Genomic Characteristics of a Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli RZ-13 Isolates from Diarrheic Calves with High Mortality in China
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation and Identification of E. coli RZ-13
2.2. Whole Genome Sequencing and Assembly of E. coli RZ-13
2.3. Bioinformatic Analysis of E. coli RZ-13
2.4. Nucleotide Sequence Accession Numbers of E. coli RZ-13
2.5. Conjugation and Transformation Experiments of E. coli RZ-13
2.6. Antibiotic Susceptibility Testing
3. Results and Discussion
3.1. The Genomic Profiles of E. coli RZ-13
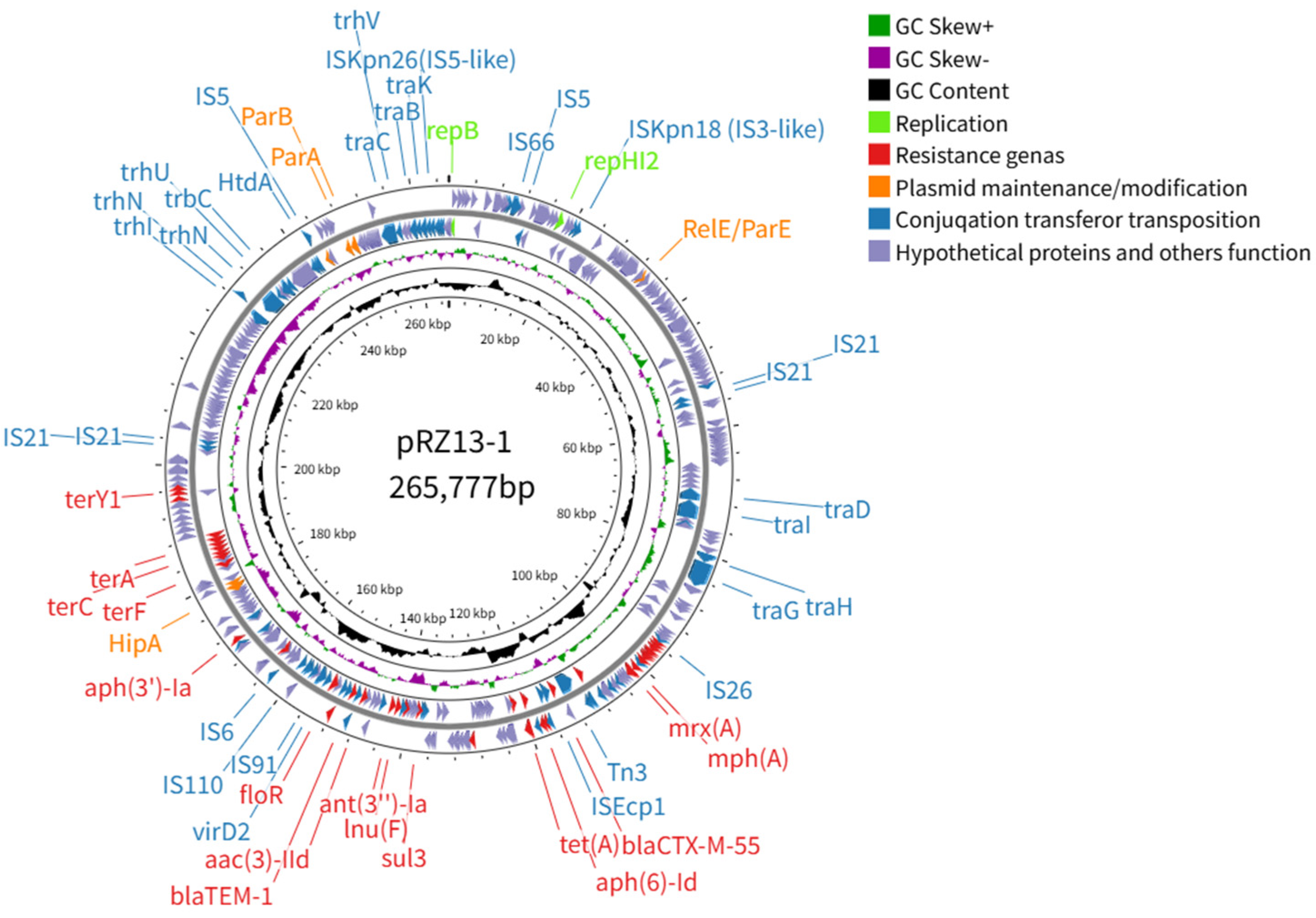
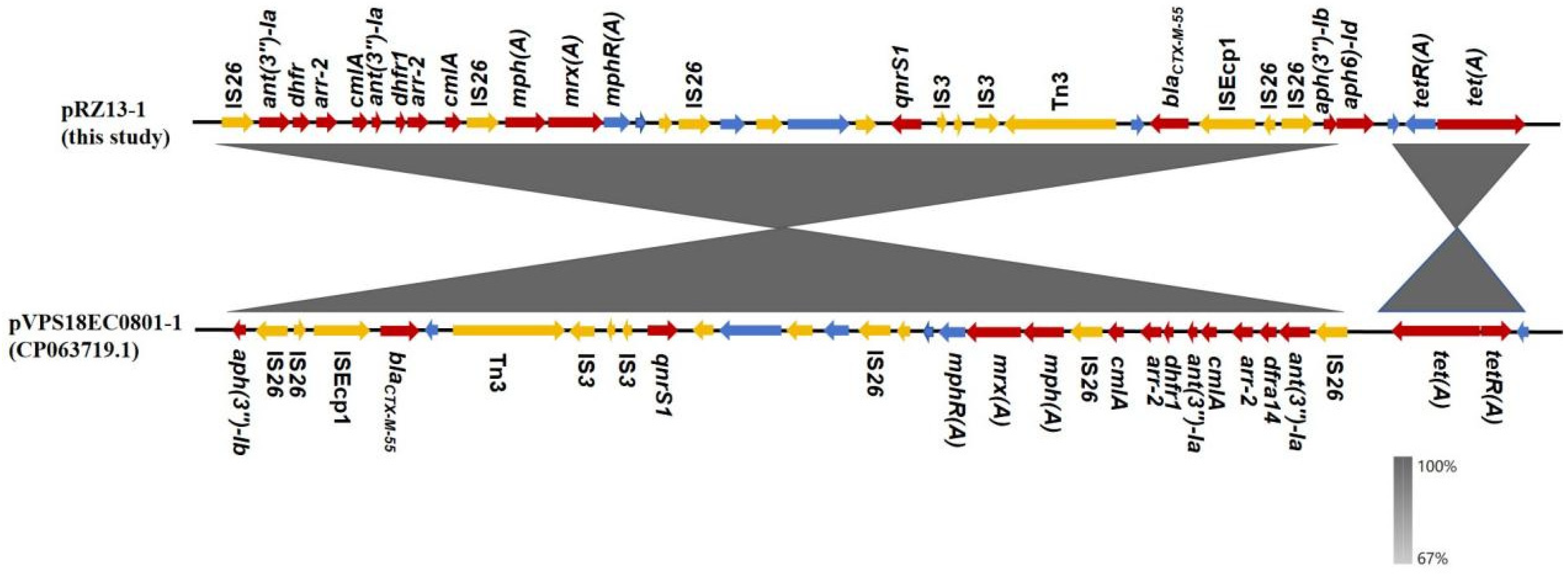
3.2. Characteristics and Transferability of the Multidrug-Resistance Plasmid pRZ13-1
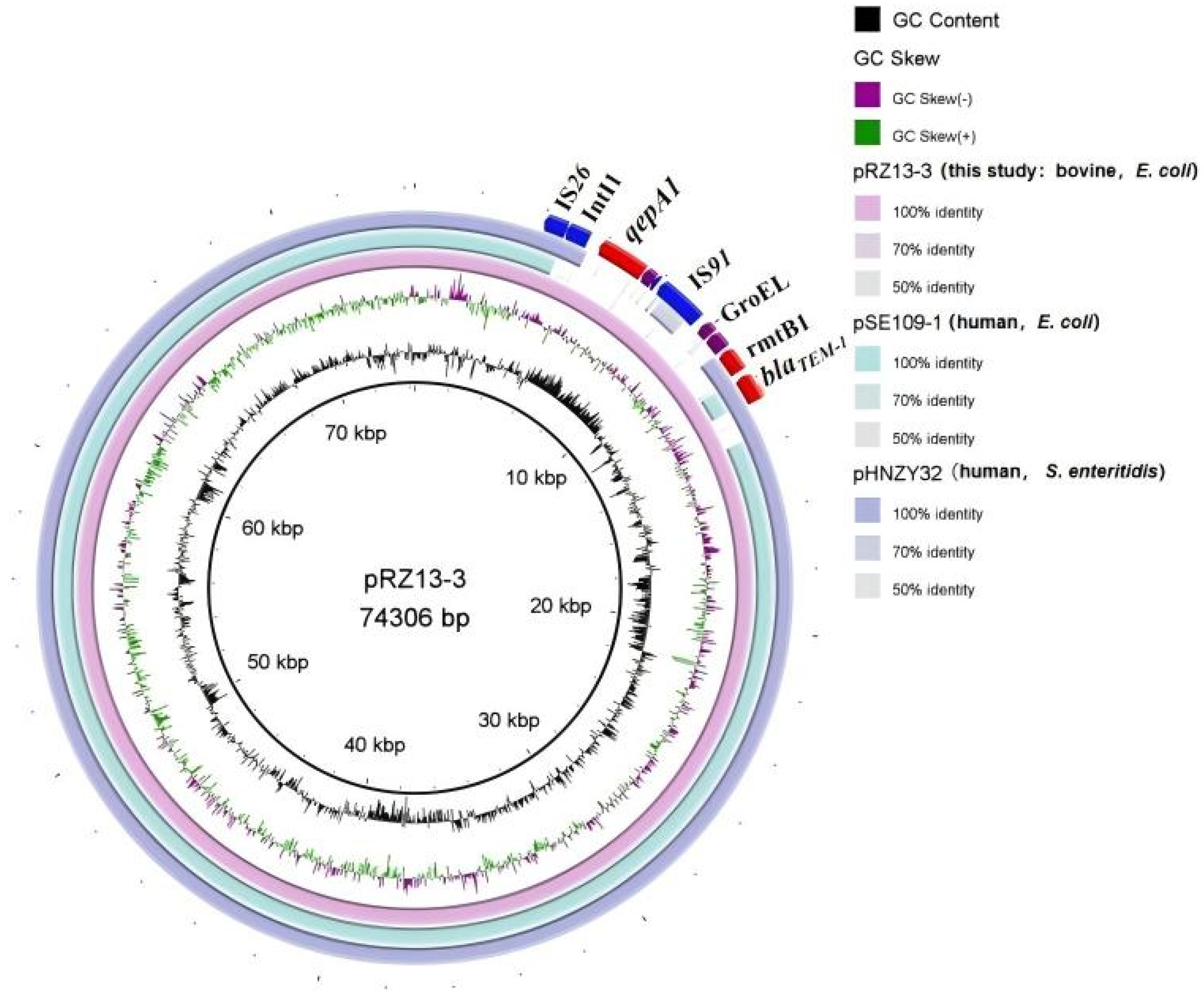
3.3. Characteristics and Transferability of the Multidrug-Resistance Plasmid pRZ13-3
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Cedeño-Muñoz, J.S.; Aransiola, S.A.; Reddy, K.V.; Ranjit, P.; Victor-Ekwebelem, M.O.; Oyedele, O.J.; Pérez-Almeida, I.B.; Maddela, N.R.; Rodríguez-Díaz, J.M. Antibiotic resistant bacteria and antibiotic resistance genes as contaminants of emerging concern: Occurrences, impacts, mitigations and future guidelines. Sci. Total. Environ. 2024, 952, 175906. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.D.; Li, X.R.; Ma, T.F.; Chen, J.Q.; Ge, C.H.; Yang, S.H.; Zhang, W.; Chen, J.; Zhang, J.J.; Qi, M.M.; et al. Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli Exhibits High Virulence in Calf Herds: A Case Report. Microbiol. Res. 2025, 16, 59. [Google Scholar] [CrossRef]
- Peymani, A.; Naserpour-Farivar, T.; Zare, E.; Azarhoosh, K.H. Distribution of blaTEM, blaSHV, and blaCTX-M. genes among ESBL-producing P. aeruginosa isolated from Qazvin and Tehran hospitals, Iran. J. Prev. Med. Hyg. 2017, 58, E155–E160. [Google Scholar]
- Wang, J.; Wang, Z.Y.; Wang, Y.; Sun, F.; Li, W.; Wu, H.; Shen, P.C.; Pan, Z.M.; Jiao, X. Emergence of 16S rRNA Methylase Gene rmtB in Salmonella Enterica Serovar London and Evolution of RmtB-Producing Plasmid Mediated by IS26. Front. Microbiol. 2021, 11, 604278. [Google Scholar] [CrossRef]
- Xia, J.; Sun, J.; Cheng, K.; Li, L.; Fang, L.X.; Zou, M.T.; Liao, X.P.; Liu, Y.H. Persistent spread of the rmtB 16S rRNA methyltransferase gene among Escherichia coli isolates from diseased food-producing animals in China. Vet. Microbiol. 2016, 188, 41–46. [Google Scholar] [CrossRef]
- Yu, F.Y.; Yao, D.; Pan, J.Y.; Chen, C.; Qin, Z.Q.; Parsons, C.; Yang, L.H.; Li, Q.Q.; Zhang, X.Q.; Qu, D.; et al. High prevalence of plasmid-mediated 16S rRNA methylase gene rmtB among Escherichia coli clinical isolates from a Chinese teaching hospital. BMC Infect. Dis. 2010, 10, 184. [Google Scholar] [CrossRef] [PubMed]
- Doi, Y.; Yokoyama, K.; Yamane, K.; Wachino, J.; Shibata, N.; Yagi, T.; Shibayama, K.; Kato, H.; Arakawa, Y. Plasmid-mediated 16S rRNA methylase in Serratia marcescens conferring high-level resistance to aminoglycosides. Antimicrob. Agents Chemother. 2004, 48, 491–496. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L.; Bonomo, R.A. Extended-spectrum beta-lactamases: A clinical update. Clin. Microbiol. Rev. 2005, 18, 657–686. [Google Scholar] [CrossRef]
- Bauernfeind, A.; Grimm, H.; Schweighart, S. A new plasmidic cefotaximase in a clinical isolate of Escherichia coli. Infection 1990, 18, 294–298. [Google Scholar] [CrossRef]
- Tabaran, A.; Soulageon, V.; Chirila, F.; Reget, O.L.; Mihaiu, M.; Borzan, M.; Dan, S.D. Pathogenic E. coli from Cattle as a Reservoir of Resistance Genes to Various Groups of Antibiotics. Antibiotics 2022, 11, 404. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control (ECDC); European Food Safety Authority (EFSA); European Medicines Agency (EMA). ECDC/EFSA/EMA second joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals: Joint Interagency Antimicrobial Consumption and Resistance Analysis (JIACRA) Report. EFSA. J. 2017, 15, e04872. [Google Scholar] [CrossRef]
- Liang, C.; Wei, D.; Yan, W.; Zhang, S.; Shi, J.; Liu, L. Fates of intracellular and extracellular antibiotic resistance genes during the cattle farm wastewater treatment process. Bioresour. Technol. 2022, 344, 126272. [Google Scholar] [CrossRef]
- Abbas, F.; Thomas, P.; Cully-Duse, B.; Andronicos, N.M.; Winter, G. Cattle-compost-soil: The transfer of antibiotic resistance in livestock agriculture. Microbiologyopen 2023, 12, e1375. [Google Scholar] [CrossRef]
- Gaballah, M.S.; Guo, J.; Sun, H.; Aboagye, D.; Sobhi, M.; Muhmood, A.; Dong, R. A review targeting veterinary antibiotics removal from livestock manure management systems and future outlook. Bioresour. Technol. 2021, 333, 125069. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, H.; Yu, S.; Li, D.; Gillings, M.R.; Ren, H.; Mao, D.; Guo, J.; Luo, Y. Inter-plasmid transfer of antibiotic resistance genes accelerates antibiotic resistance in bacterial pathogens. ISME J. 2024, 18, wrad032. [Google Scholar] [CrossRef] [PubMed]
- Helinski, D.R. A Brief History of Plasmids. EcoSal. Plus. 2022, 10, eESP00282021. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.J. Role of Plasmids in the Ecology and Evolution of “High-Risk” Extraintestinal Pathogenic Escherichia coli Clones. EcoSal. Plus. 2021, 9, eESP-0013-2020. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef]
- Turton, J.F.; Ward, M.E.; Woodford, N.; Kaufmann, M.E.; Pike, R.; Livermore, D.M.; Pitt, T.L. The role of ISAba1 in expression of OXA carbapenemase genes in Acinetobacter baumannii. FEMS Microbiol. Lett. 2006, 258, 72–77. [Google Scholar] [CrossRef]
- Yang, X.; Dong, N.; Chan, E.W.; Zhang, R.; Chen, S. Carbapenem Resistance-Encoding and Virulence-Encoding Conjugative Plasmids in Klebsiella pneumoniae. Trends. Microbiol. 2021, 29, 65–83. [Google Scholar] [CrossRef]
- Pitout, J.D.D.; Finn, T.J. The evolutionary puzzle of Escherichia coli ST131. Infect. Genet. Evol. 2020, 81, 104265. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome. Res. 2017, 27, 722–736. [Google Scholar] [CrossRef] [PubMed]
- Vaser, R.; Sović, I.; Nagarajan, N.; Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 2017, 27, 737–746. [Google Scholar] [CrossRef]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC. Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef] [PubMed]
- Brettin, T.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Olsen, G.J.; Olson, R.; Overbeek, R.; Parrello, B.; Pusch, G.D.; et al. RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 2015, 5, 8365. [Google Scholar] [CrossRef]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef]
- Chen, L.; Yang, J.; Yu, J.; Yao, Z.; Sun, L.; Shen, Y.; Jin, Q. VFDB: A reference database for bacterial virulence factors. Nucleic Acids Res. 2005, 33, D325–D328. [Google Scholar] [CrossRef]
- Stothard, P.; Grant, J.R.; Van Domselaar, G. Visualizing and comparing circular genomes using the CGView family of tools. Brief. Bioinform. 2019, 20, 1576–1582. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef]
- Yang, X.; Wai-Chi Chan, E.; Zhang, R.; Chen, S.A. Conjugative plasmid that augments virulence in Klebsiella pneumoniae. Nat. Microbiol. 2019, 4, 2039–2043. [Google Scholar] [CrossRef]
- Zhao, J.; Pu, D.; Li, Z.; Zhang, Y.; Liu, X.; Zhuo, X.; Lu, B.; Cao, B. Loss and gain of ceftazidime-avibactam susceptibility in a non-carbapenemase-producing K1-ST23 hypervirulent Klebsiella pneumoniae. Virulence 2024, 15, 2348251. [Google Scholar] [CrossRef]
- Hackel, M.A.; Tsuji, M.; Yamano, Y.; Echols, R.; Karlowsky, J.A.; Sahm, D.F. Reproducibility of broth microdilution MICs for the novel siderophore cephalosporin, cefiderocol, determined using iron-depleted cation-adjusted Mueller-Hinton broth. Diagn. Microbiol. Infect. Dis. 2019, 94, 321–325. [Google Scholar] [CrossRef] [PubMed]
- Tamma, P.D.; Heil, E.L.; Justo, J.A.; Mathers, A.J.; Satlin, M.J.; Bonomo, R.A. Infectious Diseases Society of America 2024 Guidance on the Treatment of Antimicrobial-Resistant Gram-Negative Infections. Clin. Infect. Dis. 2024, 7, ciae403. [Google Scholar] [CrossRef] [PubMed]
- Perrott, J.; Mabasa, V.H.; Ensom, M.H. Comparing outcomes of meropenem administration strategies based on pharmacokinetic and pharmacodynamic principles: A qualitative systematic review. Ann. Pharmacother. 2010, 44, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Cirillo, I.; Vaccaro, N.; Turner, K.; Solanki, B.; Natarajan, J.; Redman, R. Pharmacokinetics, safety, and tolerability of doripenem after 0.5-, 1-, and 4-hour infusions in healthy volunteers. J. Clin. Pharmacol. 2009, 49, 798–806. [Google Scholar] [CrossRef]
- Sakka, S.G.; Glauner, A.K.; Bulitta, J.B.; Kinzig-Schippers, M.; Pfister, W.; Drusano, G.L.; Sörgel, F. Population pharmacokinetics and pharmacodynamics of continuous versus short-term infusion of imipenem-cilastatin in critically ill patients in a randomized, controlled trial. Antimicrob. Agents. Chemother. 2007, 51, 3304–3310. [Google Scholar] [CrossRef]
- Tsuji, B.T.; Pogue, J.M.; Zavascki, A.P.; Paul, M.; Daikos, G.L.; Forrest, A.; Giacobbe, D.R.; Viscoli, C.; Giamarellou, H.; Karaiskos, I.; et al. International Consensus Guidelines for the Optimal Use of the Polymyxins: Endorsed by the American College of Clinical Pharmacy (ACCP), European Society of Clinical Microbiology and Infectious Diseases (ESCMID), Infectious Diseases Society of America (IDSA), International Society for Anti-infective Pharmacology (ISAP), Society of Critical Care Medicine (SCCM), and Society of Infectious Diseases Pharmacists (SIDP). Pharmacotherapy 2019, 39, 10–39. [Google Scholar] [CrossRef]
- Ramesh Kumar, U.; Nguyen, N.T.; Dewangan, N.K.; Mohiuddin, S.G.; Orman, M.A.; Cirino, P.C.; Conrad, J.C. Co-Expression of type 1 fimbriae and flagella in Escherichia coli: Consequences for adhesion at interfaces. Soft. Matter. 2024, 20, 7397–7404. [Google Scholar] [CrossRef] [PubMed]
- Saldaña, Z.; Xicohtencatl-Cortes, J.; Avelino, F.; Phillips, A.D.; Kaper, J.B.; Puente, J.L.; Girón, J.A. Synergistic role of curli and cellulose in cell adherence and biofilm formation of attaching and effacing Escherichia coli and identification of Fis as a negative regulator of curli. Environ. Microbiol. 2009, 11, 992–1006. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sun, H.; Wang, M.; Liu, Y.; Wu, P.; Yao, T.; Yang, W.; Yang, Q.; Yan, J.; Yang, B. Regulation of flagellar motility and biosynthesis in enterohemorrhagic Escherichia coli O157:H7. Gut. Microbes. 2022, 14, 2110822. [Google Scholar] [CrossRef]
- Suarez, G.; Sierra, J.C.; Erova, T.E.; Sha, J.; Horneman, A.J.; Chopra, A.K. A type VI secretion system effector protein, VgrG1, from Aeromonas hydrophila that induces host cell toxicity by ADP ribosylation of actin. J. Bacteriol. 2010, 192, 155–168. [Google Scholar] [CrossRef]
- Farooq, A.; Rafique, A.; Han, E.; Park, S.M. Global dissemination of the beta-lactam resistance gene blaTEM-1 among pathogenic bacteria. Sci. Total. Environ. 2025, 963, 178521. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Fang, C.; Zhang, J.; Hua, W.; He, R.; Zhou, M. Carbapenemase- and Colistin Resistant Escherichia coli Strains from Children in China: High Genetic Diversity and First Report of blaNDM-5, blaCTX-M-65, blaOXA-10, blaTEM-1, and mcr-1.1 Genes Co-Occurrence in E. coli ST156. Infect. Drug Resist. 2022, 15, 5315–5320. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Tang, H.; Bao, R. Comparative analysis of five type II TA systems identified in Pseudomonas aeruginosa reveals their contributions to persistence and intracellular survival. Front. Cell Infect. Microbiol. 2023, 13, 1127786. [Google Scholar] [CrossRef]
- Li, Y.G.; Kishida, K.; Ogawa-Kishida, N.; Christie, P.J. Ligand-Displaying E. coli Cells and Minicells for Programmable Delivery of Toxic Payloads via Type IV Secretion Systems. BioRxiv 2023, 14, e0214323. [Google Scholar] [CrossRef]
- Peng, W.; Wang, Y.; Fu, Y.; Deng, Z.; Lin, S.; Liang, R. Characterization of the Tellurite-Resistance Properties and Identification of the Core Function Genes for Tellurite Resistance in Pseudomonas citronellolis SJTE-3. Microorganisms 2022, 10, 95. [Google Scholar] [CrossRef]
- Ponnusamy, D.; Clinkenbeard, K.D. Role of Tellurite Resistance Operon in Filamentous Growth of Yersinia pestis in Macrophages. PLoS ONE 2015, 10, e0141984. [Google Scholar] [CrossRef]
- Muñoz-Villagrán, C.M.; Mendez, K.N.; Cornejo, F.; Figueroa, M.; Undabarrena, A.; Morales, E.H.; Arenas-Salinas, M.; Arenas, F.A.; Castro-Nallar, E.; Vásquez, C.C. Comparative genomic analysis of a new tellurite-resistant Psychrobacter strain isolated from the Antarctic Peninsula. PeerJ 2018, 6, e4402. [Google Scholar] [CrossRef]
- Farias, P.; Francisco, R.; Morais, P.V. Potential of tellurite resistance in heterotrophic bacteria from mining environments. IScience 2022, 25, 104566. [Google Scholar] [CrossRef]
- Tate, H.; Li, C.; Nyirabahizi, E.; Tyson, G.H.; Zhao, S.; Rice-Trujillo, C.; Jones, S.B.; Ayers, S.; M’ikanatha, N.M.; Hanna, S.; et al. A National Antimicrobial Resistance Monitoring System Survey of Antimicrobial-Resistant Foodborne Bacteria Isolated from Retail Veal in the United States. J. Food. Prot. 2021, 84, 1749–1759. [Google Scholar] [CrossRef]
- Marimuthu, K.; Venkatachalam, I.; Koh, V.; Harbarth, S.; Perencevich, E.; Cherng, B.P.Z.; Fong, R.K.C.; Pada, S.K.; Ooi, S.T.; Smitasin, N.; et al. Whole genome sequencing reveals hidden transmission of carbapenemase-producing Enterobacterales. Nat. Commun. 2022, 13, 3052. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Yang, J.; Abbas, M.; Yang, Q.; Li, Q.; Liu, M.; Zhu, D.; Wang, M.; Tian, B.; Cheng, A. Threats across boundaries: The spread of ESBL-positive Enterobacteriaceae bacteria and its challenge to the “one health” concept. Front. Microbiol. 2025, 16, 1496716. [Google Scholar] [CrossRef]
- Tian, D.; Zhao, M.; Zheng, S.; Jiang, X.; Zhang, B. Involvement of Tn3 transposon in formation and transmission of hypervirulent and carbapenem-resistant Klebsiella pneumoniae. Microbiol. Spectr. 2023, 11, e0303823. [Google Scholar] [CrossRef] [PubMed]
- Harmer, C.J.; Hall, R.M. IS26 and the IS26 family: Versatile resistance gene movers and genome reorganizers. Microbiol. Mol. Biol. Rev. 2024, 88, e0011922. [Google Scholar] [CrossRef]
- Alonso, J.C. Toxin-Antitoxin Systems in Pathogenic Bacteria. Toxins 2021, 13, 74. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; He, L.; Li, Y.; Zeng, Z.; Deng, Y.; Liu, Y.; Liu, J.H. Complete sequence of a F2:A-:B- plasmid pHN3A11 carrying rmtB and qepA, and its dissemination in China. Vet. Microbiol. 2014, 174, 267–271. [Google Scholar] [CrossRef]
- Bonnin, R.A.; Poirel, L.; Carattoli, A.; Nordmann, P. Characterization of an IncFII plasmid encoding NDM-1 from Escherichia coli ST131. PLoS ONE 2012, 7, e34752. [Google Scholar] [CrossRef]
- Çetiner, U.; Raz, O.; Britt, M.; Sukharev, S. Dissipation during the Gating Cycle of the Bacterial Mechanosensitive Ion Channel Approaches the Landauer Limit. Entropy 2023, 25, 779. [Google Scholar] [CrossRef] [PubMed]
- Fourie, K.R.; Wilson, H.L. Understanding GroEL and DnaK Stress Response Proteins as Antigens for Bacterial Diseases. Vaccines 2020, 8, 773. [Google Scholar] [CrossRef] [PubMed]


| Location | Size (bp) | GC% | No. of CDSs | Plasmid Type | Replicon Type | Virulence Factors | Resistance Genes |
|---|---|---|---|---|---|---|---|
| Chromosome | 486,4281 | 51.43% | 4759 | – | – | – | blaTEM-1, tet(A) |
| pRZ13-1 | 265,777 | 46.80% | 356 | lncHI2-lncHI2A | repB, repHI2 | terC | aac(3)-IId, aph(6)-Id, aph(3″)-Ia, aph(3″)-Ib, blaCTX-M-55, blaTEM-1, ant(3″)-Ia, mph(A), mef(B), mphR, lnu(F), floR, qnrS1, sul3, dfrA14, cmlA, arr-2, mrx(A), tet(A), terABCDEFZY1 |
| pRZ13-2 | 92,100 | 47.30% | 132 | ND | repB | – | – |
| pRZ13-3 | 74,304 | 53.00% | 110 | IncFII | repA | traJ, traT | qepA1, rmtB, blaTEM-1, blaCTX-M-55 |
| pRZ13-4 | 61,618 | 42.38% | 94 | lncI2 | repA | – | – |
| Plasmid | Size/bp | Query Coordinates/bp | Identity /% | Region | Isolation Source | Collection Date | Organism |
|---|---|---|---|---|---|---|---|
| EC9 | 4880,174 | 32,871~84,110 | 99 | Hangzhou | Urine | September 2021 | E. coli |
| pESBL-blaCTX-M-55 | 95,161 | 33,961~85,202 | 99 | - | Chicken | - | E. coli |
| pHNZY32 | 145,804 | 85,267~136,504 | 99 | Guangzhou | Human | - | E. coli |
| pSE109-1 | 90,315 | 29,116~80,347 | 99 | - | Human | September 2013 | S. Enteritidis |
| pSE104-1 | 106,652 | 45,447~96,684 | 99 | Shanghai | Human | July 2013 | S. Enteritidis |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhu, D.-D.; Zhang, L.; Yang, S.-H.; Ge, C.-H.; Chen, J.-Q.; Ma, T.-F.; Yang, H.-J. Genomic Characteristics of a Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli RZ-13 Isolates from Diarrheic Calves with High Mortality in China. Microorganisms 2026, 14, 521. https://doi.org/10.3390/microorganisms14030521
Zhu D-D, Zhang L, Yang S-H, Ge C-H, Chen J-Q, Ma T-F, Yang H-J. Genomic Characteristics of a Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli RZ-13 Isolates from Diarrheic Calves with High Mortality in China. Microorganisms. 2026; 14(3):521. https://doi.org/10.3390/microorganisms14030521
Chicago/Turabian StyleZhu, Di-Di, Liang Zhang, Shao-Hua Yang, Chuan-Hui Ge, Jia-Qi Chen, Teng-Fei Ma, and Hong-Jun Yang. 2026. "Genomic Characteristics of a Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli RZ-13 Isolates from Diarrheic Calves with High Mortality in China" Microorganisms 14, no. 3: 521. https://doi.org/10.3390/microorganisms14030521
APA StyleZhu, D.-D., Zhang, L., Yang, S.-H., Ge, C.-H., Chen, J.-Q., Ma, T.-F., & Yang, H.-J. (2026). Genomic Characteristics of a Multidrug-Resistant Extraintestinal Pathogenic Escherichia coli RZ-13 Isolates from Diarrheic Calves with High Mortality in China. Microorganisms, 14(3), 521. https://doi.org/10.3390/microorganisms14030521





