Lead Discovery Strategies for Identification of Chlamydia pneumoniae Inhibitors
Abstract
:1. Introduction
2. Lead Discovery Strategies
2.1. Epidemiological/Ethnopharmacological Approach
2.2. Target-Based Virtual Screening
2.3. Ligand-Based Virtual Screening
2.4. Pharmacophore-Based Design
3. Plant Phenolics as Antichlamydial Agents
3.1. Oxidative Stress
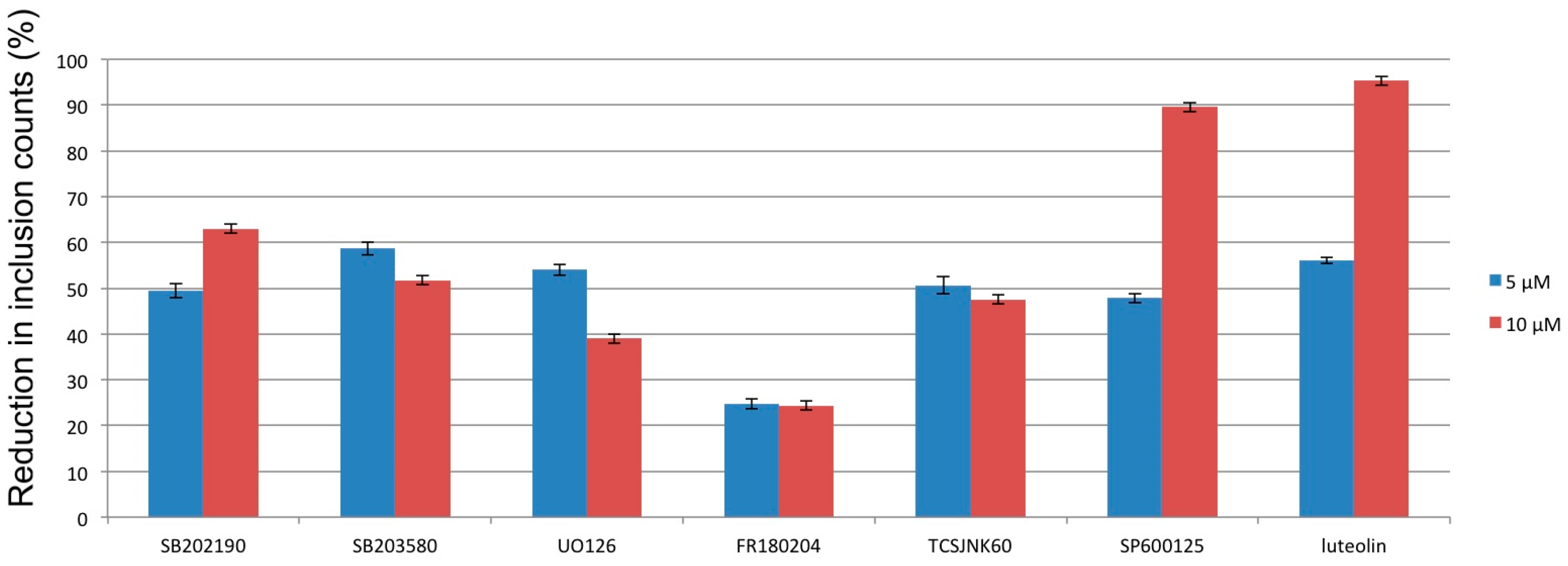
3.2. Mitogen-Activated Protein Kinases
3.3. Intracellular Calcium Levels
4. Models of Persistent Infection
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Saikku, P.; Wang, S.P.; Kleemola, M.; Brander, E.; Rusanen, E.; Grayston, J.T. An Epidemic of Mild Pneumonia due to an Unusual Strain of Chlamydia psittaci. J. Infect. Dis. 1985, 151, 832–839. [Google Scholar] [CrossRef] [PubMed]
- Grayston, J.; Kuo, C.; Want, S.; Altman, J. A New Chlamydia psittaci Strain, TWAR, Isolated in Acute Respiratory Tract Infections. N. Engl. J. Med. 1987, 315, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Heinzen, R.; Hackstadt, T. The Chlamydia trachomatis Parasitophorous Vacuolar Membrane is not Passively Permeable to Low-Molecular-Weight Compounds. Infect. Immun. 1997, 65, 1088–1094. [Google Scholar] [PubMed]
- Roblin, P.M.; Hammerschlag, M.R. Microbiologic Eefficacy of Aazithromycin and Ssusceptibilities to Aazithromycin of Iisolates of Chlamydia pneumoniae from Adults and Children with Community-Acquired Pneumonia. Antimicrob. Agents Chemother. 1998, 42, 194–196. [Google Scholar] [PubMed]
- Kohlhoff, S.A.; Hammerschlag, M.R. Treatment of Chlamydial Infections: 2014 Update. Expert Opin. Pharmacother. 2015, 16, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Borel, N.; Leonard, C.; Slade, J.; Schoborg, R.V. Chlamydial Antibiotic Resistance and Treatment Failure in Veterinary and Human Medicine. Curr. Clin. Microbiol. Rep. 2016, 3, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Hanski, L.; Vuorela, P.M. Recent Advances in Technologies for Developing Drugs against Chlamydia pneumoniae. Expert Opin. Drug Discov. 2014, 9, 791–802. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural Products as Sources of New Drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Schriever, C.; Pendland, S.L.; Mahady, G.B. Red Wine, Resveratrol, Chlamydia pneumoniae and the French Connection. Atherosclerosis 2003, 171, 379–380. [Google Scholar] [CrossRef] [PubMed]
- Vuorela, P.M.; Leinonen, M.; Saikku, P.; Tammela, P.; Rauha, J.P.; Wennberg, T.; Vuorela, H. Natural Products in the Process of Finding New Drug Candidates. Curr. Med. Chem. 2004, 11, 1375–1389. [Google Scholar] [CrossRef]
- Salin, O.; Törmäkangas, L.; Leinonen, M.; Saario, E.; Hagström, M.; Ketola, R.; Saikku, P.; Vuorela, H.; Vuorela, P. Corn Mint (Mentha arvensis) Extract Decreases Acute Chlamydia pneumoniae Infection in vitro and in vivo. J. Agric. Food Chem. 2011, 59, 12836–12842. [Google Scholar] [CrossRef] [PubMed]
- Alvesalo, J.; Vuorela, H.; Tammela, P.; Leinonen, M.; Saikku, P.; Vuorela, P. Inhibitory Effect of Dietary Phenolic Compounds on Chlamydia pneumoniae in Cell Cultures. Biochem. Pharmacol. 2006, 71, 735–741. [Google Scholar] [CrossRef] [PubMed]
- Törmäkangas, L.; Vuorela, P.; Saario, E.; Leinonen, M.; Saikku, P.; Vuorela, H. In vivo Treatment of Acute Chlamydia pneumoniae Infection with the Flavonoids Quercetin and Luteolin and an Alkyl Gallate, Octyl Gallate, in a Mouse Model. Biochem. Pharmacol. 2005, 70, 1222–1230. [Google Scholar] [CrossRef] [PubMed]
- Hakala, E.; Hanski, L.; Uvell, H.; Yrjönen, T.; Vuorela, H.; Elofsson, M.; Vuorela, P. Dibenzocyclooctadiene Lignans from Schisandra. spp. Selectively Inhibit the Growth of the Intracellular Bacteria Chlamydia pneumoniae and Chlamydia trachomatis. J. Antibiot. 2015, 68, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Chinese Pharmacopoeia Comission. Pharmacopeia of the People’s Republic of China, English edition, 9th ed.; U.S. Pharmacopoeial Convention: Rockville, MD, USA, 2010. [Google Scholar]
- Hakala, E.; Hanski, L.; Yrjönen, T.; Vuorela, H.; Vuorela, P. The Lignan-Containing Extract of Schisandra chinensis Berries Inhibits the Growth of Chlamydia pneumoniae. Nat. Prod. Commun. 2016, 10, 1001–1004. [Google Scholar]
- Heuer, D.; Brinkmann, V.; Meyer, T.F.; Szczepek, A.J. Expression and Translocation of Chlamydial Protease during Acute and Persistent Infection of the Epithelial HEp-2 Cells with Chlamydophila (Chlamydia) pneumoniae. Cell. Microbiol. 2003, 5, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Christian, J.G.; Heymann, J.; Paschen, S.A.; Vier, J.; Schauenburg, L.; Wupp, J.; Meyer, T.F.; Häcker, G.; Heuer, D. Targeting of a Chlamydial Protease Impedes Intracellular Bacterial Growth. PLoS Pathog. 2011, 7, e1002283. [Google Scholar] [CrossRef] [PubMed]
- Christian, J.; Vier, J.; Paschen, S.A.; Häcker, G. Cleavage of the NF-κB Family Protein p65/RelA by the Chlamydial Protease-like Activity Factor (CPAF) Impairs Proinflammatory Signaling in Cells Infected with Chlamydiae. J. Biol. Chem. 2010, 53, 41320–41327. [Google Scholar] [CrossRef] [PubMed]
- Savijoki, K.; Alvesalo, J.; Vuorela, P.; Leinonen, M.; Kalkkinen, N. Proteomic Analysis of Chlamydia pneumoniae-infected HL Cells Reveals Extensive Degradation of Cytoskeletal Proteins. FEMS Immunol. Med. Microbiol. 2008, 54, 375–384. [Google Scholar] [CrossRef] [PubMed]
- Alvesalo, J.; Siiskonen, A.; Vainio, M.; Tammela, P.; Vuorela, P. Similarity Based Virtual Screening: A Tool for Targeted Library Design. J. Med. Chem. 2006, 49, 2353–2356. [Google Scholar] [CrossRef] [PubMed]
- Keurulainen, L.; Salin, O.; Siiskonen, A.; Kern, J.M.; Maas, M.; Yli-Kauhaluoma, J.; Vuorela, P. Design and Synthesis of 2-Arylbenzimidazoles and Evaluation of Their Inhibitory Effect Against Chlamydia pneumoniae. J. Med. Chem. 2010, 53, 7664–7674. [Google Scholar] [CrossRef] [PubMed]
- Siiskonen, A.; Keurulainen, L.; Salin, O.; Kiuru, P.; Pohjala, L.; Vuorela, P.; Yli-Kauhaluoma, J. Conformation Study of 2-Arylbenzimidazoles as Inhibitors of Chlamydia pneumoniae Growth. Bioorg. Med. Chem. Lett. 2012, 22, 4882–4886. [Google Scholar] [CrossRef] [PubMed]
- Chopra, I. The 2012 Garrod Lecture: Discovery of Antibacterial Drugs in the 21st Century. J. Antimicrob. Chemot. 2012, 68, 496–505. [Google Scholar] [CrossRef] [PubMed]
- Karhu, E.; Isojärvi, J.; Vuorela, P.; Hanski, L.; Fallarero, A. Identification and Characterization of Privileged Antichlamydial Structures Using a Novel Ligand-based Strategy. J. Nat. Prod. 2016. submitted for publication. [Google Scholar]
- Hanski, L.; Ausbacher, D.; Tiirola, T.; Strøm, M.B.; Vuorela, P.M. Amphipathic β2,2-Amino Acid Derivatives Suppress Infectivity and Disrupt the Intracellular Replication Cycle of Chlamydia pneumoniae. PLoS ONE 2016, 11, e0157306. [Google Scholar]
- Donati, M.; Di Leo, K.; Benincasa, M.; Cavrini, F.; Accardo, S.; Moroni, A.; Gennaro, R.; Cevenini, R. Activity of Cathelicidin Peptides against Chlamydia spp. Antimicrob. Agents Chemoth. 2005, 49, 1201–1202. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.; Ausbacher, D.; Zachariassen, Z.G.; Anderssen, T.; Havelkova, M.; Strøm, M.B. Anticancer Activity of Small Amphipathic β2,2-Amino Acid Derivatives. Eur. J. Med. Chem. 2012, 58, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Hanski, L.; Genina, N.; Uvell, H.; Malinobskaha, K.; Gylfe, Å.; Laaksonen, T.; Kolakovicc, R.; Mäkilä, E.; Salonen, J.; Hirvonen, J.; et al. Inhibitory Activity of the Isoflavone Biochanin A on Intracellular Bacteria of Genus Chlamydia and Initial Development of a Buccal Formulation. PLoS ONE 2014, 9, e115115. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Lazaro, M. Distribution and Biological Activities of the Flavonoid Luteolin. Mini Rev. Med. Chem. 2009, 9, 31–59. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Shi, R.; Wang, X.; Shen, H.M. Luteolin, a Flavonoid with Potential for Cancer Prevention and Therapy. Curr. Cancer Drug Targets 2008, 8, 634–646. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Li, D.; Jiang, D. Targeting Cell Signaling and Apoptotic Pathways by Luteolin: Cardioprotective Role in Rat Cardiomyocytes Following Ischemia/Reperfusion. Nutrients 2012, 4, 2008–2019. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.B.; Moon, K.A.; Lee, K.Y.; Park, C.S.; Bae, Y.J.; Moon, H.B.; Cho, Y.S. Chlamydophila pneumoniae Triggers Release of CCL20 and Vascular Endothelial Growth Factor from Human Bronchial Epithelial Cells Through Enhanced Intracellular Oxidative Stress and MAPK Activation. J. Clin. Immunol. 2009, 29, 629–636. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, M.; Filardo, S.; De Santis, F.; Sessa, R. Chlamydia pneumoniae Infection in Atherosclerotic Lesion Development through Oxidative Stress: A Brief Overview. Int. J. Mol. Sci. 2013, 14, 15105–15120. [Google Scholar] [CrossRef] [PubMed]
- Azenabor, A.A.; Chaudhry, A.U. Effective Macrophage Redox Defense against Chlamydia pneumoniae Depends on l-type Ca2+ Channel Activation. Med. Microbiol. Immunol. 2003, 192, 99–106. [Google Scholar] [PubMed]
- Krill, M.; Kramp, J.; Petrov, T.; Klicken, A.C.; Hocke, A.C.; Walter, C.; Schmeck, B.; Seibold, J.; Maas, M.; Ludwig, S.; Kuipers, J.G.; Suttorp, N.; Hippenstiel, S. Differences in Cell Activation by Chlamydophila pneumoniae and Chlamydia trachomatis Infection in Human Endothelial Cells. Infect. Immun. 2004, 2, 6615–6621. [Google Scholar] [CrossRef] [PubMed]
- Haralambieva, I.H.; Iankov, I.D.; Ivanova, P.V.; Mitev, V.; Mitov, I.G. Chlamydophila pneumoniae Induces p44/p42 Mitogen-activated Protein Kinase Activation in Human Fibroblasts through Toll-like Receptor 4. J. Med. Microbiol. 2004, 53, 1187–1193. [Google Scholar] [CrossRef] [PubMed]
- Lego, D.; Bianco, M.; Quattrini, A.; Mancini, F.; Carollo, M.; Schiavoni, I.; Ciervo, A.; Ausiello, C.M.; Fedele, G. Chlamydia pneumoniae Modulates Human Monocyte-derived Dendritic Cells Functions Driving the Induction of a Type 1/Type 17 Inflammatory Response. Microbes Infect. 2013, 15, 105–114. [Google Scholar]
- Velma, S.A.; Krings, G.; Lopes-Virella, M.F. Chlamydophila pneumoniae Induces ICAM-1 Expression in Human Aortic Endothelial Cells via Protein Kinase C-Dependent Activation of Nuclear Factor-kappaB. Circ. Res. 2003, 92, 1130–1137. [Google Scholar] [CrossRef] [PubMed]
- Bea, F.; Puolakkainen, M.H.; McMillen, T.; Hudson, F.N.; Hackmann, N.; Kuo, C.C.; Campbell, L.A.; Rosenfeld, M.E. Chlamydia pneumoniae Induces Tissue Factor Expression in Mouse Macrophages via Activation of Egr-1 and the MEK-ERK1/2 Pathway. Circ. Res. 2003, 92, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Cheng, B.; Wu, X.; Sun, S.; Wu, Q.; Mei, C.; Xu, Q.; Wu, J.; He, P. MAPK-PPARα/γ Signal Transduction Pathways Are Involved in Chlamydia pneumoniae-Induced Macrophage-Derived Foam Cell Formation. Microb. Pathog. 2014, 69–70, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sasu, S.; LaVerda, D.; Qureshi, N.; Golenbock, D.T.; Beasley, D. Chlamydia pneumoniae and Chlamydial Heat Shock Protein 60 Stimulate Proliferation of Human Vascular Smooth Muscle Cells via Toll-like Receptor 4 and p44/p42 Mitogen-Activated Protein Kinase Activation. Circ. Res. 2001, 89, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Kern, J.M.; Maass, V.; Rupp, J.; Maass, M. Proliferative Stimulation of the Vascular Endothelin-1 Axis in vitro and ex vivo by Infection with Chlamydia pneumoniae. Thromb. Haemost. 2009, 102, 743–753. [Google Scholar] [CrossRef] [PubMed]
- Coombes, B.K.; Mahony, J.B. Identification of MEK- and Phosphoinositide 3-Kinase-dependent Signalling as Essential Events during Chlamydia pneumoniae Invasion of HEp2 Cells. Cell. Microbiol. 2002, 4, 447–460. [Google Scholar] [CrossRef] [PubMed]
- Mölleken, K.; Becker, E.; Hegemann, J.H. The Chlamydia pneumoniae Invasin Protein Pmp21 Recruits the EGF Receptor for Host Cell Entry. PLoS Pathog. 2013, 9, e1003325. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; McCarty, G.; Dong, F.; Hatch, G.M.; Pan, Z.K.; Zhong, G. Activation of Raf/MEK/ERK/cPLA2 Signaling Pathway is Essential for Chlamydial Acquisition of Host Glycerophospholipids. J. Biol. Chem. 2004, 279, 9409–9416. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Johnston, S.C.; Chou, J.; Dean, D. A Systemic Network for Chlamydia pneumoniae Entry into Human Cells. J. Bacteriol. 2010, 192, 2809–2815. [Google Scholar] [CrossRef] [PubMed]
- Azenabor, A.A.; Kennedy, P.; York, J. Free Intracellular Ca2+ Regulates Bacterial Lipopolysaccharide Induction of iNOS in Human Macrophages. Immunobiology 2009, 214, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Summanen, J.; Vuorela, P.; Rauha, J.; Tammela, P.; Marjamäki, K.; Pasternak, M.; Törnqvist, K.; Vuorela, H. Effects of Simple Aromatic Compounds and Flavonoids on Ca2+ Fluxes in Rat Pituitary GH4C1 Cells. Eur. J. Pharmacol. 2001, 414, 125–133. [Google Scholar] [CrossRef]
- Wu, S.N.; Chiang, H.T.; Shen, A.Y.; Lo, Y.K. Differential Effects of Quercetin, a Natural Polyphenolic Flavonoid, on L-type Calcium Current in Pituitary Tumor (GH3) Cells and Neuronal NG108–15 Cells. J. Cell. Physiol. 2003, 195, 298–308. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Liu, K.; Yi, J.; Zhu, D.; Liu, G.; Liu, B. Luteolin Inhibits Lysophosphatidylcholine-induced Apoptosis in Endothelial Cells by a Calcium/Mitocondrion/Caspases-dependent Pathway. Planta. Med. 2010, 76, 433–438. [Google Scholar]
- Serge, I.N.; Li, S.; Ho, C.T.; Rawson, N.E.; Dushenkov, S. Polymethoxyflavones Activate Ca2+-dependent Apoptotic Targets in Adipocytes. J. Agric. Food Chem. 2009, 57, 5771–5776. [Google Scholar] [CrossRef] [PubMed]
- Bagli, E.; Stefaniotou, M.; Morbidelli, L.; Ziche, M.; Psillas, K.; Murphy, C.; Fotsis, T. Luteolin Inhibits Vascular Endothelial Growth Factor-Induced Angiogenesis; Inhibition of Endothelial Cell Survival and Proliferation by Targeting Phosphatidylinositol 3′-Kinase Activity. Cancer Res. 2004, 64, 7936–7946. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, A.; Möller, S.; Bhattacharyya, A.; Behnen, M.; Rupp, J.; van Zandbergen, G.; Solbach, W.; Laskay, T. Mechanisms of Apoptosis Inhibition in Chlamydia pneumoniae-Infected Neutrophils. Int. J. Med. Microbiol. 2015, 305, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Verschooten, L.; Smaers, K.; van Kelst, S.; Proby, C.; Maes, D.; Declercq, L.; Agostinis, P.; Garmyn, M. The Flavonoid Luteolin Increases the Resistance of Normal, but not Malignant Keratinocytes, against UVB-induced Apoptosis. J. Investig. Dermatol. 2010, 130, 2277–2285. [Google Scholar] [CrossRef] [PubMed]
- Fischer, S.F.; Vier, J.; Kirschnek, S.; Klos, A.; Hess, S.; Ying, S.; Häcker, G. Chlamydia Inhibit Host Cell Apoptosis by Degradation of Proapoptotic BH3-Only Proteins. J. Exp. Med. 2004, 200, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Fischer, S.F.; Harlander, T.; Vier, J.; Häcker, G. Protection against CD95-Induced Apoptosis by Chlamydial Infection at a Mitochondrial Step. Infect. Immun. 2004, 72, 1107–1115. [Google Scholar] [CrossRef] [PubMed]
- Schöier, J.; Högdahl, M.; Söderlund, G.; Kihlström, E. Chlamydia (Chlamydophila) pneumonia-Induced Cell Death in Human Coronary Artery Endothelial Cells is Caspase-Independent and Accompanied by Subcellular Translocations of Bax and Apoptosis-Inducing Factor. FEMS Immunol. Med. Microbiol. 2006, 47, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Gieffers, J.; Füllgraf, H.; Jahn, J.; Klinger, M.; Dalhoff, K.; Katus, H.A.; Solbach, W.; Maass, M. Chlamydia pneumoniae Infection in Circulating Human Monocytes is Refractory to Antibiotic Treatment. Circulation 2001, 103, 351–356. [Google Scholar] [CrossRef] [PubMed]
- Haranaga, S.; Yamaguchi, H.; Ikejima, H.; Friedman, H.; Yamamoto, Y. Chlamydia pneumoniae infection of alveolar macrophages: A model. J. Infect. Dis. 2003, 187, 1107–1115. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.; Hess, S.; Endlich, K.; Thalmann, J.; Holzberg, D.; Kracht, M.; Schaefer, M.; Bartling, G.; Klos, A. Silencing or Permanent Activation: Host-cell Responses in Models of Persistent Chlamydia pneumoniae Infection. Cell. Microbiol. 2005, 7, 1099–1108. [Google Scholar] [CrossRef] [PubMed]
- Timms, P.; Good, D.; Wan, C.; Theodoropoulos, C.; Mukhopadhyay, S.; Summersgill, J.; Mathews, S. Differential transcriptional responses between the interferon-gamma-induction and iron-limitation models of persistence for Chlamydia pneumoniae. J. Microbiol. Immunol. Infect. 2009, 42, 27–37. [Google Scholar] [PubMed]
- Klos, A.; Thalmann, J.; Peters, J.; Gérard, H.C.; Hudson, A.P. The Transcript Profile of Persistent Chlamydophila. (Chlamydia) pneumoniae in vitro Depends on the Means by Which Persistence Is Induced. FEMS Microbiol. Lett. 2009, 291, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Kutlin, A.; Robin, P.M.; Hammerschlag, M.R. Effect of Prolonged Treatment with Azithromycin, Clarithromycin, or Levofloxacin on Chlamydia pneumoniae in a Continuous-Infection Model. Antimicrob. Agents Chemother. 2002, 46, 409–412. [Google Scholar] [CrossRef] [PubMed]
- Kutlin, A.; Flegg, C.; Stenzel, D.; Reznik, T.; Roblin, P.M.; Mathews, S.; Timms, P.; Hammerschlag, M.R. Ultrastructural Study of Chlamydia pneumoniae in a Continuous-Infection Model. J. Clin. Microbiol. 2001, 39, 3721–3723. [Google Scholar] [CrossRef] [PubMed]
- Silver, L. Multi-Targeting by Monotherapeutic Antibacterials. Nat. Rev. Drug Discov. 2007, 6, 41–55. [Google Scholar] [CrossRef] [PubMed]
- Emmet O’Brien, M.; Restrepo, M.I.; Martin-Loeches, I. Update on the Combination Effect of Macrolide Antibiotics in Community-Acquired Pneumonia. Respir. Investig. 2015, 53, 201–209. [Google Scholar] [CrossRef] [PubMed]

| Target | C. pneumoniae | Luteolin | References |
|---|---|---|---|
| ROS | + | − | [30,33,34] |
| NF-κB | + | − | [31,39] |
| MAP kinases | + | +/− | [32,36] |
| PI3 kinase | + | − | [53,54] |
| Cytosolic Ca2+ | + | +/− | [35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52] |
| Bcl-2 proteins | − | +/− | [56,58] |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hanski, L.; Vuorela, P. Lead Discovery Strategies for Identification of Chlamydia pneumoniae Inhibitors. Microorganisms 2016, 4, 43. https://doi.org/10.3390/microorganisms4040043
Hanski L, Vuorela P. Lead Discovery Strategies for Identification of Chlamydia pneumoniae Inhibitors. Microorganisms. 2016; 4(4):43. https://doi.org/10.3390/microorganisms4040043
Chicago/Turabian StyleHanski, Leena, and Pia Vuorela. 2016. "Lead Discovery Strategies for Identification of Chlamydia pneumoniae Inhibitors" Microorganisms 4, no. 4: 43. https://doi.org/10.3390/microorganisms4040043
APA StyleHanski, L., & Vuorela, P. (2016). Lead Discovery Strategies for Identification of Chlamydia pneumoniae Inhibitors. Microorganisms, 4(4), 43. https://doi.org/10.3390/microorganisms4040043





