recent developments and applications. FEMS Microbiol. Lett. 2008, 278, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Riedel, G.E.; Ausubel, F.M.; Cannon, F.C. Physical map of chromosomal nitrogen fixation (nif) genes of Klebsiella pneumoniae. Proc. Natl. Acad. Sci. USA 1979, 76, 2866–2870. [Google Scholar] [CrossRef] [PubMed]
- Cankar, K.; Kraigher, H.; Ravnikar, M.; Rupnik, M. Bacterial endophytes from seeds of Norway spruce (Picea abies L. Karst). FEMS Microbiol. Lett. 2005, 244, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Egener, T.; Hurek, T.; Reinhold-hurek, B.; Mikrobiologie, M.; Symbioseforschung, A. Endophytic Expression of nif Genes of Azoarcus sp. Strain BH72 in Rice Roots. Mol. Plant Microbe Interact. 1999, 12, 813–819. [Google Scholar] [CrossRef]
- Weilharter, A.; Mitter, B.; Shin, M.V.; Chain, P.S.G.; Nowak, J.; Sessitsch, A. Complete genome sequence of the plant growth-promoting endophyte Burkholderia phytofirmans strain PsJN. J. Bacteriol. 2011, 193, 3383–3384. [Google Scholar] [CrossRef] [PubMed]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; van Themaat, E.V.L.; Schulze-Lefert, P. Structure and Functions of the Bacterial Microbiota of Plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef] [PubMed]
- De Souza, R.S.C.; Okura, V.K.; Armanhi, J.S.L.; Jorrín, B.; Lozano, N.; da Silva, M.J.; González-Guerrero, M.; de Araújo, L.M.; Verza, N.C.; Bagheri, H.C.; et al. Unlocking the bacterial and fungal communities assemblages of sugarcane microbiome. Sci. Rep. 2016, 6, 28774. [Google Scholar] [CrossRef] [PubMed]
- Case, R.J.; Boucher, Y.; Dahllöf, I.; Holmström, C.; Doolittle, W.F.; Kjelleberg, S. Use of 16S rRNA and rpoB genes as molecular markers for microbial ecology studies. Appl. Environ. Microbiol. 2007, 73, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Kaul, S.; Sharma, T.; Dhar, M.K. “Omics” Tools for Better Understanding the Plant–Endophyte Interactions. Front. Plant Sci. 2016, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Isebrands, J.G.; Aronsson, P.; Carlson, M.; Ceulemans, R.; Coleman, M.; Dickinson, N.; Dimitriou, J.; Doty, S.; Gardiner, E.; Heinsoo, K.; et al. Environmental Applications of Poplars and Willows. In Poplars and willows: Trees for Society and the Environment; Isebrands, J.G., Richardson, J., Eds.; The Food and Agriculture Organization of the United Nations: Rome, Italy, 2014; pp. 258–336. ISBN 9781780641089. [Google Scholar]
- Kang, J.W.; Khan, Z.; Doty, S.L. Biodegradation of trichloroethylene by an endophyte of hybrid poplar. Appl. Environ. Microbiol. 2012, 78, 3504–3507. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L.; Dosher, M.R.; Singleton, G.L.; Moore, A.L.; Van Aken, B.; Stettler, R.F.; Strand, S.E.; Gordon, M.P. Identification of an endophytic Rhizobium in stems of Populus. Symbiosis 2005, 39, 27–35. [Google Scholar]
- Hacquard, S.; Schadt, C.W. Towards a holistic understanding of the beneficial interactions across the Populus microbiome. New Phytol. 2015, 205, 1424–1430. [Google Scholar] [CrossRef] [PubMed]
- Khan, Z.; Guelich, G.; Phan, H.; Redman, R.; Doty, S. Bacterial and Yeast Endophytes from Poplar and Willow Promote Growth in Crop Plants and Grasses. ISRN Agron. 2012, 1–11. [Google Scholar] [CrossRef]
- Kandel, S.L. Salicaceae Enophytes: Growth Promotion Potential in Rice and Maize and Bio-Control of Plant Pathogen. Ph.D. Thesis, University of Washington, Seattle, WA, USA, 2016. [Google Scholar]
- Cangelosi, G.A.; Abest, E.; Martinettii, G.; Nester, E.W. Genetic analysis of Agrobacterium. Methods Enzymol. 1991, 204, 384–397. [Google Scholar] [PubMed]
- Rho, H.; Hsieh, M.; Kandel, S.L.; Cantillo, J.; Doty, S.L.; Kim, S.H. Do Endophytes Promote Growth of Host Plants Under Stress? A Meta-Analysis on Plant Stress Mitigation by Endophytes. Microb. Ecol. 2017, 1–12. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Ye, Z.; Yang, D.; Yan, J.; Xiao, L.; Zhong, T.; Yuan, M.; Cai, X.; Fang, Z.; Jing, Y. Characterization of endophytic Rahnella sp. JN6 from Polygonum pubescens and its potential in promoting growth and Cd, Pb, Zn uptake by Brassica napus. Chemosphere 2013, 90, 1960–1965. [Google Scholar] [CrossRef] [PubMed]


| Endophyte Species | Native Host | Plant Colonized | Tissues Colonized | Effect on Plant | References |
|---|---|---|---|---|---|
| Acetobacter diazotrophicus | Sugarcane | Sugarcane | Stem | N/A | Dong et al., 1994 [27] |
| Achromobacter sp., and Acinetobacter sp. | Poaceae family (maize, wheat, pearl millet, sorghum and rice) | Wheat | Root | Growth enhancement | Patel et al., 2017 [125] |
| Azoarcus sp. | Kallar grass | Rice, and Kallar grass | Root, shoot | Growth enhancement | Hurek et al., 1994 [126] |
| Azoarcus sp. | Kallar grass | Rice | Root | N/A | Reinhold-Hurek et al., 2006 [88] |
| Azospirillum sp. | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Bacillus megaterium | Maize | Maize | Root, stem, leaf | N/A | Liu et al., 2006 [127] |
| Bacillus pumilus | Rice | Rice | Root | Growth enhancement | Bacilio-Jimenez et al., 2001 [128] |
| Bacillus sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Bacillus sp. | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Bacillus subtilis | Mulberry | Mulberry | Root, stem, leaf | Reduced bacterial wilt | Ji et al., 2008 [130] |
| Burkholderia cepacia | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Burkholderia phytofirmans | Onion | Grapevine | Root, stem, berry | N/A | Compant et al., 2008 [131] |
| Burkholderia phytofirmans | Onion | Grapevine | Root, stem, leaf | Growth enhancement | Compant et al., 2005 [81] |
| Burkholderia phytofirmans | Onion | Switchgrass | Root, leaf, sheath | Growth enhancement | Kim et al., 2012 [132] |
| Burkholderia phytofirmans | Onion | Arabidopsis thaliana | Root | Growth enhancement, increased chlorophyll content | Zuniga et al., 2013 [67] |
| Burkholderia phytofirmans | Onion | White lupin, and maize | Root, seed | N/A | Kost et al., 2014 [66] |
| Burkholderia phytofirmans | Onion | Maize | Root, stem, leaf | Growth enhancement, increased drought tolerance | Naveed et al., 2014 [60] |
| Burkholderia sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Burkholderia vietnamiensis | Poplar | Kentucky bluegrass | Root, shoot | Growth enhancement | Xin et al., 2009 [43] |
| Burkholderia vietnamiensis | Rice | Rice | Root | Growth enhancement | Govindarajan et al., 2008 [133] |
| Burkholderia vietnamiensis | Sugarcane | Sugarcane | Root | Growth enhancement, increased yield | Govindarajan et al., 2006 [134] |
| Corynebacterium flavescens | Rice | Rice | Root | Growth enhancement | Bacilio-Jimenez et al., 2001 [128] |
| Enterobacter sp. | Maize | Maize | Root, stem, leaf | Growth enhancement, increased drought tolerance | Naveed et al., 2014 [60] |
| Enterobacter sp. | Hybrid poplar (Populus deltoides × P. nigra) | Hybrid poplar | Root, leaf bud | Growth enhancement, reduced phytotoxicity of TCE, degradation of TCE | Doty et al., 2017 [135] |
| Enterobacter sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Enterobacter sp. | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Gluconacetobacter diazotrophicus | Sugarcane | Wheat, and sorghum | Root, shoot, stem, leaf | N/A | Luna et al., 2010 [136] |
| Gluconacetobacter diazotrophicus | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Gluconacetobacter diazotrophicus | Sugarcane | Rice | Root | N/A | Meneses et al., 2017 [76] |
| Gluconacetobacter diazotrophicus | Sugarcane | Sugarcane, and rice | Root, Shoot | N/A | Rouws et al., 2010 [137] |
| Herbaspirillum seropedicae | Maize | Maize | Root | N/A | Balsanelli et al., 2014 [77] |
| Herbaspirillum seropedicae | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Herbaspirillum seropedicae | Maize | Maize | Root | Increased rooting, change in gene expression | Amaral et al., 2014 [138] |
| Herbaspirillum seropedicae | Maize | Maize, wheat, rice and sorghum | Root, stem, leaf | N/A | Roncata-Maccari et al., 2003 [139] |
| Herbaspirillum seropedicae | Rice | Rice | Root, coleoptile, leaf | Growth enhancement | James et al., 2002 [99] |
| Herbaspirillum seropedicae | Sorghum | Maize | Root, leaf | N-fixation, change in metabolic profile | Brusamarello-Santos et al., 2017 [140] |
| Herbaspirillum seropedicae | Sorghum | Wheat | Root | Change in gene expression | Pankievicz et al., 2016 [87] |
| Herbaspirillum sp. | Rice (Oryza officianalis) | Rice (Oryza spp.) | Shoot, seed, leaf | Growth enhancement, N-fixation | Elbeltagy et al., 2001 [12] |
| Klebsiella pneumoniae | Maize | Alfalfa, Arabidopsis, wheat, and rice | Root, hypocotyl | N/A | Dong et al., 2003 [141] |
| Klebsiella pneumoniae | Maize | Wheat | Root | Growth enhancement, increased chlorophyll content, N-fixation | Iniguez et al. 2004 [14] |
| Klebsiella pneumoniae | Maize | Alfalfa | Root, hypocotyl | N/A | Dong et al., 2003 [141] |
| Klebsiella sp. | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Microbacterium sp. | Rape | Rape | Root | Growth enhancement, increased Pb uptake, root elongation, | Sheng et al., 2008 [142] |
| Ochrobactrum sp. | Rice | Rice | Root | N/A | Verma et al., 2004 [143] |
| Pantoea agglomerans | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Pantoea agglomerans | Rice | Rice | Root | N/A | Verma et al., 2001 [144] |
| Pantoea sp. | Rice | Rice | Root | N/A | Verma et al., 2004 [143] |
| Pseudomonas fluorescences | Miscanthus | Pea | N/A | Growth enhancement in phosphate limited conditions | Oteino et al. 2015 [48] |
| Pseudomonas fluorescens | Rape | Rape | Root | Growth enhancement, increased Pb uptake, root elongation, | Sheng et al., 2008 [142] |
| Pseudomonas fluorescens | Black nightshade | Black nightshade and tobacco | Root | Growth enhancement | Long et al., 2008 [145] |
| Pseudomonas fluorescens | Wheat | Tomato | Root | N/A | Duijff et al., 1997 [146] |
| Pseudomonas putida | Hybrid poplar | Willow | Root | Growth enhancement, reduced phytotoxicity of phenanthrene, degradation of phenanthrene | Khan et al., 2014 [147] |
| Pseudomonas putida | Potato | Potato | Root, stem | Growth enhancement, Phytophthora infestans suppression | Andreote et al., 2009 [148] |
| Pseudomonas putida | Poplar | Pea | Root, stem, leaf | Increased accumulation of and tolerance to 2,4-dichlorophenoxyacetic acid | Germaine et al., 2006 [149] |
| Pseudomonas sp. | Black nightshade | Black nightshade and tobacco | Root | Growth enhancement | Long et al., 2008 [145] |
| Pseudomonas sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Pseudomonas sp. | Poplar | Poplar | Root, stem, leaf | N/A | Germaine et al., 2004 [92] |
| Pseudomonas thivervalensis | Black nightshade | Black nightshade and tobacco | Root | Growth enhancement | Long et al., 2008 [145] |
| Ralstonia sp. | Poaceae family (maize, wheat, pearl millet, sorghum and rice) | Wheat | Root | Growth enhancement | Patel et al., 2017 [125] |
| Rhanella aquatilis | Sweet potato | Hybrid poplar | N/A | Increased rooting | Khan et al., 2009 [150] |
| Rhizobium sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Rhizobium sp. | Poaceae family (maize, wheat, pearl millet, sorghum and rice) | Wheat | Root | Growth enhancement | Patel et al., 2017 [125] |
| Rhizobium sp. | Maize | Maize | N/A | Growth enhancement | Riggs et al., 2001 [29] |
| Serratia marcescens | Rice | Rice | Root, stem, leaf | Growth enhancement | Gyaneshwar et al., 2001 [98] |
| Staphylococcus sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Stenotrophomonas sp. | Tomato | Wheat | N/A | Growth enhancement | Tian et al., 2017 [129] |
| Consortium (Gluconacetobacter diazotrophicus, Herbaspirillum seropedicae, Herbaspirillum rubrisubalbicans, Azospirillum amazonense and Burkholderia sp.) | Sugarcane | Sugarcane | Root, shoot | Growth enhancement, increased N content | Oliveira et al., 2002 [151] |
| Consortium (Burkholderia vietnamiensis, Rhanella sp., Acinetobacter sp., Herbaspirillum sp., Pseudomonas putida, Sphingomonas spp. | Poplar and willow | Sweet corn | Root, shoot | Growth enhancement, increased CO2 assimilation | Knoth et al., 2012 [19] |
| Consortium (Burkholderia vietnamiensis, Rhanella sp., Enterobacter sp., Pseudomonas graminis, Acinetobacter sp., Herbaspirillum sp., Sphingomonas yanoikuyae, Pseudomonas putida, Sphingomonas, Sphingomonas yanoikuyae) | Poplar and willow | Poplar and hybrid poplar | N/A | Growth enhancement | Knoth et al., 2014 [152] |
| Consortium (Burkholderia vietnamiensis, Rhizobium tropici, Acinetobacter calcoaceticus, Rhanella sp., Burkholderia sp., Enterobacter asburiae, Sphingomonas yanoikuyae, Pseudomonas sp., Curtobacterium sp.) | Poplar and willow | Hybrid poplar | N/A | Growth enhancement, increased drought tolerance | Khan et al., 2016 [45] |
| Consortium (Burkholderia vietnamiensis, Rhizobium tropici, Acinetobacter calcoaceticus, Rhanella sp., Burkholderia sp., Sphingomonas yanoikuyae, Pseudomonas sp., Sphingomonas sp.) | Poplar and willow | Rice | Root, shoot | Growth enhancement (N-limited conditions) | Kandel et al., 2015 [18] |
| Consortium (Gluconacetobacter diazotrophicus, Herbaspirillum seropedicae, Herbaspirillum rubrisubalbicans, Azospirillum amazonense, Burkholderia tropica) | Sugarcane | Sugarcane | Root | N/A | Oliveira et al., 2009 [153] |
| Consortium (Pseudomonas spp., Paentbacillus spp., Sphingomonas azotifigens) | Ryegrass and rice | Ryegrass | Root, stem, leaf | Growth enhancement, increased TFA | Castanheira et al., 2017 [95] |
| Consortium (Rhizobium tropici bv. populus, Acinetobacter calcoaceticus, Rhanella sp., Burkholderia sp., Sphingomonas spp.) | Poplar and willow | Douglas-fir | Root, needles | Growth enhancement (nutrient limited conditions) | Khan et al., 2015 [154] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kandel, S.L.; Joubert, P.M.; Doty, S.L. Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms 2017, 5, 77. https://doi.org/10.3390/microorganisms5040077
Kandel SL, Joubert PM, Doty SL. Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms. 2017; 5(4):77. https://doi.org/10.3390/microorganisms5040077
Chicago/Turabian StyleKandel, Shyam L., Pierre M. Joubert, and Sharon L. Doty. 2017. "Bacterial Endophyte Colonization and Distribution within Plants" Microorganisms 5, no. 4: 77. https://doi.org/10.3390/microorganisms5040077
APA StyleKandel, S. L., Joubert, P. M., & Doty, S. L. (2017). Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms, 5(4), 77. https://doi.org/10.3390/microorganisms5040077





