Ramen Consumption and Gut Microbiota Diversity in Japanese Women: Cross-Sectional Data from the NEXIS Cohort Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Food Frequency Questionnaire
2.3. Fecal Sample Collection
2.4. DNA Extraction and 16S rRNA Gene Amplicon Sequencing
2.5. Bioinformatics Analysis
2.6. Blood Biochemicals
2.7. Dichotomous Grouping for Ramen Intake
2.8. Statistical Analysis
3. Results
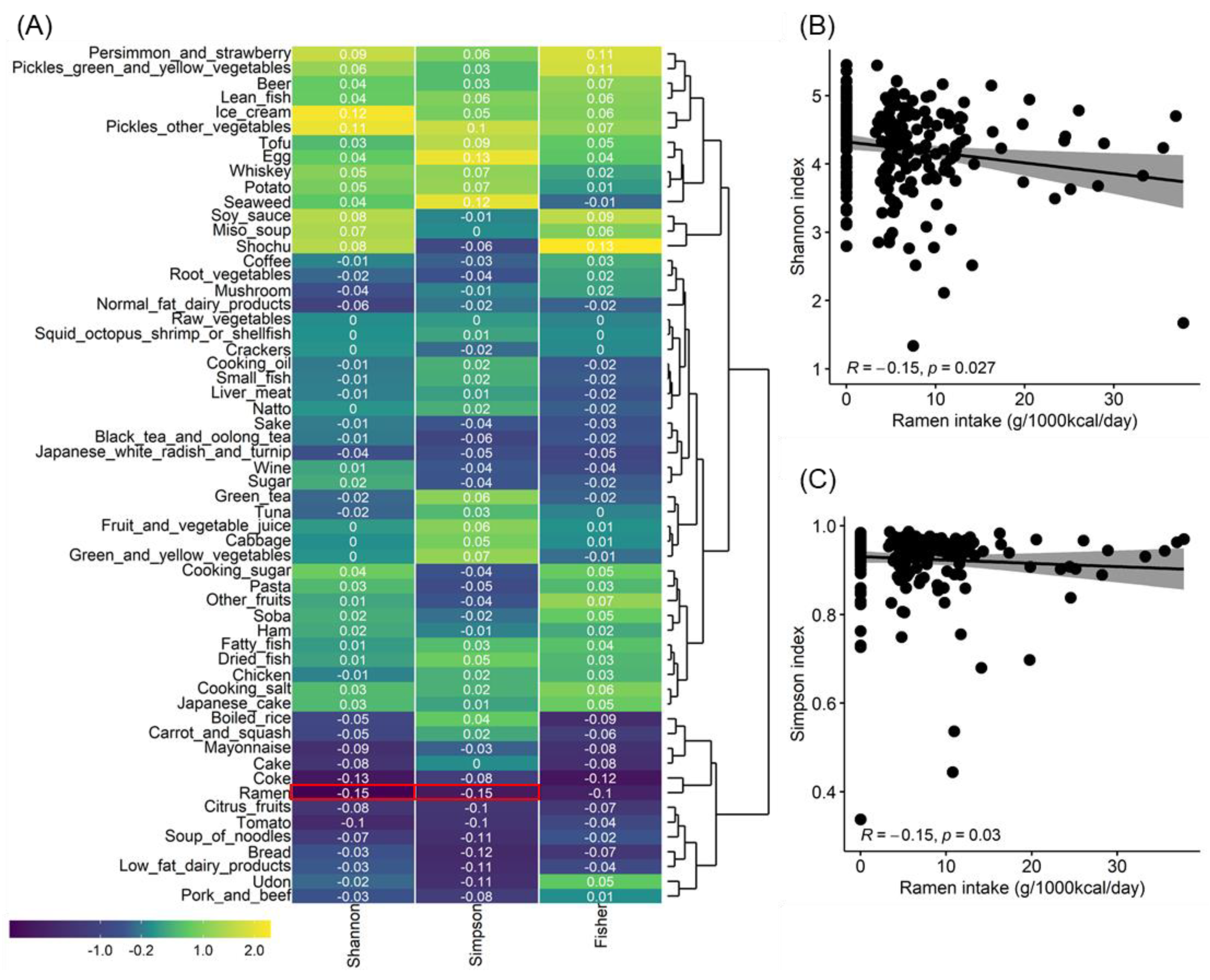
3.1. Gut Microbiota Diversity Associated with Ramen Intake
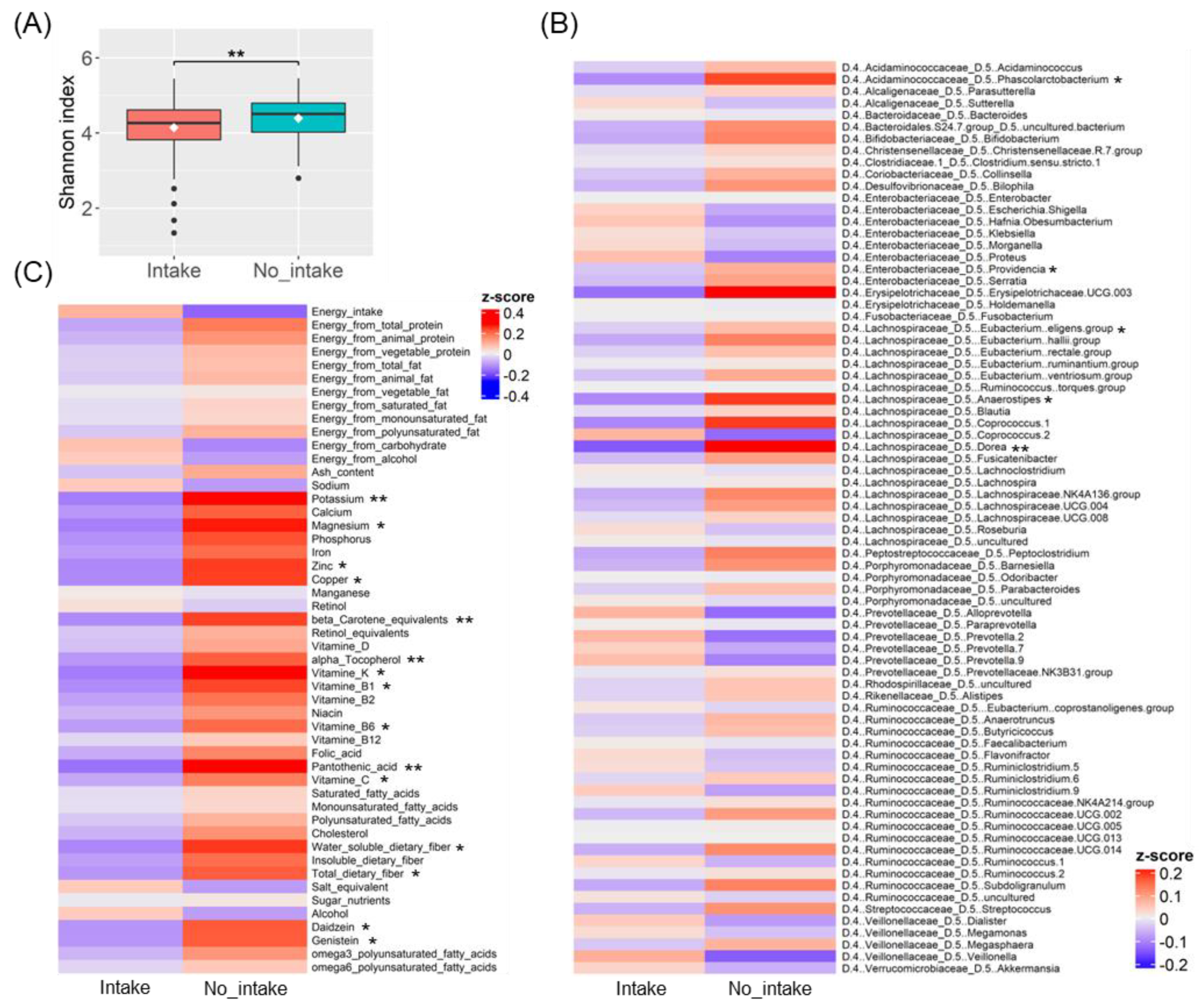
3.2. Differences in Gut Bacteria and Nutrient Intake Due to Ramen Intake
3.3. Multivariate Analysis for Ramen Intake Patterns and Nutrient Intake, Gut Microbiota, and Blood Biochemical Indices
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bradbury, K.E.; Appleby, P.N.; Key, T.J. Fruit, vegetable, and fiber intake in relation to cancer risk: Findings from the European Prospective Investigation into Cancer and Nutrition (EPIC). Am. J. Clin. Nutr. 2014, 100 (Suppl. S1), 394S–398S. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, A.; Mann, J.; Cummings, J.; Winter, N.; Mete, E.; Te Morenga, L. Carbohydrate Quality and Human Health: A Series of Systematic Reviews and Meta-Analyses. Lancet 2019, 393, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, R.; Goto, A.; Sawada, N.; Yamaji, T.; Iwasaki, M.; Noda, M.; Iso, H.; Tsugane, S. Dietary fiber intake and total and cause-specific mortality: The Japan Public Health Center-based prospective study. Am. J. Clin. Nutr. 2020, 111, 1027–1035. [Google Scholar] [CrossRef] [PubMed]
- Morton, G.J.; Meek, T.H.; Schwartz, M.W. Neurobiology of food intake in health and disease. Nat. Rev. Neurosci. 2014, 15, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Tsugane, S. Why has Japan become the world’s most long-lived country: Insights from the food and nutrition perspective? Eur. J. Clin. Nutr. 2021, 75, 921–928. [Google Scholar] [CrossRef]
- Suzuki, N.; Goto, Y.; Ota, H.; Kito, K.; Mano, F.; Joo, E.; Ikeda, K.; Inagaki, N.; Nakayama, T. Characteristics of the Japanese diet described in epidemiologic publications: A qualitative systematic review. J. Nutr. Sci. Vitaminol. 2018, 64, 129–137. [Google Scholar] [CrossRef]
- Matsuyama, S.; Sawada, N.; Tomata, Y.; Zhang, S.; Goto, A.; Yamaji, T.; Iwasaki, M.; Inoue, M.; Tsuji, I.; Tsugane, S.; et al. Association between adherence to the Japanese diet and all-cause and cause-specific mortality: The Japan Public Health Center-based Prospective Study. Eur. J. Nutr. 2021, 60, 1327–1336. [Google Scholar] [CrossRef]
- Fan, Y.; Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 2021, 19, 55–71. [Google Scholar] [CrossRef]
- Tang, W.H.W.; Kitai, T.; Hazen, S.L. Gut microbiota in cardiovascular health and disease. Circ. Res. 2017, 120, 1183–1196. [Google Scholar] [CrossRef]
- Mizutani, S.; Yamada, T.; Yachida, S. Significance of the gut microbiome in multistep colorectal carcinogenesis. Nat. Cancer Sci. 2020, 111, 766–773. [Google Scholar] [CrossRef]
- Hosomi, K.; Saito, M.; Park, J.; Murakami, H.; Shibata, N.; Ando, M.; Nagatake, T.; Konishi, K.; Ohno, H.; Tanisawa, K.; et al. Oral administration of Blautia wexlerae ameliorates obesity and type 2 diabetes via metabolic remodeling of the gut microbiota. Nat. Commun. 2022, 13, 4477. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. The influence of diet on the gut microbiome and its implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef] [PubMed]
- Maskarinec, G.; Hullar, M.A.J.; Monroe, K.R.; Shepherd, J.A.; Hunt, J.; Randolph, T.W.; Wilkens, L.R.; Boushey, C.J.; Le Marchand, L.; Lim, U.; et al. Fecal microbial diversity and structure were associated with diet quality in a multiethnic cohort adiposity phenotype study. J. Nutr. 2019, 149, 1575–1584. [Google Scholar] [CrossRef]
- Cotillard, A.; Cartier-Meheust, A.; Litwin, N.S.; Chaumont, S.; Saccareau, M.; Lejzerowicz, F.; Tap, J.; Koutnikova, H.; Lopez, D.G.; McDonald, D.; et al. The a posteriori dietary pattern better explains variations in the gut microbiome than individual markers in the American Gut Project. Am. J. Clin. Nutr. 2022, 115, 432–443. [Google Scholar] [CrossRef]
- Pickard, J.M.; Zeng, M.Y.; Caruso, R.; Núñez, G. Gut microbiota: Role in pathogen colonization, immune responses, and inflammatory disease. Immunol. Rev. 2017, 279, 70–89. [Google Scholar] [CrossRef]
- Manor, O.; Dai, C.L.; Kornilov, S.A.; Smith, B.; Price, N.D.; Lovejoy, J.C.; Gibbons, S.M.; Magis, A.T. Health and disease markers correlate with gut microbiome composition across thousands of people. Nat. Commun. 2020, 11, 5206. [Google Scholar] [CrossRef]
- Greenberg, A.S.; Youngster, I.; Cohen, A.N.; Livovsky, M.D.; Strahilevitz, J.; Israeli, E.; Melzer, E.; Paz, K.; Fliss-Isakov, N.; Maharshak, N. Five years of fecal microbiota transplantation-an update of the Israeli experience. World J. Gastroenterol. 2018, 24, 5403–5414. [Google Scholar] [CrossRef]
- Yatsuya, H.; Tsugane, S. What constitutes healthiness of Washoku or Japanese diet? Eur. J. Clin. Nutr. 2021, 75, 863–864. [Google Scholar] [CrossRef]
- Oki, K.; Toyama, M.; Banno, T.; Chonan, O.; Benno, Y.; Watanabe, K. Comprehensive analysis of the fecal microbiota of healthy Japanese adults revealed a new bacterial lineage associated with a phenotype characterized by a high frequency of bowel movements and lean body type. BMC Microbiol. 2016, 16, 284. [Google Scholar] [CrossRef]
- Park, J.; Kato, K.; Murakami, H.; Hosomi, K.; Tanisawa, K.; Nakagata, T.; Ohno, H.; Konishi, K.; Kawashima, H.; Chen, Y.A.; et al. Comprehensive analysis of the gut microbiota in a healthy population and covariates affecting microbial variation in two large Japanese cohorts. BMC Microbiol. 2021, 21, 151. [Google Scholar] [CrossRef]
- Matsuoka, T.; Hosomi, K.; Park, J.; Goto, Y.; Nishimura, M.; Maruyama, S.; Murakami, H.; Konishi, K.; Miyachi, M.; Kawashima, H.; et al. Relationship between barley consumption and gut microbiome characteristics in a healthy Japanese population: A cross-sectional study. BMC Nutr. 2022, 8, 23. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Hosomi, K.; Kawashima, H.; Chen, Y.A.; Mohsen, A.; Ohno, H.; Konishi, K.; Tanisawa, K.; Kifushi, M.; Kogawa, M.; et al. Dietary vitamin B1 intake influences the gut microbial community and the consequent production of short-chain fatty acids. Nutrients 2022, 14, 2078. [Google Scholar] [CrossRef] [PubMed]
- Takagi, T.; Naito, Y.; Inoue, R.; Kashiwagi, S.; Uchiyama, K.; Mizushima, K.; Tsuchiya, S.; Dohi, O.; Yoshida, N.; Kamada, K.; et al. Differences in the gut microbiota associated with age, sex, and stool consistency in healthy Japanese subjects. J. Gastroenterol. 2019, 54, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Yoshikata, R.; Myint, K.Z.; Ohta, H.; Ishigaki, Y. Interrelationship between diet, lifestyle habits, gut microflora, and the equol-producer phenotype: Baseline findings from a placebo-controlled intervention trial. Menopause 2019, 26, 273–285. [Google Scholar] [CrossRef]
- Farrand, C.; Charlton, K.; Crino, M.; Santos, J.; Rodriguez-Fernandez, R.; Mhurchu, N.C.; Webster, J. Know Your Noodles! As-sessing Variations in Sodium Content of Instant Noodles across Countries. Nutrients 2017, 9, 612. [Google Scholar] [CrossRef]
- Watanabe, D.; Murakami, H.; Ohno, H.; Tanisawa, K.; Konishi, K.; Tsunematsu, Y.; Sato, M.; Miyoshi, N.; Wakabayashi, K.; Watanabe, K.; et al. Association between dietary intake and the prevalence of tumourigenic bacteria in the gut microbiota of middle-aged Japanese adults. Sci. Rep. 2020, 10, 15221. [Google Scholar] [CrossRef]
- Kobayashi, S.; Murakami, K.; Sasaki, S.; Okubo, H.; Hirota, N.; Notsu, A.; Fukui, M.; Date, C. Comparison of the relative validity of food group intakes estimated by comprehensive and brief self-administered diet history questionnaires against 16 d dietary records in Japanese adults. Public. Health Nutr. 2011, 14, 1200–1211. [Google Scholar] [CrossRef]
- Kobayashi, S.; Honda, S.; Murakami, K.; Sasaki, S.; Okubo, H.; Hirota, N.; Notsu, A.; Fukui, M.; Date, C. Both comprehensive and brief self-administered diet history questionnaires satisfactorily rank nutrient intakes in Japanese adults. J. Epidemiol. 2012, 22, 151–159. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Hosomi, K.; Ohno, H.; Murakami, H.; Natsume-Kitatani, Y.; Tanisawa, K.; Hirata, S.; Suzuki, H.; Nagatake, T.; Nishino, T.; Mizuguchi, K.; et al. The method for preparing DNA from feces in guanidine thiocyanate solution affects the 16S rRNA-based profiling of human microbiota diversity. Sci. Rep. 2017, 7, 4339. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows the analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Mohsen, A.; Park, J.; Chen, Y.A.; Kawashima, H.; Mizuguchi, K. Impact of quality trimming on the efficiency of reads joining and diversity analysis of Illumina paired-end reads in the context of the QIIME1 and QIIME2 microbiome analysis frameworks. BMC Bioinform. 2019, 20, 581. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project has improved the data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Tripette, J.; Gando, Y.; Murakami, H.; Kawakami, R.; Tanisawa, K.; Ohno, H.; Konishi, K.; Tanimoto, M.; Tanaka, N.; Kawano, H.; et al. Effect of a 1-year intervention comprising brief counselling sessions and low-dose physical activity recommendations in Japanese adults, and retention of the effect at 2 years: A randomized trial. BMC Sports Sci. Med. Rehabil. 2021, 13, 133. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, R.; Asakura, K.; Uechi, K.; Masayasu, S.; Sasaki, S. Adequacy of iodine intake in three different Japanese adult dietary patterns: A nationwide study. Nutr. J. 2015, 14, 129. [Google Scholar] [CrossRef] [PubMed]
- Gominak, S.C. Vitamin D deficiency changes the intestinal microbiome, reducing vitamin B production in the gut. The resulting lack of pantothenic acid adversely affects the immune system, resulting in a “pro-inflammatory” state associated with atherosclerosis and autoimmunity. Med. Hypotheses 2016, 94, 103–107. [Google Scholar] [CrossRef]
- Weaver, C.M. Potassium and health. Adv. Nutr. 2013, 4, 368S–377S. [Google Scholar] [CrossRef]
- Platonova, K.; Kitamura, K.; Watanabe, Y.; Takachi, R.; Saito, T.; Kabasawa, K.; Takahashi, A.; Kobayashi, R.; Oshiki, R.; Solovev, A.; et al. Dietary calcium and vitamin K are associated with osteoporotic fracture risk in middle-aged and elderly Japanese women, but not men: The Murakami Cohort Study. Br. J. Nutr. 2021, 125, 319–328. [Google Scholar] [CrossRef]
- Zhang, X.; Yasuda, K.; Gilmore, R.A.; Westmoreland, S.V.; Platt, D.M.; Miller, G.M.; Vallender, E.J. Alcohol-induced changes in the gut microbiome and metabolome of rhesus macaques. Psychopharmacology 2019, 236, 1531–1544. [Google Scholar] [CrossRef]
- Gurwara, S.; Dai, A.; Ajami, N.J.; Graham, D.Y.; White, D.L.; Chen, L.; Jang, A.; Chen, E.; El-Serag, H.B.; Petrosino, J.F.; et al. Alcohol use alters the colonic mucosa-associated gut microbiota in humans. Nutr. Res. 2020, 83, 119–128. [Google Scholar] [CrossRef]
- Kimura, I.; Inoue, D.; Maeda, T.; Hara, T.; Ichimura, A.; Miyauchi, S.; Kobayashi, M.; Hirasawa, A.; Tsujimoto, G. Short-chain fatty acids and ketones directly regulate sympathetic nervous system via G protein-coupled receptor 41 (GPR41). Proc. Natl. Acad. Sci. USA 2011, 108, 8030–8035. [Google Scholar] [CrossRef]
- Kimura, I.; Ozawa, K.; Inoue, D.; Imamura, T.; Kimura, K.; Maeda, T.; Terasawa, K.; Kashihara, D.; Hirano, K.; Tani, T.; et al. Gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 2013, 4, 1829. [Google Scholar] [CrossRef]
- Prudêncio, A.P.A.; Fonseca, D.C.; Machado, N.M.; Alves, J.T.M.; Sala, P.; Fernandes, G.R.; Torrinhas, R.S.; Waitzberg, D.L. Red meat intake, indole-3-acetate, and Dorea longicatena together affect insulin resistance after gastric bypass. Nutrients 2023, 15, 1185. [Google Scholar] [CrossRef]
- Ji, Y.; Yin, W.; Liang, Y.; Sun, L.; Yin, Y.; Zhang, W. Anti-inflammatory and anti-oxidative activity of indole-3-acetic acid involves induction of HO-1 and neutralization of free radicals in RAW264.7 cells. Int. J. Mol. Sci. 2020, 21, 1579. [Google Scholar] [CrossRef]
- Garcia-Mantrana, I.; Selma-Royo, M.; Alcantara, C.; Collado, M.C. Shifts on gut microbiota associated to mediterranean diet adherence and specific dietary intakes on general adult population. Front. Microbiol. 2018, 9, 890. [Google Scholar] [CrossRef]
- Nanri, H.; Hara, M.; Nishida, Y.; Shimanoe, C.; Nakamura, K.; Higaki, Y.; Imaizumi, T.; Taguchi, N.; Sakamoto, T.; Horita, M.; et al. Dietary patterns and serum gamma-glutamyl transferase levels in Japanese men and women. J. Epidemiol. 2015, 25, 378–386. [Google Scholar] [CrossRef]
- Pereira, M.A. Sugar-sweetened and artificially sweetened beverages in relation to obesity risk. Adv. Nutr. 2014, 5, 797–808. [Google Scholar] [CrossRef]
- Ramne, S.; Brunkwall, L.; Ericson, U.; Gray, N.; Kuhnle, G.G.C.; Nilsson, P.M.; Orho-Melander, M.; Sonestedt, E. Gut microbiota composition in relation to intake of added sugar, sugar-sweetened beverages and artificially sweetened beverages in the Malmö Offspring Study. Eur. J. Nutr. 2021, 60, 2087–2097. [Google Scholar] [CrossRef]
- Whittingham, M.J.; Stephens, P.A.; Bradbury, R.B.; Freckleton, R.P. Why do we still use stepwise modelling in ecology and behaviour? J. Anim. Ecol. 2006, 75, 1182–1189. [Google Scholar] [CrossRef]

| Variables | Units | Mean ± SD | Variables | Units | Mean ± SD |
|---|---|---|---|---|---|
| Demographic and anthropometrics | Hematocrit | % | 40.2 ± 2.8 | ||
| Age | y | 59.5 ± 11.9 | Platelet | 104 count/μL | 2.39 ± 4.8 |
| Body height | cm | 156.5 ± 5.8 | Mean corpuscular volume (MCV) | fl | 93.1 ± 4.7 |
| Body weight | kg | 54.6 ± 7.4 | Mean corpuscular hemoglobin (MCH) | pg | 30.3 ± 1.9 |
| BMI | kg/m2 | 22.3 ± 2.9 | Mean corpuscular hemoglobin concentration (MCHC) | % | 32.6 ± 1.0 |
| Waist circumference | cm | 80.4 ± 9.3 | Aspartate transaminase (AST) | IU/L | 22.1 ± 5.8 |
| Hip circumference | cm | 92.1 ± 5.4 | Alanine transaminase (ALT) | IU/L | 16.7 ± 7.4 |
| Calf circumference | cm | 34.3 ± 2.3 | γ-Glutamyltransferase (γ-GTP) | IU/L | 23.9 ± 19.2 |
| Blood pressure and Blood biochemistry | Fasting glucose | mg/dL | 88.3 ± 15.3 | ||
| Systolic blood pressure | mmHg | 120.0 ± 17.1 | Hemoglobin A1c (HbA1c) | % | 5.57 ± 0.51 |
| Diastolic blood pressure | mmHg | 71.2 ± 10.5 | Triglyceride | mg/dL | 86.6 ± 49.6 |
| White blood cell (WBC) | count/μL | 4681 ± 1135 | Total cholesterol (Cho) | mg/dL | 219.4 ± 35.9 |
| Red blood cell (RBC) | 104 count/μL | 433 ± 32 | High-density lipoprotein cholesterol (HDL-cho) | mg/dL | 72.6 ± 18.1 |
| Hemoglobin | g/dL | 13.1 ± 1.1 | Fasting insulin | μU/dL | 4.09 ± 4.01 |
| Crude OR (95%CI) | Adj. OR (95%CI) | P (Wald’s Test) | |
|---|---|---|---|
| Age | 1.03 (1.01, 1.06) | 1.17 (1.05, 1.3) | 0.004 |
| Gut_habit_stool_frequency | 0.93 (0.73, 1.18) | 0.24 (0.1, 0.6) | 0.002 |
| D.1..Actinobacteria_D.2..Actinobacteria_D.3..Bifidobacteriales_D.4..Bifidobacteriaceae_D.5..Bifidobacterium | 1.0003 (0.9999, 1.0008) | 1.0013 (1, 1.0026) | 0.047 |
| D.1..Bacteroidetes_D.2..Bacteroidia_D.3..Bacteroidales_D.4..Porphyromonadaceae_D.5..Barnesiella | 1 (1, 1) | 1.01 (1, 1.02) | 0.032 |
| D.1..Bacteroidetes_D.2..Bacteroidia_D.3..Bacteroidales_D.4..Prevotellaceae_D.5..Alloprevotella | 0.95 (0.86, 1.06) | 0.72 (0.33, 1.57) | 0.405 |
| D.1..Bacteroidetes_D.2..Bacteroidia_D.3..Bacteroidales_D.4..Prevotellaceae_D.5..Prevotella.2 | 0.9983 (0.9955, 1.0011) | 0.9937 (0.9891, 0.9984) | 0.009 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5...Eubacterium..ruminantium.group | 1 (1, 1) | 1.01 (1, 1.02) | 0.016 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5...Eubacterium..ventriosum.group | 1 (1, 1.01) | 1.02 (1, 1.04) | 0.038 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5..Dorea | 1.02 (1.01, 1.03) | 1.12 (1.07, 1.17) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5..Lachnoclostridium | 1 (1, 1) | 0.98 (0.98, 0.99) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5..Lachnospiraceae.NK4A136.group | 1.0021 (0.9989, 1.0054) | 1.0056 (0.9976, 1.0137) | 0.171 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Lachnospiraceae_D.5..Roseburia | 0.9998 (0.999, 1.0006) | 0.9974 (0.9952, 0.9995) | 0.017 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Peptostreptococcaceae_D.5..Peptoclostridium | 1.01 (0.99, 1.02) | 1.13 (1.07, 1.2) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..Faecalibacterium | 1 (0.9997, 1.0003) | 0.9985 (0.9976, 0.9993) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..Ruminiclostridium.9 | 0.99 (0.97, 1.01) | 0.93 (0.89, 0.98) | 0.009 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..Ruminococcaceae.NK4A214.group | 1 (0.99, 1.01) | 0.96 (0.94, 0.98) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..Ruminococcaceae.UCG.014 | 1 (1, 1) | 1.02 (1.01, 1.03) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..Ruminococcus.1 | 0.9993 (0.9968, 1.0017) | 0.9907 (0.9857, 0.9958) | <0.001 |
| D.1..Firmicutes_D.2..Clostridia_D.3..Clostridiales_D.4..Ruminococcaceae_D.5..uncultured | 1 (1, 1) | 1.01 (1, 1.02) | 0.009 |
| D.1..Firmicutes_D.2..Erysipelotrichia_D.3..Erysipelotrichales_D.4..Erysipelotrichaceae_D.5..Erysipelotrichaceae.UCG.003 | 1.01 (1, 1.02) | 1.02 (0.99, 1.05) | 0.141 |
| D.1..Firmicutes_D.2..Erysipelotrichia_D.3..Erysipelotrichales_D.4..Erysipelotrichaceae_D.5..Holdemanella | 1 (1, 1) | 0.98 (0.98, 0.99) | <0.001 |
| D.1..Firmicutes_D.2..Negativicutes_D.3..Selenomonadales_D.4..Veillonellaceae_D.5..Megamonas | 0.9998 (0.9992, 1.0005) | 0.9991 (0.9979, 1.0003) | 0.125 |
| D.1..Proteobacteria_D.2..Alphaproteobacteria_D.3..Rhodospirillales_D.4..Rhodospirillaceae_D.5..uncultured | 1.0008 (0.998, 1.0035) | 1.0084 (1.002, 1.0149) | 0.01 |
| D.1..Proteobacteria_D.2..Gammaproteobacteria_D.3..Enterobacteriales_D.4..Enterobacteriaceae_D.5..Proteus | 0.98 (0.91, 1.05) | 0.57 (0.3, 1.1) | 0.096 |
| Energy_intake | 0.9993 (0.9986, 1) | 0.9947 (0.9922, 0.9973) | <0.001 |
| Energy_from_total_protein | 1.1 (1, 1.22) | 0.22 (0.05, 0.94) | 0.041 |
| Insoluble_dietary_fiber | 1.21 (1.01, 1.44) | 0.11 (0.03, 0.43) | 0.002 |
| Ash_content | 1.09 (0.94, 1.26) | 1.06 × 1010 (785.16, 1.43 × 1010) | 0.006 |
| Beta_Carotene_equivalents | 1.0002 (1.0001, 1.0004) | 1.002 (1.0009, 1.0031) | <0.001 |
| Calcium | 1 (1, 1.01) | 0.92 (0.89, 0.96) | <0.001 |
| Pantothenic_acid | 1.95 (1.29, 2.97) | 8.37 ×104 (352.47, 1.99 ×107) | <0.001 |
| Phosphorus | 1 (1, 1.01) | 1.05 (1, 1.1) | 0.054 |
| Potassium | 1 (1, 1) | 0.96 (0.93, 1) | 0.03 |
| Sodium | 1 (1, 1) | 0.94 (0.9, 0.98) | 0.002 |
| Sugar | 1.01 (0.94, 1.08) | 1.63 (1.27, 2.1) | <0.001 |
| Vitamine_B2 | 4.49 (1.01, 19.95) | 0 (0, 0) | <0.001 |
| Vitamine_B6 | 5.72 (1.11, 29.35) | 0 (0, 0.01) | 0.017 |
| omega3_polyunsaturated_fatty_acids | 1.64 (0.85, 3.17) | 3.70 × 104 (340.09, 4.03 × 106) | <0.001 |
| Total_cholesterol | 1 (0.99, 1.01) | 0.96 (0.94, 0.99) | 0.002 |
| Diastolic_blood_pressure | 1.01 (0.99, 1.04) | 1.08 (1, 1.16) | 0.049 |
| gamma_Glutamyltransferase | 1.01 (0.99, 1.02) | 1.07 (1.03, 1.12) | <0.001 |
| Mean_corpuscular_volume | 1.07 (1, 1.14) | 1.84 (1.39, 2.42) | <0.001 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, J.; Bushita, H.; Nakano, A.; Hara, A.; Ueno, H.M.; Ozato, N.; Hosomi, K.; Kawashima, H.; Chen, Y.-A.; Mohsen, A.; et al. Ramen Consumption and Gut Microbiota Diversity in Japanese Women: Cross-Sectional Data from the NEXIS Cohort Study. Microorganisms 2023, 11, 1892. https://doi.org/10.3390/microorganisms11081892
Park J, Bushita H, Nakano A, Hara A, Ueno HM, Ozato N, Hosomi K, Kawashima H, Chen Y-A, Mohsen A, et al. Ramen Consumption and Gut Microbiota Diversity in Japanese Women: Cross-Sectional Data from the NEXIS Cohort Study. Microorganisms. 2023; 11(8):1892. https://doi.org/10.3390/microorganisms11081892
Chicago/Turabian StylePark, Jonguk, Hiroto Bushita, Ayatake Nakano, Ai Hara, Hiroshi M. Ueno, Naoki Ozato, Koji Hosomi, Hitoshi Kawashima, Yi-An Chen, Attayeb Mohsen, and et al. 2023. "Ramen Consumption and Gut Microbiota Diversity in Japanese Women: Cross-Sectional Data from the NEXIS Cohort Study" Microorganisms 11, no. 8: 1892. https://doi.org/10.3390/microorganisms11081892
APA StylePark, J., Bushita, H., Nakano, A., Hara, A., Ueno, H. M., Ozato, N., Hosomi, K., Kawashima, H., Chen, Y.-A., Mohsen, A., Ohno, H., Konishi, K., Tanisawa, K., Nanri, H., Murakami, H., Miyachi, M., Kunisawa, J., Mizuguchi, K., & Araki, M. (2023). Ramen Consumption and Gut Microbiota Diversity in Japanese Women: Cross-Sectional Data from the NEXIS Cohort Study. Microorganisms, 11(8), 1892. https://doi.org/10.3390/microorganisms11081892







