Detection and Characterization of Alongshan Virus in Ticks and Tick Saliva from Lower Saxony, Germany with Serological Evidence for Viral Transmission to Game and Domestic Animals
Abstract
1. Introduction
2. Materials and Methods
2.1. Tick Sampling
2.2. Collection of Tick Saliva
2.3. RNA Isolation
2.4. RT-PCR for Detection of NS5-like Gene and Capsid/Membrane Gene of ALSV and RT-qPCR for Virus Quantification
2.5. Virus Cell Culture Experiments
2.6. In Vivo Tick ALSV Injection Experiments
2.7. In Vitro ALSV Feeding Experiments
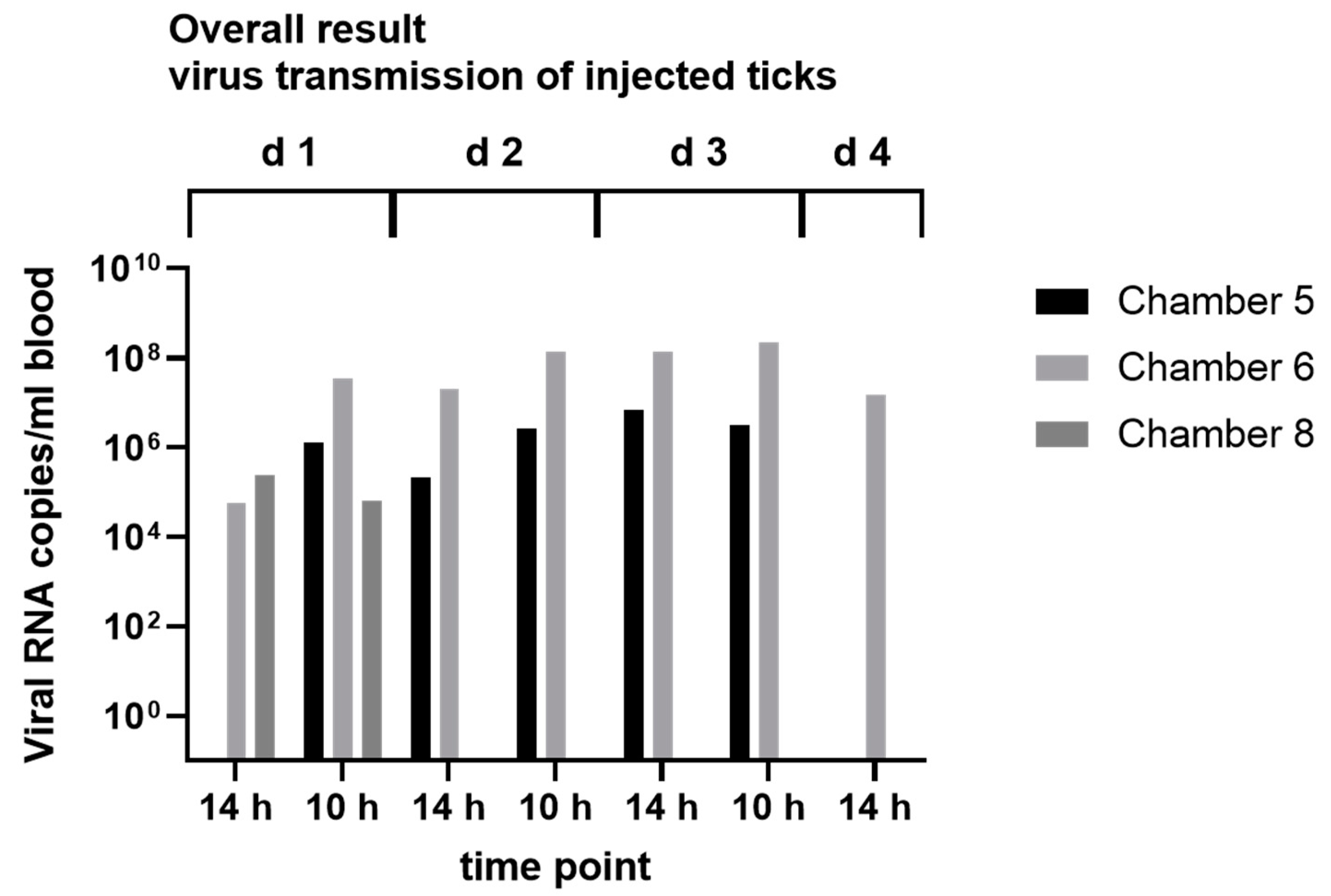
2.8. Virus Transmission Experiments
2.9. Collection of Animal Serum
2.10. Serological Screening by Luciferase Immunoprecipitation System (LIPS) Assay
3. Results
3.1. Prevalence of ALSV in Lower Saxony
3.2. Cell Culture Experiments
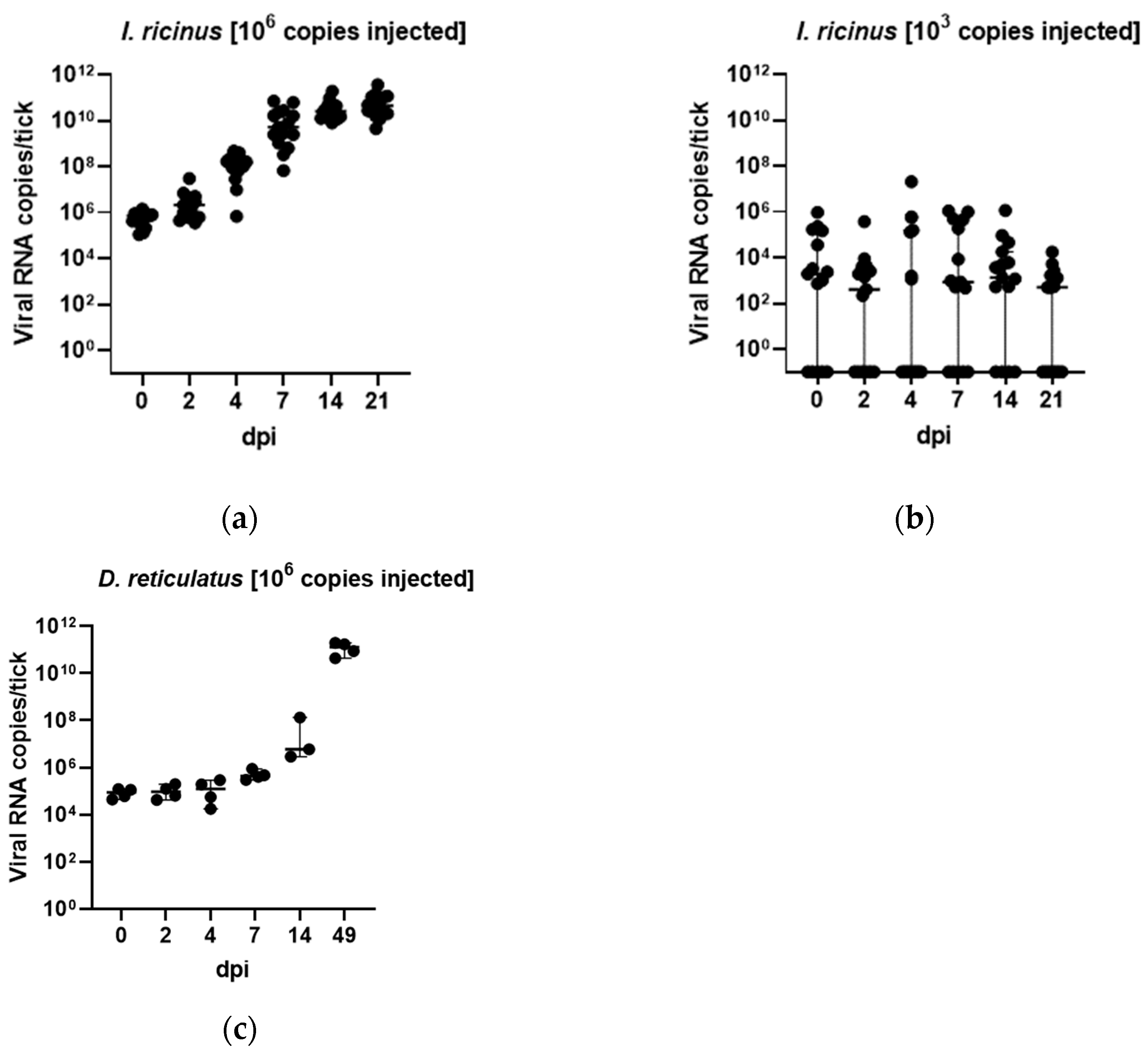
3.3. In Vivo Tick Infection Experiments
3.4. In Vitro Feeding Experiments
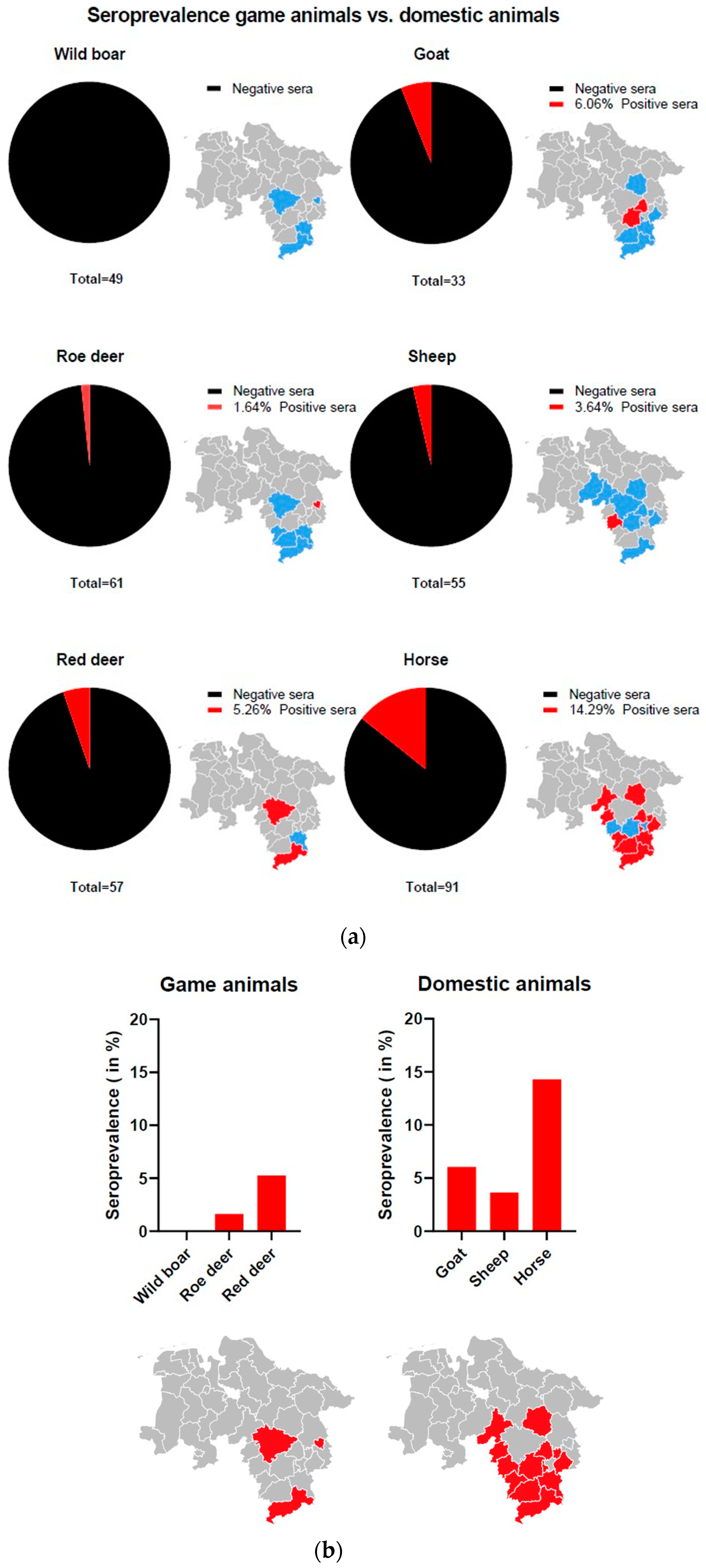
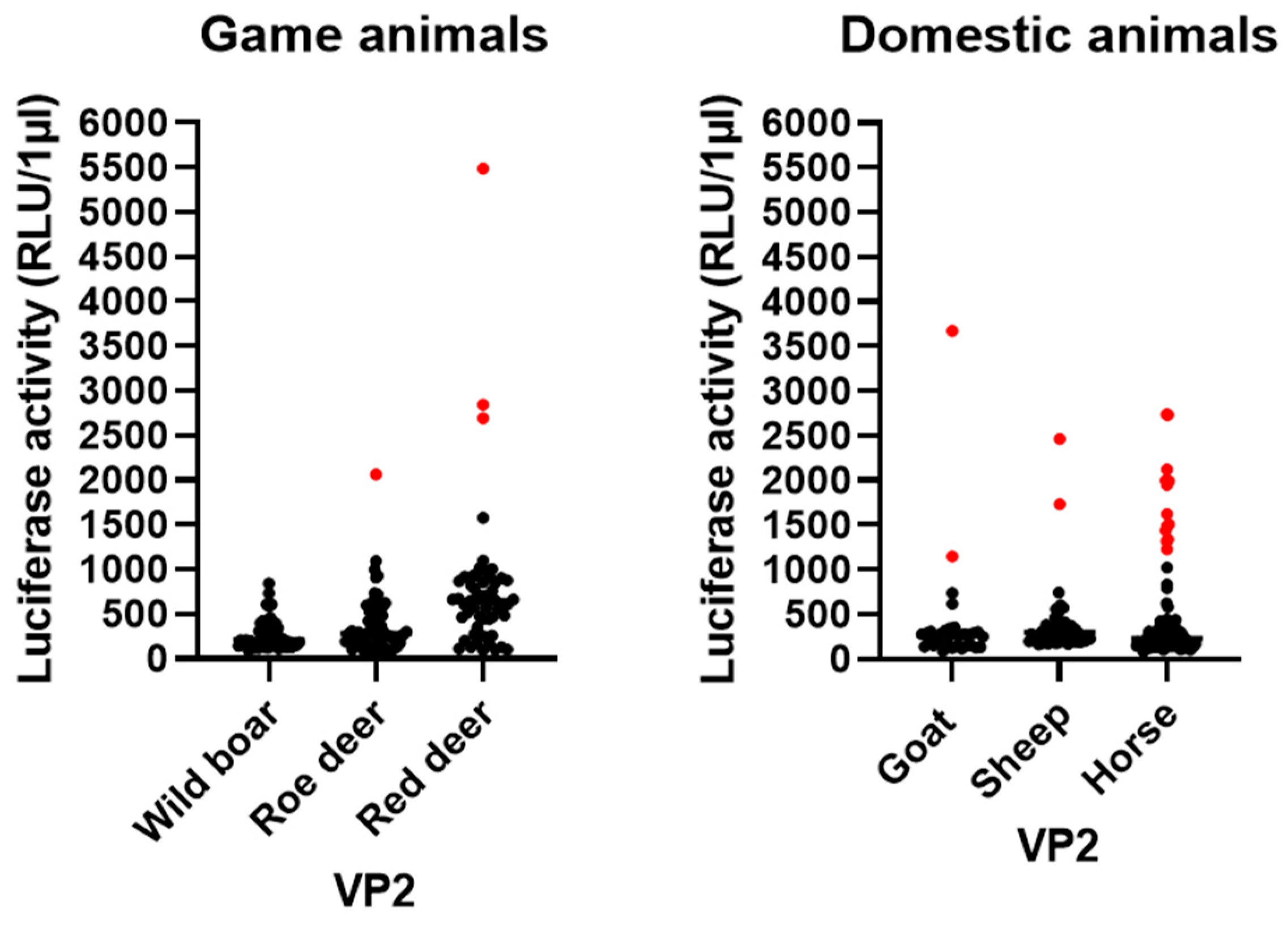
3.5. LIPS Assay
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shuman, E.D. Global climate change and infectious diseases. N. Engl. J. Med. 2010, 362, 1061–1063. [Google Scholar]
- Keesing, F.; Belden, L.K.; Daszak, P.; Dobson, A.; Harvell, C.D.; Holt, R.D.; Hudson, P.; Jolles, A.; Jones, K.E.; Mitchell, C.E. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 2010, 468, 647–652. [Google Scholar]
- Bouchard, C.; Dibernardo, A.; Koffi, J.; Wood, H.; Leighton, P.; Lindsay, L. Climate change and infectious diseases: The challenges: N increased risk of tick-borne diseases with climate and environmental changes. Can. Commun. Dis. Rep. 2019, 45, 83. [Google Scholar]
- Barzon, L. Ongoing and emerging arbovirus threats in Europe. J. Clin. Virol. 2018, 107, 38–47. [Google Scholar]
- Deplazes, P.; Joachim, A.; Mathis, A.; Strube, C.; Taubert, A.; Samson-Himmelstjerna, G.; Zahner, H. Parasitologie für Die Tiermedizin; Thieme: New York, NY, USA, 2021. [Google Scholar]
- Rizzoli, A.; Silaghi, C.; Obiegala, A.; Rudolf, I.; Hubálek, Z.; Földvári, G.; Plantard, O.; Vayssier-Taussat, M.; Bonnet, S.; Špitalská, E. Ixodes ricinus and its transmitted pathogens in urban and peri-urban areas in Europe: New hazards and relevance for public health. Front. Public Health 2014, 2, 251. [Google Scholar]
- Kilpatrick, A.M.; Randolph, S.E. Drivers, dynamics, and control of emerging vector-borne zoonotic diseases. Lancet 2012, 380, 1946–1955. [Google Scholar]
- Kuehn, B. Tickborne diseases increasing. JAMA 2019, 321, 138. [Google Scholar]
- Tokarz, R.; Williams, S.H.; Sameroff, S.; Sanchez Leon, M.; Jain, K.; Lipkin, W.I. Virome analysis of Amblyomma americanum, Dermacentor variabilis, and Ixodes scapularis ticks reveals novel highly divergent vertebrate and invertebrate viruses. J. Virol. 2014, 88, 11480–11492. [Google Scholar]
- Brinkmann, A.; Dinçer, E.; Polat, C.; Hekimoğlu, O.; Hacıoğlu, S.; Földes, K.; Özkul, A.; Öktem, İ.M.A.; Nitsche, A.; Ergünay, K. A metagenomic survey identifies Tamdy orthonairovirus as well as divergent phlebo-, rhabdo-, chu-and flavi-like viruses in Anatolia, Turkey. Ticks Tick-Borne Dis. 2018, 9, 1173–1183. [Google Scholar]
- Shi, J.; Shen, S.; Wu, H.; Zhang, Y.; Deng, F. Metagenomic profiling of viruses associated with Rhipicephalus microplus ticks in Yunnan Province, China. Virol. Sin. 2021, 36, 623–635. [Google Scholar]
- Qin, X.-C.; Shi, M.; Tian, J.-H.; Lin, X.-D.; Gao, D.-Y.; He, J.-R.; Wang, J.-B.; Li, C.-X.; Kang, Y.-J.; Yu, B. A tick-borne segmented RNA virus contains genome segments derived from unsegmented viral ancestors. Proc. Natl. Acad. Sci. USA 2014, 111, 6744–6749. [Google Scholar]
- Villa, E.C.; Maruyama, S.R.; de Miranda-Santos, I.K.; Palacios, G.; Ladner, J.T. Complete coding genome sequence for Mogiana tick virus, a Jingmenvirus isolated from ticks in Brazil. Genome Announc. 2017, 5, e00232-17. [Google Scholar]
- Zhang, X.; Wang, N.; Wang, Z.; Liu, Q. The discovery of segmented flaviviruses: Implications for viral emergence. Curr. Opin. Virol. 2020, 40, 11–18. [Google Scholar]
- Emmerich, P.; Jakupi, X.; von Possel, R.; Berisha, L.; Halili, B.; Günther, S.; Cadar, D.; Ahmeti, S.; Schmidt-Chanasit, J. Viral metagenomics, genetic and evolutionary characteristics of Crimean-Congo hemorrhagic fever orthonairovirus in humans, Kosovo. Infect. Genet. Evol. 2018, 65, 6–11. [Google Scholar]
- Ladner, J.T.; Wiley, M.R.; Beitzel, B.; Auguste, A.J.; Dupuis II, A.P.; Lindquist, M.E.; Sibley, S.D.; Kota, K.P.; Fetterer, D.; Eastwood, G. A multicomponent animal virus isolated from mosquitoes. Cell Host Microbe 2016, 20, 357–367. [Google Scholar]
- Maruyama, S.R.; Castro-Jorge, L.A.; Ribeiro, J.M.C.; Gardinassi, L.G.; Garcia, G.R.; Brandão, L.G.; Rodrigues, A.R.; Okada, M.I.; Abrão, E.P.; Ferreira, B.R. Characterisation of divergent flavivirus NS3 and NS5 protein sequences detected in Rhipicephalus microplus ticks from Brazil. Memórias Do Inst. Oswaldo Cruz 2013, 109, 38–50. [Google Scholar]
- De Oliveira Pascoal, J.; de Siqueira, S.M.; da Costa Maia, R.; Szabó, M.P.J.; Yokosawa, J. Detection and molecular characterization of Mogiana tick virus (MGTV) in Rhipicephalus microplus collected from cattle in a savannah area, Uberlândia, Brazil. Ticks Tick-Borne Dis. 2019, 10, 162–165. [Google Scholar]
- Souza, W.M.d.; Fumagalli, M.J.; Torres Carrasco, A.d.O.; Romeiro, M.F.; Modha, S.; Seki, M.C.; Gheller, J.M.; Daffre, S.; Nunes, M.R.T.; Murcia, P.R. Viral diversity of Rhipicephalus microplus parasitizing cattle in southern Brazil. Sci. Rep. 2018, 8, 16315. [Google Scholar]
- Webster, C.L.; Waldron, F.M.; Robertson, S.; Crowson, D.; Ferrari, G.; Quintana, J.F.; Brouqui, J.-M.; Bayne, E.H.; Longdon, B.; Buck, A.H. The discovery, distribution, and evolution of viruses associated with Drosophila melanogaster. PLoS Biol. 2015, 13, e1002210. [Google Scholar]
- Kuivanen, S.; Levanov, L.; Kareinen, L.; Sironen, T.; Jääskeläinen, A.J.; Plyusnin, I.; Zakham, F.; Emmerich, P.; Schmidt-Chanasit, J.; Hepojoki, J. Detection of novel tick-borne pathogen, Alongshan virus, in Ixodes ricinus ticks, south-eastern Finland, 2019. Eurosurveillance 2019, 24, 1900394. [Google Scholar]
- Temmam, S.; Bigot, T.; Chrétien, D.; Gondard, M.; Pérot, P.; Pommelet, V.; Dufour, E.; Petres, S.; Devillers, E.; Hoem, T. Insights into the host range, genetic diversity, and geographical distribution of jingmenviruses. MSphere 2019, 4, e00645-19. [Google Scholar]
- Dinçer, E.; Hacıoğlu, S.; Kar, S.; Emanet, N.; Brinkmann, A.; Nitsche, A.; Özkul, A.; Linton, Y.-M.; Ergünay, K. Survey and characterization of Jingmen tick virus variants. Viruses 2019, 11, 1071. [Google Scholar]
- Kholodilov, I.S.; Litov, A.G.; Klimentov, A.S.; Belova, O.A.; Polienko, A.E.; Nikitin, N.A.; Shchetinin, A.M.; Ivannikova, A.Y.; Bell-Sakyi, L.; Yakovlev, A.S. Isolation and characterisation of Alongshan virus in Russia. Viruses 2020, 12, 362. [Google Scholar]
- Gofton, A.W.; Blasdell, K.R.; Taylor, C.; Banks, P.B.; Michie, M.; Roy-Dufresne, E.; Poldy, J.; Wang, J.; Dunn, M.; Tachedjian, M. Metatranscriptomic profiling reveals diverse tick-borne bacteria, protozoans and viruses in ticks and wildlife from Australia. Transbound. Emerg. Dis. 2022, 69, e2389–e2407. [Google Scholar]
- Ogola, E.O.; Kopp, A.; Bastos, A.D.; Slothouwer, I.; Marklewitz, M.; Omoga, D.; Rotich, G.; Getugi, C.; Sang, R.; Torto, B. Jingmen tick virus in ticks from Kenya. Viruses 2022, 14, 1041. [Google Scholar]
- Jia, N.; Liu, H.-B.; Ni, X.-B.; Bell-Sakyi, L.; Zheng, Y.-C.; Song, J.-L.; Li, J.; Jiang, B.-G.; Wang, Q.; Sun, Y. Emergence of human infection with Jingmen tick virus in China: A retrospective study. EBioMedicine 2019, 43, 317–324. [Google Scholar]
- Wang, Z.-D.; Wang, B.; Wei, F.; Han, S.-Z.; Zhang, L.; Yang, Z.-T.; Yan, Y.; Lv, X.-L.; Li, L.; Wang, S.-C. A new segmented virus associated with human febrile illness in China. N. Engl. J. Med. 2019, 380, 2116–2125. [Google Scholar]
- Wang, Z.-D.; Wang, W.; Wang, N.-N.; Qiu, K.; Zhang, X.; Tana, G.; Liu, Q.; Zhu, X.-Q. Prevalence of the emerging novel Alongshan virus infection in sheep and cattle in Inner Mongolia, northeastern China. Parasites Vectors 2019, 12, 450. [Google Scholar]
- Wu, Z.; Chen, J.; Zhang, L.; Zhang, Y.; Liu, L.; Niu, G. Molecular evidence for potential transovarial transmission of Jingmen tick virus in Haemaphysalis longicornis fed on cattle from Yunnan Province, China. J. Med. Virol. 2022, 95, e28357. [Google Scholar]
- Zhang, Y.; Li, Z.; Pang, Z.; Wu, Z.; Lin, Z.; Niu, G. Identification of Jingmen tick virus (JMTV) in Amblyomma testudinarium from Fujian Province, southeastern China. Parasites Vectors 2022, 15, 339. [Google Scholar]
- Nuttall, P.A. Molecular characterization of tick-virus interactions. Front. Biosci. Landmark 2009, 14, 2466–2483. [Google Scholar]
- Li, L.-F.; Zhang, M.-Z.; Zhu, J.-G.; Cui, X.-M.; Zhang, C.-F.; Niu, T.-Y.; Li, J.; Sun, Y.; Wei, W.; Liu, H.-B.; et al. Dermacentor silvarum, a Medically Important Tick, May Not Be a Competent Vector to Transmit Jingmen Tick Virus. Vector-Borne Zoonotic Dis. 2022, 22, 402–407. [Google Scholar]
- Nuttall, P.A.; Jones, L.D.; Labuda, M.; Kaufman, W.R. Adaptations of arboviruses to ticks. J. Med. Entomol. 1994, 31, 1–9. [Google Scholar]
- Kobayashi, D.; Kuwata, R.; Kimura, T.; Shimoda, H.; Fujita, R.; Faizah, A.N.; Kai, I.; Matsumura, R.; Kuroda, Y.; Watanabe, S. Detection of Jingmenviruses in Japan with Evidence of Vertical Transmission in Ticks. Viruses 2021, 13, 2547. [Google Scholar]
- Boelke, M.; Bestehorn, M.; Marchwald, B.; Kubinski, M.; Liebig, K.; Glanz, J.; Schulz, C.; Dobler, G.; Monazahian, M.; Becker, S.C. First isolation and phylogenetic analyses of tick-borne encephalitis virus in Lower Saxony, Germany. Viruses 2019, 11, 462. [Google Scholar]
- Knoll, S.; Springer, A.; Hauck, D.; Schunack, B.; Pachnicke, S.; Fingerle, V.; Strube, C. Distribution of Borrelia burgdorferi s.l. and Borrelia miyamotoi in Ixodes tick populations in Northern Germany, co-infections with Rickettsiales and assessment of potential influencing factors. Med. Vet. Entomol. 2021, 35, 595–606. [Google Scholar]
- Patton, T.G.; Dietrich, G.; Brandt, K.; Dolan, M.C.; Piesman, J.; Gilmore, R.D., Jr. Saliva, salivary gland, and hemolymph collection from Ixodes scapularis ticks. JoVE 2012, e3894. [Google Scholar] [CrossRef]
- Liebig, K.; Boelke, M.; Grund, D.; Schicht, S.; Springer, A.; Strube, C.; Chitimia-Dobler, L.; Dobler, G.; Jung, K.; Becker, S.C. Tick populations from endemic and non-endemic areas in Germany show differential susceptibility to TBEV. Sci. Rep. 2020, 10, 1–10. [Google Scholar]
- Kröber, T.; Guerin, P.M. An in vitro feeding assay to test acaricides for control of hard ticks. Pest Manag. Sci. Former. Pestic. Sci. 2007, 63, 17–22. [Google Scholar]
- Burbelo, P.D.; Ching, K.H.; Klimavicz, C.M.; Iadarola, M.J. Antibody profiling by luciferase immunoprecipitation systems (LIPS). J. Vis. Exp. 2009, e1549. [Google Scholar] [CrossRef]
- Vayssier-Taussat, M.; Kazimirova, M.; Hubalek, Z.; Hornok, S.; Farkas, R.; Cosson, J.-F.; Bonnet, S.; Vourch, G.; Gasqui, P.; Mihalca, A.D. Emerging horizons for tick-borne pathogens: From the ‘one pathogen–one disease’vision to the pathobiome paradigm. Future Microbiol. 2015, 10, 2033–2043. [Google Scholar]
- Rubel, F.; Brugger, K.; Monazahian, M.; Habedank, B.; Dautel, H.; Leverenz, S.; Kahl, O. The first German map of georeferenced ixodid tick locations. Parasites Vectors 2014, 7, 477. [Google Scholar]
- Weidmann, M.; Sall, A.A.; Manuguerra, J.-C.; Koivogui, L.; Adjami, A.; Traoré, F.F.; Hedlund, K.-O.; Lindegren, G.; Mirazimi, A. Quantitative analysis of particles, genomes and infectious particles in supernatants of haemorrhagic fever virus cell cultures. Virol. J. 2011, 8, 81. [Google Scholar]
- Yezli, S.; Otter, J.A. Minimum infective dose of the major human respiratory and enteric viruses transmitted through food and the environment. Food Environ. Virol. 2011, 3, 279. [Google Scholar]
- Bonnet, S.; Liu, X. Laboratory artificial infection of hard ticks: A tool for the analysis of tick-borne pathogen transmission. Acarologia 2012, 52, 453–464. [Google Scholar]
- Belova, O.A.; Burenkova, L.A.; Karganova, G.G. Different tick-borne encephalitis virus (TBEV) prevalences in unfed versus partially engorged ixodid ticks–evidence of virus replication and changes in tick behavior. Ticks Tick-Borne Dis. 2012, 3, 240–246. [Google Scholar]
- Labuda, M.; Nuttall, P.; Kožuch, O.; Elečková, E.; Williams, T.; Žuffová, E.; Sabo, A. Non-viraemic transmission of tick-borne encephalitis virus: A mechanism for arbovirus survival in nature. Experientia 1993, 49, 802–805. [Google Scholar]
- Wolfe, N.D.; Dunavan, C.P.; Diamond, J. Origins of major human infectious diseases. Nature 2007, 447, 279–283. [Google Scholar]
| Region | Sampled Animals | Number of Ticks |
|---|---|---|
| Wolfsburg | 25 | 254 |
| Hanover | 11 | 40 |
| Harz | 48 | 690 |
| Total | 84 | 984 |
| Region | Pools | Positive Pools [Positive/Total] | Red Deer [Positive/Total] | Roe Deer [Positive/Total] | Fallow Deer [Positive/Total] |
|---|---|---|---|---|---|
| Wolfsburg | 25 | 8/25 | 0/2 | 8/22 | 0/1 |
| Harz | 43 | 27/43 | 27/41 | 0/2 | 0 |
| Hanover | 16 | 4/16 | 3/12 | 1/4 | 0 |
| Total | 84 | 39/84 | 30/55 | 9/28 | 0/1 |
| Date | Virus Copies/mL Blood | Incubation [Day]of Different Tick Samples | Positive Body [Positive/Total] | Positive Peripheral Body Part [Positive/Total] |
|---|---|---|---|---|
| July 2020 | 106 | 1, 7 | 1/2 | 0/2 |
| October 2020 | 108 | 2, 7, 7 | 2/3 | 0/3 |
| September 2022 | 108 | 5, 5, 5 | 3/3 | 3/3 |
| Total | 6/8 | 3/8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ebert, C.L.; Söder, L.; Kubinski, M.; Glanz, J.; Gregersen, E.; Dümmer, K.; Grund, D.; Wöhler, A.-S.; Könenkamp, L.; Liebig, K.; et al. Detection and Characterization of Alongshan Virus in Ticks and Tick Saliva from Lower Saxony, Germany with Serological Evidence for Viral Transmission to Game and Domestic Animals. Microorganisms 2023, 11, 543. https://doi.org/10.3390/microorganisms11030543
Ebert CL, Söder L, Kubinski M, Glanz J, Gregersen E, Dümmer K, Grund D, Wöhler A-S, Könenkamp L, Liebig K, et al. Detection and Characterization of Alongshan Virus in Ticks and Tick Saliva from Lower Saxony, Germany with Serological Evidence for Viral Transmission to Game and Domestic Animals. Microorganisms. 2023; 11(3):543. https://doi.org/10.3390/microorganisms11030543
Chicago/Turabian StyleEbert, Cara Leonie, Lars Söder, Mareike Kubinski, Julien Glanz, Eva Gregersen, Katrin Dümmer, Domenic Grund, Ann-Sophie Wöhler, Laura Könenkamp, Katrin Liebig, and et al. 2023. "Detection and Characterization of Alongshan Virus in Ticks and Tick Saliva from Lower Saxony, Germany with Serological Evidence for Viral Transmission to Game and Domestic Animals" Microorganisms 11, no. 3: 543. https://doi.org/10.3390/microorganisms11030543
APA StyleEbert, C. L., Söder, L., Kubinski, M., Glanz, J., Gregersen, E., Dümmer, K., Grund, D., Wöhler, A.-S., Könenkamp, L., Liebig, K., Knoll, S., Hellhammer, F., Topp, A.-K., Becher, P., Springer, A., Strube, C., Nagel-Kohl, U., Nordhoff, M., Steffen, I., ... Boelke, M. (2023). Detection and Characterization of Alongshan Virus in Ticks and Tick Saliva from Lower Saxony, Germany with Serological Evidence for Viral Transmission to Game and Domestic Animals. Microorganisms, 11(3), 543. https://doi.org/10.3390/microorganisms11030543







