Pan-Genome Analyses of the Genus Cohnella and Proposal of the Novel Species Cohnella silvisoli sp. nov., Isolated from Forest Soil
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strain Isolation and Cultivation
2.2. 16S rRNA Gene Sequence and Phylogenetic Analysis
2.3. Morphological, Physiological, and Biochemical Characteristics
2.4. Chemotaxonomic Properties
2.5. Genome Sequencing, Annotation, and Pan-Genomic Analysis
2.6. OGRI Calculation and Phylogenomic Analysis
3. Results
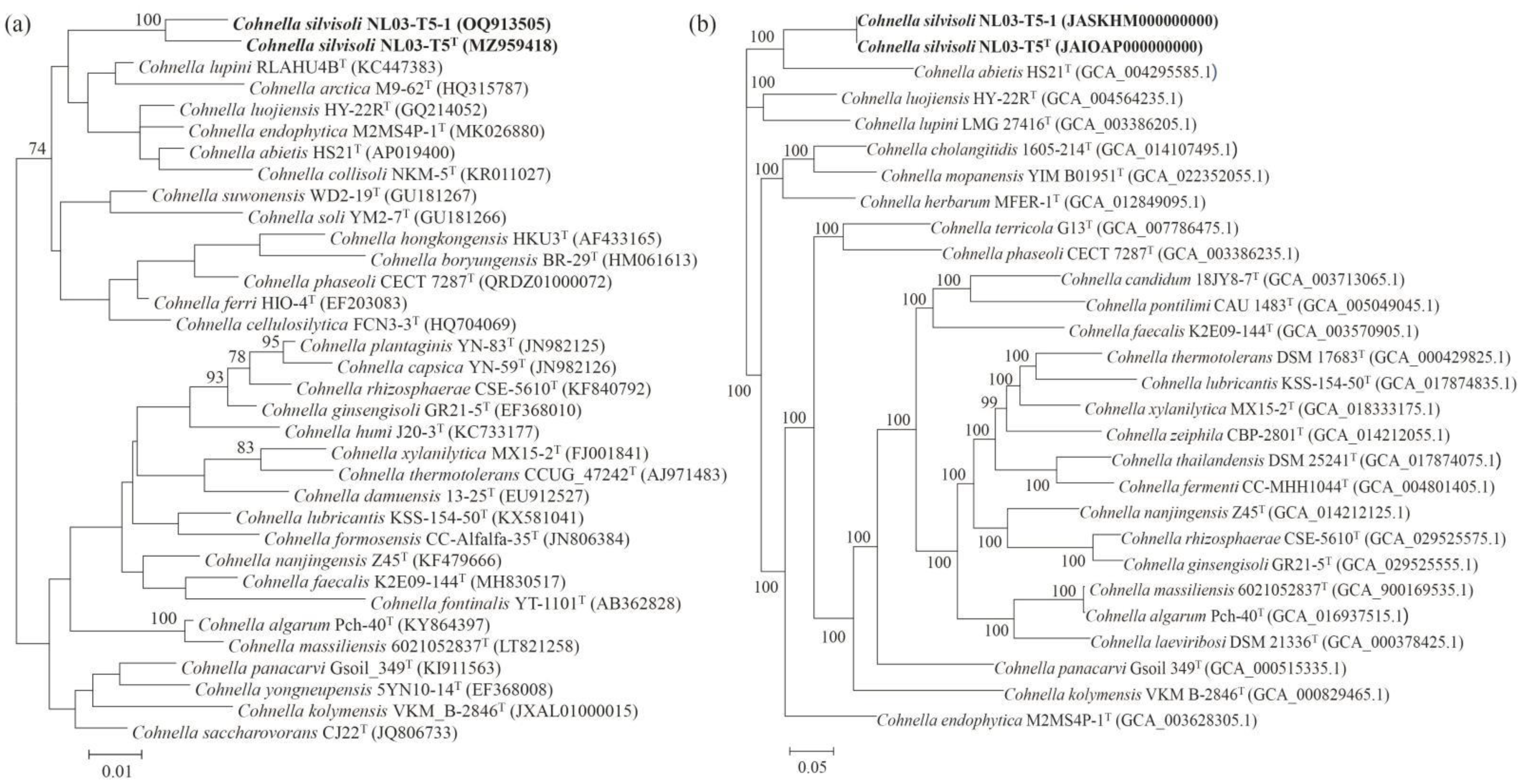
3.1. Phylogenetic Analysis Based on 16S rRNA Genes and Genomic Sequences
3.2. Physiological Characterization
3.3. Chemotaxonomic Analysis
3.4. Genomic Characteristics and OGRI Values
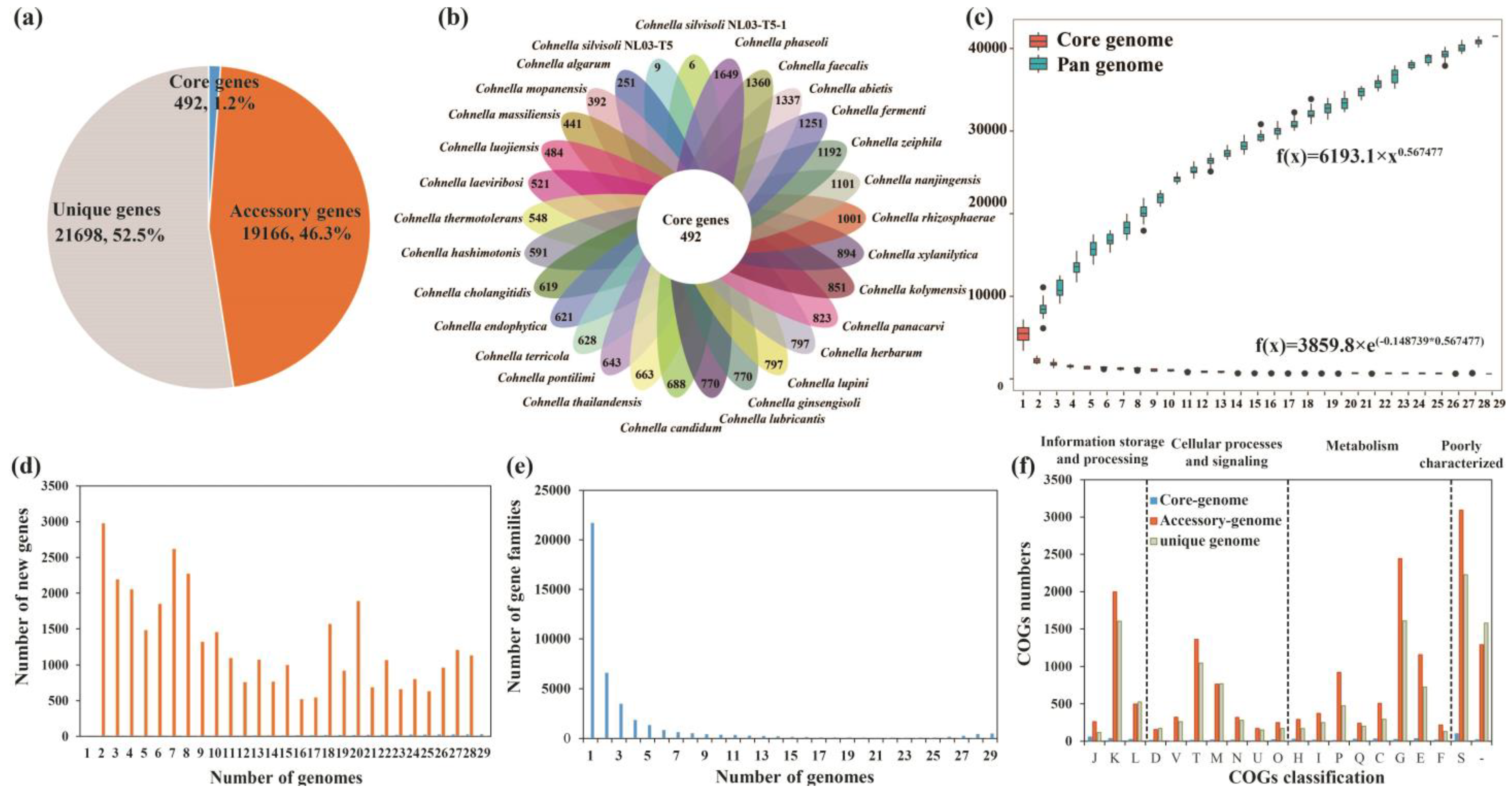
3.5. Pan-Genome Analysis of the Genus Cohnella
4. Discussion
5. Description of Cohnella silvisoli sp. nov.
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| GDMCC | Guangdong Microbial Culture Collection Center |
| JCM | Japan Collection of Microorganism |
| LMG | Belgian Co-ordinated Collections of Micro-organisms |
| NRRL | Agricultural Research Service Culture Collection |
| ANI | Average Nucleotide Identity |
| dDDH | the Digital DNA-DNA Hybridization |
| UBCG | Up-to-date Bacterial Core Gene set |
| DPG | diphosphatidylglycerol |
| PE | phosphatidylethanolamine |
| PG | phosphatidylglycerol |
| APL | unidentified aminophospholipid |
References
- Kämpfer, P.; Rosselló-Mora, R.; Falsen, E.; Busse, H.-J.; Tindall, B.J. Cohnella thermotolerans gen. nov., sp. nov., and classification of ‘Paenibacillus hongkongensis’ as Cohnella hongkongensis sp. nov. Int. J. Syst. Evol. Microbiol. 2006, 56, 781–786. [Google Scholar] [CrossRef]
- Garcia-Fraile, P.; Velazquez, E.; Mateos, P.F.; Martinez-Molina, E.; Rivas, R. Cohnella phaseoli sp. nov., isolated from root nodules of Phaseolus coccineus in Spain, and emended description of the genus Cohnella. Int. J. Syst. Evol. Microbiol. 2008, 58, 1855–1859. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Dai, J.; Wang, Y.; Xue, X.; Xu, M.; Li, W.; Fang, C.; Peng, F. Cohnella arctica sp. nov., isolated from Arctic tundra soil. Int. J. Syst. Evol. Microbiol. 2012, 62, 817–821. [Google Scholar] [CrossRef]
- Cai, F.; Wang, Y.; Qi, H.; Dai, J.; Yu, B.; An, H.; Rahman, E.; Fang, C. Cohnella luojiensis sp. nov., isolated from soil of a Euphrates poplar forest. Int. J. Syst. Evol. Microbiol. 2010, 60, 1605–1608. [Google Scholar] [CrossRef] [PubMed]
- Flores-Félix, J.D.; Carro, L.; Ramírez-Bahena, M.-H.; Tejedor, C.; Igual, J.M.; Peix, A.; Velázquez, E. Cohnella lupini sp. nov., an endophytic bacterium isolated from root nodules of Lupinus albus. Int. J. Syst. Evol. Microbiol. 2014, 64, 83–87. [Google Scholar]
- Shiratori, H.; Tagami, Y.; Beppu, T.; Ueda, K. Cohnella fontinalis sp. nov., a xylanolytic bacterium isolated from fresh water. Int. J. Syst. Evol. Microbiol. 2010, 60, 1344–1348. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Jeon, C.O. Cohnella algarum sp. nov., isolated from a freshwater green alga Paulinella chromatophora. Int. J. Syst. Evol. Microbiol. 2017, 67, 4767–4772. [Google Scholar] [CrossRef]
- Zhu, H.Z.; Liu, X.D.; Jiang, C.Y.; Liu, S.J. Cohnella faecalis sp. nov., isolated from animal faeces in a karst cave. Int. J. Syst. Evol. Microbiol. 2019, 69, 572–577. [Google Scholar] [CrossRef]
- Kudryashova, E.B.; Karlyshev, A.V.; Ariskina, E.V.; Streshinskaya, G.M.; Vinokurova, N.G.; Kopitsyn, D.S.; Evtushenko, L.I. Cohnella kolymensis sp. nov., a novel bacillus isolated from Siberian permafrost. Int. J. Syst. Evol. Microbiol. 2018, 68, 2912–2917. [Google Scholar] [CrossRef]
- Lin, S.Y.; Chen, W.M.; Hameed, A.; Huang, G.H.; Hung, M.H.; Young, C.C. Cohnella fermenti sp. nov., isolated from a fermentation process. Int. J. Syst. Evol. Microbiol. 2020, 70, 2602–2610. [Google Scholar] [CrossRef]
- Aliabadi, N.; Aminzadeh, S.; Karkhane, A.A.; Haghbeen, K. Thermostable chitinase from Cohnella sp. A01: Isolation and product optimization. Braz. J. Microbiol. 2016, 47, 931–940. [Google Scholar] [CrossRef]
- Cho, E.A.; Lee, D.W.; Cha, Y.H.; Lee, S.J.; Jung, H.C.; Pan, J.G.; Pyun, Y.R. Characterization of a novel D-lyxose isomerase from Cohnella laevoribosii RI-39 sp. nov. J. Bacteriol. 2007, 189, 1655–1663. [Google Scholar] [CrossRef]
- Khianngam, S.; Tanasupawat, S.; Akaracharanya, A.; Kim, K.K.; Lee, K.C.; Lee, J.S. Cohnella cellulosilytica sp. nov., isolated from buffalo faeces. Int. J. Syst. Evol. Microbiol. 2012, 62, 1921–1925. [Google Scholar] [CrossRef]
- Kai, A.K.L.; Cheung, Y.K.; Yeung, R.K.K.; Wong, J.T.Y. Development of single-cell PCR methods for the Raphidophyceae. Harmful Algae. 2006, 5, 649–657. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The Neighbour-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Fitch, W.M. Toward defining course of evolution-minimum change for a specific tree topology. Syst. Zool. 1971, 20, 406–416. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparatives studies of nucleotide-sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Lv, Y.Y.; Wang, J.; Chen, M.H.; You, J.; Qiu, L.H. Dinghuibacter silviterrae gen. nov., sp. nov., isolated from forest soil. Int. J. Syst. Evol. Microbiol. 2016, 66, 1785–1791. [Google Scholar] [CrossRef]
- Bernardet, J.F.; Nakagawa, Y.; Holmes, B. Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int. J. Syst. Evol. Microbiol. 2002, 52, 1049–1070. [Google Scholar]
- Son, H.M.; Kook, M.; Kim, J.H.; Yi, T.H. Taibaiella koreensis sp. nov., isolated from soil of a ginseng field. Int. J. Syst. Evol. Microbiol. 2014, 64, 1018–1023. [Google Scholar] [CrossRef]
- Sasser, M. Identification of Bacteria by Gas Chromatography of Cellular Fatty Acids; MIDI Inc.: Newark, DE, USA, 1990; pp. 1–6. [Google Scholar]
- Tindall, B.; Sikorski, J.; Smibert, R.; Krieg, N. Phenotypic characterization and the principles of comparative systematics. In Methods for General and Molecular Microbiology; Reddy, C.A., Beveridge, T.J., Breznak, J.A., Marzluf, G.A., Schmidt, T.M., Snyder, L.R., Eds.; American Society for Microbiology: Washington, DC, USA, 2007; pp. 384–385. [Google Scholar]
- Minnikin, D.E.; Odonnell, A.G.; Goodfellow, M.; Alderson, G.; Athalye, M.; Schaal, A.; Parlett, J.H. An integrated procedure for the extraction of bacterial isoprendoid quinones and polar lipids. J. Microbiol. Meth. 1984, 2, 233–241. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Chaudhari, N.M.; Gupta, V.K.; Dutta, C. BPGA—An ultra-fast pan-genome analysis pipeline. Sci. Rep. 2016, 6, 24373. [Google Scholar] [CrossRef]
- Cantalapiedra, C.P.; Hernandez-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Auch, A.F.; Klenk, H.P.; Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013, 14, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.H.; Ha, S.M.; Lim, J.; Kwon, S.; Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 2017, 110, 1281–1286. [Google Scholar] [CrossRef] [PubMed]
- Na, S.I.; Kim, Y.O.; Yoon, S.H.; Ha, S.M.; Baek, I.; Chun, J. UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J. Microbiol. 2018, 56, 280–285. [Google Scholar] [CrossRef]
- Jiang, L.; Pheng, S.; Lee, K.C.; Kang, S.W.; Jeong, J.C.; Kim, C.Y.; Park, H.C.; Kim, D.H.; Kim, S.W.; Kim, S.G.; et al. Cohnella abietis sp. nov., isolated from Korean fir (Abies koreana) rhizospheric soil of Halla mountain. J. Microbiol. 2019, 57, 953–958. [Google Scholar]
- Chun, J.; Oren, A.; Ventosa, A.; Christensen, H.; Arahal, D.R.; da Costa, M.S.; Rooney, A.P.; Yi, H.; Xu, X.W.; De Meyer, S.; et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 2018, 68, 461–466. [Google Scholar] [CrossRef] [PubMed]
- Hurtado, R.; Carhuaricra, D.; Soares, S.; Viana, M.V.C.; Azevedo, V.; Maturrano, L.; Aburjaile, F. Pan-genomic approach shows insight of genetic divergence and pathogenic-adaptation of Pasteurella multocida. Gene 2018, 670, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Tettelin, H.; Riley, D.; Cattuto, C.; Medini, D. Comparative genomics: The bacterial pan-genome. Curr. Opin. Microbiol. 2008, 11, 472–477. [Google Scholar] [CrossRef] [PubMed]

| Characteristics | NL03-T5T | NL03-T5-1 | C. abietis HS21T [32] | C. lupini LMG 27416T | C. arctica NRRL B-59459T |
|---|---|---|---|---|---|
| Isolation source | soil | soil | soil | root nodules of Lupinus albus | soil |
| Colony colour | white-cream | white-cream | white | orange | white-cream |
| Oxidase | − | − | + | + | + |
| Catalase | − | − | − | + | + |
| Growth at 37 °C | w | + | − | − | − |
| Temperature range | 10–37 | 10–37 | 4–30 | 10–35 | 10–35 |
| pH range | 5.0–8.5 | 5.0–8.5 | 6–8 | 5.5–8.5 | 5.0–9.0 |
| Growth on media: | |||||
| Reasoner’s 2A | + | + | + | − | + |
| NA | − | − | + | + | − |
| Hydrolysis of: | |||||
| Gelatin | + | + | − | + | + |
| Tweens 20 | + | + | ND | − | − |
| Assimilation of: | |||||
| Mannitol | + | + | + | − | + |
| N-Acetyl-D-glucosamine | + | + | + | − | + |
| Gluconate | + | + | − | − | + |
| Capric acid | − | + | − | − | + |
| Enzyme activities: | |||||
| Esterase (C8) | + | + | ND | − | + |
| β-Glucosidase | + | − | ND | + | + |
| α-Mannosidase | + | + | ND | − | − |
| GC content (mol%) | 49.2 | 49.2 | 44.8 | 50.7 | 50.3 |
| Fatty Acid | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| anteiso-C13:0 | 1.6 | 4.3 | - | 2.1 |
| iso-C14:0 | 3.7 | 2.7 | 1.1 | 2.6 |
| C14:0 | 1.3 | 1.6 | 2.3 | 3.3 |
| iso-C15:0 | 5.5 | 6.0 | 2.1 | 2.1 |
| anteiso-C15:0 | 50.7 | 50.9 | 42.2 | 44.7 |
| C16:0 | 7.2 | 9.9 | 22.0 | 14.5 |
| alcohol-C16:1 ω7c | - | - | 6.5 | - |
| C16:1 ω11c | - | - | 7.0 | - |
| iso-C16:0 | 21.2 | 13.7 | 6.9 | 18.0 |
| iso-C17:0 | 1.9 | 2.4 | 1.2 | - |
| anteiso-C17:0 | 4.0 | 3.2 | 5.2 | 4.1 |
| C17:0 | - | - | - | 2.1 |
| C18:0 | - | - | - | 1.4 |
| Summed feature 8 | - | - | - | 1.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.; Mao, L.; Bao, G.; Zhu, H. Pan-Genome Analyses of the Genus Cohnella and Proposal of the Novel Species Cohnella silvisoli sp. nov., Isolated from Forest Soil. Microorganisms 2023, 11, 2726. https://doi.org/10.3390/microorganisms11112726
Wang C, Mao L, Bao G, Zhu H. Pan-Genome Analyses of the Genus Cohnella and Proposal of the Novel Species Cohnella silvisoli sp. nov., Isolated from Forest Soil. Microorganisms. 2023; 11(11):2726. https://doi.org/10.3390/microorganisms11112726
Chicago/Turabian StyleWang, Chunling, Lutian Mao, Gegen Bao, and Honghui Zhu. 2023. "Pan-Genome Analyses of the Genus Cohnella and Proposal of the Novel Species Cohnella silvisoli sp. nov., Isolated from Forest Soil" Microorganisms 11, no. 11: 2726. https://doi.org/10.3390/microorganisms11112726
APA StyleWang, C., Mao, L., Bao, G., & Zhu, H. (2023). Pan-Genome Analyses of the Genus Cohnella and Proposal of the Novel Species Cohnella silvisoli sp. nov., Isolated from Forest Soil. Microorganisms, 11(11), 2726. https://doi.org/10.3390/microorganisms11112726






