Coinfection of Dermal Fibroblasts by Human Cytomegalovirus and Human Herpesvirus 6 Can Boost the Expression of Fibrosis-Associated MicroRNAs
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Cultures and Viruses
2.2. Virus Infection
2.3. Nucleic Acid Extraction
2.4. Virus Quantitation in Infected Cells
2.5. miRNA Analysis
2.6. Statistical Analyses
3. Results
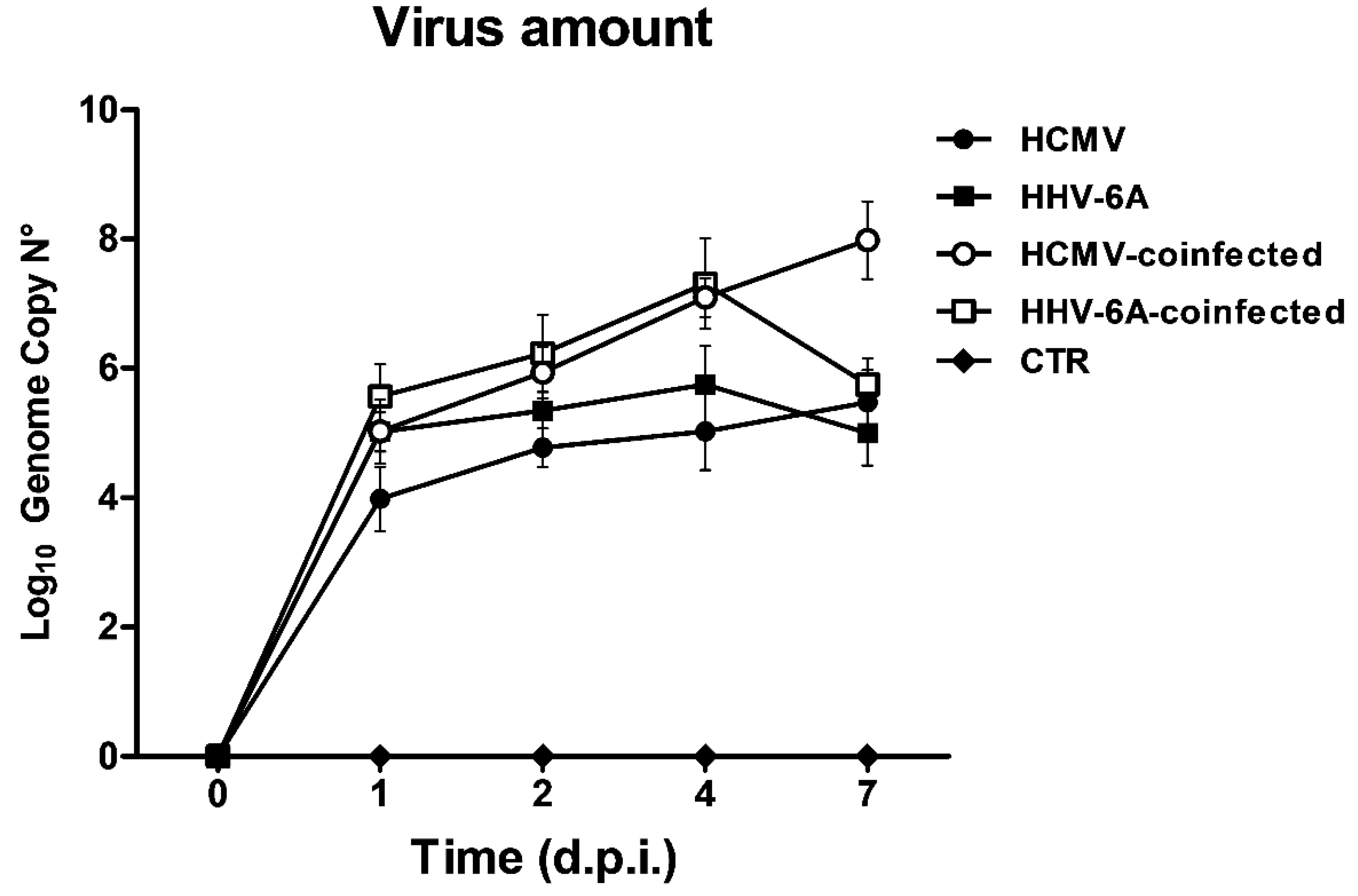
3.1. HCMV and HHV-6A Coinfection in Primary Human Dermal Fibroblasts
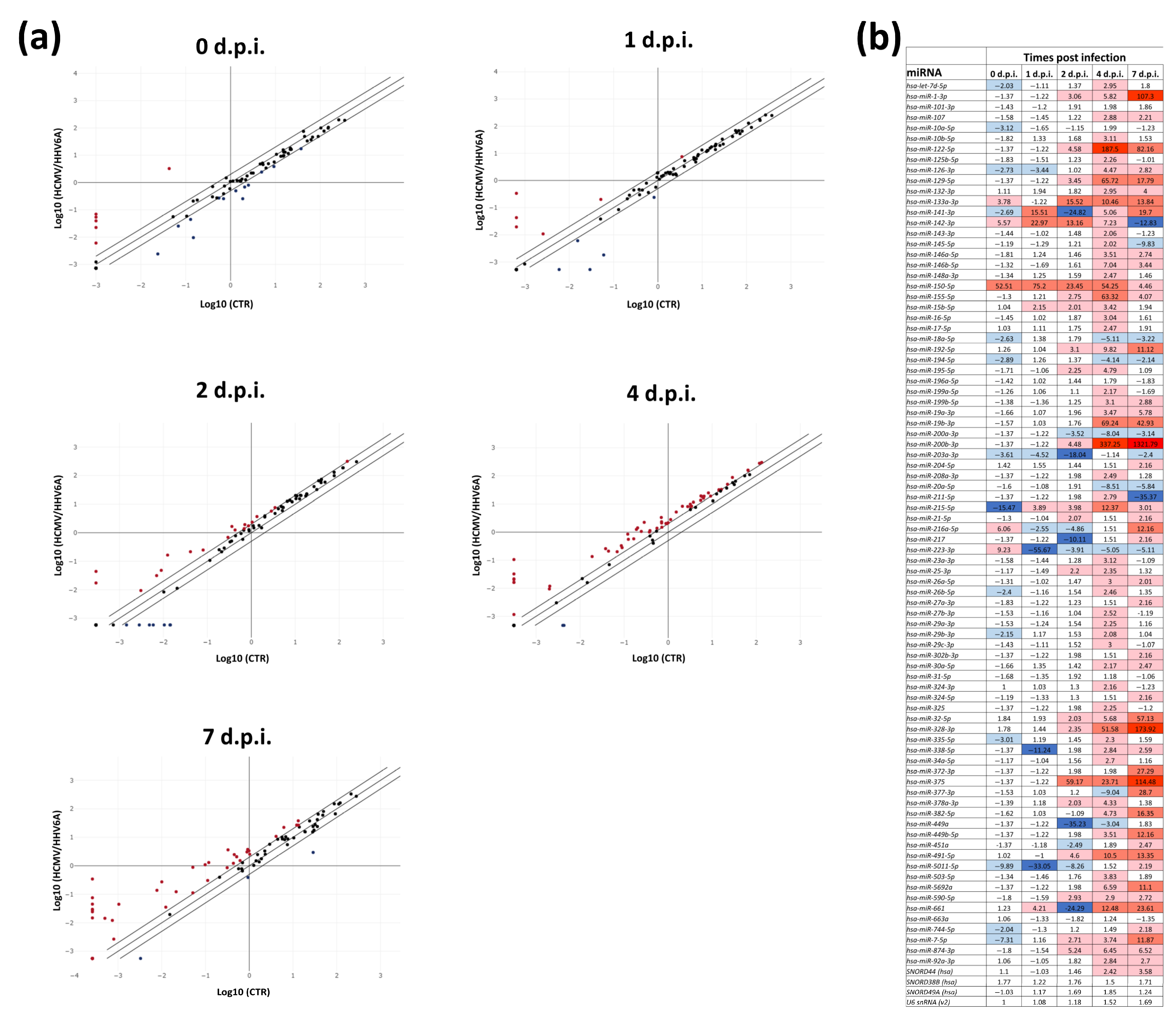
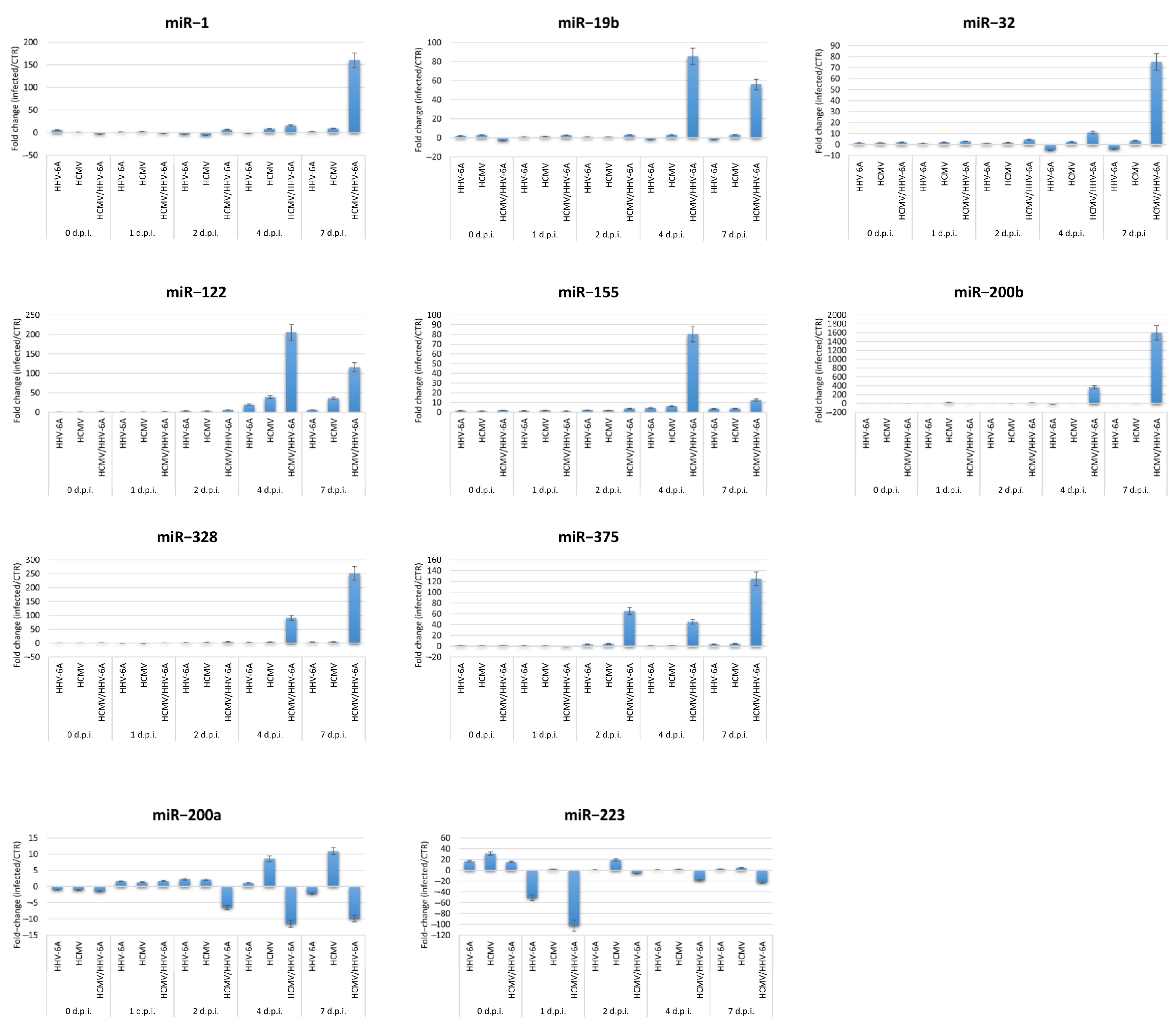
3.2. Effect of HCMV and HHV-6A Coinfection on the Expression of Fibrosis-Associated miRNAs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wynn, T.A.; Ramalingam, T.R. Mechanisms of Fibrosis: Therapeutic Translation for Fibrotic Disease. Nat. Med. 2012, 18, 1028–1040. [Google Scholar] [CrossRef]
- Bergamasco, A.; Hartmann, N.; Wallace, L.; Verpillat, P. Epidemiology of Systemic Sclerosis and Systemic Sclerosis-Associated Interstitial Lung Disease. Clin. Epidemiol. 2019, 11, 257. [Google Scholar] [CrossRef]
- Denton, C.P.; Khanna, D. Systemic Sclerosis. Lancet 2017, 390, 1685–1699. [Google Scholar] [CrossRef]
- Wigley, F.M.; Boin, F. Clinical Features and Treatment of Scleroderma. In Kelley and Firestein’s Textbook of Rheumatology, 10th ed.; Elsevier-Health Sciences Division; Riverport Lane: Maryland Heights, MO, USA, 2017; pp. 1424–1460. [Google Scholar]
- Ferri, C.; Sebastiani, M.; Lo Monaco, A.; Iudici, M.; Giuggioli, D.; Furini, F.; Manfredi, A.; Cuomo, G.; Spinella, A.; Colaci, M.; et al. Systemic Sclerosis Evolution of Disease Pathomorphosis and Survival. Our Experience on Italian Patients’ Population and Review of the Literature. Autoimmun. Rev. 2014, 13, 1026–1034. [Google Scholar] [CrossRef] [PubMed]
- Ferri, C.; Arcangeletti, M.-C.; Caselli, E.; Zakrzewska, K.; Maccari, C.; Calderaro, A.; D’accolti, M.; Soffritti, I.; Arvia, R.; Sighinolfi, G.; et al. Insights into the Knowledge of Complex Diseases: Environmental Infectious/Toxic Agents as Potential Etiopathogenetic Factors of Systemic Sclerosis. J. Autoimmun. 2021, 124, 102727. [Google Scholar] [CrossRef] [PubMed]
- Murdaca, G.; Contatore, M.; Gulli, R.; Mandich, P.; Puppo, F. Genetic Factors and Systemic Sclerosis. Autoimmun. Rev. 2016, 15, 427–432. [Google Scholar] [CrossRef]
- Arcangeletti, M.C.; D’accolti, M.; Maccari, C.; Soffritti, I.; De Conto, F.; Chezzi, C.; Calderaro, A.; Ferri, C.; Caselli, E. Impact of Human Cytomegalovirus and Human Herpesvirus 6 Infection on the Expression of Factors Associated with Cell Fibrosis and Apoptosis: Clues for Implication in Systemic Sclerosis Development. Int. J. Mol. Sci. 2020, 21, 6397. [Google Scholar] [CrossRef]
- Arcangeletti, M.C.; Maccari, C.; Vescovini, R.; Volpi, R.; Giuggioli, D.; Sighinolfi, G.; De Conto, F.; Chezzi, C.; Calderaro, A.; Ferri, C. A Paradigmatic Interplay between Human Cytomegalovirus and Host Immune System: Possible Involvement of Viral Antigen-Driven CD8+ T Cell Responses in Systemic Sclerosis. Viruses 2018, 10, 508. [Google Scholar] [CrossRef] [PubMed]
- Caselli, E.; Soffritti, I.; D’Accolti, M.; Bortolotti, D.; Rizzo, R.; Sighinolfi, G.; Giuggioli, D.; Ferri, C. HHV-6A Infection and Systemic Sclerosis: Clues of a Possible Association. Microorganisms 2019, 8, 39. [Google Scholar] [CrossRef]
- Broccolo, F.; Drago, F.; Paolino, S.; Cassina, G.; Gatto, F.; Fusetti, L.; Matteoli, B.; Zaccaria, E.; Parodi, A.; Lusso, P.; et al. Reactivation of Human Herpesvirus 6 (HHV-6) Infection in Patients with Connective Tissue Diseases. J. Clin. Virol. 2009, 46, 43–46. [Google Scholar] [CrossRef]
- Broccolo, F.; Drago, F.; Cassina, G.; Fava, A.; Fusetti, L.; Matteoli, B.; Ceccherini-Nelli, L.; Sabbadini, M.G.; Lusso, P.; Parodi, A.; et al. Selective Reactivation of Human Herpesvirus 6 in Patients with Autoimmune Connective Tissue Diseases. J. Med. Virol. 2013, 85, 1925–1934. [Google Scholar] [CrossRef] [PubMed]
- Soffritti, I.; D’Accolti, M.; Ravegnini, G.; Arcangeletti, M.C.; Maccari, C.; De Conto, F.; Calderaro, A.; Caselli, E. Modulation of Micrornome by Human Cytomegalovirus and Human Herpesvirus 6 Infection in Human Dermal Fibroblasts: Possible Significance in the Induction of Fibrosis in Systemic Sclerosis. Cells 2021, 10, 1060. [Google Scholar] [CrossRef] [PubMed]
- Soffritti, I.; D’Accolti, M.; Maccari, C.; Bini, F.; Mazziga, E.; de Conto, F.; Calderaro, A.; Arcangeletti, M.C.; Caselli, E. Human Cytomegalovirus and Human Herpesvirus 6 Coinfection of Dermal Fibroblasts Enhances the Pro-Inflammatory Pathway Predisposing to Fibrosis: The Possible Impact on Systemic Sclerosis. Microorganisms 2022, 10, 1600. [Google Scholar] [CrossRef]
- Álvarez-Lafuente, R.; Fernández-Gutiérrez, B.; De Miguel, S.; Jover, J.A.; Rollin, R.; Loza, E.; Clemente, D.; Lamas, J.R. Potential Relationship between Herpes Viruses and Rheumatoid Arthritis: Analysis with Quantitative Real Time Polymerase Chain Reaction. Ann. Rheum. Dis. 2005, 64, 1357–1359. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Lafuente, R.; Martinez, A.; Garcia-Montojo, M.; Mas, A.; De Las Heras, V.; Dominguez-Mozo, M.I.; Maria Del Carmen, C.; López-Cavanillas, M.; Bartolome, M.; Gomez De La Concha, E.; et al. MHC2TA Rs4774C and HHV-6A Active Replication in Multiple Sclerosis Patients. Eur. J. Neurol. 2010, 17, 129–135. [Google Scholar] [CrossRef]
- Caselli, E.; Zatelli, M.C.; Rizzo, R.; Benedetti, S.; Martorelli, D.; Trasforini, G.; Cassai, E.; degli Uberti, E.C.; Di Luca, D.; Dolcetti, R. Virologic and Immunologic Evidence Supporting an Association between HHV-6 and Hashimoto’s Thyroiditis. PLoS Pathog. 2012, 8, e1002951. [Google Scholar] [CrossRef]
- Magro, C.M.; Crowson, A.N.; Ferri, C. Cytomegalovirus-Associated Cutaneous Vasculopathy and Scleroderma sans Inclusion Body Change. Hum. Pathol. 2007, 38, 42–49. [Google Scholar] [CrossRef]
- Ferri, C.; Cazzato, M.; Giuggioli, D.; Sebastiani, M.; Magro, C. Systemic Sclerosis Following Human Cytomegalovirus Infection. Ann. Rheum. Dis. 2002, 61, 937–938. [Google Scholar] [CrossRef]
- Lunardi, C.; Dolcino, M.; Peterlana, D.; Bason, C.; Navone, R.; Tamassia, N.; Beri, R.; Corrocher, R.; Puccetti, A. Antibodies against Human Cytomegalovirus in the Pathogenesis of Systemic Sclerosis: A Gene Array Approach. PLoS Med. 2006, 3, 94–108. [Google Scholar] [CrossRef]
- Arnson, Y.; Amital, H.; Guiducci, S.; Matucci-Cerinic, M.; Valentini, G.; Barzilai, O.; Maya, R.; Shoenfeld, Y. The Role of Infections in the Immunopathogensis of Systemic Sclerosis-Evidence from Serological Studies. Ann. N. Y. Acad. Sci. 2009, 1173, 627–632. [Google Scholar] [CrossRef]
- Marou, E.; Liaskos, C.; Efthymiou, G.; Dardiotis, E.; Daponte, A.; Scheper, T.; Meyer, W.; Hadjigeorgiou, G.; Bogdanos, D.P.; Sakkas, L.I. Increased Immunoreactivity against Human Cytomegalovirus UL83 in Systemic Sclerosis. Clin. Exp. Rheumatol. 2017, 35, S31–S34. [Google Scholar]
- Efthymiou, G.; Dardiotis, E.; Liaskos, C.; Marou, E.; Scheper, T.; Meyer, W.; Daponte, A.; Daoussis, D.; Hadjigeorgiou, G.; Bogdanos, D.P.; et al. A Comprehensive Analysis of Antigen-Specific Antibody Responses against Human Cytomegalovirus in Patients with Systemic Sclerosis. Clin. Immunol. 2019, 207, 87–96. [Google Scholar] [CrossRef]
- Lunardi, C.; Bason, C.; Navone, R.; Millo, E.; Damonte, G.; Corrocher, R.; Puccetti, A. Systemic Sclerosis Immunoglobulin G Autoantibodies Bind the Human Cytomegalovirus Late Protein UL94 and Induce Apoptosis in Human Endothelial Cells. Nat. Med. 2000, 6, 1183–1186. [Google Scholar] [CrossRef] [PubMed]
- Rotola, A.; Di Luca, D.; Cassai, E.; Ricotta, D.; Giulio, A.; Turano, A.; Caruso, A.; Muneretto, C. Human Herpesvirus 6 Infects and Replicates in Aortic Endothelium. J. Clin. Microbiol. 2000, 38, 3135–3136. [Google Scholar] [CrossRef] [PubMed]
- Caruso, A.; Rotola, A.; Comar, M.; Favilli, F.; Galvan, M.; Tosetti, M.; Campello, C.; Caselli, E.; Alessandri, G.; Grassi, M.; et al. HHV-6 Infects Human Aortic and Heart Microvascular Endothelial Cells, Increasing Their Ability to Secrete Proinflammatory Chemokines. J. Med. Virol. 2002, 67, 528–533. [Google Scholar] [CrossRef]
- Mostmans, Y.; Cutolo, M.; Giddelo, C.; Decuman, S.; Melsens, K.; Declercq, H.; Vandecasteele, E.; De Keyser, F.; Distler, O.; Gutermuth, J.; et al. The Role of Endothelial Cells in the Vasculopathy of Systemic Sclerosis: A Systematic Review. Autoimmun. Rev. 2017, 16, 774–786. [Google Scholar] [CrossRef]
- Sinzger, C.; Grefte, A.; Plachter, B.; Gouw, A.S.H.; Hauw The, T.; Jahn, G. Fibroblasts, Epithelial Cells, Endothelial Cells and Smooth Muscle Cells Are Major Targets of Human Cytomegalovirus Infection in Lung and Gastrointestinal Tissues. J. Gen. Virol. 1995, 76, 741–750. [Google Scholar] [CrossRef]
- Altorok, N.; Almeshal, N.; Wang, Y.; Kahaleh, B. Epigenetics, the Holy Grail in the Pathogenesis of Systemic Sclerosis. Rheumatology 2015, 54, 1759–1770. [Google Scholar] [CrossRef]
- Szabo, I.; Muntean, L.; Crisan, T.; Rednic, V.; Sirbe, C.; Rednic, S. Novel Concepts in Systemic Sclerosis Pathogenesis: Role for MiRNAs. Biomedicines 2021, 9, 1471. [Google Scholar] [CrossRef]
- Henry, T.W.; Mendoza, F.A.; Jimenez, S.A. Role of MicroRNA in the Pathogenesis of Systemic Sclerosis Tissue Fibrosis and Vasculopathy. Autoimmun. Rev. 2019, 18, 102396. [Google Scholar] [CrossRef]
- Rizzo, R.; Soffritti, I.; D’Accolti, M.; Bortolotti, D.; Di Luca, D.; Caselli, E. HHV-6A/6B Infection of NK Cells Modulates the Expression of MiRNAs and Transcription Factors Potentially Associated to Impaired NK Activity. Front. Microbiol. 2017, 8, 2143. [Google Scholar] [CrossRef]
- Caselli, E.; Bortolotti, D.; Marci, R.; Rotola, A.; Gentili, V.; Soffritti, I.; D’Accolti, M.; Lo Monte, G.L.; Sicolo, M.; Barao, I.; et al. HHV-6A Infection of Endometrial Epithelial Cells Induces Increased Endometrial NK Cell-Mediated Cytotoxicity. Front. Microbiol. 2017, 8, 2525. [Google Scholar] [CrossRef]
- Doridot, L.; Jeljeli, M.; Chêne, C.; Batteux, F. Implication of Oxidative Stress in the Pathogenesis of Systemic Sclerosis via Inflammation, Autoimmunity and Fibrosis. Redox Biol. 2019, 25, 101122. [Google Scholar] [CrossRef]
- Caselli, E.; Bracci, A.; Galvan, M.; Boni, M.; Rotola, A.; Bergamini, C.; Cermelli, C.; Dal Monte, P.; Gompels, U.A.; Cassai, E.; et al. Human Herpesvirus 6 (HHV-6) U94/REP Protein Inhibits Betaherpesvirus Replication. Virology 2006, 346, 402–414. [Google Scholar] [CrossRef] [PubMed]
- DesJardin, J.A.; Gibbons, L.; Cho, E.; Supran, S.E.; Falagas, M.E.; Werner, B.G.; Snydman, D.R. Human Herpesvirus 6 Reactivation Is Associated with Cytomegalovirus Infection and Syndromes in Kidney Transplant Recipients at Risk for Primary Cytomegalovirus Infection. J. Infect. Dis. 1998, 178, 1783–1786. [Google Scholar] [CrossRef] [PubMed]
- Van Leer-Buter, C.C.; Sanders, J.S.F.; Vroom, H.E.J.; Riezebos-Brilman, A.; Niesters, H.G.M. Human Herpesvirus-6 DNAemia Is a Sign of Impending Primary CMV Infection in CMV Sero-Discordant Renal Transplantations. J. Clin. Virol. 2013, 58, 422–426. [Google Scholar] [CrossRef]
- Handous, I.; Achour, B.; Marzouk, M.; Rouis, S.; Hazgui, O.; Brini, I.; Khelif, A.; Hannachi, N.; Boukadida, J. Co-Infections of Human Herpesviruses (CMV, HHV-6, HHV-7 and EBV) in Non-Transplant Acute Leukemia Patients Undergoing Chemotherapy. Virol. J. 2020, 17, 37. [Google Scholar] [CrossRef] [PubMed]
- Roa, P.L.; Hill, J.A.; Kirby, K.A.; Leisenring, W.M.; Huang, M.L.; Santo, T.K.; Jerome, K.R.; Boeckh, M.; Limaye, A.P. Coreactivation of Human Herpesvirus 6 and Cytomegalovirus Is Associated With Worse Clinical Outcome in Critically Ill Adults. Crit. Care Med. 2015, 43, 1415–1422. [Google Scholar] [CrossRef]
- Von Kietzell, K.; Pozzuto, T.; Heilbronn, R.; Grössl, T.; Fechner, H.; Weger, S. Antibody-Mediated Enhancement of Parvovirus B19 Uptake into Endothelial Cells Mediated by a Receptor for Complement Factor C1q. J. Virol. 2014, 88, 8102–8115. [Google Scholar] [CrossRef]
- Mendez, J.C.; Dockrell, D.H.; Espy, M.J.; Smith, T.F.; Wilson, J.A.; Harmsen, W.S.; Ilstrup, D.; Paya, C.V. Human β-Herpesvirus Interactions in Solid Organ Transplant Recipients. J. Infect. Dis. 2001, 183, 179–184. [Google Scholar] [CrossRef]
- Humar, A.; Malkan, G.; Moussa, G.; Greig, P.; Levy, G.; Mazzulli, T. Human Herpesvirus—6 Is Associated with Cytomegalovirus Reactivation in Liver Transplant Recipients. J. Infect. Dis. 2000, 181, 1450–1453. [Google Scholar] [CrossRef]
- Ding, R.; Liu, X.; Zhang, J.; Yuan, J.; Zheng, S.; Cheng, X.; Jia, J. Downregulation of MiR-1-3p Expression Inhibits the Hypertrophy and Mineralization of Chondrocytes in DDH. J. Orthop. Surg. Res. 2021, 16, 512. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Z.; Gao, L.; Qiao, Z.; Yu, M.; Yu, B.; Yang, T. MiR-1-3p Suppresses Proliferation of Hepatocellular Carcinoma through Targeting SOX9. Onco. Targets. Ther. 2019, 12, 2149–2157. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Sun, Y.; Tang, Y.; Li, H.; Wang, X.; Pan, X.; Liu, W.; Zhang, X.; Zhang, F.; Xu, Y.; et al. Circulating Mir-19b-3p as a Novel Prognostic Biomarker for Acute Heart Failure. J. Am. Heart Assoc. 2021, 10, 22304. [Google Scholar] [CrossRef]
- Liu, Y.; Song, J.W.; Lin, J.Y.; Miao, R.; Zhong, J.C. Roles of MicroRNA-122 in Cardiovascular Fibrosis and Related Diseases. Cardiovasc. Toxicol. 2020, 20, 463–473. [Google Scholar] [CrossRef] [PubMed]
- Gómez, E.; Laurés, A.; Baltar, J.M.; Melón, S.; Díez, B.; De Oña, M. Cytomegalovirus Replication and “Herpesvirus Burden” as Risk Factor of Cardiovascular Events in the First Year After Renal Transplantation. Transplant. Proc. 2005, 37, 3760–3763. [Google Scholar] [CrossRef]
- Horváth, R.; Černý, J.; Benedík, J.; Hökl, J.; Jelínková, I.; Benedík, J. The Possible Role of Human Cytomegalovirus (HCMV) in the Origin of Atherosclerosis. J. Clin. Virol. 2000, 16, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Comar, M.; D’Agaro, P.; Campello, C.; Poli, A.; Breinholt, J.P.; Towbin, J.A.; Vatta, M. Human Herpes Virus 6 in Archival Cardiac Tissues from Children with Idiopathic Dilated Cardiomyopathy or Congenital Heart Disease. J. Clin. Pathol. 2009, 62, 80–83. [Google Scholar] [CrossRef]
- Hao, X.; Luan, J.; Jiao, C.; Ma, C.; Feng, Z.; Zhu, L.; Zhang, Y.; Fu, J.; Lai, E.; Zhang, B.; et al. LNA-Anti-MiR-150 Alleviates Renal Interstitial Fibrosis by Reducing pro-Inflammatory M1/M2 Macrophage Polarization. Front. Immunol. 2022, 13, 3415. [Google Scholar] [CrossRef]
- Eissa, M.G.; Artlett, C.M. The MicroRNA MiR-155 Is Essential in Fibrosis. Non-Coding RNA 2019, 5, 23. [Google Scholar] [CrossRef]
- Chung, A.C.K.; Huang, X.R.; Meng, X.; Lan, H.Y. MiR-192 Mediates TGF-β/Smad3-Driven Renal Fibrosis. J. Am. Soc. Nephrol. 2010, 21, 1317–1325. [Google Scholar] [CrossRef]
- Ren, F.J.; Yao, Y.; Cai, X.Y.; Fang, G.Y. Emerging Role of MiR-192-5p in Human Diseases. Front. Pharmacol. 2021, 12, 160. [Google Scholar] [CrossRef]
- Ye, M.; Wang, S.; Sun, P.; Qie, J. Integrated MicroRNA Expression Profile Reveals Dysregulated MiR-20a-5p and MiR-200a-3p in Liver Fibrosis. Biomed Res. Int. 2021, 2021, 9583932. [Google Scholar] [CrossRef] [PubMed]
- Lan, W.; Chen, S.; Tong, L. MicroRNA-215 Regulates Fibroblast Function: Insights from a Human Fibrotic Disease. Cell Cycle 2015, 14, 1973–1984. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Li, Z.; Lin, Y.; Che, H.; Hu, Y.; Kang, X.; Zhang, Y.; Wang, L.; Zhang, Y. High glucose promotes hepatic fibrosis via miR-32/MTA3-mediated epithelial-to-mesenchymal transition Corrigendum in/10.3892/mmr. 2022.12827. Mol. Med. Rep. 2022, 26, 3190–3200. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Liang, H.; Gao, X.; Li, X.; Zhang, Y.; Pan, Z.; Li, C.; Wang, Y.; Liu, Y.; Yuan, W.; et al. MicroRNA-328, a Potential Anti-Fibrotic Target in Cardiac Interstitial Fibrosis. Cell. Physiol. Biochem. 2016, 39, 827–836. [Google Scholar] [CrossRef]
- He, W.; Zhuang, J.; Zhao, Z.G.; Luo, H.; Zhang, J. MiR-328 Prevents Renal Fibrogenesis by Directly Targeting TGF-Β2. Bratisl. Lek. Listy 2018, 119, 434–440. [Google Scholar] [CrossRef]
- Zhuang, Y.; Yang, D.; Shi, S.; Wang, L.; Yu, M.; Meng, X.; Fan, Y.; Zhou, R.; Wang, F. MiR-375-3p Promotes Cardiac Fibrosis by Regulating the Ferroptosis Mediated by GPX4. Comput. Intell. Neurosci. 2022, 2022, 9629158. [Google Scholar] [CrossRef]
- Pommier, A.; Varilh, J.; Bleuse, S.; Delétang, K.; Bonini, J.; Bergougnoux, A.; Brochiero, E.; Koenig, M.; Claustres, M.; Taulan-Cadars, M. MiRNA Repertoires of Cystic Fibrosis Ex Vivo Models Highlight MiR-181a and MiR-101 That Regulate WISP1 Expression. J. Pathol. 2021, 253, 186–197. [Google Scholar] [CrossRef]
- Xie, T.; Liang, J.; Guo, R.; Liu, N.; Noble, P.W.; Jiang, D. Comprehensive MicroRNA Analysis in Bleomycin-Induced Pulmonary Fibrosis Identifies Multiple Sites of Molecular Regulation. Physiol. Genomics 2011, 43, 479–487. [Google Scholar] [CrossRef]
- Bissey, P.A.; Law, J.H.; Bruce, J.P.; Shi, W.; Renoult, A.; Chua, M.L.K.; Yip, K.W.; Liu, F.F. Dysregulation of the MiR-449b Target TGFBI Alters the TGFβ Pathway to Induce Cisplatin Resistance in Nasopharyngeal Carcinoma. Oncogenesis 2018, 7, 40. [Google Scholar] [CrossRef] [PubMed]
- Chung, A.C.K.; Lan, H.Y. MicroRNAs in Renal Fibrosis. Front. Physiol. 2015, 6, 50. [Google Scholar] [CrossRef]
- Wu, F.; He, H.; Chen, Y.; Zhu, D.; Jiang, T.; Wang, J. CircPDE7B/MiR-661 Axis Accelerates the Progression of Human Keloid Fibroblasts by Upregulating Fibroblast Growth Factor 2 (FGF2). Mol. Cell. Biochem. 2022, 477, 1113–1126. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Chen, M.; Wang, B.; Han, Y.; Shang, H.; Chen, J.; Chen, J. MiR-7-5p Promotes Hepatic Stellate Cell Activation by Targeting Fibroblast Growth Factor Receptor 4. Gastroenterol. Res. Pract. 2020, 2020, 5346573. [Google Scholar] [CrossRef]
- Wei, P.; Xie, Y.; Abel, P.W.; Huang, Y.; Ma, Q.; Li, L.; Hao, J.; Wolff, D.W.; Wei, T.; Tu, Y. Transforming Growth Factor (TGF)-Β1-Induced MiR-133a Inhibits Myofibroblast Differentiation and Pulmonary Fibrosis. Cell Death Dis. 2019, 10, 670. [Google Scholar] [CrossRef]
- Li, G.; Zhao, C.; Fang, S. SGLT2 Promotes Cardiac Fibrosis Following Myocardial Infarction and Is Regulated by MiR-141. Exp. Ther. Med. 2021, 22, 715. [Google Scholar] [CrossRef] [PubMed]
- Song, T.; Zhou, M.; Li, W.; Lv, M.; Zheng, L.; Zhao, M. The Anti-Inflammatory Effect of Vasoactive Peptides from Soybean Protein Hydrolysates by Mediating Serum Extracellular Vesicles-Derived MiRNA-19b/CYLD/TRAF6 Axis in the Vascular Microenvironment of SHRs. Food Res. Int. 2022, 160, 111742. [Google Scholar] [CrossRef]
- Singh, V.; Kushwaha, S.; Ansari, J.A.; Gangopadhyay, S.; Mishra, S.K.; Dey, R.K.; Giri, A.K.; Patnaik, S.; Ghosh, D. MicroRNA-129-5p-Regulated Microglial Expression of the Surface Receptor CD200R1 Controls Neuroinflammation. J. Biol. Chem. 2022, 298, 101521. [Google Scholar] [CrossRef]
- Xia, S.; Huang, J.; Yan, L.; Han, J.; Zhang, W.; Shao, H.; Shen, H.; Wang, J.; Wang, J.; Tao, C.; et al. MiR-150 Promotes Progressive T Cell Differentiation via Inhibiting FOXP1 and RC3H1. Hum. Immunol. 2022, 83, 778–788. [Google Scholar] [CrossRef]
- Yang, L.-L.; Liu, J.-Q.; Bai, X.-Z.; Fan, L.; Han, F.; Jia, W.-B.; Su, L.-L.; Shi, J.-H.; Tang, C.-W.; Hu, D.-H. Acute Downregulation of MiR-155 at Wound Sites Leads to a Reduced Fibrosis through Attenuating Inflammatory Response. Biochem. Biophys. Res. Commun. 2014, 453, 153–159. [Google Scholar] [CrossRef]
- Pottier, N.; Maurin, T.; Chevalier, B.; Puisségur, M.-P.; Lebrigand, K.; Robbe-Sermesant, K.; Bertero, T.; Lino Cardenas, C.L.; Courcot, E.; Rios, G.; et al. Identification of Keratinocyte Growth Factor as a Target of MicroRNA-155 in Lung Fibroblasts: Implication in Epithelial-Mesenchymal Interactions. PLoS ONE 2009, 4, e6718. [Google Scholar] [CrossRef]
- Qin, S.; Li, B.; Chen, M.; Qin, M.; Liu, J.; Lv, Q. MiR-32-5p promoted epithelial-to-mesenchymal transition of oral squamous cell carcinoma cells via regulating the KLF2/CXCR4 pathway. Kaohsiung J. Med. Sci. 2022, 38, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Zheng, Y.; Hong, W.; Chen, B.; Dong, P.; Zheng, J. MicroRNA-200a Suppresses Epithelial-to-Mesenchymal Transition in Rat Hepatic Stellate Cells via GLI Family Zinc Finger 2. Mol. Med. Rep. 2015, 12, 8121–8128. [Google Scholar] [CrossRef] [PubMed]
- Jimenez Calvente, C.; Del Pilar, H.; Tameda, M.; Johnson, C.D.; Feldstein, A.E. MicroRNA 223 3p Negatively Regulates the NLRP3 Inflammasome in Acute and Chronic Liver Injury. Mol. Ther. 2020, 28, 653–663. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, H.; Wang, H.; Du, J.; Dong, P.; Liu, M.; Lin, Y. Long Non-Coding RNA DUXAP8 Promotes the Cell Proliferation, Migration, and Invasion of Papillary Thyroid Carcinoma via MiR-223-3p Mediated Regulation of CXCR4. Bioengineered 2021, 12, 496–506. [Google Scholar] [CrossRef]
- Leone, V.; D’Angelo, D.; Rubio, I.; de Freitas, P.M.; Federico, A.; Colamaio, M.; Pallante, P.; Medeiros-Neto, G.; Fusco, A. MiR-1 Is a Tumor Suppressor in Thyroid Carcinogenesis Targeting CCND2, CXCR4, and SDF-1α. J. Clin. Endocrinol. Metab. 2011, 96, E1388–E1398. [Google Scholar] [CrossRef]
- Yang, R.Q.; Teng, H.; Xu, X.H.; Liu, S.Y.; Wang, Y.H.; Guo, F.J.; Liu, X.J. Microarray Analysis of MicroRNA Deregulation and Angiogenesis-Related Proteins in Endometriosis. Genet. Mol. Res. 2016, 15, 1–8. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soffritti, I.; D’Accolti, M.; Maccari, C.; Bini, F.; Mazziga, E.; Arcangeletti, M.-C.; Caselli, E. Coinfection of Dermal Fibroblasts by Human Cytomegalovirus and Human Herpesvirus 6 Can Boost the Expression of Fibrosis-Associated MicroRNAs. Microorganisms 2023, 11, 412. https://doi.org/10.3390/microorganisms11020412
Soffritti I, D’Accolti M, Maccari C, Bini F, Mazziga E, Arcangeletti M-C, Caselli E. Coinfection of Dermal Fibroblasts by Human Cytomegalovirus and Human Herpesvirus 6 Can Boost the Expression of Fibrosis-Associated MicroRNAs. Microorganisms. 2023; 11(2):412. https://doi.org/10.3390/microorganisms11020412
Chicago/Turabian StyleSoffritti, Irene, Maria D’Accolti, Clara Maccari, Francesca Bini, Eleonora Mazziga, Maria-Cristina Arcangeletti, and Elisabetta Caselli. 2023. "Coinfection of Dermal Fibroblasts by Human Cytomegalovirus and Human Herpesvirus 6 Can Boost the Expression of Fibrosis-Associated MicroRNAs" Microorganisms 11, no. 2: 412. https://doi.org/10.3390/microorganisms11020412
APA StyleSoffritti, I., D’Accolti, M., Maccari, C., Bini, F., Mazziga, E., Arcangeletti, M.-C., & Caselli, E. (2023). Coinfection of Dermal Fibroblasts by Human Cytomegalovirus and Human Herpesvirus 6 Can Boost the Expression of Fibrosis-Associated MicroRNAs. Microorganisms, 11(2), 412. https://doi.org/10.3390/microorganisms11020412








