Identification and Characterization of Antiyeast Organic Acids Produced by Lactiplantibacillus plantarum 3121M0s Derived from Mongolian Traditional Fermented Milk, Airag
Abstract
1. Introduction
2. Materials and Methods
2.1. Microbial Strains and Culture Conditions
2.2. Antiyeast Activity Assay
2.3. Cocultivation in Skim Milk
2.4. Species Identification
2.5. Determination of Optimal Culture Conditions
2.6. Evaluation of Effects of pH, Heating, and Enzymes on Antiyeast Activity
2.7. Ethyl Acetate Extraction
2.8. HPLC and MS
2.9. Evaluation of Effects of Adding Tyrosine and Phenylalanine into Culture Medium
2.10. Statistical Analysis
3. Results
3.1. Selection of Strain 3121M0s with High Antiyeast Activity
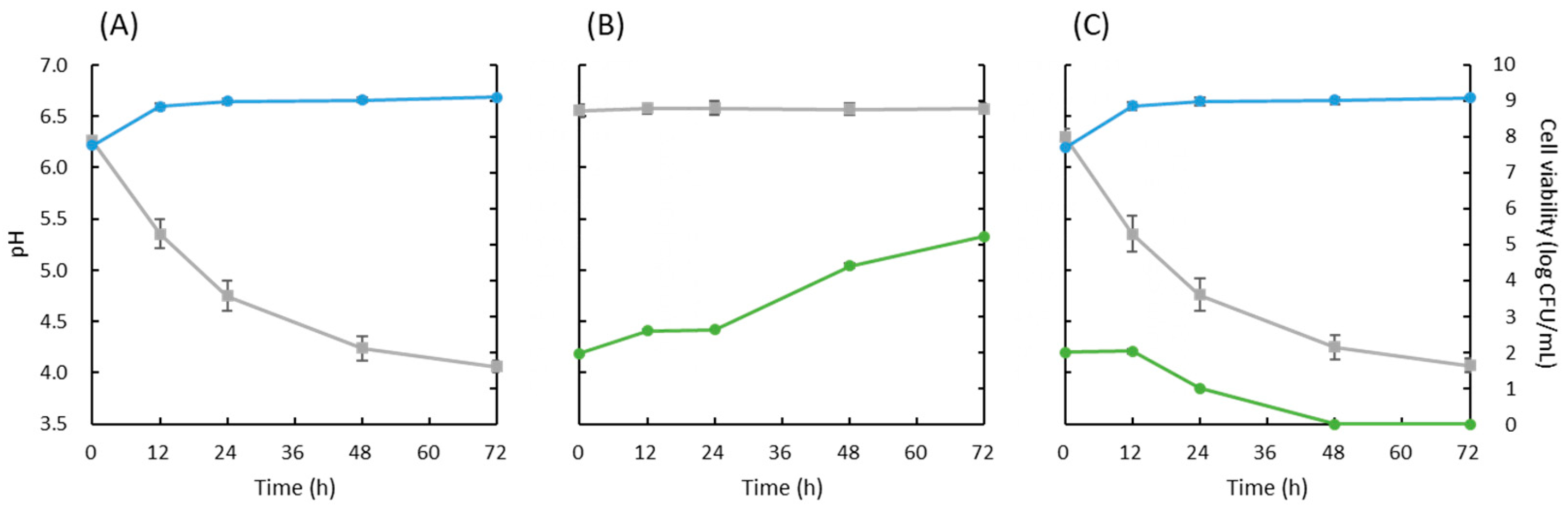
3.2. Evaluation of Antiyeast Effect of Strain 3121M0s in Skim Milk
3.3. Species Identification of Strain 3121M0s
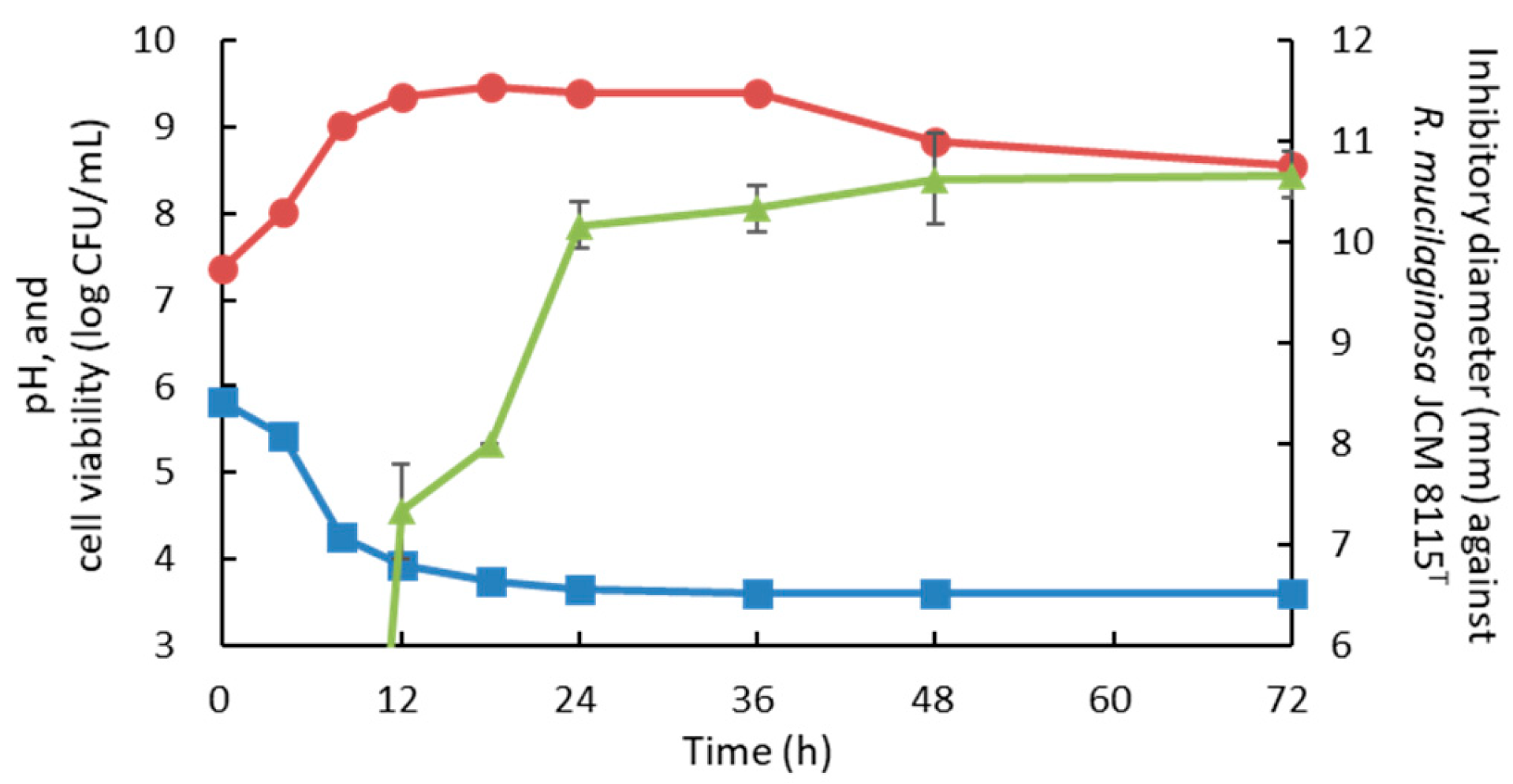
3.4. Determination of Optimal Culture Conditions of Strain 3121M0s
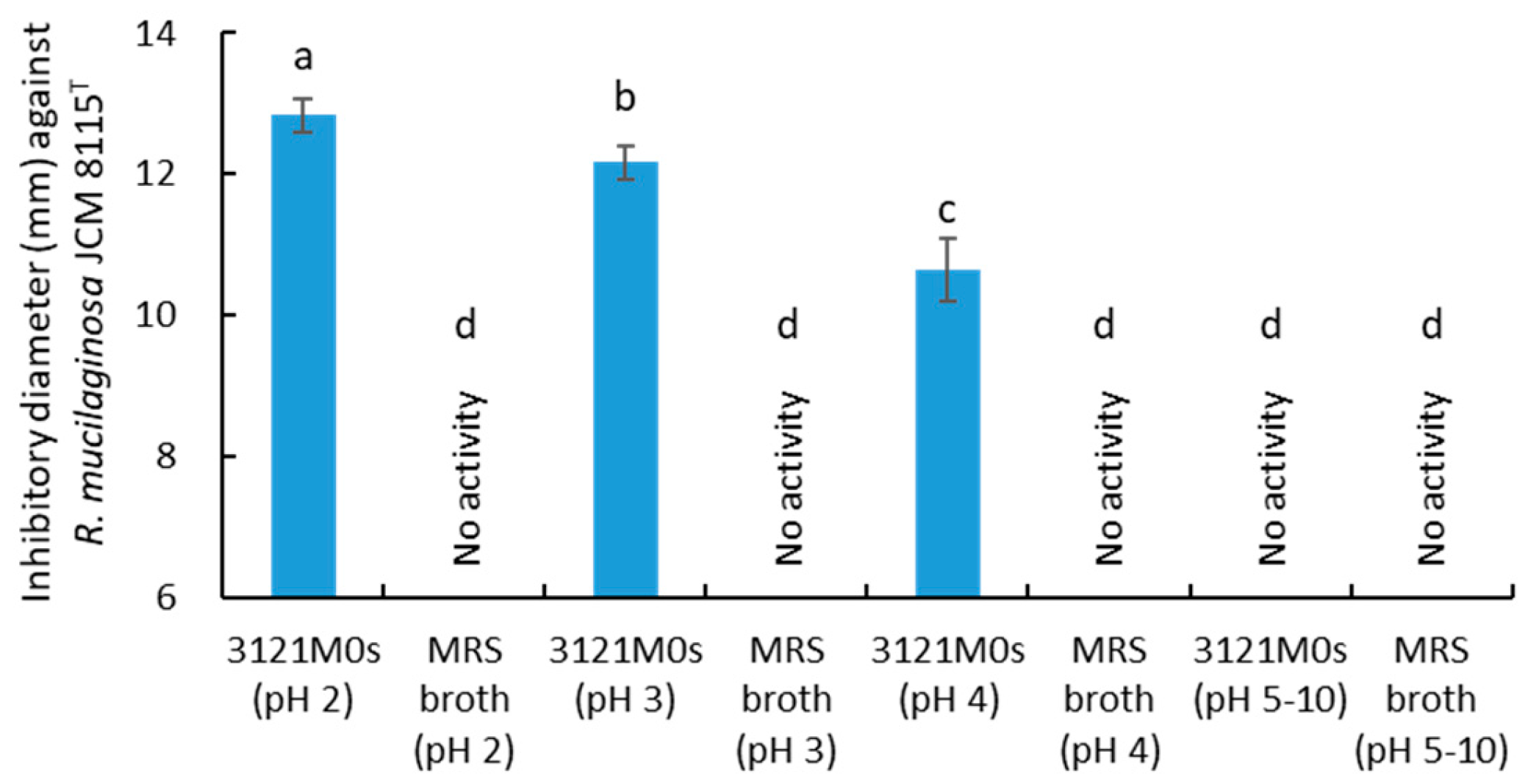
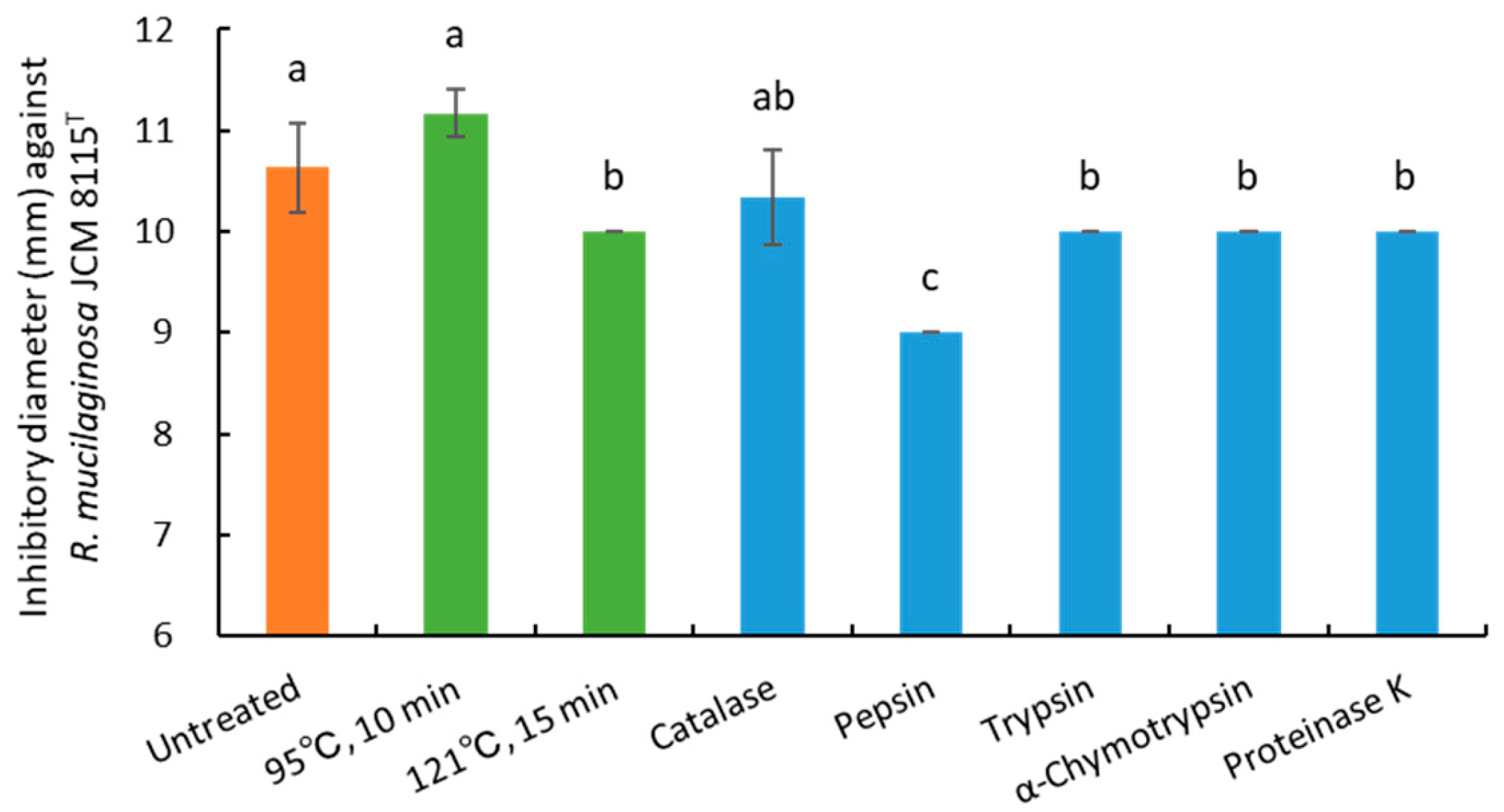
3.5. Evaluation of pH, Heating, and Enzymes on Antiyeast Activity of Strain 3121M0s
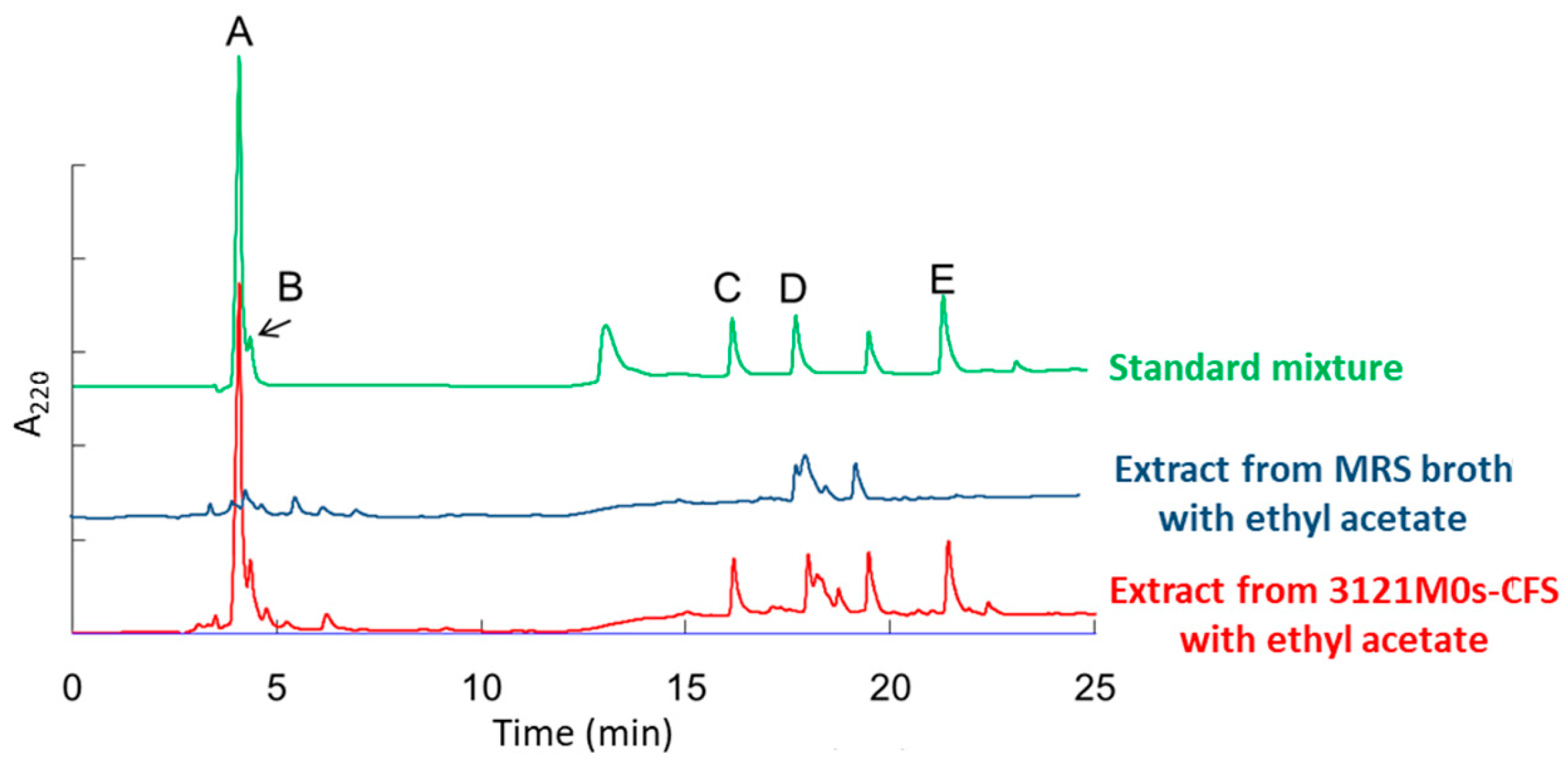
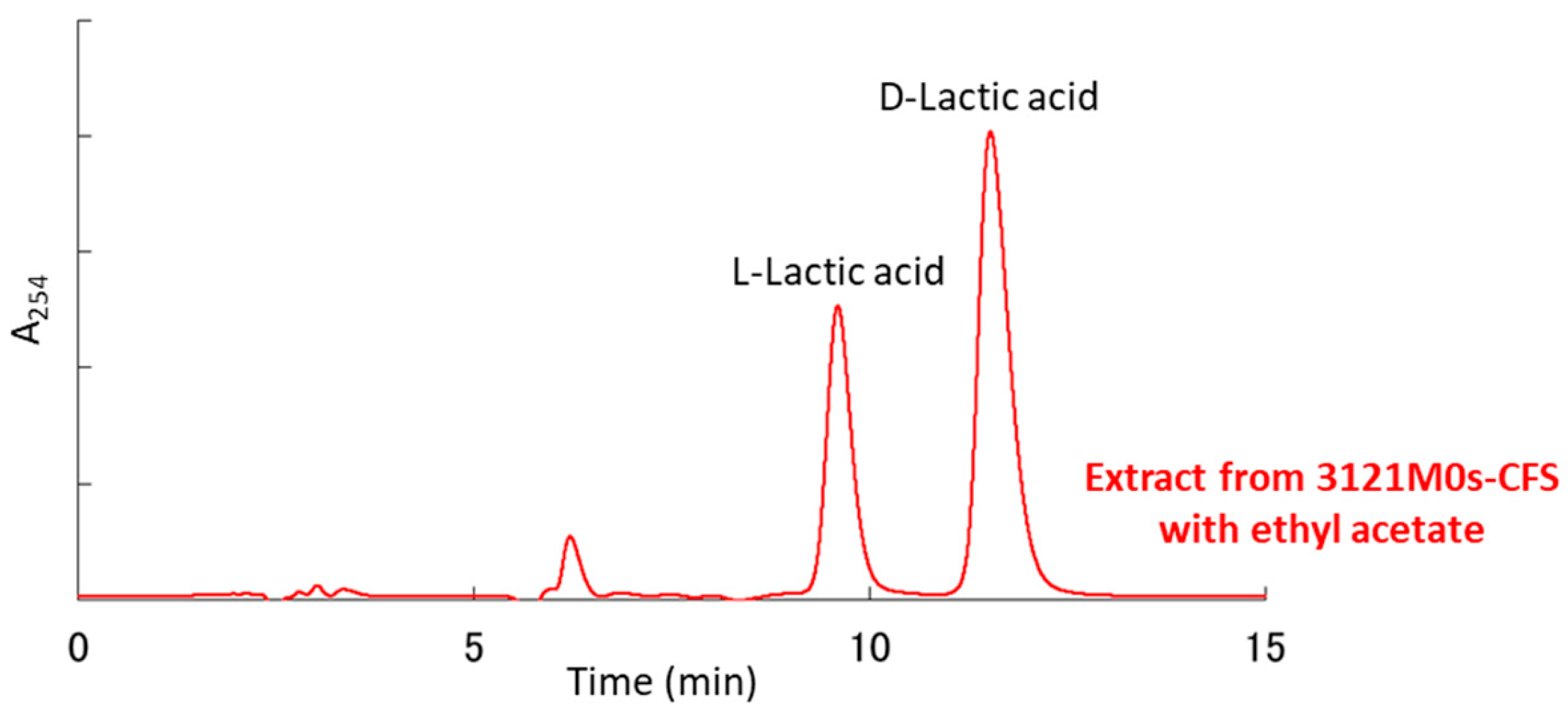
3.6. Identification and Quantification of Organic Acids Produced by Strain 3121M0s
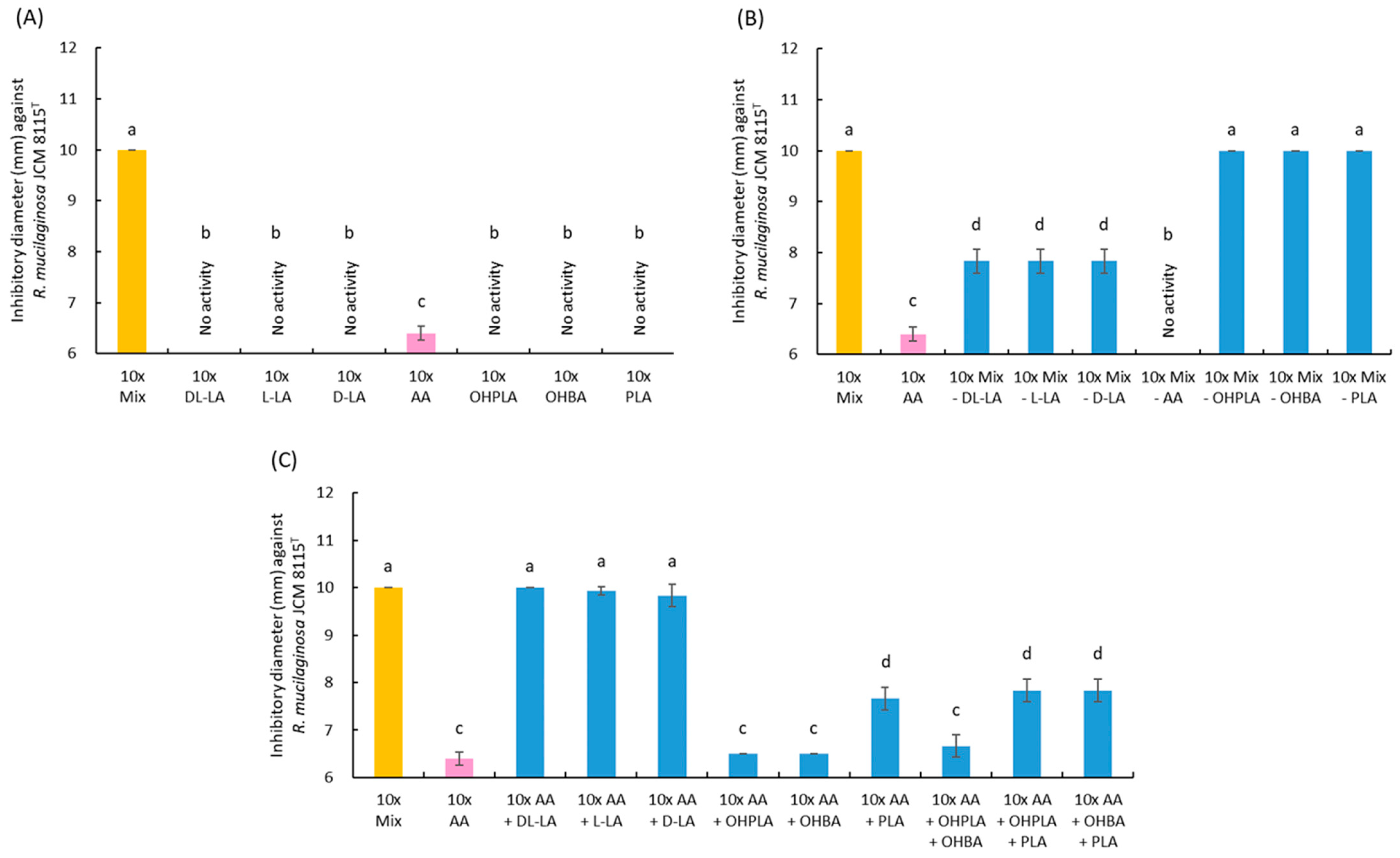
3.7. Antiyeast Activity of Organic Acids Produced by Strain 3121M0s
3.8. Comparison of Organic Acid Productivity and Antiyeast Activity Between Strains 3121M0s and JCM 1149T
3.9. Effects of L-Tyrosine and L-Phenylalanine on Organic Acid Productivity and Antiyeast Activity of Strain 3121M0s
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| LAB | Lactic acid bacteria |
| AA | Acetic acid |
| PLA | 3-Phenyllactic acid |
| MRS | De Man, Rogosa, and Sharp |
| YM | Yeast and Mold |
| RSM | Reconstituted skim milk |
| S.D. | Standard deviation |
| CFU | Colony forming unit |
| CFS | Cell-free culture supernatant |
| 10× CFS | Tenfold concentrated cell-free culture supernatant |
| OAs | Organic acids |
| rRNA | Ribosomal ribonucleic acid |
| PCR | Polymerase chain reaction |
| DNA | Deoxyribonucleic acid |
| BLAST | Basic Local Alignment Search Tool |
| NCBI | National Center for Biotechnology Information |
| DDBJ | DNA Data Bank Japan |
| OD | Optical density |
| HPLC | High performance liquid chromatography |
| OHBA | 4-Hydroxybenzoic acid |
| OHPLA | 4-hydroxyphenyllactic acid |
| LA | Lactic acid |
| MS | Mass spectrometry |
| FAB-MS | Fast atom bombardment mass spectrometry |
| EI-MS | Electron ionization mass spectrometry |
| Tyr | Tyrosine |
| Phe | Phenylalanine |
| ANOVA | Analysis of variance |
| NADH | Reduced nicotinamide adenine dinucleotide |
| CcpA | catabolite control protein A |
| ATP | Adenosine triphosphate |
References
- Snyder, A.B.; Worobo, R.W. Fungal Spoilage in Food Processing. J. Food Prot. 2018, 81, 1035–1040. [Google Scholar] [CrossRef] [PubMed]
- Gallo, M.; Ferrara, L.; Calogero, A.; Montesano, D.; Naviglio, D. Relationships between food and diseases: What to know to ensure food safety. Food Res. Int. 2020, 137, 109414. [Google Scholar] [CrossRef] [PubMed]
- Hymery, N.; Vasseur, V.; Coton, M.; Mounier, J.; Jany, J.-L.; Barbier, G.; Coton, E. Filamentous Fungi and Mycotoxins in Cheese: A Review. Compr. Rev. Food Sci. Food Saf. 2014, 13, 437–456. [Google Scholar] [CrossRef]
- Khan, R.; Anwar, F.; Mohamad Ghazali, F. A comprehensive review of mycotoxins: Toxicology, detection, and effective mitigation approaches. Heliyon 2024, 10, e28361. [Google Scholar] [CrossRef]
- Mafe, A.N.; Büsselberg, D. Mycotoxins in Food: Cancer Risks and Strategies for Control. Foods 2024, 13, 3502. [Google Scholar] [CrossRef]
- Bintsis, T. Yeasts in different types of cheese. AIMS Microbiol. 2021, 7, 447–470. [Google Scholar] [CrossRef]
- Libkind, D.; Peris, D.; Cubillos, F.A.; Steenwyk, J.L.; Opulente, D.A.; Langdon, Q.K.; Rokas, A.; Hittinger, C.T. Into the wild: New yeast genomes from natural environments and new tools for their analysis. FEMS Yeast Res. 2020, 20, foaa008. [Google Scholar] [CrossRef]
- Kovács, M.; Pomázi, A.; Taczman-Brückner, A.; Kiskó, G.; Dobó, V.; Kocsis, T.; Mohácsi-Farkas, C.; Belák, Á. Detection and Identification of Food-Borne Yeasts: An Overview of the Relevant Methods and Their Evolution. Microorganisms 2025, 13, 981. [Google Scholar] [CrossRef]
- Fleet, G.H. Yeasts in Dairy Products. J. Appl. Bacteriol. 1990, 68, 199–211. [Google Scholar] [CrossRef]
- Viljoen, B.C.; Greyling, T. Yeasts associated with Cheddar and Gouda making. Int. J. Food Microbiol. 1995, 28, 79–88. [Google Scholar] [CrossRef]
- Fleet, G.H. Yeasts in foods and beverages: Impact on product quality and safety. Curr. Opin. Biotechnol. 2007, 18, 170–175. [Google Scholar] [CrossRef]
- Sagdic, O.; Ozturk, I.; Bayram, O.; Kesmen, Z.; Yilmaz, M.T. Characterization of Butter Spoiling Yeasts and Their Inhibition by Some Spices. J. Food Sci. 2010, 75, M597–M603. [Google Scholar] [CrossRef]
- Melville, P.A.; Benites, N.R.; Ruz-Peres, M.; Yokoya, E. Proteinase and phospholipase activities and development at different temperatures of yeasts isolated from bovine milk. J. Dairy Res. 2011, 78, 385–390. [Google Scholar] [CrossRef] [PubMed]
- Moubasher, A.H.; Abdel-Sater, M.A.; Soliman, Z.S.M. Yeasts and filamentous fungi associated with some dairy products in Egypt. J. Mycol. Med. 2018, 28, 76–86. [Google Scholar] [CrossRef] [PubMed]
- Hernández, A.; Pérez-Nevado, F.; Ruiz-Moyano, S.; Serradilla, M.J.; Villalobos, M.C.; Martín, A.; Córdoba, M.G. Spoilage yeasts: What are the sources of contamination of foods and beverages? Int. J. Food Microbiol. 2018, 286, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Riešutė, R.; Šalomskienė, J.; Saez Moreno, D.; Gustienė, S. Effect of yeasts on food quality and safety and possibilities of their inhibition. Trends Food Sci. Technol. 2021, 108, 1–10. [Google Scholar] [CrossRef]
- Geronikou, A.; Srimahaeak, T.; Rantsiou, K.; Triantafillidis, G.; Larsen, N.; Jespersen, L. Occurrence of Yeasts in White-Brined Cheeses: Methodologies for Identification, Spoilage Potential and Good Manufacturing Practices. Front. Microbiol. 2020, 11, 582778. [Google Scholar] [CrossRef]
- Geronikou, A.; Larsen, N.; Lillevang, S.K.; Jespersen, L. Occurrence and Identification of Yeasts in Production of White-Brined Cheese. Microorganisms 2022, 10, 1079. [Google Scholar] [CrossRef]
- Geronikou, A.; Larsen, N.; Lillevang, S.K.; Jespersen, L. Diversity and succession of contaminating yeasts in white-brined cheese during cold storage. Food Microbiol. 2023, 113, 104266. [Google Scholar] [CrossRef]
- de Melo Pereira, G.V.; Maske, B.L.; de Carvalho Neto, D.P.; Karp, S.G.; De Dea Lindner, J.; Martin, J.G.P.; Hosken, B.O.; Soccol, C.R. What Is Candida Doing in My Food? A Review and Safety Alert on Its Use as Starter Cultures in Fermented Foods. Microorganisms 2022, 10, 1855. [Google Scholar] [CrossRef]
- Wirth, F.; Goldani, L.Z. Epidemiology of Rhodotorula: An emerging pathogen. Interdiscip. Perspect. Infect. Dis. 2012, 2012, 465717. [Google Scholar] [CrossRef]
- Enache-Angoulvant, A.; Hennequin, C. Invasive Saccharomyces infection: A comprehensive review. Clin. Infect. Dis. 2005, 41, 1559–1568. [Google Scholar] [CrossRef] [PubMed]
- Shen, D.; He, X.; Weng, P.; Liu, Y.; Wu, Z. A review of yeast: High cell-density culture, molecular mechanisms of stress response and tolerance during fermentation. FEMS Yeast Res. 2022, 22, foac050. [Google Scholar] [CrossRef]
- Rohm, H.; Lechner, F.; Lehner, M. Microflora of Austrian natural-set yogurt. J. Food Prot. 1990, 53, 478–480. [Google Scholar] [CrossRef] [PubMed]
- Ayoub, M.-J.; Bechara, P.; Habchi, M.; Hosry, R.; Akl, M.; Haj Hassan, S.; Abi Nakhoul, P. Raw goat’s milk fermented Anbaris from Lebanon: Insights into the microbial dynamics and chemical changes occurring during artisanal production, with a focus on yeasts. J. Dairy Res. 2022, 89, 440–448. [Google Scholar] [CrossRef] [PubMed]
- Karssa, T.H.; Kussaga, J.B.; Semedo-Lemsaddek, T.; Mugula, J.K. Insights on the microbiology of Ethiopian fermented milk products: A review. Food Sci. Nutr. 2024, 12, 6990–7003. [Google Scholar] [CrossRef]
- Chandran, A.; Beena, A.K. Impact of climate and traditional practice on quality of homemade dahi. J. Dairy Res. 2024, 91, 339–342. [Google Scholar] [CrossRef]
- Garnier, L.; Valence, F.; Pawtowski, A.; Auhustsinava-Galerne, L.; Frotté, N.; Baroncelli, R.; Deniel, F.; Coton, E.; Mounier, J. Diversity of spoilage fungi associated with various French dairy products. Int. J. Food Microbiol. 2017, 241, 191–197. [Google Scholar] [CrossRef]
- Bento de Carvalho, T.; Nunes Silva, B.; Tomé, E.; Teixeira, P. Preventing fungal spoilage from raw materials to final product: Innovative preservation techniques for fruit fillings. Foods 2024, 13, 2669. [Google Scholar] [CrossRef]
- Crowley, S.; Mahony, J.; van Sinderen, D. Current perspectives on antifungal lactic acid bacteria as natural bio-preservatives. Trends Food Sci. Technol. 2013, 33, 93–109. [Google Scholar] [CrossRef]
- Abouloifa, H.; Hasnaoui, I.; Rokni, Y.; Bellaouchi, R.; Ghabbour, N.; Karboune, S.; Brasca, M.; Abousalham, A.; Jaouadi, B.; Saalaoui, E.; et al. Chapter Two—Antifungal activity of lactic acid bacteria and their application in food biopreservation. Adv. Appl. Microbiol. 2022, 120, 33–77. [Google Scholar] [CrossRef] [PubMed]
- Nasrollahzadeh, A.; Mokhtari, S.; Khomeiri, M.; Saris, P.E.J. Antifungal preservation of food by lactic acid bacteria. Foods 2022, 11, 395. [Google Scholar] [CrossRef] [PubMed]
- Liang, N.; Zhao, Z.; Curtis, J.M.; Gänzle, M.G. Antifungal cultures and metabolites of lactic acid bacteria for use in dairy fermentations. Int. J. Food Microbiol. 2022, 383, 109938. [Google Scholar] [CrossRef]
- Ström, K.; Sjögren, J.; Broberg, A.; Schnürer, J. Lactobacillus plantarum MiLAB 393 produces the antifungal cyclic dipeptides cyclo(L-Phe-L-Pro) and cyclo(L-Phe-trans-4-OH-L-Pro) and 3-phenyllactic acid. Appl. Environ. Microbiol. 2002, 68, 4322–4327. [Google Scholar] [CrossRef]
- Yépez, A.; Luz, C.; Meca, G.; Vignolo, G.; Mañes, J.; Aznar, R. Biopreservation potential of lactic acid bacteria from Andean fermented food of vegetal origin. Food Control 2017, 78, 393–400. [Google Scholar] [CrossRef]
- Sjögren, J.; Magnusson, J.; Broberg, A.; Schnürer, J.; Kenne, L. Antifungal 3-hydroxy fatty acids from Lactobacillus plantarum MiLAB 14. Appl. Environ. Microbiol. 2003, 69, 7554–7557. [Google Scholar] [CrossRef]
- Vimont, A.; Fernandez, B.; Ahmed, G.; Pilote Fortin, H.; Fliss, I. Quantitative antifungal activity of reuterin against food isolates of yeasts and moulds and its potential application in yogurt. Int. J. Food Microbiol. 2019, 289, 182–188. [Google Scholar] [CrossRef]
- Zheng, X.; Nie, W.; Xu, J.; Zhang, H.; Liang, X.; Chen, Z. Characterization of antifungal cyclic dipeptides of Lacticaseibacillus paracasei ZX1231 and active packaging film prepared with its cell-free supernatant and bacterial nanocellulose. Food Res. Int. 2022, 162, 112024. [Google Scholar] [CrossRef]
- Belguesmia, Y.; Choiset, Y.; Rabesona, H.; Baudy-Floc’h, M.; Le Blay, G.; Haertlé, T.; Chobert, J.-M. Antifungal properties of durancins isolated from Enterococcus durans A5-11 and of its synthetic fragments. Lett. Appl. Microbiol. 2013, 56, 237–244. [Google Scholar] [CrossRef]
- Okkers, D.J.; Dicks, L.M.; Silvester, M.; Joubert, J.J.; Odendaal, H.J. Characterization of pentocin TV35b, a bacteriocin-like peptide isolated from Lactobacillus pentosus with a fungistatic effect on Candida albicans. J. Appl. Microbiol. 1999, 87, 726–734. [Google Scholar] [CrossRef]
- Yang, E.; Fan, L.; Jiang, Y.; Doucette, C.; Fillmore, S. Antimicrobial activity of bacteriocin-producing lactic acid bacteria isolated from cheeses and yogurts. AMB Express 2012, 2, 48. [Google Scholar] [CrossRef]
- Magnusson, J.; Schnürer, J. Lactobacillus coryniformis subsp. coryniformis strain Si3 produces a broad-spectrum proteinaceous antifungal compound. Appl. Environ. Microbiol. 2001, 67, 1–5. [Google Scholar] [CrossRef]
- Watanabe, K.; Fujimoto, J.; Sasamoto, M.; Dugersuren, J.; Tumursuh, T.; Demberel, S. Diversity of lactic acid bacteria and yeasts in Airag and Tarag, traditional fermented milk products of Mongolia. World J. Microbiol. Biotechnol. 2008, 24, 1313–1325. [Google Scholar] [CrossRef]
- Sun, Z.H.; Liu, W.J.; Zhang, J.C.; Yu, J.; Gao, W.; Jiri, M.; Menghe, B.; Sun, T.S.; Zhang, H.P. Identification and characterization of the dominant lactic acid bacteria isolated from traditional fermented milk in Mongolia. Folia Microbiol. 2010, 55, 270–276. [Google Scholar] [CrossRef]
- Takeda, S.; Yamasaki, K.; Takeshita, M.; Kikuchi, Y.; Tsend-Ayush, C.; Dashnyam, B.; Ahhmed, A.M.; Kawahara, S.; Muguruma, M. The investigation of probiotic potential of lactic acid bacteria isolated from traditional Mongolian dairy products. Anim. Sci. J. 2011, 82, 571–579. [Google Scholar] [CrossRef]
- Choi, S.-H. Characterization of airag collected in Ulaanbaatar, Mongolia with emphasis on isolated lactic acid bacteria. J. Anim. Sci. Technol. 2016, 58, 10. [Google Scholar] [CrossRef] [PubMed]
- Oyundelger, G.; Sumisa, F.; Batjargal, B.; Yoshida, T. Isolation and identification of new lactic acid bacteria with potent biological activity and yeasts in Airag, a traditional Mongolian fermented beverage. Food Sci. Technol. Res. 2016, 22, 575–582. [Google Scholar] [CrossRef]
- Oki, K.; Dugersuren, J.; Demberel, S.; Watanabe, K. Pyrosequencing analysis of the microbial diversity of airag, khoormog and tarag, traditional fermented dairy products of Mongolia. Biosci. Microbiota Food Health 2014, 33, 53–64. [Google Scholar] [CrossRef]
- Shinoda, A.; Koga, Y.; Tsuchiya, R.; Tserenpurev, B.-O.; Battsetseg, B.; Morinaga, Y.; Nakayama, J. Impact of container type on the microbiome of airag, a Mongolian fermented mare’s milk. Biosci. Microbiota Food Health 2025, 44, 90–99. [Google Scholar] [CrossRef]
- Sudun; Wulijideligen; Arakawa, K.; Miyamoto, M.; Miyamoto, T. Interaction between lactic acid bacteria and yeasts in airag, an alcoholic fermented milk. Anim. Sci. J. 2013, 84, 66–74. [Google Scholar] [CrossRef]
- Miyamoto, M.; Ueno, H.M.; Watanabe, M.; Tatsuma, Y.; Seto, Y.; Miyamoto, T.; Nakajima, H. Distinctive proteolytic activity of cell envelope proteinase of Lactobacillus helveticus isolated from airag, a traditional Mongolian fermented mare’s milk. Int. J. Food Microbiol. 2015, 197, 65–71. [Google Scholar] [CrossRef]
- Ganzorig, B.; Zayabaatar, E.; Pham, M.T.; Marito, S.; Huang, C.-M.; Lee, Y.-H. Lactobacillus plantarum Generate Eelectricity through Flavin Mononucleotide-Mediated Extracellular Electron Transfer to Upregulate Epithelial Type I Collagen Expression and Thereby Promote Microbial Adhesion to Intestine. Biomedicines 2023, 11, 667. [Google Scholar] [CrossRef]
- Galindev, U.; Erdenebold, U.; Batnasan, G.; Ganzorig, O.; Batdorj, B. Anti-obesity effects of potential probiotic Lactobacillus strains isolated from Mongolian fermented dairy products in high-fat diet-induced obese rodent model. Braz. J. Microbiol. 2024, 55, 2501–2509. [Google Scholar] [CrossRef]
- Batdorj, B.; Dalgalarrondo, M.; Choiset, Y.; Pedroche, J.; Métro, F.; Prévost, H.; Chobert, J.-M.; Haertlé, T. Purification and characterization of two bacteriocins produced by lactic acid bacteria isolated from Mongolian airag. J. Appl. Microbiol. 2006, 101, 837–848. [Google Scholar] [CrossRef] [PubMed]
- Wulijideligen; Asahina, T.; Hara, K.; Arakawa, K.; Nakano, H.; Miyamoto, T. Production of bacteriocin by Leuconostoc mesenteroides 406 isolated from Mongolian fermented mare’s milk, airag. Anim. Sci. J. 2012, 83, 704–711. [Google Scholar] [CrossRef] [PubMed]
- Arakawa, K.; Yoshida, S.; Aikawa, H.; Hano, C.; Bolormaa, T.; Burenjargal, S.; Miyamoto, T. Production of a bacteriocin-like inhibitory substance by Leuconostoc mesenteroides subsp. dextranicum 213M0 isolated from Mongolian fermented mare milk, airag. Anim. Sci. J. 2016, 87, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Hasiqimuge; Hano, C.; Arakawa, K.; Yoshida, S.; Zhao, J.; Toh, H.; Morita, H.; Miyamoto, T. A novel C-terminal truncated bacteriocin found by comparison between Leuconostoc mesenteroides 406 and 213M0 isolated from Mongolian traditional fermented milk, airag. Microorganisms 2024, 12, 1781. [Google Scholar] [CrossRef]
- Arakawa, K.; Kawai, Y.; Iioka, H.; Tanioka, M.; Nishimura, J.; Kitazawa, H.; Koga, M.; Saito, T.; Itoh, T. Microbial Community Analysis of Food-Spoilage Bacteria in Commercial Custard Creams Using Culture-Dependent and Independent Methods. J. Dairy Sci. 2008, 91, 2938–2946. [Google Scholar] [CrossRef]
- Klaenhammer, T.R. Genetics of Bacteriocins Produced by Lactic Acid Bacteria. FEMS Microbiol. Rev. 1993, 12, 39–85. [Google Scholar] [CrossRef]
- Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 2000, 17, 540–552. [Google Scholar] [CrossRef]
- Torriani, S.; Felis, G.E.; Dellaglio, F. Differentiation of Lactobacillus plantarum, L. pentosus, and L. paraplantarum by recA Gene Sequence Analysis and Multiplex PCR Assay with recA Gene-Derived Primers. Appl. Environ. Microbiol. 2001, 67, 3450–3454. [Google Scholar] [CrossRef]
- Valerio, F.; Lavermicocca, P.; Pascale, M.; Visconti, A. Production of Phenyllactic Acid by Lactic Acid Bacteria: An Approach to the Selection of Strains Contributing to Food Quality and Preservation. FEMS Microbiol. Lett. 2004, 233, 289–295. [Google Scholar] [CrossRef]
- Crowley, S.; Mahony, J.; van Sinderen, D. Comparative analysis of two antifungal Lactobacillus plantarum isolates and their application as bioprotectants in refrigerated foods. J. Appl. Microbiol. 2012, 113, 1417–1427. [Google Scholar] [CrossRef] [PubMed]
- Leyva Salas, M.; Thierry, A.; Lemaître, M.; Garric, G.; Harel-Oger, M.; Chatel, M.; Lê, S.; Mounier, J.; Valence, F.; Coton, E. Antifungal Activity of Lactic Acid Bacteria Combinations in Dairy Mimicking Models and Their Potential as Bioprotective Cultures in Pilot Scale Applications. Front. Microbiol. 2018, 9, 1787. [Google Scholar] [CrossRef] [PubMed]
- Afzali, S.; Edalatian Dovom, M.R.; Habibi Najafi, M.B.; Mazaheri Tehrani, M. Determination of the anti-yeast activity of Lactobacillus spp. isolated from traditional Iranian cheeses in vitro and in yogurt drink (Doogh). Sci. Rep. 2020, 10, 6291. [Google Scholar] [CrossRef] [PubMed]
- Makki, G.M.; Kozak, S.M.; Jencarelli, K.G.; Alcaine, S.D. Evaluation of the efficacy of commercial protective cultures against mold and yeast in queso fresco. J. Dairy Sci. 2020, 103, 9946–9957. [Google Scholar] [CrossRef]
- Makki, G.M.; Kozak, S.M.; Jencarelli, K.G.; Alcaine, S.D. Evaluation of the efficacy of commercial protective cultures to inhibit mold and yeast in cottage cheese. J. Dairy Sci. 2021, 104, 2709–2718. [Google Scholar] [CrossRef]
- de Vries, M.C.; Vaughan, E.E.; Kleerebezem, M.; de Vos, W.M. Lactobacillus plantarum—Survival, functional and potential probiotic properties in the human intestinal tract. Int. Dairy J. 2006, 16, 1018–1028. [Google Scholar] [CrossRef]
- Siezen, R.J.; van Hylckama Vlieg, J.E.T. Genomic diversity and versatility of Lactobacillus plantarum, a natural metabolic engineer. Microb. Cell Fact. 2011, 10, S3. [Google Scholar] [CrossRef]
- Fidanza, M.; Panigrahi, P.; Kollmann, T.R. Lactiplantibacillus plantarum—Nomad and Ideal Probiotic. Front. Microbiol. 2021, 12, 712236. [Google Scholar] [CrossRef]
- Yilmaz, B.; Bangar, S.P.; Echegaray, N.; Suri, S.; Tomasevic, I.; Lorenzo, J.M.; Melekoglu, E.; Rocha, J.M.; Ozogul, F. The Impacts of Lactiplantibacillus plantarum on the Functional Properties of Fermented Foods: A Review of Current Knowledge. Microorganisms 2022, 10, 826. [Google Scholar] [CrossRef]
- Bintsis, T.; Papademas, P. The Evolution of Fermented Milks, from Artisanal to Industrial Products: A Critical Review. Fermentation 2022, 8, 679. [Google Scholar] [CrossRef]
- Ganzorig, O.; Batdorj, B.; Ishii, S. Characterization of volatile compound profile in Mongolian traditional fermented mare’s milk, as Airag. Anim. Sci. J. 2025, 96, e70024. [Google Scholar] [CrossRef]
- Mbandi, E.; Shelef, L.A. Enhanced inhibition of Listeria monocytogenes and Salmonella enteritidis in meat by combinations of sodium lactate and diacetate. J. Food Prot. 2001, 64, 640–644. [Google Scholar] [CrossRef] [PubMed]
- Ji, Q.-Y.; Wang, W.; Yan, H.; Qu, H.; Liu, Y.; Qian, Y.; Gu, R. The Effect of Different Organic Acids and Their Combination on the Cell Barrier and Biofilm of Escherichia coli. Foods 2023, 12, 3011. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Wang, Y.; Guo, M.; Kong, Y.; Fan, X.; Sun, S.; Du, C.; Gong, H. Antifungal mechanisms of phenyllactic acid against Mucor racemosus: Insights from spore growth suppression and proteomic analysis. Food Chem. 2025, 475, 143309. [Google Scholar] [CrossRef]
- Lorquet, F.; Goffin, P.; Muscariello, L.; Baudry, J.-B.; Ladero, V.; Sacco, M.; Kleerebezem, M.; Hols, P. Characterization and functional analysis of the poxB gene, which encodes pyruvate oxidase in Lactobacillus plantarum. J. Bacteriol. 2004, 186, 3749–3759. [Google Scholar] [CrossRef]
- Goffin, P.; Lorquet, F.; Kleerebezem, M.; Hols, P. Major Role of NAD-Dependent Lactate Dehydrogenases in Aerobic Lactate Utilization in Lactobacillus plantarum during Early Stationary Phase. J. Bacteriol. 2004, 186, 6661–6666. [Google Scholar] [CrossRef]
- Zotta, T.; Parente, E.; Ricciardi, A. Aerobic Metabolism in the Genus Lactobacillus: Impact on Stress Response and Potential Applications in the Food Industry. J. Appl. Microbiol. 2017, 122, 857–869. [Google Scholar] [CrossRef]
- Lu, Y.; Song, S.; Tian, H.; Yu, H.; Zhao, J.; Chen, C. Functional Analysis of the Role of CcpA in Lactobacillus plantarum Grown on Fructooligosaccharides or Glucose: A Transcriptomic Perspective. Microb. Cell Fact. 2018, 17, 201. [Google Scholar] [CrossRef]
- Dallagnol, A.M.; Catalán, C.A.N.; Mercado, M.I.; Font de Valdez, G.; Rollán, G.C. Effect of biosynthetic intermediates and citrate on the phenyllactic and hydroxyphenyllactic acids production by Lactobacillus plantarum CRL 778. J. Appl. Microbiol. 2011, 111, 1447–1455. [Google Scholar] [CrossRef]
- Li, M.; Meng, X.; Sun, Z.; Zhu, C.; Ji, H. Effects of NADH availability on 3-phenyllactic acid production by Lactobacillus plantarum expressing formate dehydrogenase. Curr. Microbiol. 2019, 76, 706–712. [Google Scholar] [CrossRef]
- Dao, Y.; Zhang, K.; Lu, X.; Lu, Z.; Liu, C.; Liu, M.; Luo, Y. Role of glucose and 2-oxoglutarate/malate translocator (OMT1) in the production of phenyllactic acid and p-hydroxyphenyllactic acid, two food-borne pathogen inhibitors. J. Agric. Food Chem. 2019, 67, 5820–5826. [Google Scholar] [CrossRef] [PubMed]
- Abouloifa, H.; Rokni, Y.; Hasnaoui, I.; Bellaouchi, R.; Gaamouche, S.; Ghabbour, N.; Karboune, S.; Ben Salah, R.; Brasca, M.; D’hallewin, G.; et al. Characterization of antimicrobial compounds obtained from the potential probiotic Lactiplantibacillus plantarum S61 and their application as a biopreservative agent. Braz. J. Microbiol. 2022, 53, 1501–1513. [Google Scholar] [CrossRef]
- Plumed-Ferrer, C.; Koistinen, K.M.; Tolonen, T.L.; Lehesranta, S.J.; Kärenlampi, S.O.; Mäkimattila, E.; Joutsjoki, V.; Virtanen, V.; von Wright, A. Comparative study of sugar fermentation and protein expression patterns of two Lactobacillus plantarum strains grown in three different media. Appl. Environ. Microbiol. 2008, 74, 5349–5358. [Google Scholar] [CrossRef]

| Strain | Overlay Assay * Against | Agar–Well Diffusion Assay ** Against | |||||
|---|---|---|---|---|---|---|---|
| Rhodotorula mucilaginosa | Candida parapsilosis | Saccharomyces cerevisiae | Rhodotorula mucilaginosa | Candida parapsilosis | Saccharomyces cerevisiae | ||
| JCM 8115T | JCM 1612T | 4C | JCM 8115T | JCM 1612T | 4C | ||
| 3121M0s | +++ | ++ | - | 10.63 ± 0.45 | - | - | |
| 3272B0 | +++ | ++ | - | - | - | - | |
| 3273B0 | +++ | ++ | - | - | - | - | |
| 3461M0 | +++ | ++ | - | - | - | - | |
| 123 | +++ | ± | - | N.D. | N.D. | N.D. | |
| IB3781 | +++ | ± | - | N.D. | N.D. | N.D. | |
| NMCB2521 | +++ | ± | - | N.D. | N.D. | N.D. | |
| NMCM2521 | +++ | ± | - | N.D. | N.D. | N.D. | |
| 3070 | +++ | - | - | N.D. | N.D. | N.D. | |
| 301102S | +++ | - | - | N.D. | N.D. | N.D. | |
| 425 | +++ | - | - | N.D. | N.D. | N.D. | |
| IM376 | ++ | + | - | N.D. | N.D. | N.D. | |
| IB374 | + | + | - | N.D. | N.D. | N.D. | |
| JCM 1120T | - | + | - | N.D. | N.D. | N.D. | |
| 2253RB0 | - | + | - | N.D. | N.D. | N.D. | |
| 5890 | - | - | ± | N.D. | N.D. | N.D. | |
| 50 strains | ++ | - | - | N.D. | N.D. | N.D. | |
| 77 strains | + | - | - | N.D. | N.D. | N.D. | |
| 93 strains | - | - | - | N.D. | N.D. | N.D. | |
| Assay | Debaryomyces hansenii | Hanseniaspora uvarum | Issatchenkia orientalis | Kluyveromyces marxianus | Pichia kudriavzevii | Saccharomyces cerevisiae | Wickerhamomyces anomalus | Zygosaccharomyces rouxii |
|---|---|---|---|---|---|---|---|---|
| JCM 1990T | JCM 2184T | JCM 1710T | JCM 1617T | JCM 3536T | JCM 7255T | JCM 3585T | JCM 2325T | |
| Overlay * | ++ | + | - | + | - | + | - | + |
| Agar–well diffusion ** | 10.83 ± 0.24 | 8.67 ± 0.24 | - | 7.00 ± 0.00 | - | - | - | 7.33 ± 0.24 |
| Ingredient | Concentration (mg/L) | |
|---|---|---|
| 3121M0s-CFS | JCM 1149T-CFS | |
| DL-Lactic acid | 16,053.0 | 16,244.4 |
| (D-form:L-form) | (10,981.0:5072.0) | (11,571.9:4672.5) |
| Acetic acid | 1437.9 | 902.2 |
| 4-Hydroxyphenyllactic acid | 28.7 | 3.0 |
| 4-Hydroxybenzoic acid | 26.9 | 34.4 |
| 3-Phenyllactic acid | 99.1 | 69.2 |
| Solution | Antiyeast Activity (mm; Mean ± S.D.) * | ||
|---|---|---|---|
| 3121M0s | JCM 1149T | MRS Broth | |
| Tenfold culture supernatant ** | 10.63 ± 0.45 a | 7.00 ± 0.00 c | - d |
| Tenfold organic acids mixture *** | 10.00 ± 0.00 b | 6.83 ± 0.24 c | N.D. |
| Additive to MRS Broth | Concentration (mg/L) in 3121M0s-CFS | Antiyeast Activity (mm; Mean ± S.D.) of 10-Fold 3121M0s-CFS (pH 4.0) * | |
|---|---|---|---|
| 4-Hydroxyphenyllactic Acid | 3-Phenyllactic Acid | ||
| No added | 28.7 | 99.1 | 10.63 ± 0.45 a |
| L-Tyrosine | 125.4 | Trace | 11.00 ± 0.00 a |
| L-Phenylalanine | Trace | 126.5 | 11.00 ± 0.00 a |
| L-Tyrosin and L-phenylalanine | 84.9 | 88.9 | 11.00 ± 0.00 a |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lijon, M.B.; Matsu-ura, Y.; Ukita, T.; Arakawa, K.; Miyamoto, T. Identification and Characterization of Antiyeast Organic Acids Produced by Lactiplantibacillus plantarum 3121M0s Derived from Mongolian Traditional Fermented Milk, Airag. Microorganisms 2025, 13, 2017. https://doi.org/10.3390/microorganisms13092017
Lijon MB, Matsu-ura Y, Ukita T, Arakawa K, Miyamoto T. Identification and Characterization of Antiyeast Organic Acids Produced by Lactiplantibacillus plantarum 3121M0s Derived from Mongolian Traditional Fermented Milk, Airag. Microorganisms. 2025; 13(9):2017. https://doi.org/10.3390/microorganisms13092017
Chicago/Turabian StyleLijon, Md. Bakhtiar, Yuko Matsu-ura, Takumi Ukita, Kensuke Arakawa, and Taku Miyamoto. 2025. "Identification and Characterization of Antiyeast Organic Acids Produced by Lactiplantibacillus plantarum 3121M0s Derived from Mongolian Traditional Fermented Milk, Airag" Microorganisms 13, no. 9: 2017. https://doi.org/10.3390/microorganisms13092017
APA StyleLijon, M. B., Matsu-ura, Y., Ukita, T., Arakawa, K., & Miyamoto, T. (2025). Identification and Characterization of Antiyeast Organic Acids Produced by Lactiplantibacillus plantarum 3121M0s Derived from Mongolian Traditional Fermented Milk, Airag. Microorganisms, 13(9), 2017. https://doi.org/10.3390/microorganisms13092017






