Personalized Response of Parkinson’s Disease Gut Microbiota to Nootropic Medicinal Herbs In Vitro: A Proof of Concept
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Participants and Sample Collection
2.2. Nervine Medicinal Herbs Examined in the Current Microbiome Study
2.3. Anaerobic Fecal Cultures
2.4. Chemically Defined Medium
2.5. Microbial DNA Isolation
2.6. 16S rRNA Sequence Analysis
2.7. Genome-Wide Metabolic Reconstruction
3. Results
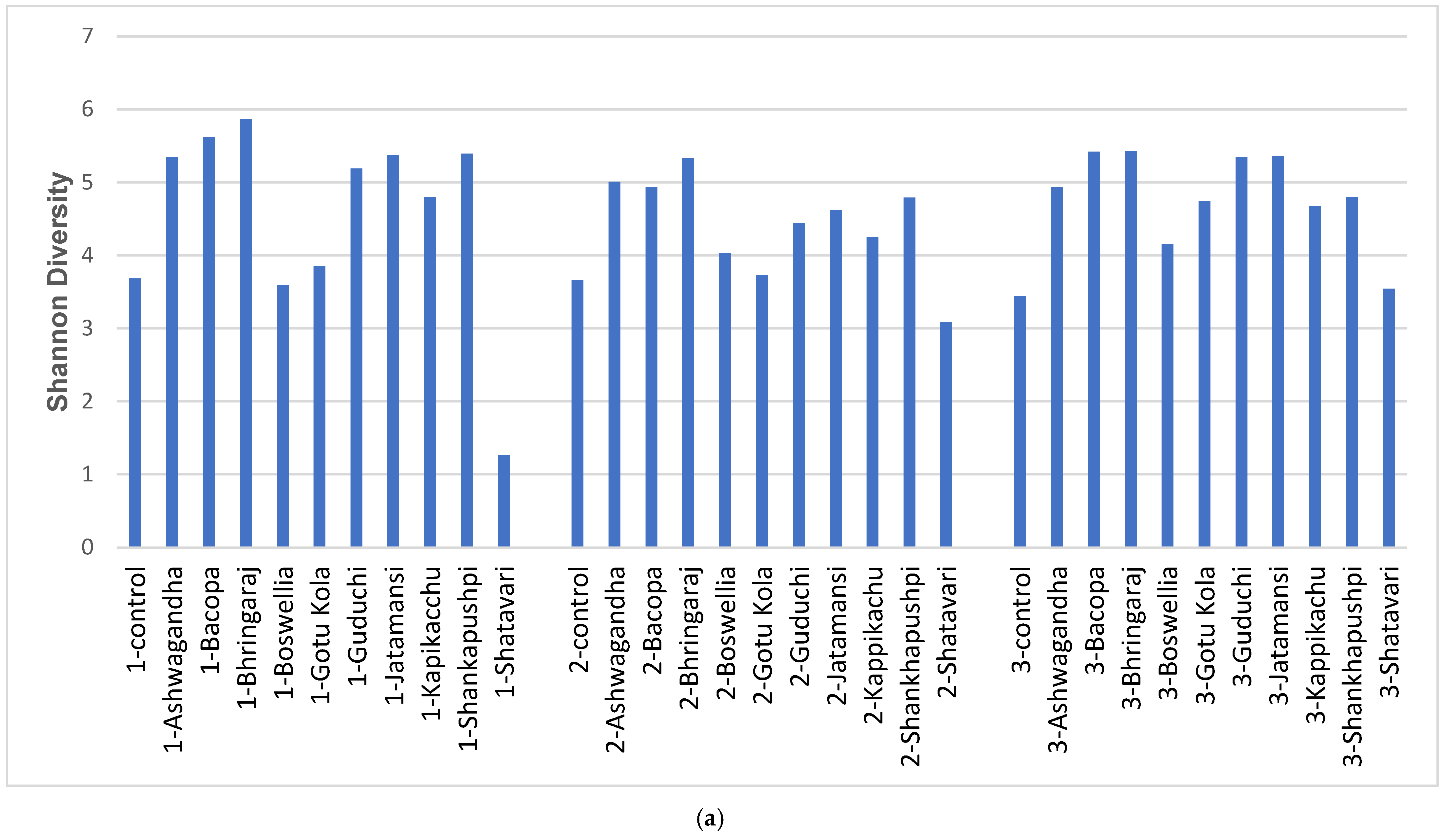
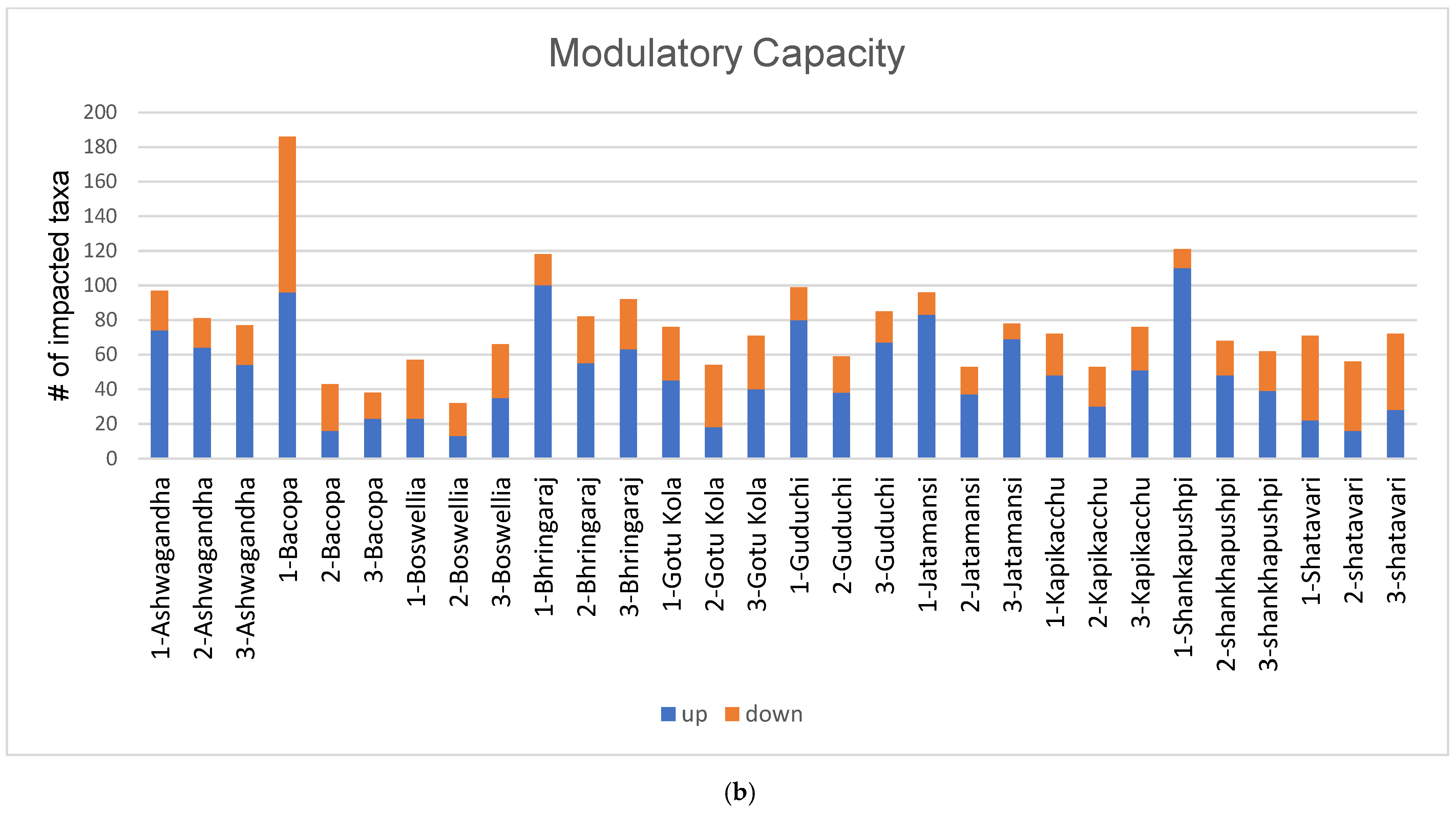
3.1. Diversity and Modulatory Effects of Medicinal Herbs
3.2. Phylogenetic Analysis and Inter-Personal Variable Responses to Nootropic Herbs
3.3. Nootropic Herb Supplementation Corrects Genus-Level Dysbioses Associated with PD
3.4. Interpersonal Variability of PD Microbiota to Medicinal Herbs
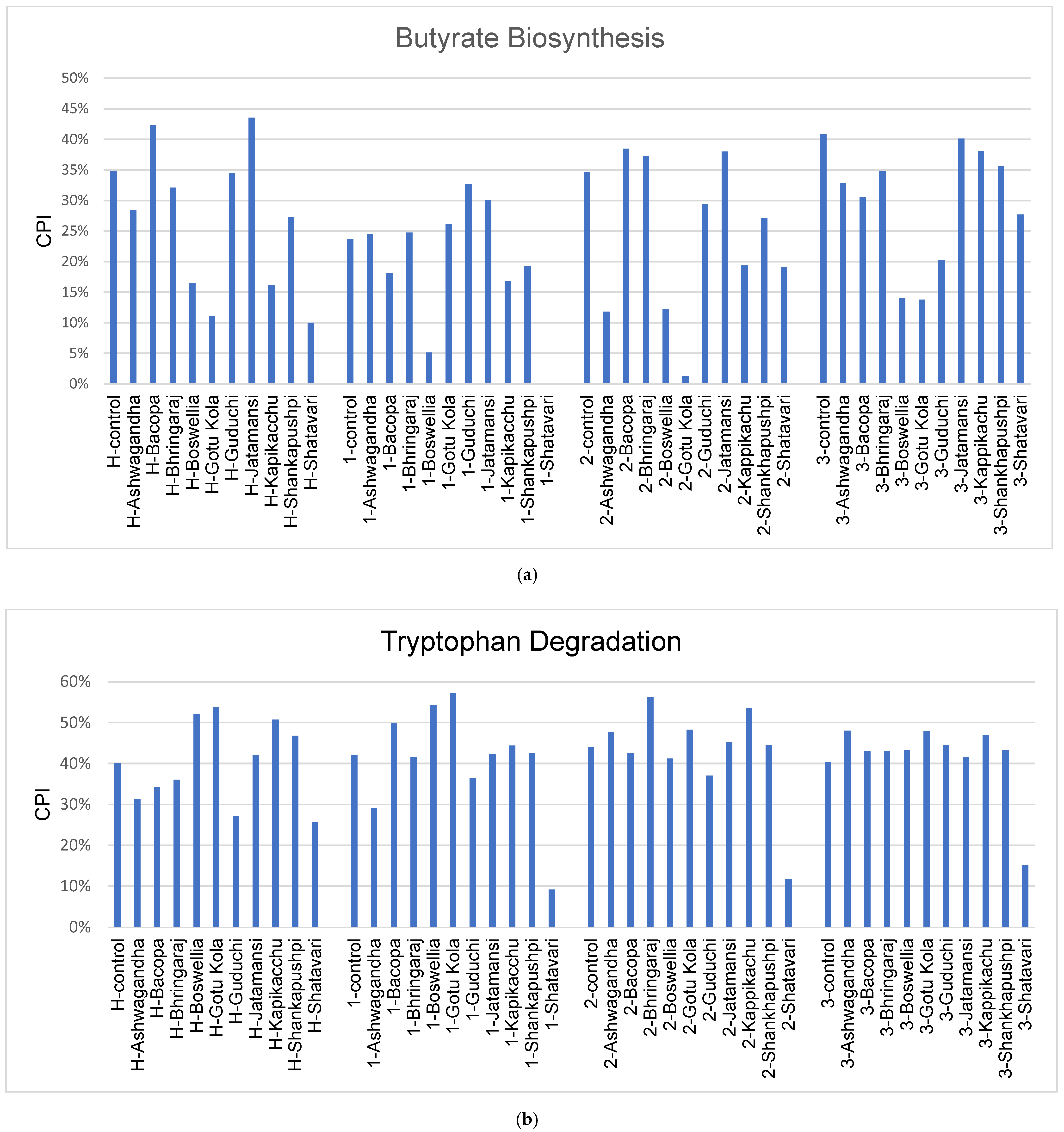
3.5. Gut Permeability and PD Microbiome
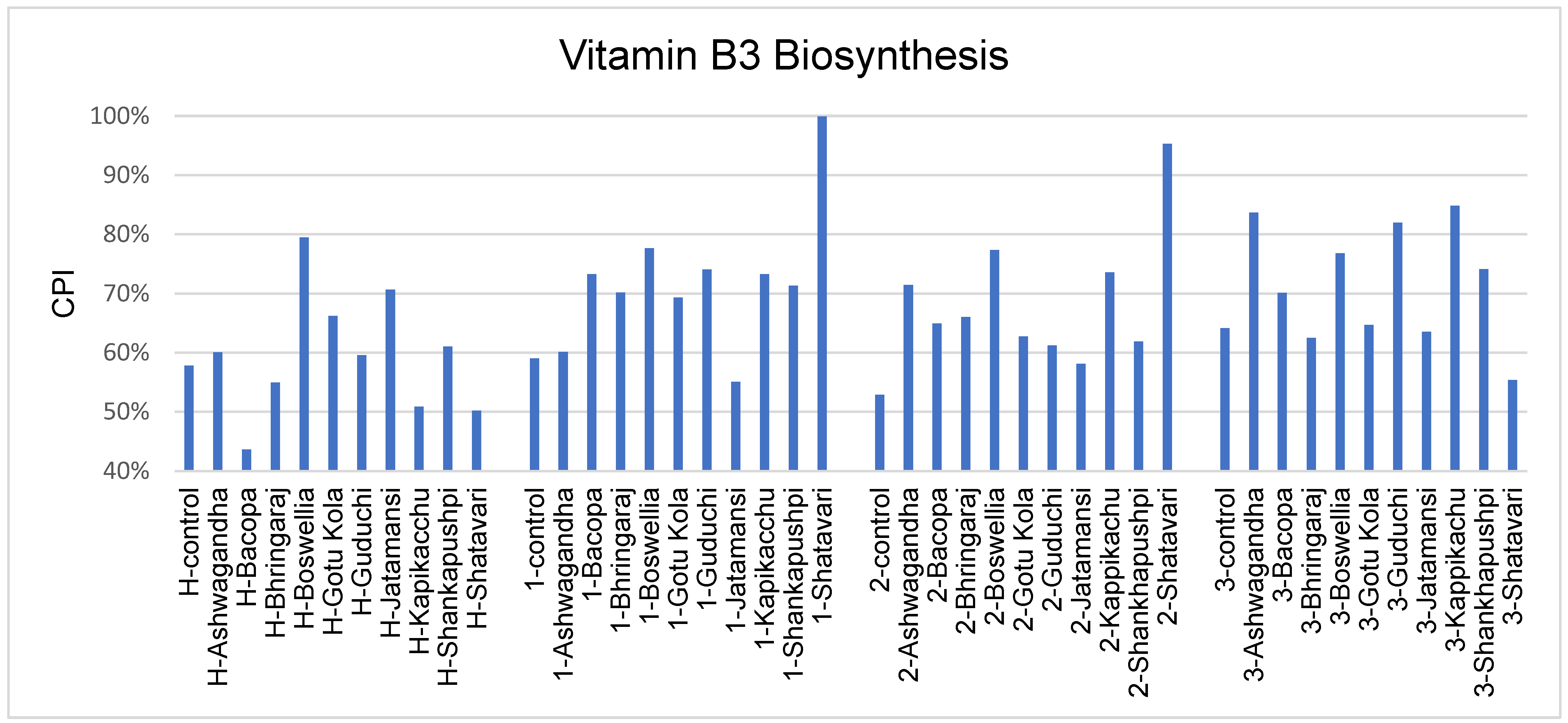
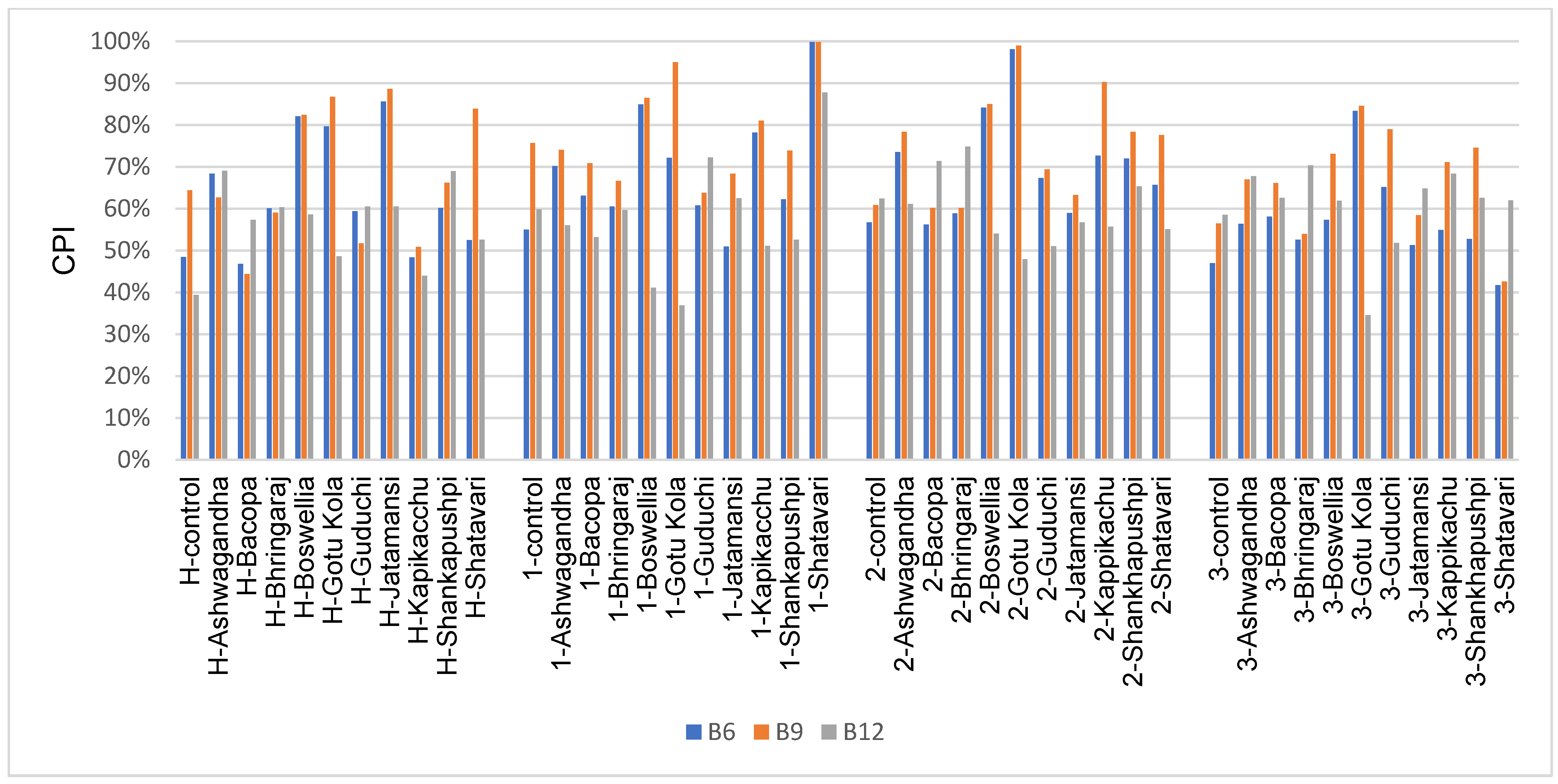
3.6. Vitamin Biosynthesis Potential
4. Discussion
4.1. Diversity and Modulatory Effects of Medicinal Herbs
4.2. Nootropic Herb Supplementation Corrects Genus-Level Dysbioses Associated with PD
4.3. Gut Permeability and PD Microbiome
4.4. Vitamin Biosynthesis Potential
4.5. Limitations and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Scheperjans, F.; Aho, V.; Pereira, P.A.B.; Koskinen, K.; Paulin, L.; Pekkonen, E.; Haapaniemi, E.; Kaakkola, S.; Eerola-Rautio, J.; Pohja, M.; et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov. Disord. 2015, 30, 350–358. [Google Scholar] [CrossRef]
- Unger, M.M.; Spiegel, J.; Dillmann, K.-U.; Grundmann, D.; Philippeit, H.; Bürmann, J.; Faßbender, K.; Schwiertz, A.; Schäfer, K.H. Short chain fatty acids and gut microbiota differ between patients with Parkinson’s disease and age-matched controls. Parkinsonism Relat. Disord. 2016, 32, 66–72. [Google Scholar] [CrossRef] [PubMed]
- Petrov, V.A.; Saltykova, I.V.; Zhukova, I.A.; Alifirova, V.M.; Zhukova, N.G.; Dorofeeva, Y.B.; Tyakht, A.V.; Kovarsky, B.A.; Alekseev, D.G.; Kostryukova, E.S.; et al. Analysis of Gut Microbiota in Patients with Parkinson’s Disease. Bull. Exp. Biol. Med. 2017, 162, 734–737. [Google Scholar] [CrossRef]
- Minato, T.; Maeda, T.; Fujisawa, Y.; Tsuji, H.; Nomoto, K.; Ohno, K.; Hirayama, M. Progression of Parkinson’s disease is associated with gut dysbiosis: Two-year follow-up study. PLoS ONE 2017, 12, e0187307. [Google Scholar] [CrossRef]
- Felice, V.D.; Quigley, E.M.; Sullivan, A.M.; O’Keeffe, G.W.; O’Mahony, S.M. Microbiota-gut-brain signalling in Parkinson’s disease: Implications for non-motor symptoms. Park. Relat. Disord. 2016, 27, 1–8. [Google Scholar] [CrossRef]
- Li, S.S.; Zhu, A.; Benes, V.; Costea, P.I.; Hercog, R.; Hildebrand, F.; Huerta-Cepas, J.; Nieuwdorp, M.; Salojärvi, J.; Voigt, A.Y.; et al. Durable coexistence of donor and recipient strains after fecal microbiota transplantation. Science 2016, 352, 586–589. [Google Scholar] [CrossRef]
- Chen, T.; Noto, D.; Hoshino, Y.; Mizuno, M.; Miyake, S. Butyrate suppresses demyelination and enhances remyelination. J. Neuroinflammation 2019, 16, 165. [Google Scholar] [CrossRef]
- Srivastav, S.; Neupane, S.; Bhurtel, S.; Katila, N.; Maharjan, S.; Choi, H.; Hong, J.T.; Choi, D.-Y. Probiotics mixture increases butyrate, and subsequently rescues the nigral dopaminergic neurons from MPTP and rotenone-induced neurotoxicity. J. Nutr. Biochem. 2019, 69, 73–86. [Google Scholar] [CrossRef]
- Perez-Pardo, P.; Kliest, T.; Dodiya, H.B.; Broersen, L.M.; Garssen, J.; Keshavarzian, A.; Kraneveld, A.D. The gut-brain axis in Parkinson’s disease: Possibilities for food-based therapies. Eur. J. Pharmacol. 2017, 817, 86–95. [Google Scholar] [CrossRef] [PubMed]
- Hill-Burns, E.M.; Debelius, J.W.; Morton, J.T.; Wissemann, W.T.; Lewis, M.R.; Wallen, Z.D.; Peddada, S.D.; Factor, S.A.; Molho, E.; Zabetian, C.P.; et al. Parkinson’s disease and Parkinson’s disease medications have distinct signatures of the gut microbiome. Mov. Disord. 2017, 32, 739–749. [Google Scholar] [CrossRef]
- Keshavarzian, A.; Green, S.J.; Engen, P.A.; Voigt, R.M.; Naqib, A.; Forsyth, C.B.; Mutlu, E.; Shannon, K.M. Colonic bacterial composition in Parkinson’s disease. Mov. Disord. 2015, 30, 1351–1360. [Google Scholar] [CrossRef]
- Aho, V.T.E.; Houser, M.C.; Pereira, P.A.B.; Chang, J.; Rudi, K.; Paulin, L.; Hertzberg, V.; Auvinen, P.; Tansey, M.G.; Scheperjans, F. Relationships of gut microbiota, short-chain fatty acids, inflammation, and the gut barrier in Parkinson’s disease. Mol. Neurodegener. 2021, 16, 6. [Google Scholar] [CrossRef]
- Wallen, Z.D.; Demirkan, A.; Twa, G.; Cohen, G.; Dean, M.N.; Standaert, D.G.; Sampson, T.R.; Payami, H. Metagenomics of Parkinson’s disease implicates the gut microbiome in multiple disease mechanisms. Nat. Commun. 2022, 13, 6958. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.H.; Chen, C.C.; Chiang, H.L.; Liou, J.M.; Chang, C.M.; Lu, T.P.; Chuang, E.Y.; Tai, Y.C.; Cheng, C.; Lin, H.Y.; et al. Altered gut microbiota and inflammatory cytokine responses in patients with Parkinson’s disease. J. Neuroinflammation 2019, 16, 129. [Google Scholar] [CrossRef] [PubMed]
- Cirstea, M.S.; Yu, A.C.; Golz, E.; Sundvick, K.; Kliger, D.; Radisavljevic, N.; Foulger, L.H.; Mackenzie, M.; Huan, T.; Finlay, B.; et al. Microbiota Composition and Metabolism Are Associated with Gut Function in Parkinson’s Disease. Mov. Disord. 2020, 35, 1208–1217. [Google Scholar] [CrossRef]
- Yan, Z.; Yang, F.; Cao, J.; Ding, W.; Yan, S.; Shi, W.; Wen, S.; Yao, L. Alterations of gut microbiota and metabolome with Parkinson’s disease. Microb. Pathog. 2021, 160, 105187. [Google Scholar] [CrossRef] [PubMed]
- Angot, E.; Brundin, P. Dissecting the potential molecular mechanisms underlying α-synuclein cell-to-cell transfer in Parkinson’s disease. Park. Relat. Disord. 2009, 15 (Suppl. S3), S143–S147. [Google Scholar] [CrossRef]
- Braak, H.; Rüb, U.; Gai, W.P.; Del Tredici, K. Idiopathic Parkinson’s disease: Possible routes by which vulnerable neuronal types may be subject to neuroinvasion by an unknown pathogen. J. Neural. Transm. 2003, 110, 517–536. [Google Scholar] [CrossRef]
- Sampson, T.R.; Debelius, J.W.; Thron, T.; Janssen, S.; Shastri, G.G.; Ilhan, Z.E.; Challis, C.; Schretter, C.E.; Rocha, S.; Gradinaru, V.; et al. Gut Microbiota Regulate Motor Deficits and Neuroinflammation in a Model of Parkinson’s Disease. Cell 2016, 167, 1469–1480.e1412. [Google Scholar] [CrossRef] [PubMed]
- Schwiertz, A.; Spiegel, J.; Dillmann, U.; Grundmann, D.; Bürmann, J.; Faßbender, K.; Schäfer, K.-H.; Unger, M.M. Fecal markers of intestinal inflammation and intestinal permeability are elevated in Parkinson’s disease. Park. Relat. Disord. 2018, 50, 104–107. [Google Scholar] [CrossRef]
- Clairembault, T.; Leclair-Visonneau, L.; Coron, E.; Bourreille, A.; Le Dily, S.; Vavasseur, F.; Heymann, M.-F.; Neunlist, M.; Derkinderen, P. Structural alterations of the intestinal epithelial barrier in Parkinson’s disease. Acta Neuropathol. Commun. 2015, 3, 12. [Google Scholar] [CrossRef] [PubMed]
- Fang, X. Potential role of gut microbiota and tissue barriers in Parkinson’s disease and amyotrophic lateral sclerosis. Int. J. Neurosci. 2016, 126, 771–776. [Google Scholar] [CrossRef]
- Gorecki, A.M.; Preskey, L.; Bakeberg, M.C.; Kenna, J.E.; Gildenhuys, C.; MacDougall, G.; Dunlop, S.A.; Mastaglia, F.L.; Akkari, P.A.; Koengten, F.; et al. Altered gut microbiome in Parkinson’s disease and the influence of lipopolysaccharide in a human α-synuclein over-expressing mouse model. Front. Neurosci. 2019, 13, 839. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.-P.; Wang, J.-F.; Xue, W.-J.; Liu, H.-M.; Liu, B.-R.; Zeng, Y.-L.; Li, S.-N.; Huang, B.-X.; Lv, Q.-K.; Wang, W.; et al. Anti-inflammatory effects of BHBA in both in vivo and in vitro Parkinson’s disease models are mediated by GPR109A-dependent mechanisms. J. Neuroinflammation 2015, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Tieu, K.; Perier, C.; Caspersen, C.; Teismann, P.; Wu, D.C.; Yan, S.D.; Naini, A.; Vila, M.; Jackson-Lewis, V.; Ramasamy, R.; et al. D-beta-hydroxybutyrate rescues mitochondrial respiration and mitigates features of Parkinson disease. J. Clin. Investig. 2003, 112, 892–901. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Osto, M.; Geurts, L.; Everard, A. Involvement of gut microbiota in the development of low-grade inflammation and type 2 diabetes associated with obesity. Gut Microbes 2012, 3, 279–288. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef]
- Skelly, A.; Sato, Y.; Kearney, S.; Honda, K. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat. Rev. Immunol. 2019, 19, 305–323. [Google Scholar] [CrossRef]
- Mathur, D.; Goyal, K.; Koul, V.; Anand, A. The Molecular Links of Re-Emerging Therapy: A Review of Evidence of Brahmi (Bacopa monniera). Front. Pharmacol. 2016, 7, 44. [Google Scholar] [CrossRef]
- Chakravarty, A.K.; Sarkar, T.; Masuda, K.; Shiojima, K.; Nakane, T.; Kawahara, N. Bacopaside I and II: Two pseudojujubogenin glycosides from Bacopa monniera. Phytochemistry 2001, 58, 553–556. [Google Scholar] [CrossRef]
- Thomas, R.B.; Joy, S.; Ajayan, M.S.; Paulose, C.S. Neuroprotective Potential of Bacopa monnieri and Bacoside A Against Dopamine Receptor Dysfunction in the Cerebral Cortex of Neonatal Hypoglycaemic Rats. Cell. Mol. Neurobiol. 2013, 33, 1065–1074. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.; Pandey, S.; Yadav, S.K.; Verma, R.; Singh, S.P.; Mahdi, A.A. Role of ethanolic extract of Bacopa monnieri against 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) induced mice model via inhibition of apoptotic pathways of dopaminergic neurons. Brain Res. Bull. 2017, 135, 120–128. [Google Scholar] [CrossRef]
- Singh, B.; Pandey, S.; Verma, R.; Ansari, J.A.; Mahdi, A.A. Comparative evaluation of extract of Bacopa monnieri and Mucuna pruriens as neuroprotectant in MPTP model of Parkinson’s disease. Experiment 2016, 54, 758–766. [Google Scholar]
- Peterson, C.T.; Sharma, V.; Iablokov, S.N.; Albayrak, L.; Khanipov, K.; Uchitel, S.; Chopra, D.; Mills, P.J.; Fofanov, Y.; Rodionov, D.A.; et al. 16S rRNA gene profiling and genome reconstruction reveal community metabolic interactions and prebiotic potential of medicinal herbs used in neurodegenerative disease and as nootropics. PLoS ONE 2019, 14, e0213869. [Google Scholar] [CrossRef]
- Peterson, C.T.; Vaughn, A.R.; Sharma, V.; Chopra, D.; Mills, P.J.; Peterson, S.N.; Sivamani, R.K. Effects of Turmeric and Curcumin Dietary Supplementation on Human Gut Microbiota: A Double-Blind, Randomized, Placebo-Controlled Pilot Study. J. Evidence-Based Integr. Med. 2018, 23. [Google Scholar] [CrossRef]
- Peterson, C.T.; Pourang, A.; Dhaliwal, S.; Kohn, J.N.; Uchitel, S.; Singh, H.; Mills, P.J.; Peterson, S.N.; Sivamani, R.K. Modulatory Effects of Triphala and Manjistha Dietary Supplementation on Human Gut Microbiota: A Double-Blind, Randomized, Placebo-Controlled Pilot Study. J. Altern. Complement. Med. 2020, 26, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Stoddard, S.F.; Smith, B.J.; Hein, R.; Roller, B.R.; Schmidt, T.M. rrnDB: Improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 2015, 43, D593–D598. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef]
- Ashniev, G.A.; Petrov, S.N.; Iablokov, S.N.; Rodionov, D.A. Genomics-Based Reconstruction and Predictive Profiling of Amino Acid Biosynthesis in the Human Gut Microbiome. Microorganisms 2022, 10, 740. [Google Scholar] [CrossRef]
- Rodionov, D.A.; Arzamasov, A.A.; Khoroshkin, M.S.; Iablokov, S.N.; Leyn, S.A.; Peterson, S.N.; Novichkov, P.S.; Osterman, A.L. Micronutrient Requirements and Sharing Capabilities of the Human Gut Microbiome. Front. Microbiol. 2019, 10, 1316. [Google Scholar] [CrossRef]
- Frolova, M.S.; Suvorova, I.A.; Iablokov, S.N.; Petrov, S.N.; Rodionov, D.A. Genomic reconstruction of short-chain fatty acid production by the human gut microbiota. Front. Mol. Biosci. 2022, 9, 949563. [Google Scholar] [CrossRef]
- Conlon, J.M.; Domin, J.; Thim, L.; DiMarzo, V.; Morris, H.R.; Bloom, S.R. Primary Structure of Neuromedin U from the Rat. J. Neurochem. 1988, 51, 988–991. [Google Scholar] [CrossRef] [PubMed]
- Bedarf, J.R.; Hildebrand, F.; Coelho, L.P.; Sunagawa, S.; Bahram, M.; Goeser, F.; Bork, P.; Wüllner, U. Functional implications of microbial and viral gut metagenome changes in early stage L-DOPA-naïve Parkinson’s disease patients. Genome Med. 2017, 9, 39. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Cui, L.; Yang, Y.; Miao, J.; Zhao, X.; Zhang, J.; Cui, G.; Zhang, Y. Gut Microbiota Differs Between Parkinson’s Disease Patients and Healthy Controls in Northeast China. Front. Mol. Neurosci. 2019, 12, 171. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Yue, L.; Fang, X.; Wang, G.; Li, C.; Sun, X.; Jia, X.; Yang, J.; Song, J.; Zhang, Y.; et al. Altered gut microbiota in Parkinson’s disease patients/healthy spouses and its association with clinical features. Park. Relat. Disord. 2020, 81, 84–88. [Google Scholar] [CrossRef]
- Li, Z.; Liang, H.; Hu, Y.; Lu, L.; Zheng, C.; Fan, Y.; Wu, B.; Zou, T.; Luo, X.; Zhang, X.; et al. Gut bacterial profiles in Parkinson’s disease: A systematic review. CNS Neurosci. Ther. 2023, 29, 140–157. [Google Scholar] [CrossRef]
- Alonso, J.; Pavón, S.; Vidal, J.; Delgado, M. Advanced Comparison of Phased Array and X-rays in the Inspection of Metallic Welding. Materials 2022, 15, 7108. [Google Scholar] [CrossRef]
- Sharma, S.; Awasthi, A.; Singh, S. Altered gut microbiota and intestinal permeability in Parkinson’s disease: Pathological highlight to management. Neurosci. Lett. 2019, 712, 134516. [Google Scholar] [CrossRef]
- Karunaratne, T.B.; Okereke, C.; Seamon, M.; Purohit, S.; Wakade, C.; Sharma, A. Niacin and Butyrate: Nutraceuticals Targeting Dysbiosis and Intestinal Permeability in Parkinson’s Disease. Nutrients 2020, 13, 28. [Google Scholar] [CrossRef]
- Deng, Y.; Zhou, M.; Wang, J.; Yao, J.; Yu, J.; Liu, W.; Wu, L.; Wang, J.; Gao, R. Involvement of the microbiota-gut-brain axis in chronic restraint stress: Disturbances of the kynurenine metabolic pathway in both the gut and brain. Gut Microbes 2021, 13, 1869501. [Google Scholar] [CrossRef]
- Scott, S.A.; Fu, J.; Chang, P.V. Microbial tryptophan metabolites regulate gut barrier function via the aryl hydrocarbon receptor. Proc. Natl. Acad. Sci. USA 2020, 117, 19376–19387. [Google Scholar] [CrossRef] [PubMed]
- Wakade, C.; Chong, R.; Bradley, E.; Morgan, J.C. Low-dose niacin supplementation modulates GPR109A, niacin index and ameliorates Parkinson’s disease symptoms without side effects. Clin. Case Rep. 2015, 3, 635–637. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, C.G.; Guth, N.; Helmschmied, K.; Armstrong, V.W.; Paulus, W.; Happe, S. Homocysteine in restless legs syndrome. Sleep Med. 2008, 9, 388–392. [Google Scholar] [CrossRef]
- Periñán, M.T.; Macías-García, D.; Jesús, S.; Martín-Rodríguez, J.F.; Muñoz-Delgado, L.; Jimenez-Jaraba, M.V.; Buiza-Rueda, D.; Bonilla-Toribio, M.; Adarmes-Gómez, A.D.; Gómez-Garre, P.; et al. Homocysteine levels, genetic background, and cognitive impairment in Parkinson’s disease. J. Neurol. 2023, 270, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Pallavicini, F.B.; Bosatra, M.G.; Verde, G.; Emmi, V.; Braschi, A.; Raimondi, F.; Iotti, G.; Villa, S. Indications, clinical implications and limitations in the use of a severity of illness index in intensive therapy. Minerva Anestesiol. 1987, 53, 565–569. [Google Scholar]
- Hawkes, C.H.; Del Tredici, K.; Braak, H. Parkinson’s disease: A dual-hit hypothesis. Neuropathol. Appl. Neurobiol. 2007, 33, 599–614. [Google Scholar] [CrossRef] [PubMed]
- Borghammer, P.; Just, M.K.; Horsager, J.; Skjærbæk, C.; Raunio, A.; Kok, E.H.; Savola, S.; Murayama, S.; Saito, Y.; Myllykangas, L.; et al. A postmortem study suggests a revision of the dual-hit hypothesis of Parkinson’s disease. Npj Park. Dis. 2022, 8, 166. [Google Scholar] [CrossRef]
- Vatanen, T.; Kostic, A.D.; D’hennezel, E.; Siljander, H.; Franzosa, E.A.; Yassour, M.; Kolde, R.; Vlamakis, H.; Arthur, T.D.; Hämäläinen, A.-M.; et al. Variation in Microbiome LPS Immunogenicity Contributes to Autoimmunity in Humans. Cell 2016, 165, 842–853. [Google Scholar] [CrossRef]
- Ho, M.S. Microglia in Parkinson’s Disease. Adv. Exp. Med. Biol. 2019, 1175, 335–353. [Google Scholar] [CrossRef]
- Wakade, C.; Chong, R.; Bradley, E.; Thomas, B.; Morgan, J. Upregulation of GPR109A in Parkinson’s Disease. PLoS ONE 2014, 9, e109818. [Google Scholar] [CrossRef] [PubMed]
- Wakade, C.; Giri, B.; Malik, A.; Khodadadi, H.; Morgan, J.C.; Chong, R.K.; Baban, B. Niacin modulates macrophage polarization in Parkinson’s disease. J. Neuroimmunol. 2018, 320, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Peterson, C.T.; Pérez-Santiago, J.; Iablokov, S.N.; Rodionov, D.A.; Peterson, S.N. Alteration of Community Metabolism by Prebiotics and Medicinal Herbs. Microorganisms 2023, 11, 868. [Google Scholar] [CrossRef] [PubMed]
| Species | Common Name | Family |
|---|---|---|
| Asparagus racemosus | Shatavari | Asparagaceae |
| Bacopa monnieri | brahmi or waterhyssop | Plantaginaceae |
| Boswellia serrata | Frankincense | Burseraceae |
| Centella asiatica | gotu kola or pennywort | Apiaceae |
| Eclipta alba | bhringaraj or false daisy | Asteraceae |
| Evolvulus alsinoides | shankhapushpi | Convolvulaceae |
| Mucuna pruriens | kapikacchu or velvet bean | Fabaceae |
| Nardostachys jatamansi | Jatamansi | Valerianaceae |
| Tinospora cordifolia | Guduchi | Menispermaceae |
| Withania somnifera | Ashwagandha | Solanaceae |
| Taxa | PD>HC | PD<HC | Ashwagandha | Bacopa | Boswellia | Bhringaraj | Gotu kola | Guduchi | Jatamansi | Kapikacchu | Shankhapushpi | Shatavari | 1 | 2 | 3 | H | Average |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Akkermansia | 13 | 0 | 2,3,H | 2,3,H | H | 2,3,H | H | 2,3,H | 2,H | H | 2,3,H | 2,3,H | 0% | 70% | 60% | 100% | 58% |
| Alistipes | 5 | 0 | 3 | 2 | 1,2.3,H | none | 1,2 | 2 | 2 | 2,3,H | 2 | 1,2,3 | 30% | 80% | 40% | 20% | 43% |
| Anaerotruncus | 2 | 0 | 1,H | none | none | none | 1,H | H | none | 1,H | H | 1,H | 40% | 0% | 0% | 60% | 25% |
| Anaerotruncus/C. Soleaferrea/other | 2 | 0 | none | none | none | none | H | none | none | H | H | H | 0% | 0% | 0% | 40% | 10% |
| Bacteroides | 1 | 2 | 1,2,3 | 1,2,3 | 1,2,3,H | 1,2,3 | 1,2,3,H | 1,2,3 | 1,3 | 1,2,3 | 1,2,3,H | 1,H | 100% | 80% | 80% | 40% | 75% |
| Bifidobacterium | 10 | 0 | 2 | 2 | 2 | none | 2,H | none | none | none | none | 2 | 0% | 50% | 0% | 0% | 13% |
| Bilophila | 4 | 0 | none | none | none | none | none | 2 | none | none | none | 2 | 0% | 20% | 0% | 0% | 5% |
| Blautia | 0 | 7 | 2 | none | none | none | none | 2 | 2 | none | none | none | 0% | 30% | 0% | 0% | 8% |
| Butyrivibrio/Fusicatenibacter/Pseudobutyrivibrio | 2 | 1 | none | none | none | none | none | none | none | none | none | none | 0% | 0% | 0% | 0% | 0% |
| Catabacter | 2 | 0 | none | none | none | none | none | none | none | none | none | H | 0% | 0% | 0% | 0% | 0% |
| Christensenella | 3 | 0 | none | none | H | H | none | none | none | H | none | H | 0% | 0% | 0% | 40% | 10% |
| Coprococcus | 1 | 2 | 1,2,3,H | 3,H | 2,3 | 1,3,H | 1 | 1,3,H | 1,3 | 2,3,H | 1,3 | none | 60% | 30% | 80% | 50% | 55% |
| Coprococcus/Frisingicoccus | 1 | 2 | 1 | 1 | none | none | none | none | none | none | none | none | 20% | 0% | 0% | 0% | 5% |
| Desulfovibrio | 3 | 0 | none | none | none | none | none | none | none | none | none | none | 0% | 0% | 0% | 0% | 0% |
| Dorea | 0 | 2 | 2,3 | 2,3 | none | 2,3 | 3 | 2,3 | 2,3 | 2,3 | none | none | 0% | 60% | 70% | 0% | 33% |
| Enterobacter | 2 | 0 | none | none | none | none | none | none | none | none | none | none | 0% | 0% | 0% | 0% | 0% |
| Enterococcus | 3 | 0 | none | none | none | none | 3 | 3 | 3 | none | 3 | none | 0% | 0% | 40% | 0% | 10% |
| Escherichia | 3 | 0 | 1,2,3,H | 1,3,H | H | 1,2,3,H | H | 1,H | 1,H | 1,2,3,H | 1,H | 1,2,3,H | 80% | 40% | 50% | 100% | 68% |
| Faecalibacterium | 0 | 8 | 1,H | 1,3,H | 1,H | 1,H | 1,H | 1,H | 1,H | 1,H | 1,H | 1 | 100% | 0% | 10% | 90% | 50% |
| Fusicatenibacter | 0 | 3 | 1,H | H | none | H | H | H | none | 1,2,H | none | none | 20% | 10% | 0% | 60% | 23% |
| Gemmiger | 0 | 2 | 2,H | H | 2,H | 2,H | H | H | 2,H | 2,H | 2,H | none | 0% | 60% | 0% | 90% | 38% |
| Holdemania | 2 | 0 | 1,3 | none | 1,3 | none | 1 | none | none | 3 | none | 1,3 | 40% | 0% | 40% | 0% | 20% |
| Lachnospira | 1 | 2 | 1,H | 1,H | none | 1,H | none | 1,H | 1,H | 1,H | 1,H | 1 | 80% | 0% | 0% | 70% | 38% |
| Lactobacillus | 9 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 0% | 0% | 100% | 0% | 25% |
| Parabacteroides | 3 | 1 | none | none | none | none | none | none | none | none | none | none | 0% | 0% | 0% | 0% | 0% |
| Roseburia | 1 | 9 | 1,H | 1,3,H | none | 1,H | none | 1,H | 1,3,H | 1,3,H | 1,H | none | 70% | 0% | 30% | 70% | 43% |
| Ruminococcus | 1 | 2 | none | 1 | none | 1 | none | 1 | none | 1 | 1 | none | 50% | 0% | 0% | 0% | 13% |
| Average | 26% | 20% | 22% | 31% | 25% | ||||||||||||
| Total | 690% | 530% | 600% | 830% | 663% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peterson, C.T.; Iablokov, S.N.; Rodionov, D.A.; Peterson, S.N. Personalized Response of Parkinson’s Disease Gut Microbiota to Nootropic Medicinal Herbs In Vitro: A Proof of Concept. Microorganisms 2023, 11, 1979. https://doi.org/10.3390/microorganisms11081979
Peterson CT, Iablokov SN, Rodionov DA, Peterson SN. Personalized Response of Parkinson’s Disease Gut Microbiota to Nootropic Medicinal Herbs In Vitro: A Proof of Concept. Microorganisms. 2023; 11(8):1979. https://doi.org/10.3390/microorganisms11081979
Chicago/Turabian StylePeterson, Christine Tara, Stanislav N. Iablokov, Dmitry A. Rodionov, and Scott N. Peterson. 2023. "Personalized Response of Parkinson’s Disease Gut Microbiota to Nootropic Medicinal Herbs In Vitro: A Proof of Concept" Microorganisms 11, no. 8: 1979. https://doi.org/10.3390/microorganisms11081979
APA StylePeterson, C. T., Iablokov, S. N., Rodionov, D. A., & Peterson, S. N. (2023). Personalized Response of Parkinson’s Disease Gut Microbiota to Nootropic Medicinal Herbs In Vitro: A Proof of Concept. Microorganisms, 11(8), 1979. https://doi.org/10.3390/microorganisms11081979









