Assessment of the Factors Contributing to the Growth or Spoilage of Meyerozyma guilliermondii in Organic Yogurt: Comparison of Methods for Strain Differentiation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Isolation and Culture Conditions
2.2. Identification
2.3. Physiological Analysis
2.4. Methods for Strain Differentiation (Typing)
2.5. Sugar Fermentation
| Isolated Strains | Identification a | Origin | Spoilage | CFU/g | Standard Deviation (Std. dev.) | RFLPs mtDNA | RFELP IGS | |
|---|---|---|---|---|---|---|---|---|
| HinfI | Hha I (B) | Hae III (H) | ||||||
| Mi1 | M. guilliermondii | Strawberry jam | Bubbles | ND | ND | A | B1 | H1 |
| Mi2 | M. guilliermondii | Strawberry jam | Bubbles | A | B1 | H1 | ||
| Mi3 | M. guilliermondii | Strawberry jam | Bubbles | 3.58 × 107 | 1.28 × 107 | A | B1 | H1 |
| Mi4 | M. guilliermondii | Strawberry yogurt | Swollen | 6.03 × 107 | 1.30 × 107 | A | B1 | H1 |
| Mi5 | M. guilliermondii | Strawberry yogurt | Swollen | A | B1 | H1 | ||
| Mi6 | M. guilliermondii | Strawberry yogurt | Swollen | 3.43 × 107 | 8.24 × 106 | A | B1 | H1 |
| Mi7 | M. guilliermondii | Berry yogurt | Swollen | 3.53 × 107 | 9.27 × 106 | A | B1 | H1 |
| Mi8 | M. guilliermondii | Berry yogurt | Swollen | 1.52 × 108 | 5.37 × 106 | A | B1 | H1 |
| YN2 | M. guilliermondii | Plain yogurt | Not swollen | 6.52 × 106 | 1.99 × 106 | A | B1 | H1 |
| YN5 | M. guilliermondii | Plain yogurt | Not swollen | 2.47 × 107 | 2.31 × 106 | A | B1 | H1 |
| YN8 | M. guilliermondii | Plain yogurt | Not swollen | 1.40 × 108 | 8.88 × 106 | A | B1 | H1 |
| YF6 | M. guilliermondii | Strawberry yogurt | Swollen | 9.72 × 107 | 6.57 × 106 | A | B1 | H1 |
| YF9 | M. guilliermondii | Strawberry yogurt | Swollen | 1.15 × 108 | 1.63 × 107 | A | B1 | H1 |
| YFB1 | M. guilliermondii | Berry yogurt | Swollen | 1.26 × 108 | 3.16 × 106 | A | B1 | H1 |
| YFB4 | M. guilliermondii | Berry yogurt | Swollen | 5.80 × 107 | 6.16 × 106 | A | B1 | H1 |
| YFB7 | M. guilliermondii | Berry yogurt | Swollen, bubbles | 1.14 × 108 | 3.85 × 106 | A | B1 | H1 |
| YA10 | M. guilliermondii | Apricot yogurt | Swollen, bubbles | 1.19 × 108 | 1.45 × 107 | A | B1 | H1 |
| MA11.1 | M. guilliermondii | Apricot jam | Bubbles | 4.60 × 106 | 2.38 × 105 | A | B1 | H1 |
| MA11.2 | W. anomalus | Apricot jam | - | - | - | |||
| Strains from: Spanish type culture collection | ||||||||
| CECT 1456 T | M. guilliermondii | Insect frass on Ulmus americana | - | - | B | B2 | H1 | |
| CECT 1019 | M. guilliermondii | Flower of Gentiana imbricata | - | - | F | B1 | H2 | |
| CECT 1438 | M. guilliermondii | Pozol, Mexican fermented maize dough | - | - | - | A | B1 | H2 |
| CECT 10157 | M. guilliermondii | Fruit in syrup | - | - | - | C | B1 | H2 |
| CECT 1021 | M. guilliermondii | Soil from drilling care | - | - | - | D | B3 | H2 |
| CECT 12791 | M. guilliermondii | Soil | - | - | - | D | B1 | H2 |
| CECT 12839 | M. guilliermondii | Beer var. garrafal | - | - | - | E | B1 | H2 |
| Isolated Strains | Type Strain a | |
|---|---|---|
| Assimilation-Growth | ||
| d-Glucose | + | (+, D) |
| d-Galactose | + | (−, D) |
| Sucrose | + | (+, D) |
| Lactose | − | − |
| dl-Lactate | + | − |
| Fermentation | ||
| d-Glucose | + | + |
| d-Galactose | + | W |
| Sucrose | + | + |
| Lactose | − | − |
2.6. Assessment of Gas Production in Lab-Contaminated Organic Yogurt
| Yoghurt Type | Inoculum | Incubation Temperature | |||||
|---|---|---|---|---|---|---|---|
| 8 °C | 28 °C | ||||||
| CFU/g | Std. dev. | Gas | CFU/g | Std. dev. | Gas | ||
| Plain | Low | 4.30 × 106 | 4.24 × 105 | No | 4.28 × 106 | 4.83 × 105 | No |
| High | 5.41 × 106 | 5.55 × 105 | No | 1.56 × 106 | 3.44 × 105 | No | |
| Past. plain | Low | 2.99 × 106 | 2.40 × 105 | No | 1.00 × 107 | 5.85 × 105 | No |
| High | 9.29 × 106 | 1.33 × 106 | No | 1.24 × 107 | 1.36 × 105 | No | |
| Fruit | Low | 1.21 × 107 | 3.12 × 105 | Low | 5.32 × 106 | 3.92 × 105 | High |
| High | 8.56 × 106 | 8.97 × 105 | Low | 5.52 × 106 | 8.91 × 105 | High | |
| Past. fruit | Low | 2.96 × 107 | 5.43 × 106 | Low | 9.32 × 106 | 7.55 × 105 | High |
| High | 5.57 × 106 | 1.32 × 106 | Low | 1.08 × 107 | 1.10 × 106 | High | |
3. Results
3.1. Strain Isolation and Identification
3.2. Comparison of Typing Techniques
3.3. Analysis of the Survival, Growth, and Spoiling Abilities of the Strains
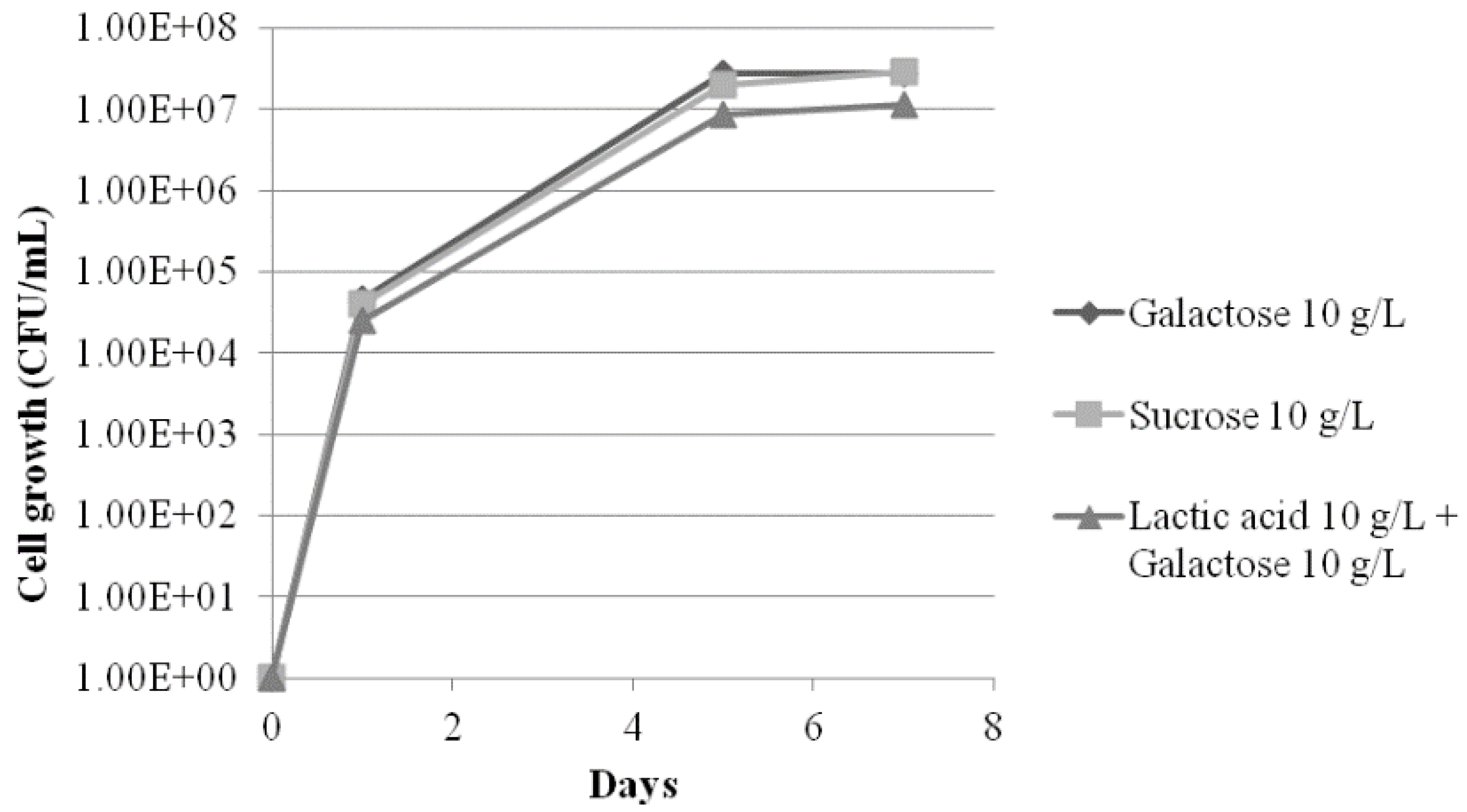
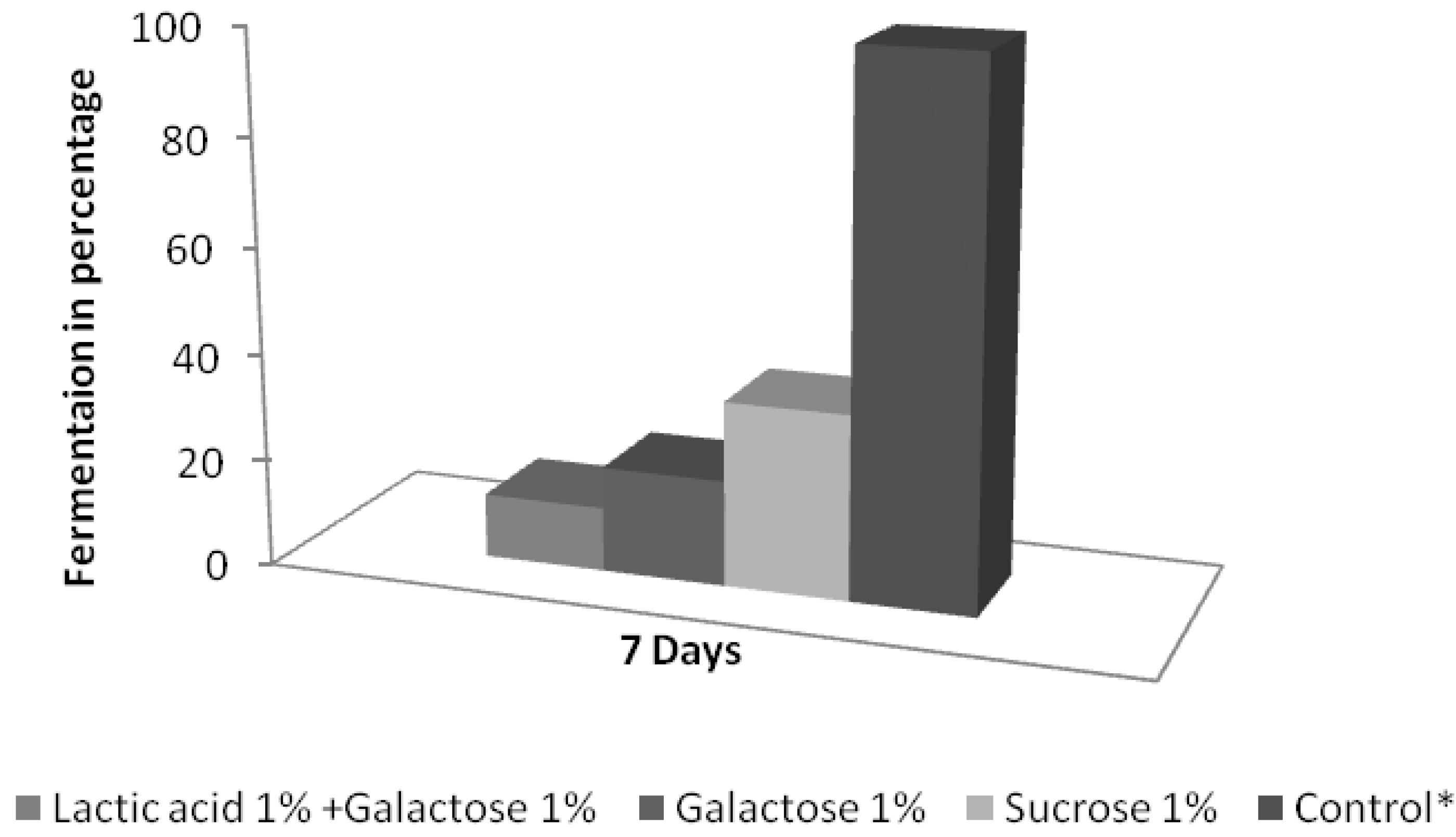
4. Discussion
5. Conclusions
Acknowledgements
Author Contributions
Conflicts of Interest
References
- Kurtzman, C.P.; Suzuki, M. Phylogenetic analysis of ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces, and Scheffersomyces. Micoscience 2010, 51, 2–14. [Google Scholar] [CrossRef]
- Kurtzman, C.P. Meyerozyma Kurtzman & M. Suzuki (2010). In The Yeasts: A Taxonomic Study, 5th ed.; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier Science: London, UK, 2011; Volume 2, pp. 621–624. [Google Scholar]
- Cooper, C.R., Jr. Yeast pathogenic to humans. In The Yeasts: A Taxonomic Study, 5th ed.; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier Science: London, UK, 2011; Volume 1, pp. 9–19. [Google Scholar]
- Wszelaki, A.L.; Mitcham, E.J. Effect of combinations of hot water dips, biological control and controlled atmospheres for control of gray mold on harvested strawberries. Postharvest Biol. Technol. 2003, 27, 255–264. [Google Scholar] [CrossRef]
- Petersson, S.; Schnurer, J. Biocontrol of Mold Growth in High-Moisture Wheat Stored under Airtight Conditions by Pichia anomala, Pichia guilliermondii, and Saccharomyces cerevisiae. Appl. Environ. Microbiol. 1995, 61, 1027–1032. [Google Scholar] [PubMed]
- Droby, S.; Wisniewski, M.E.; Cohen, L.; Weiss, B.; Touitou, D.; Eilam, Y.; Chalutz, E. Influence of CaCl2 on Penicillium digitatum, grapefruit peel tissue, and biocontrol activity of Pichia guilliermondii. Phytopathology 1997, 87, 310–315. [Google Scholar] [CrossRef] [PubMed]
- Lahlali, R.; Hamadi, Y.; El Guilli, M.; Jijakli, M.H. Efficacy assessment of Pichia guilliermondii strain Z1, a new biocontrol agent, against citrus blue mould in Morocco under the influence of temperature and relative humidity. Biol. Control 2011, 56, 217–224. [Google Scholar] [CrossRef]
- Zhao, Y.; Tu, K.; Shao, X.F.; Jing, W.; Yang, J.L.; Su, Z.P. Biological control of the post-harvest pathogens Alternaria solani, Rhizopus stolonifer, and Botrytis cinerea on tomato fruit by Pichia guilliermondii. J. Horticult. Sci. Biotechnol. 2008, 83, 132–136. [Google Scholar]
- Kinay, P.; Yildiz, M. The shelf life and effectiveness of granular formulations of Metschnikowia pulcherrima and Pichia guilliermondii yeast isolates that control postharvest decay of citrus fruit. Biol. Control 2008, 45, 433–440. [Google Scholar] [CrossRef]
- Mataragas, M.; Dimitriou, V.; Skandamis, P.N.; Drosinos, E.H. Quantifying the spoilage and shelf-life of yoghurt with fruits. Food Microbiol. 2011, 28, 611–616. [Google Scholar] [CrossRef] [PubMed]
- Miles, A.A.; Misra, S.S.; Irwin, J.O. The estimation of the bactericidal power of the blood. J. Hyg. 1938, 38, 732–749. [Google Scholar] [CrossRef] [PubMed]
- Corry, J.E.T.; Curtis, G.D.W.; Baird, R.M. Testing methods for use in quality assurance of culture media. In Handbook of Culture Media for Food and Water Microbiology, 3rd ed.; Royal Society of Chemistry: Cambridge, UK, 2011. [Google Scholar]
- Esteve-Zarzoso, B.; Belloch, C.; Uruburu, F.; Querol, A. Identification of yeasts by RFLP analysis of the 5.8S rRNA gene and the two ribosomal internal transcribed spacers. Int. J. Syst. Bacteriol. 1999, 49, 329–337. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Espinar, M.T.; Esteve-Zarzoso, B.; Querol, A.; Barrio, E. RFLP analysis of the ribosomal internal transcribed spacers and the 5.8S rRNA gene region of the genus Saccharomyces: A fast method for species identification and the differentiation of flor yeasts. Antonie van Leeuwenhoek 2000, 78, 87–97. [Google Scholar] [CrossRef]
- Romi, W.; Keisam, S.; Ahmed, G.; Jeyaram, K. Reliable differentiation of Meyerozyma guilliermondii from Meyerozyma caribbica by internal transcribed spacer restriction fingerprinting. BMC Microbiol. 2014, 14, 52. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Barnett, J.A.; Payne, R.W.; Yarrow, D. Yeasts: Characteristics and Identification, 2nd ed.; Cambridge University Press: Cambridge, UK, 1990. [Google Scholar]
- Kurtzman, C.P.; Fell, J.W.; Boekhout, T. The Yeasts: A Taxonomic Study, 5th ed.; Elsevier Science: London, UK, 2011; Volume 2. [Google Scholar]
- Querol, A.; Barrio, E.; Huerta, T.; Ramon, D. Molecular monotoring of wine fermentations coducted by active dry yeast strains. Appl. Environ. Microbiol. 1992, 58, 2948–2953. [Google Scholar] [PubMed]
- Appel, D.J.; Gordon, T.R. Intraspecific variation within populations of Fusarium oxysporum based on RFLP analysis of the Intergenic Spacer Region of the rDNA. Exp. Mycol. 1995, 19, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Romero, P.; Patiño, B.; Quirós, M.; González-Jaén, M.T.; Valderrama, M.J.; de Silóniz, M.I.; Peinado, J.M. Differential detection of Debaryomyces hansenii isolated from intermediate-moisture foods by PCR-RFLP of the IGS region of rDNA. FEMS Yeast Res. 2005, 5, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Quirós, M.; Martorell, P.; Valderrama, M.J.; Querol, A.; Peinado, J.M.; Silóniz, M.I. PCR-RFLP analysis of the IGS region of rDNA: A useful tool for the practical discrimination between species of the genus Debaryomyces. Anton. Leeuw. Int. J. G. 2006, 90, 211–219. [Google Scholar] [CrossRef] [PubMed]
- López, V.; Querol, A.; Ramón, D.; Fernández-Espinar, M.T. A simplified procedure to analyse mitochondrial DNA from industrial yeasts. Int. J. Food Microbiol. 2001, 68, 75–81. [Google Scholar] [CrossRef]
- Casas, E.; de Ancos, B.; Valderrama, M.J.; Canob, P.; Peinado, J.M. Pentadiene production from potassium sorbate by osmotolerant yeasts. Int. J. Food Microbiol. 2004, 94, 93–96. [Google Scholar] [CrossRef] [PubMed]
- Desnos-Ollivier, M.; Ragon, M.; Robert, V.; Raoux, D.; Gantier, J.C.; Dromer, F. Debaryomyces hansenii (Candida famata), a rare human fungal pathogen often misidentified as Pichia guilliermondii (Candida guilliermondii). J. Clin. Microbiol. 2008, 46, 3237–3242. [Google Scholar] [CrossRef] [PubMed]
- Deák, T. Handbook of Food Spoilage Yeasts, 2nd ed.; CRC Press: Boca Ratón, FL, USA, 2008. [Google Scholar]
- Fleet, G.H. Yeasts in Dairy Products. J. Appl. Bacteriol. 1990, 68, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Viljoen, B.C.; Lourens-Hattingh, A.; Ikalafeng, B.; Peter, G. Temperature abuse initiating yeast growth in yoghurt. Food Res. Int. 2003, 36, 193–197. [Google Scholar] [CrossRef]
- Martorell, P.; Barata, A.; Malfeito-Ferreira, M.; Fernández-Espinar, M.T.; Loureiro, V.; Querol, A. Molecular typing of the yeast species Dekkera bruxellensis and Pichia guilliermondii recovered from wine related sources. Int. J. Food Microbiol. 2006, 106, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Wrent, P.; Rivas, E.M.; Peinado, J.M.; de Silóniz, M.I. Strain typing of Zygosaccharomyces yeast species using a single molecular method based on polymorphism of the intergenic spacer region (IGS). Int. J. Food Microbiol. 2010, 142, 89–96. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marquina, D.; Llorente, P.; Santos, A.; Peinado, J.M. Characterization of the yeast population in low water activity foods. Adv. Food Sci. 2001, 23, 63–67. [Google Scholar]
- Yamamura, M.; Makimura, K.; Fujisaki, R.; Satoh, K.; Kawakami, S.; Nishiya, H.; Ota, Y. Polymerase chain reaction assay for specific identification of Candida guilliermondii (Pichia guilliermondii). J. Infect. Chemother. 2009, 15, 214–218. [Google Scholar] [CrossRef] [PubMed]
- Fleet, G.H. Spoilage yeasts. Crit. Rev. Biotechnol. 1992, 12, 1–44. [Google Scholar] [CrossRef] [PubMed]
- Fleet, G.H.; Balia, R. The public health and probiotic significance in foods and beverages. In Yeast in Foods and beverages; Querol, A., Fleet, G.H., Eds.; Springer-Verlang: Berlin, Germany, 2006; pp. 381–389. [Google Scholar]
- Corte, L.; di Cagno, R.; Groenewald, M.; Roscini, L.; Colabella, C.; Gobbetti, M.; Cardinali, G. Phenotypic and molecular diversity of Meyerozyma guilliermondii strains isolated from food and other environmental niches, hints for an incipient speciation. Food Microbiol. 2015, 48, 206–215. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wrent, P.; Rivas, E.-M.; De Prado, E.G.; Peinado, J.M.; De Silóniz, M.-I. Assessment of the Factors Contributing to the Growth or Spoilage of Meyerozyma guilliermondii in Organic Yogurt: Comparison of Methods for Strain Differentiation. Microorganisms 2015, 3, 428-440. https://doi.org/10.3390/microorganisms3030428
Wrent P, Rivas E-M, De Prado EG, Peinado JM, De Silóniz M-I. Assessment of the Factors Contributing to the Growth or Spoilage of Meyerozyma guilliermondii in Organic Yogurt: Comparison of Methods for Strain Differentiation. Microorganisms. 2015; 3(3):428-440. https://doi.org/10.3390/microorganisms3030428
Chicago/Turabian StyleWrent, Petra, Eva-María Rivas, Elena Gil De Prado, José M. Peinado, and María-Isabel De Silóniz. 2015. "Assessment of the Factors Contributing to the Growth or Spoilage of Meyerozyma guilliermondii in Organic Yogurt: Comparison of Methods for Strain Differentiation" Microorganisms 3, no. 3: 428-440. https://doi.org/10.3390/microorganisms3030428





