Melanisation of Aspergillus terreus—Is Butyrolactone I Involved in the Regulation of Both DOPA and DHN Types of Pigments in Submerged Culture?
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strain, Chemicals and Culture Conditions
Addition of Butyrolactone I
2.2. Gene Expression Analysis Using Microarrays
2.3. Strand-Specific Transcriptome Sequencing and Analysis
2.4. Data Availability
3. Results
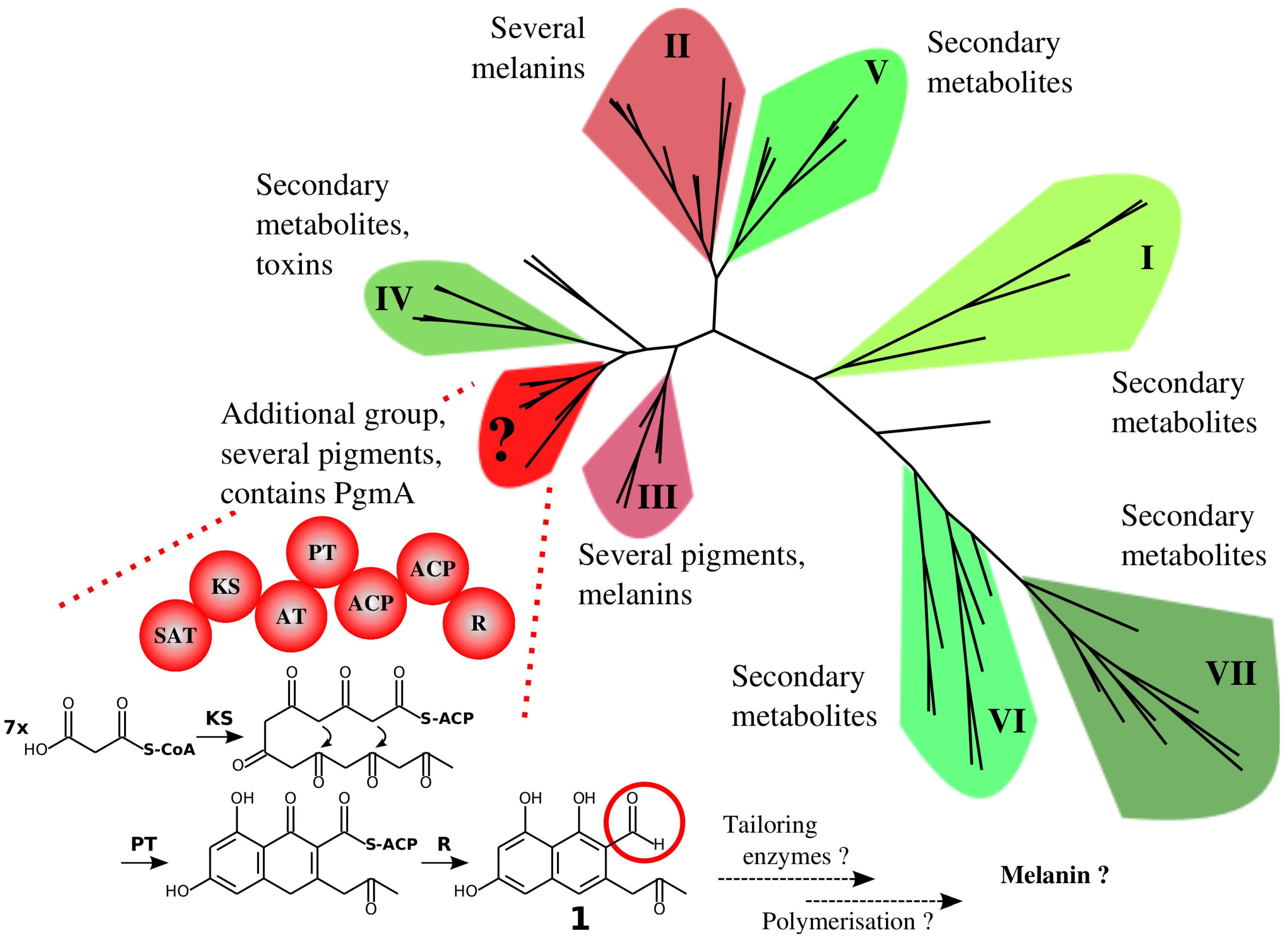
3.1. A Non-Canonical PKS Gene Cluster in A. terreus
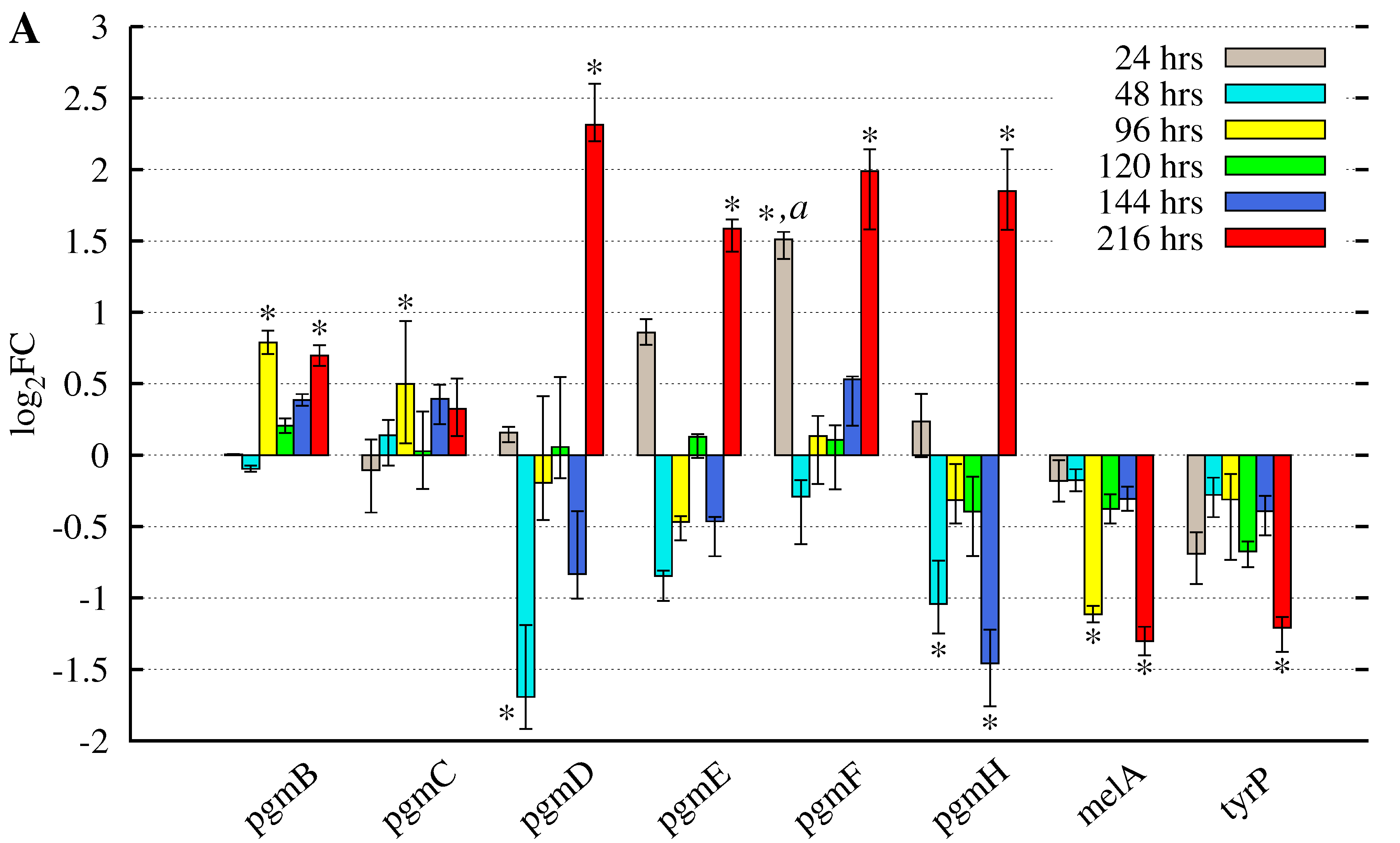
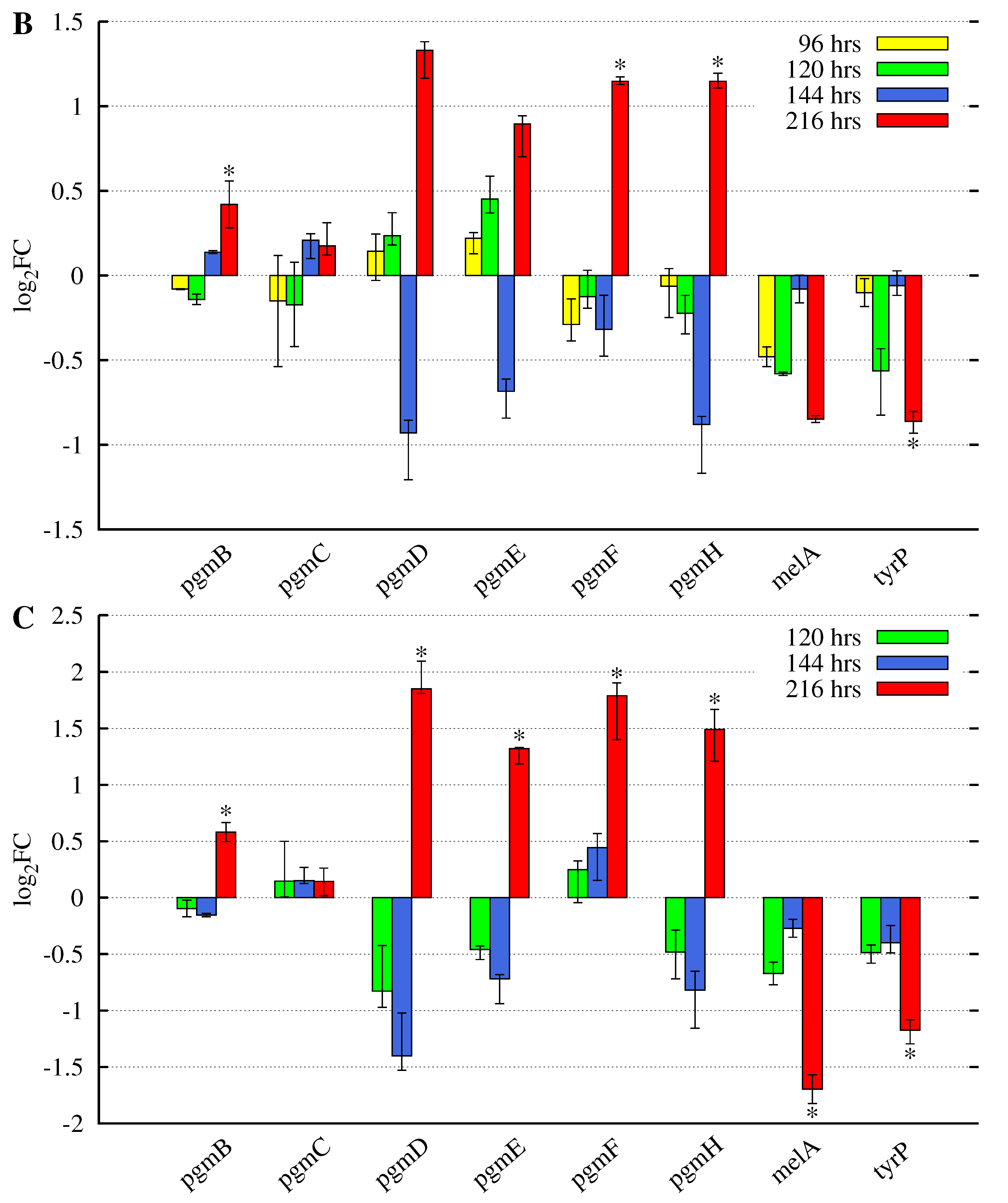
3.2. Butyrolactone I Addition Reveals Opposite Gene Expression Profiles for the Pgm Cluster Genes and DOPA-Type Melanin Biosynthesis Genes
4. Discussion
4.1. The Similarity of A. terreus Pgm Cluster with Pigment Biosynthesis Genes of Aspergilli and F. fujikuroi
4.2. Hypothesised A. terreus DHN-Type of Pigment and the Known DOPA-Type of Melanin May Be Biosynthesised under Different Growth Conditions
4.3. The Suggested DHN-Like Pigment of A. terreus May Be Related to Conidiophores as Well
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| DHN | 1,8-dihydroxynaphthalene |
| DOPA | 3,4-dihydroxyphenylalanine |
| FPKM | fragments per kilobase of exon per million reads mapped |
| ORF | open reading frame |
| p.i. | post inoculation |
| FC | fold change |
| PKS | polyketide synthase |
| SAT | starter unit: ACP transacylase |
| KS | Beta-ketoacyl synthase |
| AT | Acyl transferase |
| PT | Polyketide product template |
| ACP | Acyl carrier |
| R | Thioester reductase |
References
- Volling, K.; Thywissen, A.; Brakhage, A.A.; Saluz, H.P. Phagocytosis of melanized Aspergillus conidia by macrophages exerts cytoprotective effects by sustained PI3K/Akt signalling. Cell Microbiol. 2011, 13, 1130–1148. [Google Scholar] [CrossRef] [PubMed]
- Rosa, L.H.; de Lourdes Almeida Vieira, M.; Santiago, I.F.; Rosa, C.A. Endophytic fungi community associated with the dicotyledonous plant Colobanthus quitensis (Kunth) Bartl. (Caryophyllaceae) in Antarctica. FEMS Microbiol. Ecol. 2010, 73, 178. [Google Scholar] [CrossRef] [PubMed]
- Zalar, P.; Novak, M.; de Hoog, G.; Gunde-Cimerman, N. Dishwashers—A man-made ecological niche accommodating human opportunistic fungal pathogens. Fungal Biol. 2011, 115, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Lass-Flörl, C.; Kofler, G.; Kropshofer, G.; Hermans, J.; Kreczy, A.; Dierich, M.P.; Niederwieser, D. In-vitro testing of susceptibility to amphotericin B is a reliable predictor of clinical outcome in invasive aspergillosis. J. Antimicrob. Chemother. 1998, 42, 497. [Google Scholar] [CrossRef] [PubMed]
- Lass-Flörl, C.; Griff, K.; Mayr, A.; Petzer, A.; Gastl, G.; Bonatti, H.; Freund, M.; Kropshofer, G.; Dierich, M.P.; Nachbaur, D. Epidemiology and outcome of infections due to Aspergillus terreus: 10-Year single centre experience. Br. J. Haematol. 2005, 131, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Eisenman, H.C.; Casadevall, A. Synthesis and assembly of fungal melanin. Appl. Microbiol. Biotechnol. 2011, 93, 931–940. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, M.; Klich, M. The effects of tricyclazole, pyroquilon, phthalide, and related fungicides on the production of conidial wall pigments by Penicillium and Aspergillus species. Pestic. Biochem. Physiol. 1995, 52, 125–136. [Google Scholar] [CrossRef]
- Cabanes, J.; Chazarra, S.; Garcia-Carmona, F. Kojic acid, a cosmetic skin whitening agent, is a slow-binding inhibitor of catecholase activity of tyrosinase. J. Pharm. Pharmacol. 1994, 46, 982–985. [Google Scholar] [CrossRef] [PubMed]
- Espín, J.C.; Wichers, H.J. Slow-binding inhibition of mushroom (Agaricus bisporus) tyrosinase isoforms by tropolone. J. Agric. Food Chem. 1999, 47, 2638–2644. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.S. An updated review of tyrosinase inhibitors. Int. J. Mol. Sci. 2009, 10, 2440–2475. [Google Scholar] [CrossRef] [PubMed]
- Mayorga, M.E.; Timberlake, W.E. Isolation and molecular characterization of the Aspergillus nidulans wA gene. Genetics 1990, 126, 73–79. [Google Scholar] [PubMed]
- Cary, J.W.; Harris-Coward, P.Y.; Ehrlich, K.C.; Mavungu, J.D.D.; Malysheva, S.V.; Saeger, S.D.; Dowd, P.F.; Shantappa, S.; Martens, S.L.; Calvo, A.M. Functional characterization of a veA-dependent polyketide synthase gene in Aspergillus flavus necessary for the synthesis of asparasone, a sclerotium-specific pigment. Fungal Genet. Biol. 2014, 64, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, R.C.R.; Lisboa, H.C.F.; Pombeiro-Sponchiado, S.R. Characterization of melanin pigment produced by Aspergillus nidulans. World J. Microbiol. Biotechnol. 2011, 28, 1467–1474. [Google Scholar] [CrossRef] [PubMed]
- Pal, A.K.; Gajjar, D.U.; Vasavada, A.R. DOPA and DHN pathway orchestrate melanin synthesis in Aspergillus species. Med. Mycol. 2014, 52, 10–18. [Google Scholar] [PubMed]
- Zaehle, C.; Gressler, M.; Shelest, E.; Geib, E.; Hertweck, C.; Brock, M. Terrein biosynthesis in Aspergillus terreus and its impact on phytotoxicity. Chem. Biol. 2014, 21, 719–731. [Google Scholar] [CrossRef] [PubMed]
- Gressler, M.; Meyer, F.; Heine, D.; Hortschansky, P.; Hertweck, C.; Brock, M. Phytotoxin production in Aspergillus terreus is regulated by independent environmental signals. eLife 2015, 4, e07861. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.J.; Knox, B.P.; Sanchez, J.F.; Chiang, Y.M.; Bruno, K.S.; Wang, C.C.C. Application of an efficient gene targeting system linking secondary metabolites to their biosynthetic genes in Aspergillus terreus. Org. Lett. 2013, 15, 3562–3565. [Google Scholar] [CrossRef] [PubMed]
- Geib, E.; Gressler, M.; Viediernikova, I.; Hillmann, F.; Jacobsen, I.D.; Nietzsche, S.; Hertweck, C.; Brock, M. A non-canonical melanin biosynthesis pathway protects Aspergillus terreus conidia from environmental stress. Cell Chem. Biol. 2016, 23, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Schimmel, T.G.; Coffman, A.D.; Parsons, S.J. Effect of butyrolactone I on the producing fungus, Aspergillus terreus. Appl. Environ. Microbiol. 1998, 64, 3707–3712. [Google Scholar] [PubMed]
- Raina, S.; de Vizio, D.; Palonen, E.K.; Odell, M.; Brandt, A.M.; Soini, J.T.; Keshavarz, T. Is quorum sensing involved in lovastatin production in the filamentous fungus Aspergillus terreus? Process. Biochem. 2012, 47, 843–852. [Google Scholar] [CrossRef]
- Palonen, E.K.; Neffling, M.R.; Raina, S.; Brandt, A.; Keshavarz, T.; Meriluoto, J.; Soini, J. Butyrolactone I quantification from lovastatin producing Aspergillus terreus using tandem mass spectrometry—Evidence of signalling functions. Microorganisms 2014, 2, 111–127. [Google Scholar] [CrossRef] [PubMed]
- Palonen, E.K.; Raina, S.; Brandt, A.; Meriluoto, J.; Keshavarz, T.; Soini, J.T. Transcriptomic complexity of Aspergillus terreus velvet gene family under the influence of butyrolactone I. Microorganisms 2017, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Birren, B.; Lander, E.; Galagan, J.; Nusbaum, C.; Devon, K.; Henn, M.; Ma, L.J.; Jaffe, D.; Butler, J.; Alvarez, P.; et al. Annotation of the Aspergillus terreus NIH2624 Genome. Available online: http://www.broadinstitute.org/ftp/pub/annotation/fungi/aspergillus/genomes/ (accessed on 9 March 2007).
- Nucleotide-Nucleotide BLAST Version 2.2.29+. Available online: ftp://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/2.2.29/ (accessed on 10 October 2014).
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Marioni, J.C.; Mason, C.E.; Mane, S.M.; Stephens, M.; Gilad, Y. RNA-seq: An assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 2008, 18, 1509–1517. [Google Scholar] [CrossRef] [PubMed]
- Parkhomchuk, D.; Borodina, T.; Amstislavskiy, V.; Banaru, M.; Hallen, L.; Krobitsch, S.; Lehrach, H.; Soldatov, A. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 2009, 37, e123. [Google Scholar] [CrossRef] [PubMed]
- Levin, J.Z.; Yassour, M.; Adiconis, X.; Nusbaum, C.; Thompson, D.A.; Friedman, N.; Gnirke, A.; Regev, A. Comprehensive comparative analysis of strand-specific RNA sequencing methods. Nat. Methods 2010, 7, 709–715. [Google Scholar] [CrossRef] [PubMed]
- Integrative Genomics Viewer Software. Version IGV_2.3.26. Available online: http://software.broadinstitute.org/software/igv/ (accessed on 31 January 2014).
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative genomics viewer (IGV): High-performance genomics data visualization and exploration. Brief Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- The GENSCAN Web Server at MIT. Available online: http://genes.mit.edu/GENSCAN.html (accessed on 1 January 2015).
- ExPASy—Translate Tool. Available online: http://web.expasy.org/translate/ (accessed on 1 January 2015).
- InterProScan Sequence Search. Available online: http://www.ebi.ac.uk/interpro/search/sequence-search/ (accessed on 1 January 2015).
- Basic Local Alignment Tool, NCBI BLAST Sequence Database. Available online: http://blast.ncbi.nlm.nih.gov/ (accessed on 1 January 2015).
- Studt, L.; Wiemann, P.; Kleigrewe, K.; Humpf, H.U.; Tudzynski, B. Biosynthesis of fusarubins accounts for pigmentation of Fusarium fujikuroi perithecia. Appl. Environ. Microbiol. 2012, 78, 4468–4480. [Google Scholar] [CrossRef] [PubMed]
- Awakawa, T.; Kaji, T.; Wakimoto, T.; Abe, I. A heptaketide naphthaldehyde produced by a polyketide synthase from Nectria haematococca. Bioorg. Med. Chem. Lett. 2012, 22, 4338–4340. [Google Scholar] [CrossRef] [PubMed]
- Mulder, N.J.; Apweiler, R.; Attwood, T.K.; Bairoch, A.; Bateman, A.; Binns, D.; Bork, P.; Buillard, V.; Cerutti, L.; Copley, R.; et al. New developments in the InterPro database. Nucleic Acids Res. 2007, 35, D224–D228. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Wheeler, M.H.; Chang, Y.C.; Kwon-Chung, K.J. A developmentally regulated gene cluster involved in conidial pigment biosynthesis in Aspergillus fumigatus. J. Bacteriol. 1999, 181, 6469–6477. [Google Scholar] [PubMed]
- Tsai, H.F.; Fujii, I.; Watanabe, A.; Wheeler, M.H.; Chang, Y.C.; Yasuoka, Y.; Ebizuka, Y.; Kwon-Chung, K.J. Pentaketide melanin biosynthesis in Aspergillus fumigatus requires chain-length shortening of a heptaketide precursor. J. Biol. Chem. 2001, 276, 29292–29298. [Google Scholar] [CrossRef] [PubMed]
- Fujii, I.; Yasuoka, Y.; Tsai, H.F.; Chang, Y.C.; Kwon-Chung, K.J.; Ebizuka, Y. Hydrolytic polyketide shortening by Ayg1p, a novel enzyme involved in fungal melanin biosynthesis. J. Biol. Chem. 2004, 279, 44613–44620. [Google Scholar] [CrossRef] [PubMed]
- Keller, S.; Macheleidt, J.; Scherlach, K.; Schmaler-Ripcke, J.; Jacobsen, I.D.; Heinekamp, T.; Brakhage, A.A. Pyomelanin formation in Aspergillus fumigatus requires HmgX and the transcriptional activator HmgR but is dispensable for virulence. PLoS ONE 2011, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ahuja, M.; Chiang, Y.M.; Chang, S.L.; Praseuth, M.B.; Entwistle, R.; Sanchez, J.F.; Lo, H.C.; Yeh, H.H.; Oakley, B.R.; Wang, C.C.C. Illuminating the diversity of aromatic polyketide synthases in Aspergillus nidulans. J. Am. Chem. Soc. 2012, 134, 8212–8221. [Google Scholar] [CrossRef] [PubMed]
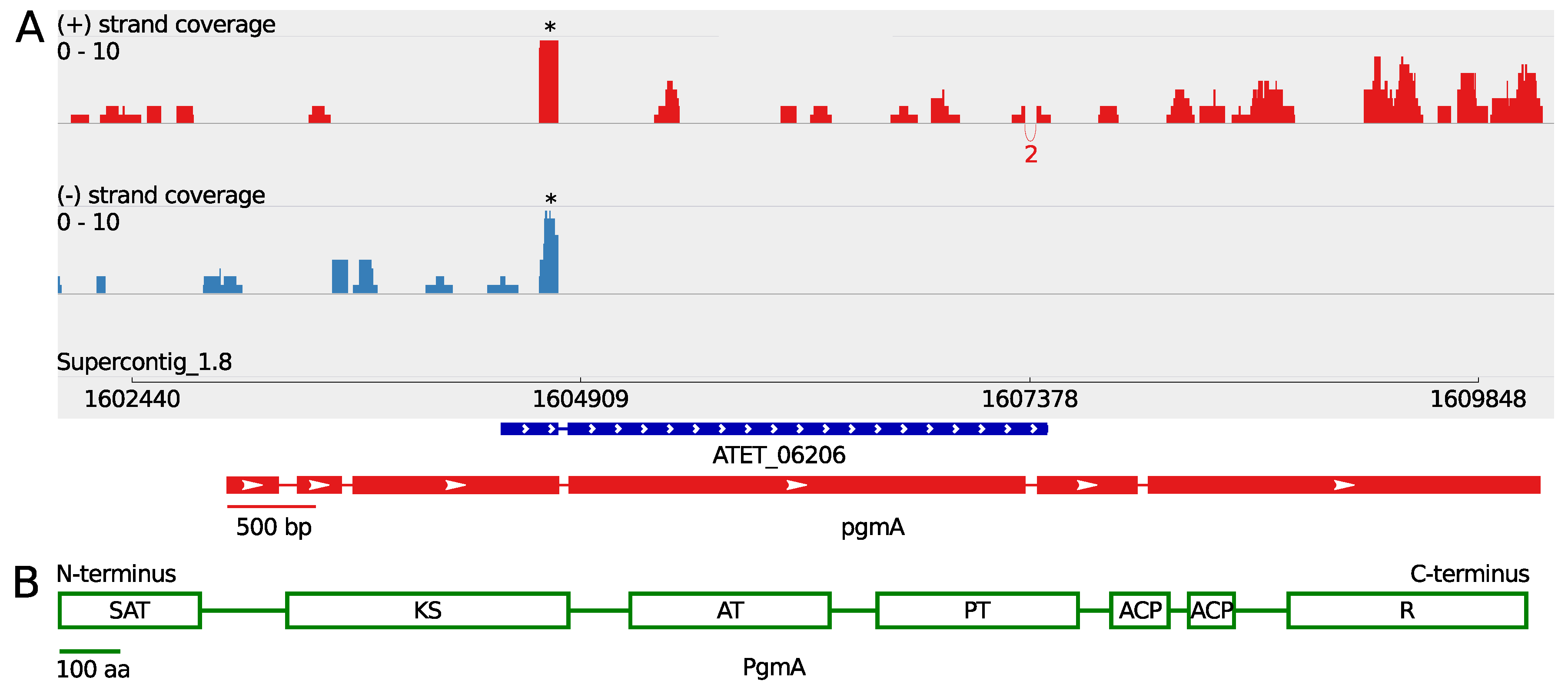

| Gene | Gene Length (bp) | ORF Length (bp) | Pooled FPKM | Coverage Max. Sense Strand |
|---|---|---|---|---|
| pgmB | 1448 | 1329 | 3.9 | 23 |
| pgmC | 1443 | 1443 | 1.1 | 6 |
| pgmR | 1284 e | 1284 e | 0.13 | 2 |
| pgmA | 7232 e | 6897 e | 0.65 | 8 |
| pgmD | 1003 | 939 | 4.7 | 44 |
| pgmE | 1008 | 1008 | 15 | 91 |
| pgmF | 1071 | 1071 | 290 | 1634 |
| pgmG | 1967 | 1683 | 56 | 333 |
| pgmH | 1714 | 1479 | 12 | 81 |
| melA | 2779 | 2779 | 6.0 | 38 |
| tyrP | 1230 | 1071 | 1.3 | 14 |
| Gene | Predicted Molecular Function | InterPRO Annotation | Domains | ID % | Fusarubin Gene |
|---|---|---|---|---|---|
| PgmB | O-methyltransferase | IPR016461 | O-methyltransferase, COMT-type | 41 | Fsr2 |
| IPR011991 | |||||
| IPR029063 | |||||
| IPR001077 | |||||
| PgmC | Cytochrome p450 monooxygenase | IPR001128 | Cytochrome P450 E-class, group 1 | NA e | Fsr3 |
| IPR002401 | |||||
| IPR017972 | |||||
| PgmR | Aflatoxin biosynthesis regulatory protein-like | IPR002409 | Aflatoxin biosynthesis regulatory protein | 31 | Fsr6 |
| IPR001138 d | |||||
| PgmA | nonreducing polyketide synthase; NR-PKS | IPR032088 IPR020841 d IPR020801 d IPR030918 d IPR009081 d IPR013120 d | Starter unit:ACP transacylase Beta-ketoacyl synthase Acyl transferase Polyketide product template Acyl carrier protein-like Thioester reductase-like | 59 | Fsr1 |
| PgmD | short-chain | IPR002347 | Short-chain | NA e | Fsr5 |
| dehydrogenase/reductase | IPR016040 | dehydrogenase/reductase | |||
| PgmE | SAM-dependent | IPR029063 | SAM-dependent | NA | NA |
| methyltransferase | methyltransferase | ||||
| PgmF | quinone reductase | IPR002085 IPR011032 IPR020843 IPR013154 IPR016040 | Alcohol dehydrogenase GroES-like Enoylreductase domain Alcohol dehydrogenase | 26 | Fsr4 |
| PgmG | MFS family permease | IPR011701 | Major facilitator superfamily | NA e | NA e |
| IPR020846 | |||||
| PgmH | FAD/FMN-binding CO dehydrogenase | IPR016167 | FAD-binding, type 2 | NA e | NA e |
| IPR016166 | FAD linked oxidase | ||||
| IPR006094 | CO dehydrogenase flavoprotein- | ||||
| IPR016169 | like, FAD-binding, subdomain 2 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Palonen, E.K.; Raina, S.; Brandt, A.; Meriluoto, J.; Keshavarz, T.; Soini, J.T. Melanisation of Aspergillus terreus—Is Butyrolactone I Involved in the Regulation of Both DOPA and DHN Types of Pigments in Submerged Culture? Microorganisms 2017, 5, 22. https://doi.org/10.3390/microorganisms5020022
Palonen EK, Raina S, Brandt A, Meriluoto J, Keshavarz T, Soini JT. Melanisation of Aspergillus terreus—Is Butyrolactone I Involved in the Regulation of Both DOPA and DHN Types of Pigments in Submerged Culture? Microorganisms. 2017; 5(2):22. https://doi.org/10.3390/microorganisms5020022
Chicago/Turabian StylePalonen, Elina K., Sheetal Raina, Annika Brandt, Jussi Meriluoto, Tajalli Keshavarz, and Juhani T. Soini. 2017. "Melanisation of Aspergillus terreus—Is Butyrolactone I Involved in the Regulation of Both DOPA and DHN Types of Pigments in Submerged Culture?" Microorganisms 5, no. 2: 22. https://doi.org/10.3390/microorganisms5020022
APA StylePalonen, E. K., Raina, S., Brandt, A., Meriluoto, J., Keshavarz, T., & Soini, J. T. (2017). Melanisation of Aspergillus terreus—Is Butyrolactone I Involved in the Regulation of Both DOPA and DHN Types of Pigments in Submerged Culture? Microorganisms, 5(2), 22. https://doi.org/10.3390/microorganisms5020022







