The Gut Microbiome of an Indigenous Agropastoralist Population in a Remote Area of Colombia with High Rates of Gastrointestinal Infections and Dysbiosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Previously Existing Diagnostic Information about the Samples
2.3. Library Preparation and Next Generation Sequencing (NGS) Analysis
2.4. Bioinformation Assessment of the NGS Reads
2.5. Data Deposit
2.6. Ethical Clearance
3. Results
3.1. Brief Demographic Sample Background
3.2. General Features of the Microbiome Dataset
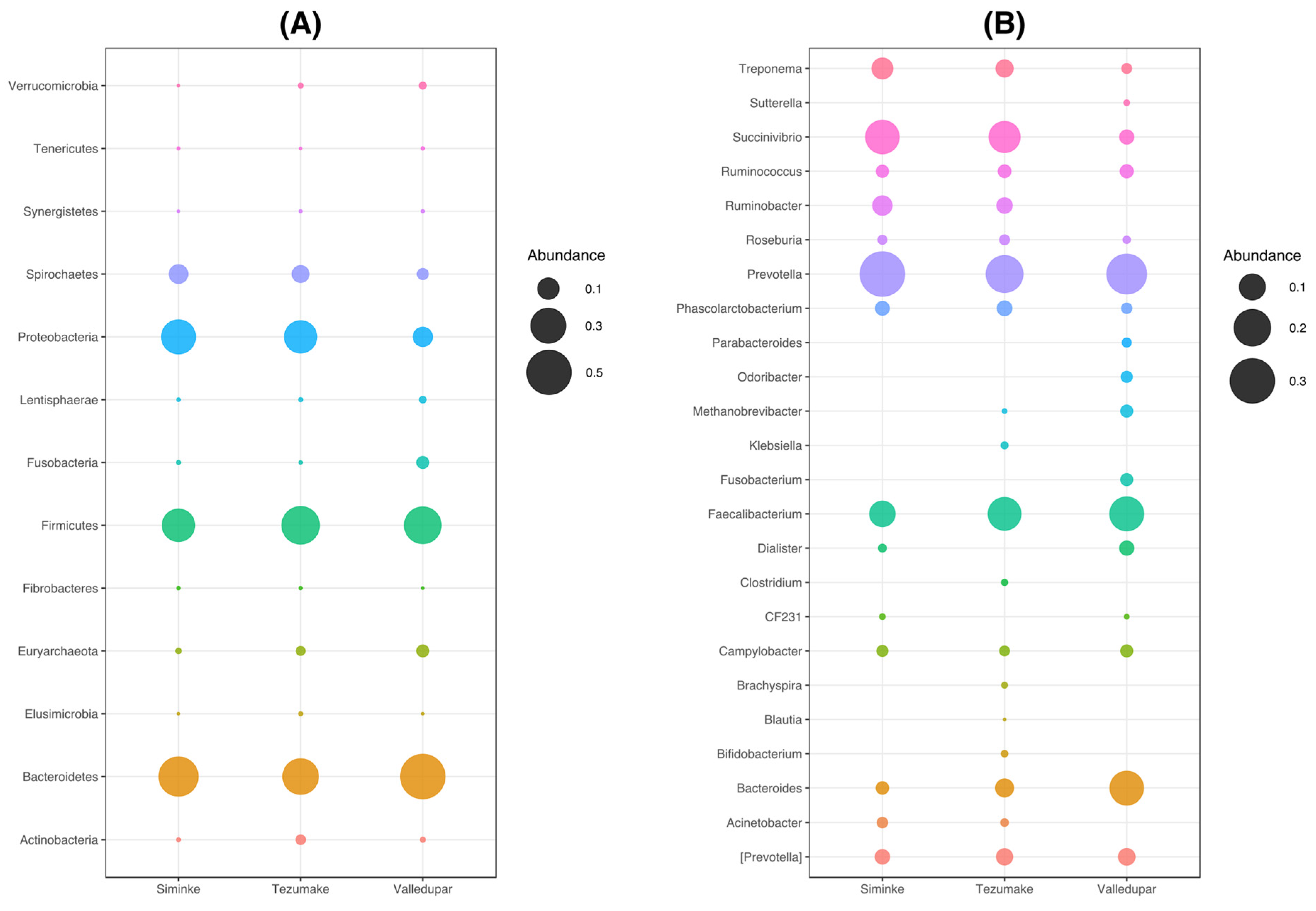
3.3. Composition of the Bacterial Communities by Location
| Comparison | Identifier | Phylum | Genus | LFC | p-Adjusted |
|---|---|---|---|---|---|
| VI | 403,701 | Firmicutes | Dialister | 25.12 | 6.98 × 10−4 |
| VI | 4,454,586 | Bacteroidetes | Odoribacter | 5.52 | 8.42 × 10−3 |
| VI | 314,915 | Bacteroidetes | Parabacteroides | −3.87 | 8.42 × 10−3 |
| VI | 562,244 | Proteobacteria | Sutterella | −4.15 | 6.98 × 10−4 |
| VT | 403,701 | Firmicutes | Dialister | 24.59 | 3.34 × 10−5 |
| VT | 4,454,586 | Bacteroidetes | Odoribacter | 5.51 | 6.53 × 10−4 |
| VT | 562,244 | Proteobacteria | Sutterella | −4.43 | 1.47 × 10−4 |
| VS | 314,915 | Bacteroidetes | Parabacteroides | −3.83 | 5.99 × 10−3 |
| VS | 567,226 | Bacteroidetes | Prevotella | −4.01 | 8.24 × 10−3 |
| VS | 531,614 | Bacteroidetes | Prevotella | −4.05 | 2.38 × 10−4 |
| VS | 335,827 | Firmicutes | Butyrivibrio | −4.16 | 1.13 × 10−3 |
| VS | 324,283 | Bacteroidetes | Prevotella | −5.50 | 3.80 × 10−4 |
| TS | 173,726 | Proteobacteria | Sutterella | 2.89 | 3.56 × 10−3 |
| TS | 179,291 | Firmicutes | Faecalibacterium | 2.09 | 7.54 × 10−3 |
| TS | 353,173 | Firmicutes | Clostridium | 1.99 | 3.56 × 10−3 |
| TS | 324,283 | Bacteroidetes | Prevotella | −3.14 | 8.17 × 10−4 |
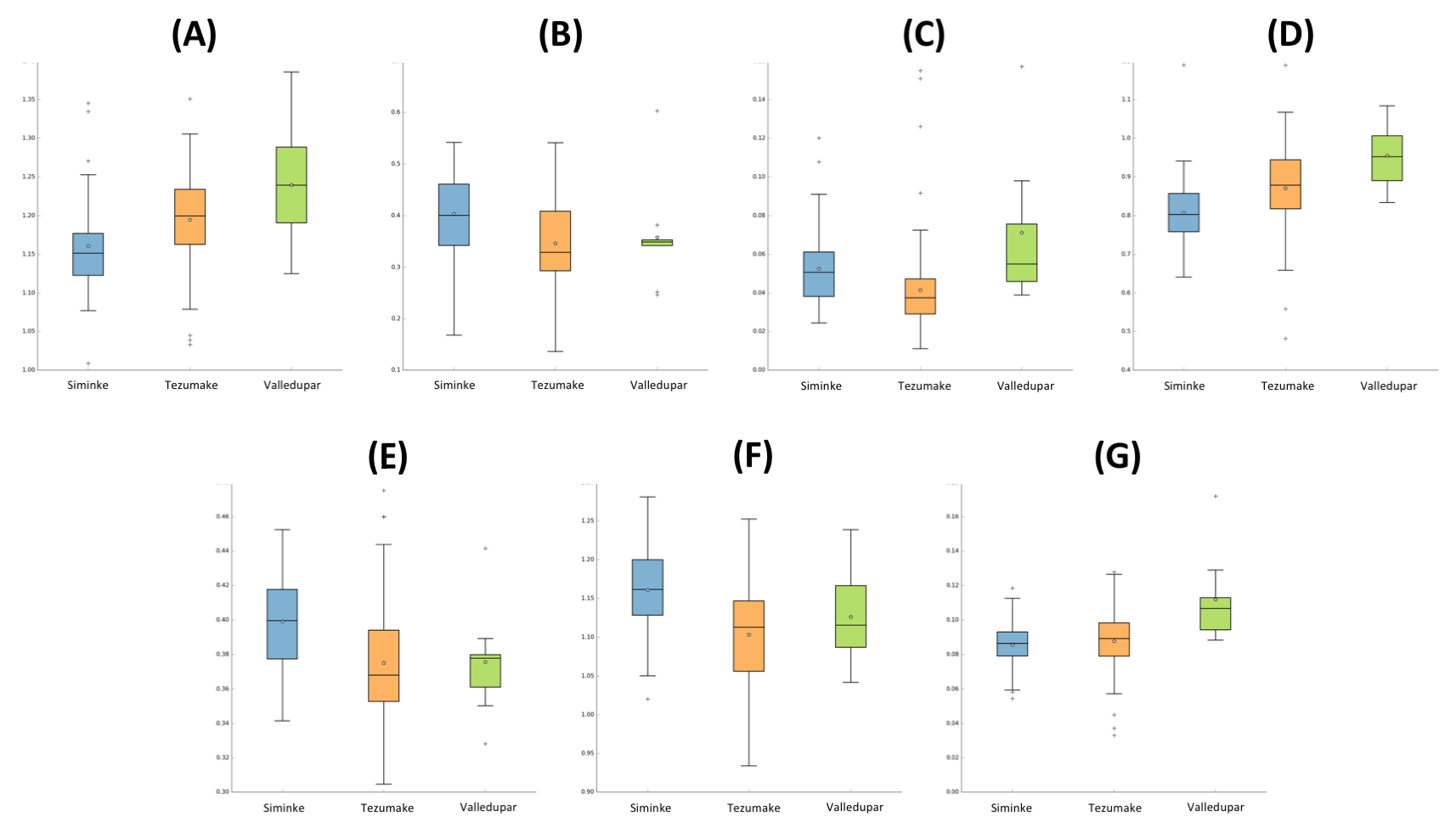
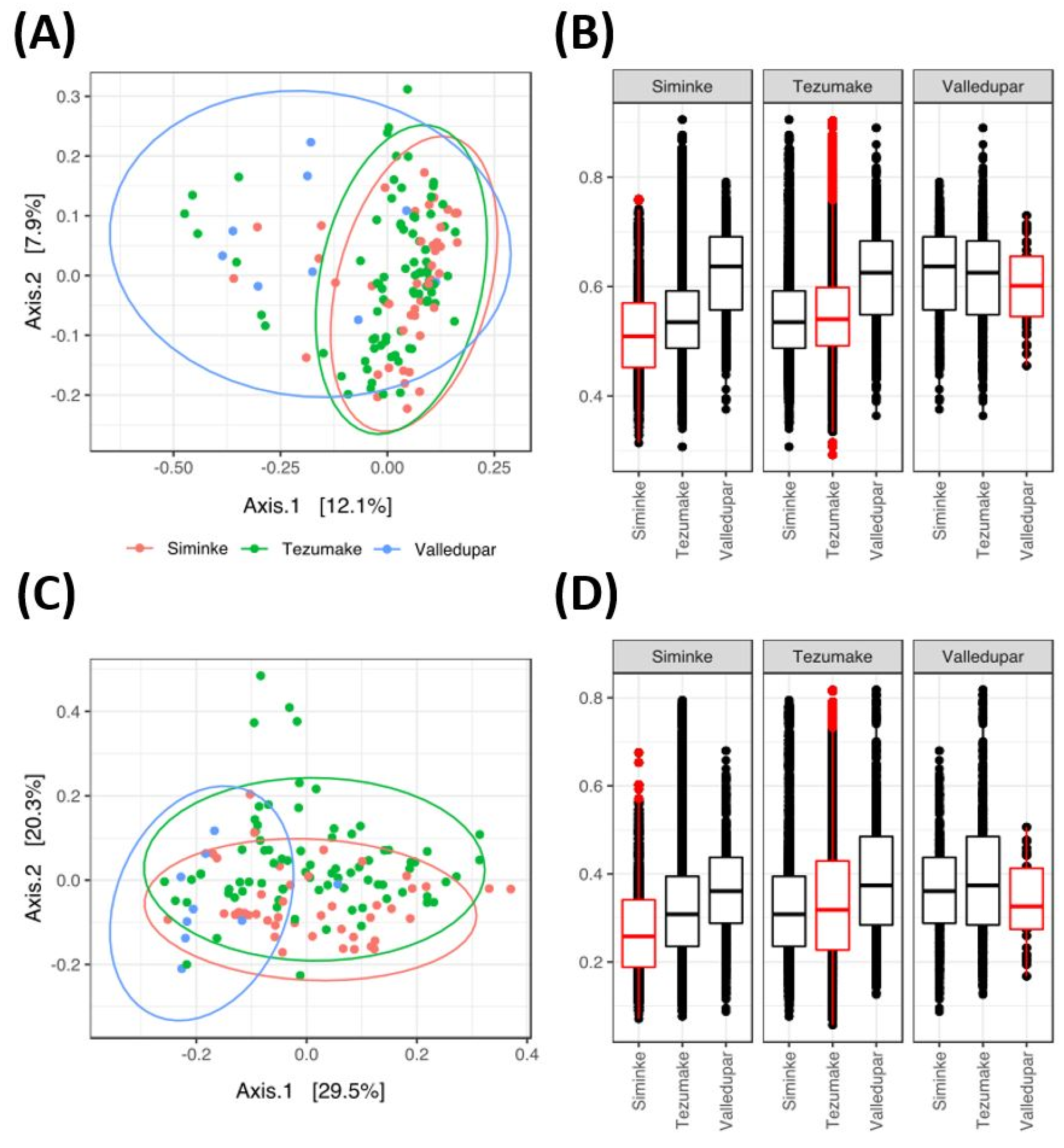
3.4. Microbial Gut Diversity of the Indigenous and Urban Populations
3.5. Functional Composition of the Microbial Communities by Location
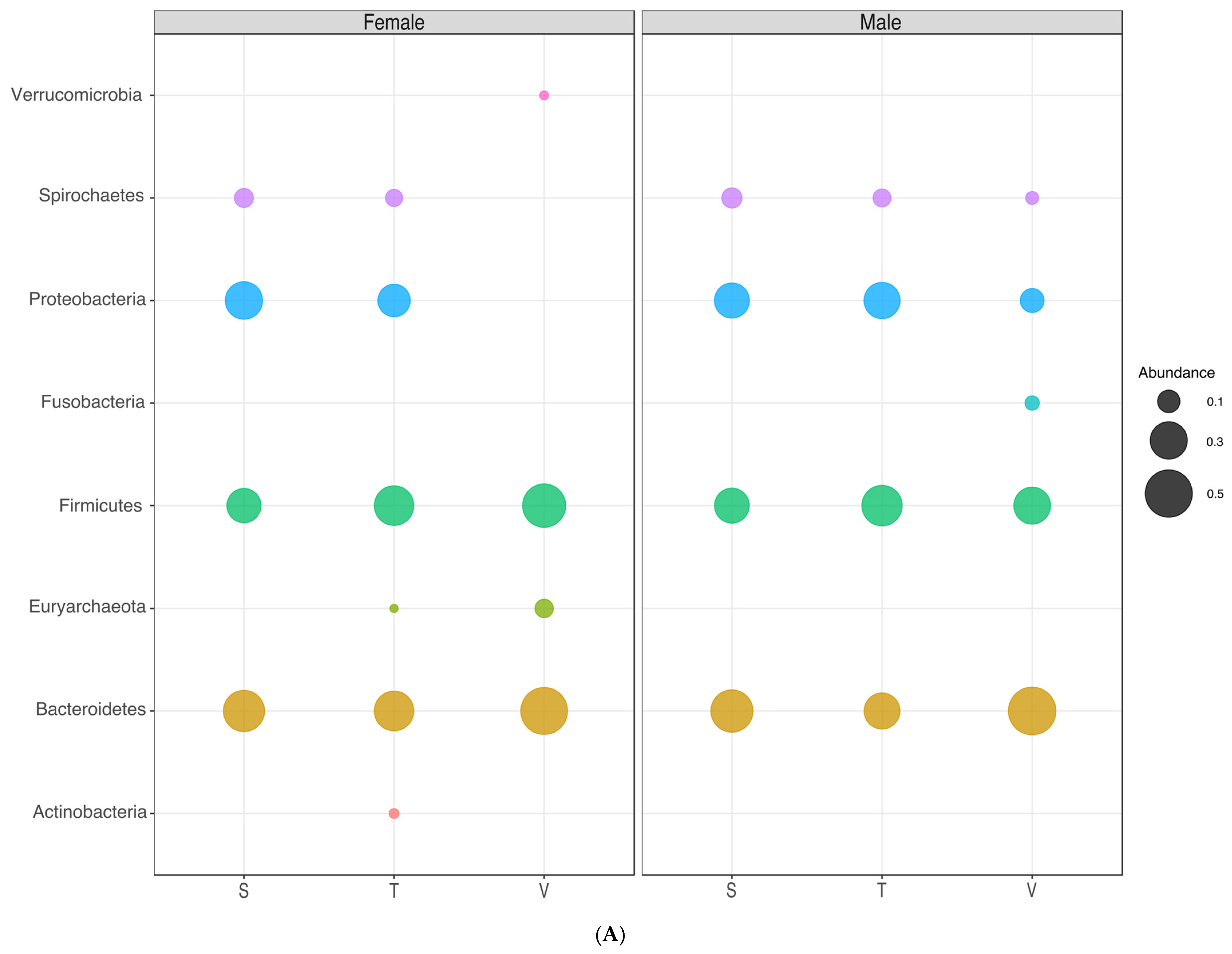
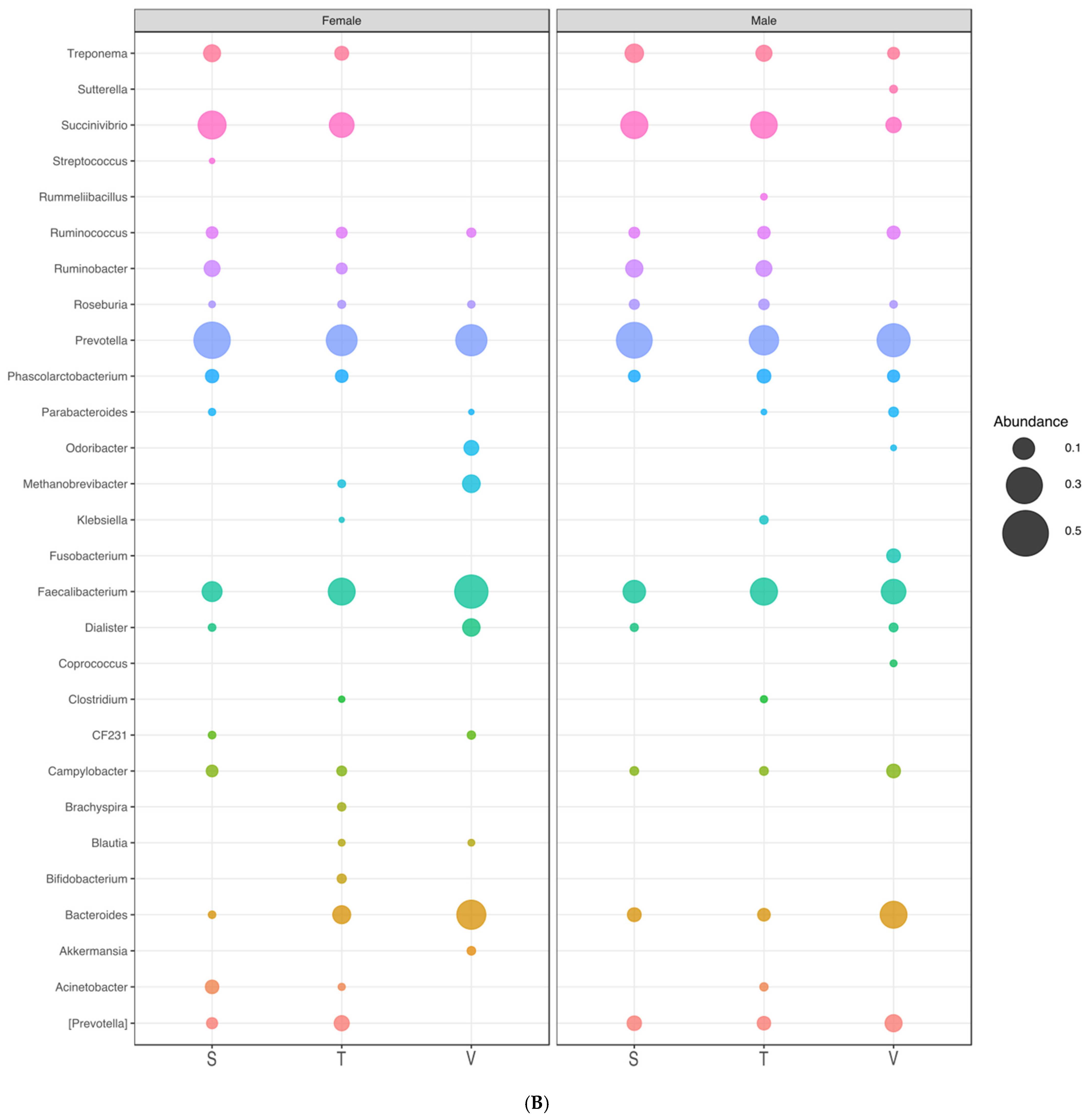
3.6. Microbial Communities in the Different Gender Groups
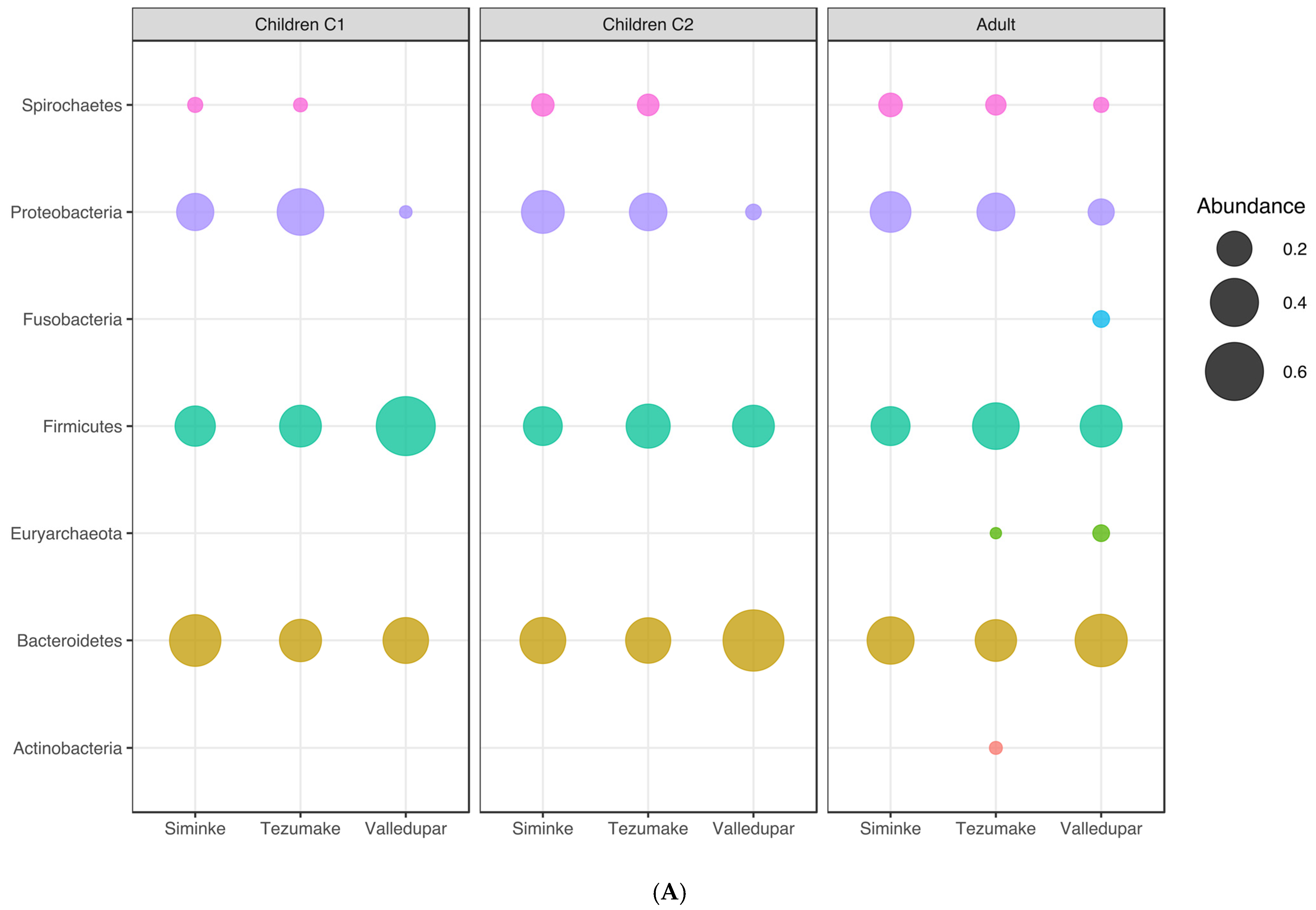
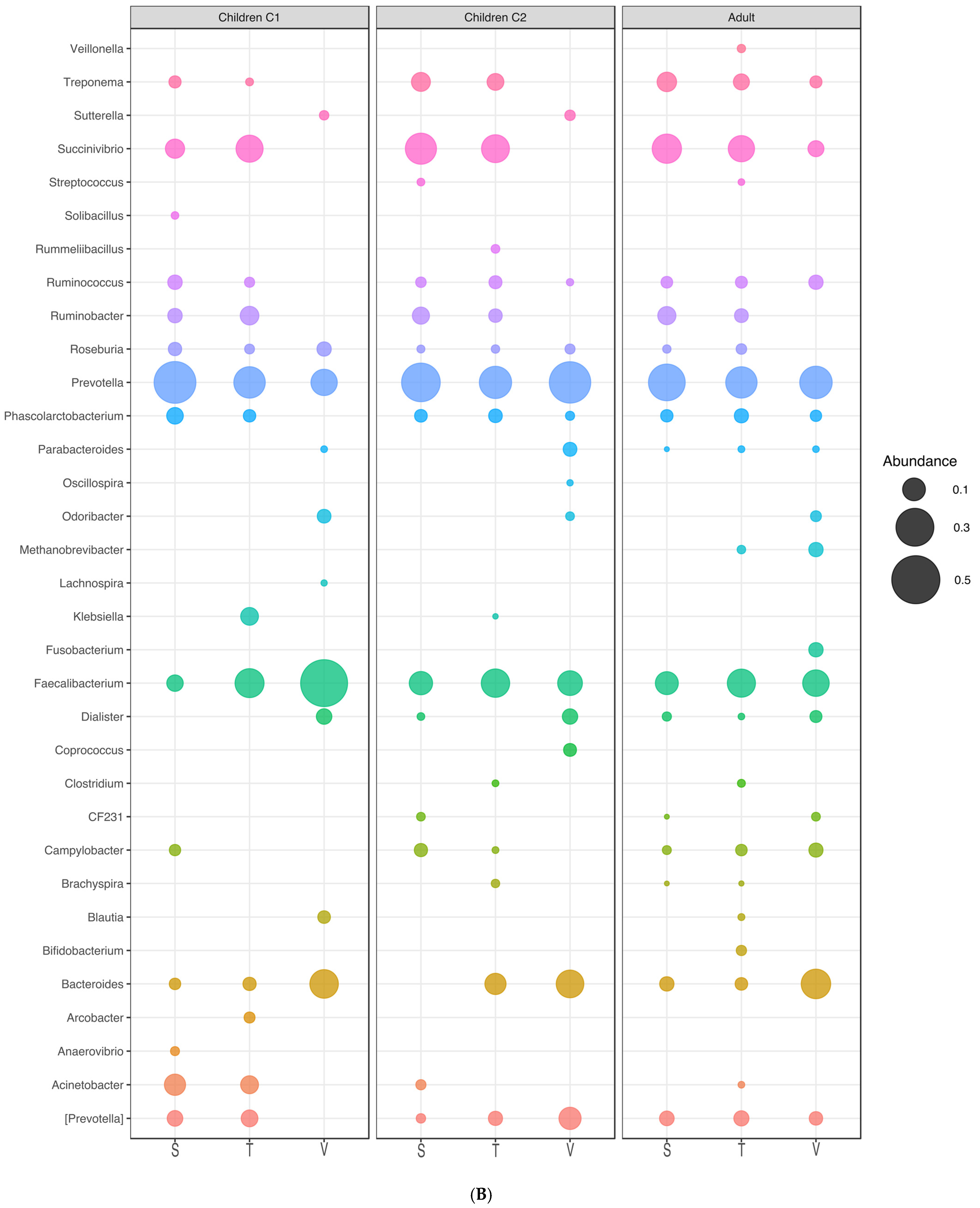
3.7. Microbial Communities Associated with Different Age Groups
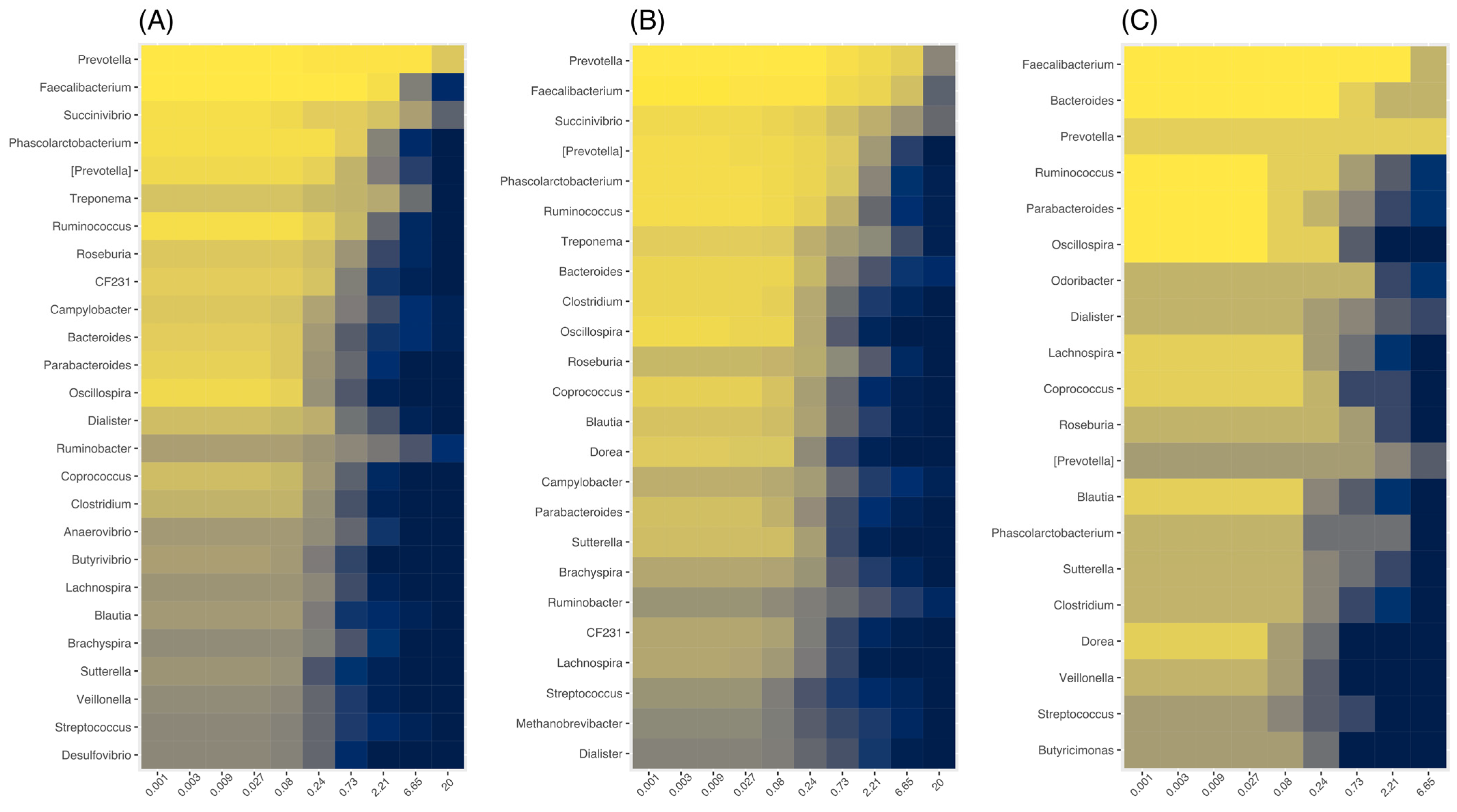
3.8. Association between Specific Genera and Demographic and Clinical Metadata
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Phylum | Siminke | Tezhumake | Valledupar |
|---|---|---|---|
| Bacteroidetes | 38.42% | 31.18% | 50.48% |
| Proteobacteria | 28.25% | 25.20% | 7.92% |
| Firmicutes | 25.51% | 35.04% | 33.31% |
| Spirochaetes | 7.44% | 5.84% | 2.06% |
| Euryarchaeota | 0.22% | 1.10% | 2.52% |
| Fusobacteria | 0.06% | 0.02% | 2.51% |
| Actinobacteria | 0.05% | 1.33% | 0.17% |
| Lentisphaerae | 0.03% | 0.06% | 0.46% |
| Fibrobacteres | 0.02% | 0.01% | 0.00% |
| Tenericutes | 0.01% | 0.00% | 0.01% |
| Verrucomicrobia | 0.00% | 0.15% | 0.54% |
| Elusimicrobia | 0.00% | 0.06% | 0.00% |
| Synergistetes | 0.00% | 0.01% | 0.01% |
| F/B ratio | 0.66 | 1.12 | 0.66 |
| Genus | Siminke | Tezhumake | Valledupar |
|---|---|---|---|
| Prevotella | 30.09% | 20.43% | 23.93% |
| Succinivibrio | 16.80% | 14.40% | 3.22% |
| Faecalibacterium | 9.70% | 16.06% | 17.21% |
| Treponema | 6.54% | 4.54% | 1.88% |
| Ruminobacter | 5.68% | 3.78% | 0.00% |
| [Prevotella] | 3.35% | 4.12% | 4.29% |
| Phascolarctobacterium | 3.12% | 3.48% | 1.99% |
| Bacteroides | 2.64% | 4.93% | 16.82% |
| Ruminococcus | 2.55% | 2.73% | 2.86% |
| Campylobacter | 2.20% | 1.87% | 2.51% |
| Acinetobacter | 2.03% | 1.44% | 0.00% |
| Roseburia | 1.69% | 1.89% | 1.40% |
| Dialister | 1.48% | 0.00% | 3.25% |
| CF231 | 1.19% | 0.00% | 1.11% |
| Klebsiella | 0.00% | 1.34% | 0.00% |
| Bifidobacterium | 0.00% | 1.29% | 0.00% |
| Clostridium | 0.00% | 1.25% | 0.00% |
| Brachyspira | 0.00% | 1.21% | 0.00% |
| Methanobrevibacter | 0.00% | 1.10% | 2.51% |
| Blautia | 0.00% | 1.03% | 0.00% |
| Fusobacterium | 0.00% | 0.00% | 2.51% |
| Odoribacter | 0.00% | 0.00% | 2.23% |
| Parabacteroides | 0.00% | 0.00% | 1.70% |
| Sutterella | 0.00% | 0.00% | 1.19% |

| Identifier | Level 1 | Level 3 | Corrected p-Values | Unique Genera |
|---|---|---|---|---|
| KO00330 | Metabolism | Arginine and proline metabolism | 0.0266 | 52 |
| KO00540 | Metabolism | Lipopolysaccharide biosynthesis | 0.0379 | 134 |
| KO00604 | Metabolism | Glycosphingolipid biosynthesis—ganglio series | 0.0247 | 143 |
| KO00860 | Metabolism | Porphyrin and chlorophyll metabolism | 0.0102 | 38 |
| KO01003 | Metabolism | Glycosyltransferases | 0.0297 | 12 |
| KO02042 | Environmental Information Processing | Bacterial toxins | 0.0237 | 56 |
| KO03110 | Genetic Information Processing | Chaperones and folding catalysts | 0.0017 | 145 |
| Phylum | Female | Male | ||||
|---|---|---|---|---|---|---|
| Siminke | Tezhumake | Valledupar | Siminke | Tezhumake | Valledupar | |
| Bacteroidetes | 37.39% | 34.44% | 49.32% | 39.36% | 27.82% | 51.06% |
| Proteobacteria | 30.33% | 22.25% | 0.00% | 26.36% | 28.23% | 11.40% |
| Firmicutes | 24.97% | 34.23% | 41.35% | 26.00% | 35.88% | 29.29% |
| Spirochaetes | 6.91% | 5.54% | 0.00% | 7.92% | 6.14% | 3.08% |
| Actinobacteria | 0.00% | 1.87% | 0.00% | 0.00% | 0.00% | 0.00% |
| Euryarchaeota | 0.00% | 1.45% | 6.46% | 0.00% | 0.00% | 0.00% |
| Verrucomicrobia | 0.00% | 0.00% | 1.63% | 0.00% | 0.00% | 0.00% |
| Fusobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| F/B ratio | 0.67 | 0.99 | 0.84 | 0.66 | 1.29 | 0.57 |
| Genus | Female | Male | ||||
|---|---|---|---|---|---|---|
| S | T | V | S | T | V | |
| Prevotella | 30.68% | 21.47% | 21.97% | 29.55% | 19.36% | 24.91% |
| Succinivibrio | 17.39% | 13.27% | 0.00% | 16.26% | 15.56% | 4.82% |
| Faecalibacterium | 8.39% | 15.94% | 25.34% | 10.90% | 16.18% | 13.15% |
| Treponema | 5.93% | 3.84% | 0.00% | 7.10% | 5.26% | 2.82% |
| Ruminobacter | 5.19% | 2.43% | 0.00% | 6.14% | 5.17% | 0.00% |
| Acinetobacter | 3.63% | 1.32% | 0.00% | 0.00% | 1.57% | 0.00% |
| Phascolarctobacterium | 3.53% | 3.17% | 0.00% | 2.75% | 3.79% | 2.95% |
| Campylobacter | 2.76% | 2.03% | 0.00% | 1.69% | 1.71% | 3.77% |
| Ruminococcus | 2.74% | 2.39% | 1.80% | 2.38% | 3.07% | 3.39% |
| [Prevotella] | 2.47% | 4.62% | 0.00% | 4.16% | 3.61% | 5.98% |
| Dialister | 1.42% | 0.00% | 6.27% | 1.53% | 0.00% | 1.73% |
| CF231 | 1.41% | 0.00% | 1.57% | 0.00% | 0.00% | 0.00% |
| Bacteroides | 1.39% | 6.65% | 18.80% | 3.78% | 3.17% | 15.83% |
| Parabacteroides | 1.33% | 0.00% | 1.08% | 0.00% | 1.10% | 2.01% |
| Roseburia | 1.24% | 1.50% | 1.38% | 2.10% | 2.29% | 1.41% |
| Streptococcus | 1.08% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Bifidobacterium | 0.00% | 1.82% | 0.00% | 0.00% | 0.00% | 0.00% |
| Brachyspira | 0.00% | 1.59% | 0.00% | 0.00% | 0.00% | 0.00% |
| Methanobrevibacter | 0.00% | 1.45% | 6.46% | 0.00% | 0.00% | 0.00% |
| Blautia | 0.00% | 1.27% | 1.26% | 0.00% | 0.00% | 0.00% |
| Clostridium | 0.00% | 1.19% | 0.00% | 0.00% | 1.31% | 0.00% |
| Klebsiella | 0.00% | 1.06% | 0.00% | 0.00% | 1.63% | 0.00% |
| Odoribacter | 0.00% | 0.00% | 4.46% | 0.00% | 0.00% | 1.12% |
| Akkermansia | 0.00% | 0.00% | 1.63% | 0.00% | 0.00% | 0.00% |
| Coprococcus | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.27% |
| Fusobacterium | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| Rummeliibacillus | 0.00% | 0.00% | 0.00% | 0.00% | 1.23% | 0.00% |
| Sutterella | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.47% |
| Phylum | Children (C1) | Children (C2) | Adult (C3) | |||
|---|---|---|---|---|---|---|
| Indigenous | Valledupar | Indigenous | Valledupar | Indigenous | Valledupar | |
| Firmicutes | 28.73% | 62.21% | 30.63% | 29.58% | 32.84% | 29.74% |
| Bacteroidetes | 35.74% | 35.60% | 35.31% | 66.43% | 32.43% | 47.64% |
| Proteobacteria | 32.28% | 1.96% | 25.78% | 3.26% | 25.37% | 10.47% |
| Spirochaetes | 2.63% | 0.00% | 6.99% | 0.00% | 6.84% | 2.94% |
| Actinobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 1.30% | 0.00% |
| Euryarchaeota | 0.00% | 0.00% | 0.00% | 0.00% | 1.06% | 3.78% |
| Fusobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| F/B ratio | 0.80 | 1.72 | 0.87 | 0.45 | 1.01 | 0.62 |
| Genus | Children (C1) | Children (C2) | Adult (C3) | |||
|---|---|---|---|---|---|---|
| Indigenous | Valledupar | Indigenous | Valledupar | Indigenous | Valledupar | |
| Prevotella | 26.17% | 13.71% | 24.45% | 36.36% | 23.07% | 21.49% |
| Succinivibrio | 11.84% | 0.00% | 16.84% | 0.00% | 15.01% | 4.72% |
| Faecalibacterium | 12.66% | 47.52% | 14.37% | 11.95% | 13.67% | 13.92% |
| Treponema | 1.88% | 0.00% | 5.71% | 0.00% | 5.68% | 2.70% |
| Ruminobacter | 5.56% | 0.00% | 4.01% | 0.00% | 4.51% | 0.00% |
| [Prevotella] | 4.94% | 0.00% | 3.06% | 9.39% | 4.10% | 3.30% |
| Phascolarctobacterium | 3.66% | 0.00% | 3.26% | 1.69% | 3.34% | 2.42% |
| Bacteroides | 2.94% | 16.24% | 5.98% | 15.18% | 3.23% | 17.46% |
| Ruminococcus | 2.64% | 0.00% | 2.79% | 1.30% | 2.60% | 3.74% |
| Campylobacter | 1.50% | 0.00% | 1.85% | 0.00% | 2.17% | 3.65% |
| Roseburia | 2.39% | 3.74% | 1.50% | 1.96% | 1.89% | 0.00% |
| Dialister | 0.00% | 4.44% | 0.00% | 4.36% | 1.40% | 2.67% |
| Bifidobacterium | 0.00% | 0.00% | 0.00% | 0.00% | 1.27% | 0.00% |
| Parabacteroides | 0.00% | 1.21% | 0.00% | 3.45% | 1.15% | 1.20% |
| Blautia | 0.00% | 2.94% | 0.00% | 0.00% | 1.13% | 0.00% |
| Veillonella | 0.00% | 0.00% | 0.00% | 0.00% | 1.12% | 0.00% |
| Clostridium | 0.00% | 0.00% | 1.00% | 0.00% | 1.07% | 0.00% |
| Methanobrevibacter | 0.00% | 0.00% | 0.00% | 0.00% | 1.05% | 3.76% |
| Acinetobacter | 6.87% | 0.00% | 0.00% | 0.00% | 1.05% | 0.00% |
| Brachyspira | 0.00% | 0.00% | 1.23% | 0.00% | 1.04% | 0.00% |
| Fusobacterium | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| Odoribacter | 0.00% | 3.46% | 0.00% | 1.60% | 0.00% | 2.24% |
| CF231 | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.62% |
| Arcobacter | 1.46% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Coprococcus | 0.00% | 0.00% | 0.00% | 3.05% | 0.00% | 0.00% |
| Klebsiella | 3.80% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Lachnospira | 0.00% | 1.15% | 0.00% | 0.00% | 0.00% | 0.00% |
| Oscillospira | 0.00% | 0.00% | 0.00% | 1.16% | 0.00% | 0.00% |
| Rummeliibacillus | 0.00% | 0.00% | 1.11% | 0.00% | 0.00% | 0.00% |
| Sutterella | 0.00% | 1.79% | 0.00% | 2.10% | 0.00% | 0.00% |
| Phylum | Children (C1) | Children (C2) | Adult (C3) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Siminke | Tezhumake | Valledupar | Siminke | Tezhumake | Valledupar | Siminke | Tezhumake | Valledupar | |
| Bacteroidetes | 46.17% | 29.95% | 35.60% | 36.09% | 34.95% | 66.43% | 38.10% | 28.93% | 47.64% |
| Proteobacteria | 22.91% | 37.49% | 1.96% | 31.09% | 23.32% | 3.26% | 27.86% | 23.82% | 10.47% |
| Firmicutes | 27.32% | 29.52% | 62.21% | 25.17% | 33.16% | 29.58% | 25.33% | 37.48% | 29.74% |
| Spirochaetes | 2.95% | 2.45% | 0.00% | 7.39% | 6.81% | 0.00% | 8.32% | 5.91% | 2.94% |
| Actinobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 2.08% | 0.00% |
| Euryarchaeota | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.56% | 3.78% |
| Fusobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| F/B ratio | 0.59 | 0.99 | 1.75 | 0.70 | 0.95 | 0.45 | 0.66 | 1.30 | 0.62 |
| Genus | Children (C1) | Children (C2) | Adult (C3) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| S | T | V | S | T | V | S | T | V | |
| Prevotella | 37.13% | 20.09% | 13.71% | 31.36% | 21.25% | 36.36% | 28.10% | 19.96% | 21.49% |
| Succinivibrio | 6.95% | 14.56% | 0.00% | 19.65% | 15.53% | 0.00% | 17.26% | 13.61% | 4.72% |
| Faecalibacterium | 5.02% | 16.90% | 47.52% | 10.54% | 16.15% | 11.95% | 10.19% | 15.82% | 13.92% |
| Treponema | 2.74% | 1.40% | 0.00% | 6.73% | 5.24% | 0.00% | 7.17% | 4.75% | 2.70% |
| Ruminobacter | 3.81% | 6.53% | 0.00% | 5.45% | 3.34% | 0.00% | 6.16% | 3.49% | 0.00% |
| [Prevotella] | 4.49% | 5.19% | 0.00% | 1.78% | 3.65% | 9.39% | 3.92% | 4.20% | 3.30% |
| Bacteroides | 2.45% | 3.20% | 16.24% | 0.00% | 8.53% | 15.18% | 3.75% | 2.90% | 17.46% |
| Phascolarctobacterium | 5.10% | 2.85% | 0.00% | 2.96% | 3.40% | 1.69% | 2.82% | 3.66% | 2.42% |
| Ruminococcus | 3.81% | 1.99% | 0.00% | 2.11% | 3.11% | 1.30% | 2.53% | 2.63% | 3.74% |
| Dialister | 0.00% | 0.00% | 4.44% | 1.37% | 0.00% | 4.36% | 1.73% | 1.20% | 2.67% |
| Campylobacter | 2.42% | 0.00% | 0.00% | 3.20% | 1.23% | 0.00% | 1.66% | 2.48% | 3.65% |
| Roseburia | 3.27% | 1.90% | 3.74% | 1.43% | 1.53% | 1.96% | 1.52% | 2.13% | 0.00% |
| Parabacteroides | 0.00% | 0.00% | 1.21% | 0.00% | 0.00% | 3.45% | 1.03% | 1.23% | 1.20% |
| CF231 | 0.00% | 0.00% | 0.00% | 1.59% | 0.00% | 0.00% | 1.03% | 0.00% | 1.62% |
| Brachyspira | 0.00% | 0.00% | 0.00% | 0.00% | 1.54% | 0.00% | 1.03% | 1.05% | 0.00% |
| Bifidobacterium | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 2.04% | 0.00% |
| Methanobrevibacter | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.55% | 3.76% |
| Veillonella | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.51% | 0.00% |
| Clostridium | 0.00% | 0.00% | 0.00% | 0.00% | 1.22% | 0.00% | 0.00% | 1.42% | 0.00% |
| Blautia | 0.00% | 0.00% | 2.94% | 0.00% | 0.00% | 0.00% | 0.00% | 1.24% | 0.00% |
| Acinetobacter | 8.53% | 5.94% | 0.00% | 2.00% | 0.00% | 0.00% | 0.00% | 1.21% | 0.00% |
| Streptococcus | 0.00% | 0.00% | 0.00% | 1.38% | 0.00% | 0.00% | 0.00% | 1.20% | 0.00% |
| Fusobacterium | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.77% |
| Odoribacter | 0.00% | 0.00% | 3.46% | 0.00% | 0.00% | 1.60% | 0.00% | 0.00% | 2.24% |
| Anaerovibrio | 1.67% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Arcobacter | 0.00% | 2.27% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Coprococcus | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 3.05% | 0.00% | 0.00% | 0.00% |
| Klebsiella | 0.00% | 5.88% | 0.00% | 0.00% | 1.07% | 0.00% | 0.00% | 0.00% | 0.00% |
| Lachnospira | 0.00% | 0.00% | 1.15% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Oscillospira | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 1.16% | 0.00% | 0.00% | 0.00% |
| Rummeliibacillus | 0.00% | 0.00% | 0.00% | 0.00% | 1.60% | 0.00% | 0.00% | 0.00% | 0.00% |
| Solibacillus | 1.35% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| Sutterella | 0.00% | 0.00% | 1.79% | 0.00% | 0.00% | 2.10% | 0.00% | 0.00% | 0.00% |
| Phylum | Children (C1) | Children (C2) | Adult (C3) | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Risk of Overweight | Overweight | Normal | Overweight | Obesity | Normal | Pre-Obesity | |||||||||
| S | T | S | T | S | T | S | T | S | T | S | T | V | S | T | |
| Bacteroidetes | 50.49% | 35.98% | 46.97% | 36.03% | 38.28% | 30.00% | 31.71% | 40.50% | 25.30% | 37.94% | 39.71% | 29.73% | 50.15% | 32.56% | 35.20% |
| Proteobacteria | 14.06% | 29.11% | 14.02% | 25.19% | 27.94% | 25.95% | 38.00% | 21.92% | 30.24% | 24.90% | 28.79% | 20.68% | 16.58% | 23.63% | 26.17% |
| Firmicutes | 34.18% | 32.50% | 25.87% | 35.29% | 25.36% | 34.52% | 25.84% | 28.16% | 31.29% | 33.48% | 23.51% | 40.48% | 16.49% | 33.61% | 28.36% |
| Spirochaetes | 0.00% | 0.00% | 12.28% | 3.32% | 8.16% | 7.52% | 4.10% | 6.05% | 13.02% | 3.53% | 7.53% | 6.21% | 4.52% | 9.91% | 9.06% |
| Euryarchaeota | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 2.45% | 0.00% | 0.00% | 0.00% | 2.20% | 0.00% | 0.00% | 1.11% |
| Fusobacteria | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 11.30% | 0.00% | 0.00% |
| Elusimicrobia | 0.00% | 1.41% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| F/B ratio | 0.68 | 0.90 | 0.55 | 0.98 | 0.66 | 1.15 | 0.81 | 0.70 | 1.24 | 0.88 | 0.59 | 1.36 | 0.33 | 1.03 | 0.81 |
References
- Reyes, A.; Semenkovich, N.P.; Whiteson, K.; Rohwer, F.; Gordon, J.I. Going viral: Next-generation sequencing applied to phage populations in the human gut. Nat. Rev. Microbiol. 2012, 10, 607–617. [Google Scholar] [CrossRef] [PubMed]
- Hamad, I.; Sokhna, C.; Raoult, D.; Bittar, F. Molecular detection of eukaryotes in a single human stool sample from Senegal. PLoS ONE 2012, 7, e40888. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, C.; Dollive, S.; Grunberg, S.; Chen, J.; Li, H.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Archaea and fungi of the human gut microbiome: Correlations with diet and bacterial residents. PLoS ONE 2013, 8, e66019. [Google Scholar] [CrossRef] [PubMed]
- Dusko Ehrlich, S.; MetaHIT Consortium. Métagénomique du microbiote intestinal: Les applications potentielles [Metagenomics of the intestinal microbiota: Potential applications]. Gastroenterol. Clin. Biol. 2010, 34 Suppl. 1, S23–S28. [Google Scholar] [CrossRef] [PubMed]
- Kau, A.L.; Ahern, P.P.; Griffin, N.W.; Goodman, A.L.; Gordon, J.I. Human nutrition, the gut microbiome and the immune system. Nature 2011, 474, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
- Kamada, N.; Seo, S.U.; Chen, G.Y.; Núñez, G. Role of the gut microbiota in immunity and inflammatory disease. Nat. Rev. Immunol. 2013, 13, 321–335. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, A.; Harty, S.; Lehto, S.M.; Moeller, A.H.; Dinan, T.G.; Dunbar, R.I.M.; Cryan, J.F.; Burnet, P.W.J. The Microbiome in Psychology and Cognitive Neuroscience. Trends Cogn. Sci. 2018, 22, 611–636. [Google Scholar] [CrossRef] [PubMed]
- Rampelli, S.; Turroni, S.; Mallol, C.; Hernandez, C.; Galván, B.; Sistiaga, A.; Biagi, E.; Astolfi, A.; Brigidi, P.; Benazzi, S.; et al. Components of a Neanderthal gut microbiome recovered from fecal sediments from El Salt. Commun. Biol. 2021, 4, 169. [Google Scholar] [CrossRef] [PubMed]
- Obregon-Tito, A.J.; Tito, R.Y.; Metcalf, J.; Sankaranarayanan, K.; Clemente, J.C.; Ursell, L.K.; Zech Xu, Z.; Van Treuren, W.; Knight, R.; Gaffney, P.M.; et al. Subsistence strategies in traditional societies distinguish gut microbiomes. Nat. Commun. 2015, 6, 6505. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.K.; Paul, S.; Dutta, C. Geography, Ethnicity or Subsistence-Specific Variations in Human Microbiome Composition and Diversity. Front. Microbiol. 2017, 8, 1162. [Google Scholar] [CrossRef]
- Hansen, M.E.B.; Rubel, M.A.; Bailey, A.G.; Ranciaro, A.; Thompson, S.R.; Campbell, M.C.; Beggs, W.; Dave, J.R.; Mokone, G.G.; Mpoloka, S.W.; et al. Population structure of human gut bacteria in a diverse cohort from rural Tanzania and Botswana. Genome Biol. 2019, 20, 16. [Google Scholar] [CrossRef] [PubMed]
- Rubel, M.A.; Abbas, A.; Taylor, L.J.; Connell, A.; Tanes, C.; Bittinger, K.; Ndze, V.N.; Fonsah, J.Y.; Ngwang, E.; Essiane, A.; et al. Lifestyle and the presence of helminths is associated with gut microbiome composition in Cameroonians. Genome Biol. 2020, 21, 122. [Google Scholar] [CrossRef] [PubMed]
- Dubois, G.; Girard, C.; Lapointe, F.J.; Shapiro, B.J. The Inuit gut microbiome is dynamic over time and shaped by traditional foods. Microbiome 2017, 5, 151. [Google Scholar] [CrossRef] [PubMed]
- Girard, C.; Tromas, N.; Amyot, M.; Shapiro, B.J. Gut Microbiome of the Canadian Arctic Inuit. mSphere 2017, 2, e00297-16. [Google Scholar] [CrossRef]
- De Filippo, C.; Di Paola, M.; Ramazzotti, M.; Albanese, D.; Pieraccini, G.; Banci, E.; Miglietta, F.; Cavalieri, D.; Lionetti, P. Diet, Environments, and Gut Microbiota. A Preliminary Investigation in Children Living in Rural and Urban Burkina Faso and Italy. Front. Microbiol. 2017, 8, 1979. [Google Scholar] [CrossRef]
- Afolayan, A.O.; Ayeni, F.A.; Moissl-Eichinger, C.; Gorkiewicz, G.; Halwachs, B.; Högenauer, C. Impact of a Nomadic Pastoral Lifestyle on the Gut Microbiome in the Fulani Living in Nigeria. Front. Microbiol. 2019, 10, 2138. [Google Scholar] [CrossRef]
- Conteville, L.C.; Oliveira-Ferreira, J.; Vicente, A.C.P. Gut Microbiome Biomarkers and Functional Diversity Within an Amazonian Semi-Nomadic Hunter-Gatherer Group. Front. Microbiol. 2019, 10, 1743. [Google Scholar] [CrossRef] [PubMed]
- Gomez, A.; Petrzelkova, K.J.; Burns, M.B.; Yeoman, C.J.; Amato, K.R.; Vlckova, K.; Modry, D.; Todd, A.; Jost Robinson, C.A.; Remis, M.J.; et al. Gut Microbiome of Coexisting BaAka Pygmies and Bantu Reflects Gradients of Traditional Subsistence Patterns. Cell Rep. 2016, 14, 2142–2153. [Google Scholar] [CrossRef]
- Sánchez-Quinto, A.; Cerqueda-García, D.; Falcón, L.I.; Gaona, O.; Martínez-Correa, S.; Nieto, J.; Santoyo, I.G. Gut Microbiome in Children from Indigenous and Urban Communities in México: Different Subsistence Models, Different Microbiomes. Microorganisms 2020, 8, 1592. [Google Scholar] [CrossRef]
- Escobar, J.S.; Klotz, B.; Valdes, B.E.; Agudelo, G.M. The gut microbiota of Colombians differs from that of Americans, Europeans and Asians. BMC Microbiol. 2014, 14, 311. [Google Scholar] [CrossRef]
- Kann, S.; Bruennert, D.; Hansen, J.; Mendoza, G.A.C.; Gonzalez, J.J.C.; Quintero, C.L.A.; Hanke, M.; Hagen, R.M.; Backhaus, J.; Frickmann, H. High Prevalence of Intestinal Pathogens in Indigenous in Colombia. J. Clin. Med. 2020, 9, 2786. [Google Scholar] [CrossRef] [PubMed]
- Frickmann, H.; Schwarz, N.G.; Rakotozandrindrainy, R.; May, J.; Hagen, R.M. PCR for enteric pathogens in high-prevalence settings. What does a positive signal tell us? Infect. Dis. 2015, 47, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Krumkamp, R.; Sarpong, N.; Schwarz, N.G.; Adlkofer, J.; Loag, W.; Eibach, D.; Hagen, R.M.; Adu-Sarkodie, Y.; Tannich, E.; May, J. Gastrointestinal infections and diarrheal disease in Ghanaian infants and children: An outpatient case-control study. PLoS Negl. Trop. Dis. 2015, 9, e0003568. [Google Scholar]
- Eibach, D.; Krumkamp, R.; Hahn, A.; Sarpong, N.; Adu-Sarkodie, Y.; Leva, A.; Käsmaier, J.; Panning, M.; May, J.; Tannich, E. Application of a multiplex PCR assay for the detection of gastrointestinal pathogens in a rural African setting. BMC Infect. Dis. 2016, 16, 150. [Google Scholar] [CrossRef]
- Zautner, A.E.; Groß, U.; Emele, M.F.; Hagen, R.M.; Frickmann, H. More Pathogenicity or Just More Pathogens?-On the Interpretation Problem of Multiple Pathogen Detections with Diagnostic Multiplex Assays. Front. Microbiol. 2017, 8, 1210. [Google Scholar] [CrossRef]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef] [PubMed]
- Clemente, J.C.; Pehrsson, E.C.; Blaser, M.J.; Sandhu, K.; Gao, Z.; Wang, B.; Magris, M.; Hidalgo, G.; Contreras, M.; Noya-Alarcón, Ó.; et al. The microbiome of uncontacted Amerindians. Sci. Adv. 2015, 1, e1500183. [Google Scholar] [CrossRef]
- Wiemer, D.; Loderstaedt, U.; von Wulffen, H.; Priesnitz, S.; Fischer, M.; Tannich, E.; Hagen, R.M. Real-time multiplex PCR for simultaneous detection of Campylobacter jejuni, Salmonella, Shigella and Yersinia species in fecal samples. Int. J. Med. Microbiol. 2011, 301, 577–584. [Google Scholar] [CrossRef]
- Köller, T.; Hahn, A.; Altangerel, E.; Verweij, J.J.; Landt, O.; Kann, S.; Dekker, D.; May, J.; Loderstädt, U.; Podbielski, A.; et al. Comparison of commercial and in-house real-time PCR platforms for 15 parasites and microsporidia in human stool samples without a gold standard. Acta Trop. 2020, 207, 105516. [Google Scholar] [CrossRef]
- Hahn, A.; Luetgehetmann, M.; Landt, O.; Schwarz, N.G.; Frickmann, H. Comparison of one commercial and two in-house TaqMan multiplex real-time PCR assays for detection of enteropathogenic, enterotoxigenic and enteroaggregative Escherichia coli. Trop. Med. Int. Health 2017, 22, 1371–1376. [Google Scholar] [CrossRef] [PubMed]
- Herlemann, D.P.; Labrenz, M.; Jürgens, K.; Bertilsson, S.; Waniek, J.J.; Andersson, A.F. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 2011, 5, 1571–1579. [Google Scholar] [CrossRef] [PubMed]
- Masella, A.P.; Bartram, A.K.; Truszkowski, J.M.; Brown, D.G.; Neufeld, J.D. PANDAseq: Paired-end assembler for illumina sequences. BMC Bioinform. 2012, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460-1. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef]
- Comeau, A.M.; Douglas, G.M.; Langille, M.G. Microbiome Helper: A Custom and Streamlined Workflow for Microbiome Research. mSystems 2017, 2, e00127-16. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Salonen, A.; Salojärvi, J.; Lahti, L.; de Vos, W.M. The adult intestinal core microbiota is determined by analysis depth and health status. Clin. Microbiol. Infect. 2012, 18 Suppl. 4, 16–20. [Google Scholar] [CrossRef]
- Shetty, S.A.; Hugenholtz, F.; Lahti, L.; Smidt, H.; de Vos, W.M. Intestinal microbiome landscaping: Insight in community assemblage and implications for microbial modulation strategies. FEMS Microbiol. Rev. 2017, 41, 182–199. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Furumichi, M.; Tanabe, M.; Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 2010, 38, D355–D360. [Google Scholar] [CrossRef]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef] [PubMed]
- Yun, Y.; Kim, H.N.; Kim, S.E.; Heo, S.G.; Chang, Y.; Ryu, S.; Shin, H.; Kim, H.L. Comparative analysis of gut microbiota associated with body mass index in a large Korean cohort. BMC Microbiol. 2017, 17, 151. [Google Scholar] [CrossRef]
- Stanislawski, M.A.; Lozupone, C.A.; Wagner, B.D.; Eggesbø, M.; Sontag, M.K.; Nusbacher, N.M.; Martinez, M.; Dabelea, D. Gut microbiota in adolescents and the association with fatty liver: The EPOCH study. Pediatr. Res. 2018, 84, 219–227. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef]
- Paniagua Voirol, L.R.; Weinhold, A.; Johnston, P.R.; Fatouros, N.E.; Hilker, M. Legacy of a Butterfly’s Parental Microbiome in Offspring Performance. Appl. Environ. Microbiol. 2020, 86, e00596-20. [Google Scholar] [CrossRef]
- García-Vega, Á.S.; Corrales-Agudelo, V.; Reyes, A.; Escobar, J.S. Diet Quality, Food Groups and Nutrients Associated with the Gut Microbiota in a Nonwestern Population. Nutrients 2020, 12, 2938. [Google Scholar] [CrossRef]
- Sharma, A.K.; Petrzelkova, K.; Pafco, B.; Jost Robinson, C.A.; Fuh, T.; Wilson, B.A.; Stumpf, R.M.; Torralba, M.G.; Blekhman, R.; White, B.; et al. Traditional Human Populations and Nonhuman Primates Show Parallel Gut Microbiome Adaptations to Analogous Ecological Conditions. mSystems 2020, 5, e00815-20. [Google Scholar] [CrossRef]
- Jha, A.R.; Davenport, E.R.; Gautam, Y.; Bhandari, D.; Tandukar, S.; Ng, K.M.; Fragiadakis, G.K.; Holmes, S.; Gautam, G.P.; Leach, J.; et al. Gut microbiome transition across a lifestyle gradient in Himalaya. PLoS Biol. 2018, 16, e2005396. [Google Scholar] [CrossRef]
- Horwood, P.F.; Tarantola, A.; Goarant, C.; Matsui, M.; Klement, E.; Umezaki, M.; Navarro, S.; Greenhill, A.R. Health Challenges of the Pacific Region: Insights From History, Geography, Social Determinants, Genetics, and the Microbiome. Front. Immunol. 2019, 10, 2184. [Google Scholar] [CrossRef]
- Ruggles, K.V.; Wang, J.; Volkova, A.; Contreras, M.; Noya-Alarcon, O.; Lander, O.; Caballero, H.; Dominguez-Bello, M.G. Changes in the Gut Microbiota of Urban Subjects during an Immersion in the Traditional Diet and Lifestyle of a Rainforest Village. mSphere 2018, 3, e00193-18. [Google Scholar] [CrossRef] [PubMed]
- Afolayan, A.O.; Adebusoye, L.A.; Cadmus, E.O.; Ayeni, F.A. Insights into the gut microbiota of Nigerian elderly with type 2 diabetes and non-diabetic elderly persons. Heliyon 2020, 6, e03971. [Google Scholar] [CrossRef] [PubMed]
- Martínez, I.; Stegen, J.C.; Maldonado-Gómez, M.X.; Eren, A.M.; Siba, P.M.; Greenhill, A.R.; Walter, J. The gut microbiota of rural papua new guineans: Composition, diversity patterns, and ecological processes. Cell. Rep. 2015, 11, 527–538. [Google Scholar] [CrossRef]
- Smits, S.A.; Leach, J.; Sonnenburg, E.D.; Gonzalez, C.G.; Lichtman, J.S.; Reid, G.; Knight, R.; Manjurano, A.; Changalucha, J.; Elias, J.E.; et al. Seasonal cycling in the gut microbiome of the Hadza hunter-gatherers of Tanzania. Science 2017, 357, 802–806. [Google Scholar] [CrossRef] [PubMed]
- Kann, S.; Concha, G.; Hartmann, M.; Köller, T.; Alker, J.; Schotte, U.; Kreienbrock, L.; Frickmann, H.; Warnke, P. Only Low Effects of Water Filters on the Enteric Carriage of Gastrointestinal Pathogen DNA in Colombian Indigenous People. Microorganisms 2022, 10, 658. [Google Scholar] [CrossRef]
- Weinreich, F.; Hahn, A.; Eberhardt, K.A.; Kann, S.; Köller, T.; Warnke, P.; Dupke, S.; Dekker, D.; May, J.; Frickmann, H.; et al. Multicentric Evaluation of SeeGene Allplex Real-Time PCR Assays Targeting 28 Bacterial, Microsporidal and Parasitic Nucleic Acid Sequences in Human Stool Samples. Diagnostics 2022, 12, 1007. [Google Scholar] [CrossRef]
- Kann, S.; Concha, G.; Köller, T.; Alker, J.; Schotte, U.; Hahn, A.; Frickmann, H.; Warnke, P. Enteric Bacteria and Parasites with Pathogenic Potential in Individuals of the Colombian Indigenous Tribe Kogui. Microorganisms 2022, 10, 1862. [Google Scholar] [CrossRef]
- Rosen, C.E.; Palm, N.W. Functional Classification of the Gut Microbiota: The Key to Cracking the Microbiota Composition Code: Functional classifications of the gut microbiota reveal previously hidden contributions of indigenous gut bacteria to human health and disease. Bioessays 2017, 39, 12. [Google Scholar] [CrossRef] [PubMed]
- Mir, R.A.; Schaut, R.G.; Allen, H.K.; Looft, T.; Loving, C.L.; Kudva, I.T.; Sharma, V.K. Cattle intestinal microbiota shifts following Escherichia coli O157:H7 vaccination and colonization. PLoS ONE 2019, 14, e0226099. [Google Scholar] [CrossRef]
- Wang, X.; Wu, X.; Cong, X.; Ren, J.; Li, J.; Zhu, J.; Dai, M.; Hrabchenko, N.; Du, Y.; Qi, J. The functional role of fecal microbiota transplantation on Salmonella Enteritidis infection in chicks. Vet. Microbiol. 2022, 269, 109449. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Song, Q.; Liu, J.; Chen, F.; Zhang, Y.; Wu, Z.; Sun, X.; Wu, X. Potential Gut Microbiota Features for Non-Invasive Detection of Schistosomiasis. Front. Immunol. 2022, 13, 941530. [Google Scholar] [CrossRef] [PubMed]
- Paz, E.A.; Chua, E.G.; Hassan, S.U.; Greeff, J.C.; Palmer, D.G.; Liu, S.; Lamichhane, B.; Sepúlveda, N.; Liu, J.; Tay, C.Y.; et al. Bacterial communities in the gastrointestinal tract segments of helminth-resistant and helminth-susceptible sheep. Anim. Microbiome 2022, 4, 23. [Google Scholar] [CrossRef] [PubMed]
- Latorre, J.D.; Adhikari, B.; Park, S.H.; Teague, K.D.; Graham, L.E.; Mahaffey, B.D.; Baxter, M.F.A.; Hernandez-Velasco, X.; Kwon, Y.M.; Ricke, S.C.; et al. Evaluation of the Epithelial Barrier Function and Ileal Microbiome in an Established Necrotic Enteritis Challenge Model in Broiler Chickens. Front. Vet. Sci. 2018, 5, 199. [Google Scholar] [CrossRef] [PubMed]
- Morton, E.R.; Lynch, J.; Froment, A.; Lafosse, S.; Heyer, E.; Przeworski, M.; Blekhman, R.; Ségurel, L. Variation in Rural African Gut Microbiota Is Strongly Correlated with Colonization by Entamoeba and Subsistence. PLoS Genet. 2015, 11, e1005658. [Google Scholar] [CrossRef]
- López-Colom, P.; Castillejos, L.; Rodríguez-Sorrento, A.; Puyalto, M.; Mallo, J.J.; Martín-Orúe, S.M. Efficacy of medium-chain fatty acid salts distilled from coconut oil against two enteric pathogen challenges in weanling piglets. J. Anim. Sci. Biotechnol. 2019, 10, 89. [Google Scholar] [CrossRef]
| Genus | Category | p-Value |
|---|---|---|
| Butyrivibrio | BMI:Overweight | 0.0401 |
| Butyrivibrio | location:Tezhumake | 0.0007 |
| Butyrivibrio | location:Valledupar | 0.0009 |
| Butyrivibrio | Salmonella:Yes | 0.0204 |
| Clostridium | Age:Child Under 5 | 0.0244 |
| Clostridium | location:Tezhumake | 0.0011 |
| Dialister | location:Tezhumake | 0.0209 |
| Dialister | location:Valledupar | 0.0451 |
| Dialister | symptoms.presence:Yes | 0.0480 |
| Faecalibacterium | EPEC:Yes | 0.0134 |
| Faecalibacterium | Hymenolepis:Yes | 0.0261 |
| Faecalibacterium | location:Tezhumake | 0.0276 |
| Faecalibacterium | location:Valledupar | 0.0301 |
| Faecalibacterium | Salmonella:Yes | 0.0009 |
| Odoribacter | BMI:Obesity | 0.0494 |
| Odoribacter | BMI:Overweight | 0.0169 |
| Odoribacter | location:Valledupar | 1.26 × 10−14 |
| Parabacteroides | Enterobius:Yes | 0.0349 |
| Parabacteroides | location:Valledupar | 0.0093 |
| Prevotella | EPEC:Yes | 0.0134 |
| Prevotella | location:Tezhumake | 0.0392 |
| Sutterella | EHEC:Yes | 0.0431 |
| Sutterella | EPEC:Yes | 0.0464 |
| Sutterella | location:Tezhumake | 0.0020 |
| Sutterella | location:Valledupar | 0.0133 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kann, S.; Eberhardt, K.; Hinz, R.; Schwarz, N.G.; Dib, J.C.; Aristizabal, A.; Mendoza, G.A.C.; Hagen, R.M.; Frickmann, H.; Barrantes, I.; et al. The Gut Microbiome of an Indigenous Agropastoralist Population in a Remote Area of Colombia with High Rates of Gastrointestinal Infections and Dysbiosis. Microorganisms 2023, 11, 625. https://doi.org/10.3390/microorganisms11030625
Kann S, Eberhardt K, Hinz R, Schwarz NG, Dib JC, Aristizabal A, Mendoza GAC, Hagen RM, Frickmann H, Barrantes I, et al. The Gut Microbiome of an Indigenous Agropastoralist Population in a Remote Area of Colombia with High Rates of Gastrointestinal Infections and Dysbiosis. Microorganisms. 2023; 11(3):625. https://doi.org/10.3390/microorganisms11030625
Chicago/Turabian StyleKann, Simone, Kirsten Eberhardt, Rebecca Hinz, Norbert Georg Schwarz, Juan Carlos Dib, Andres Aristizabal, Gustavo Andrés Concha Mendoza, Ralf Matthias Hagen, Hagen Frickmann, Israel Barrantes, and et al. 2023. "The Gut Microbiome of an Indigenous Agropastoralist Population in a Remote Area of Colombia with High Rates of Gastrointestinal Infections and Dysbiosis" Microorganisms 11, no. 3: 625. https://doi.org/10.3390/microorganisms11030625
APA StyleKann, S., Eberhardt, K., Hinz, R., Schwarz, N. G., Dib, J. C., Aristizabal, A., Mendoza, G. A. C., Hagen, R. M., Frickmann, H., Barrantes, I., & Kreikemeyer, B. (2023). The Gut Microbiome of an Indigenous Agropastoralist Population in a Remote Area of Colombia with High Rates of Gastrointestinal Infections and Dysbiosis. Microorganisms, 11(3), 625. https://doi.org/10.3390/microorganisms11030625






