Comparative Analysis of Rumen Microbiota Composition in Dairy Cows with Simple Indigestion and Healthy Cows
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal, Management, and Clinical Diagnosis
2.2. Rumen Fluid and Data Collection
2.3. DNA Extraction, PCR Amplification, and 16S rDNA Sequencing
2.4. Sequence Analysis
2.5. Statistical Analyses
3. Results
3.1. Apparent Characteristics between Healthy and Simple Indigestion Cows
3.2. Diversity of Ruminal Microbiota
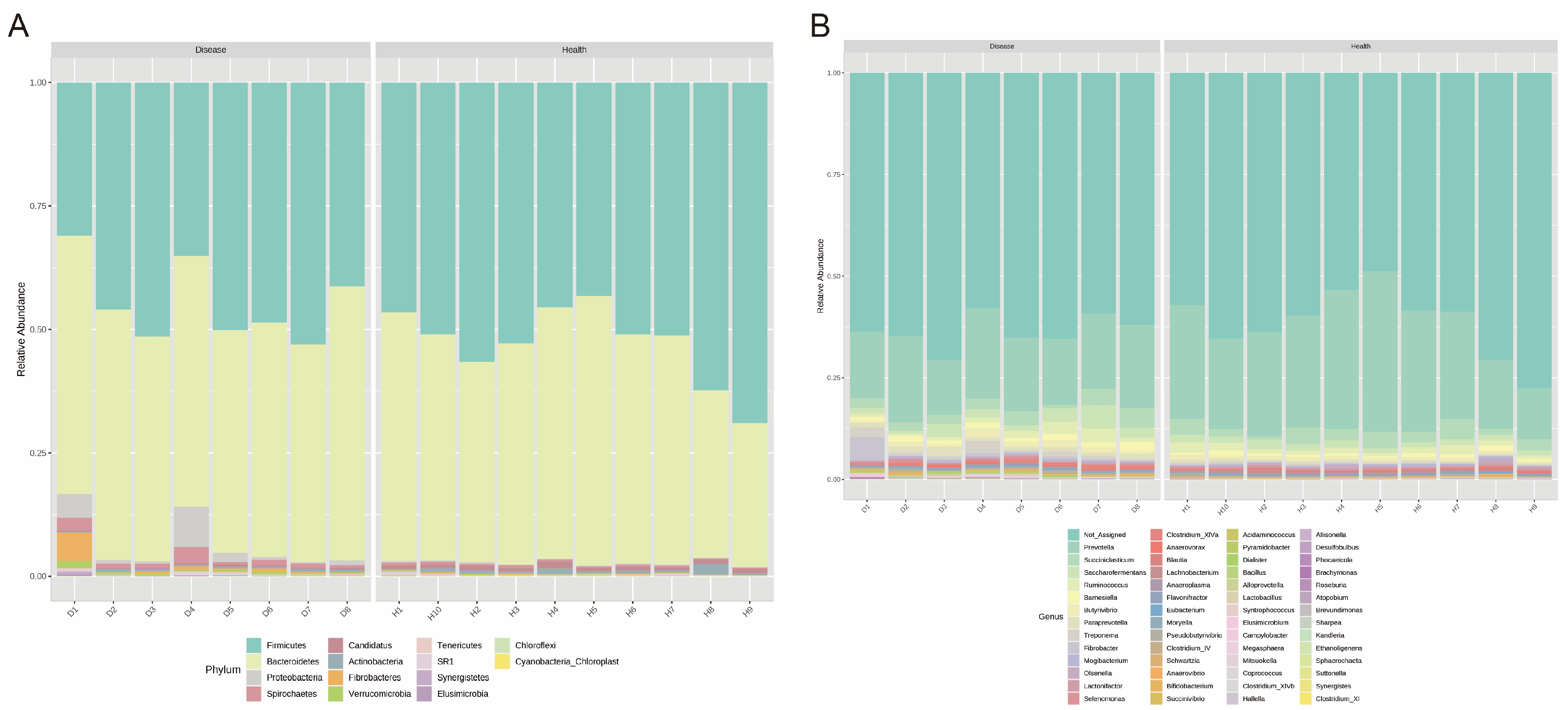
3.3. Bacterial Composition of Ruminal Microbiota
3.4. PICRUSt2 Function Prediction
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Fubini, S.L.; Yeager, A.E.; Divers, T.J. 5—Noninfectious Diseases of the Gastrointestinal Tract. In Rebhun’s Diseases of Dairy Cattle, 3rd ed.; Peek, S.F., Divers, T.J., Eds.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 168–248. [Google Scholar]
- Abutarbush, S.M. Veterinary Medicine—A Textbook of the Diseases of Cattle, Horses, Sheep, Pigs and Goats. Can. Vet. J. 2010, 51, 541. [Google Scholar]
- Van Metre, D.C.; Fecteau, G.; House, J.K.; George, L.W. Indigestion of late pregnancy in a cow. J. Am. Vet. Med. Assoc. 1995, 206, 625–627; discussion 627–628. [Google Scholar] [PubMed]
- Kinde, M.Z.; Asfaw, E. Assessment of Ruminal Disorders and Their Therapeutic Management at Three Veterinary Clinics in Gondar Town, North Western Ethiopia: A Retrospective Study. Vet. Med. 2021, 12, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Grieve, D.G.; Curtis, R.A.; Stone, J.B.; Macleod, G.K. All silage forage programs for dairy cattle. III. Health, survival, and reproduction. J. Dairy Sci. 1980, 63, 601–607. [Google Scholar] [CrossRef] [PubMed]
- Enemark, J.M.; Jørgensen, R.J.; Enemark, P.S. Rumen acidosis with a special emphasis on diagnostic aspects of subclinical rumen acidosis: A review. Vet. Zootech. 2002, 20, 16–29. [Google Scholar]
- Enemark, J.M.; Jørgensen, R.J.; Kristensen, N.B. An evaluation of parameters for the detection of subclinical rumen acidosis in dairy herds. Vet. Res. Commun. 2004, 28, 687–709. [Google Scholar] [CrossRef]
- Oetzel, G.R. Diagnosis and Management of Subacute Ruminal Acidosis in Dairy Herds. Vet. Clin. N. Am. Food Anim. Pract. 2017, 33, 463–480. [Google Scholar] [CrossRef]
- Plaizier, J.C.; Mulligan, F.J.; Neville, E.W.; Guan, L.L.; Steele, M.A.; Penner, G.B. Invited review: Effect of subacute ruminal acidosis on gut health of dairy cows. J. Dairy Sci. 2022, 105, 7141–7160. [Google Scholar] [CrossRef]
- Girard, C.L.; Graulet, B. Methods and approaches to estimate B vitamin status in dairy cows: Knowledge, gaps and advances. Methods 2021, 186, 52–58. [Google Scholar] [CrossRef]
- Kirchman, S.E.; Pinedo, P.J.; Maunsell, F.P.; Risco, C.A.; Donovan, G.A. Evaluation of milk components as diagnostic indicators for rumen indigestion in dairy cows. J. Am. Vet. Med. Assoc. 2017, 251, 580–586. [Google Scholar] [CrossRef]
- Hu, X.; Li, S.; Mu, R.; Guo, J.; Zhao, C.; Cao, Y.; Zhang, N.; Fu, Y. The Rumen Microbiota Contributes to the Development of Mastitis in Dairy Cows. Microbiol. Spectr. 2022, 10, e0251221. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Hu, X.; Bao, L.; Wu, K.; Zhao, Y.; Xiang, K.; Li, S.; Wang, Y.; Qiu, M.; Feng, L.; et al. Gut dysbiosis induces the development of mastitis through a reduction in host anti-inflammatory enzyme activity by endotoxemia. Microbiome 2022, 10, 205. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.Y.; Sun, H.Z.; Wu, X.H.; Liu, J.X.; Guan, L.L. Multi-omics reveals that the rumen microbiome and its metabolome together with the host metabolome contribute to individualized dairy cow performance. Microbiome 2020, 8, 64. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.Y.; Xie, Y.Y.; Zhong, Y.; Ma, X.J.; Sun, H.Z.; Liu, J.X. Integrated meta-omics reveals new ruminal microbial features associated with feed efficiency in dairy cattle. Microbiome 2022, 10, 32. [Google Scholar] [CrossRef]
- DePeters, E.J.; George, L.W. Rumen transfaunation. Immunol. Lett. 2014, 162, 69–76. [Google Scholar] [CrossRef]
- Liu, J.; Li, H.; Zhu, W.; Mao, S. Dynamic changes in rumen fermentation and bacterial community following rumen fluid transplantation in a sheep model of rumen acidosis: Implications for rumen health in ruminants. FASEB J. 2019, 33, 8453–8467. [Google Scholar] [CrossRef]
- Arik, H.D.; Gulsen, N.; Hayirli, A.; Alatas, M.S. Efficacy of Megasphaera elsdenii inoculation in subacute ruminal acidosis in cattle. J. Anim. Physiol. Anim. Nutr. 2019, 103, 416–426. [Google Scholar] [CrossRef]
- Sato, S. Subacute ruminal acidosis (SARA) challenge, ruminal condition and cellular immunity in cattle. Jpn. J. Vet. Res. 2015, 63 (Suppl. S1), S25–S36. [Google Scholar]
- Han, L.; Xue, W.; Cao, H.; Chen, X.; Qi, F.; Ma, T.; Tu, Y.; Diao, Q.; Zhang, C.; Cui, K. Comparison of Rumen Fermentation Parameters and Microbiota of Yaks From Different Altitude Regions in Tibet, China. Front. Microbiol. 2021, 12, 807512. [Google Scholar] [CrossRef]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Zhou, G.; Ewald, J.; Pang, Z.; Shiri, T.; Xia, J. MicrobiomeAnalyst 2.0: Comprehensive statistical, functional and integrative analysis of microbiome data. Nucleic Acids Res. 2023, 51, W310–W318. [Google Scholar] [CrossRef]
- Douglas, G.M.; Maffei, V.J.; Zaneveld, J.R.; Yurgel, S.N.; Brown, J.R.; Taylor, C.M.; Huttenhower, C.; Langille, M.G.I. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 2020, 38, 685–688. [Google Scholar] [CrossRef] [PubMed]
- Stangaferro, M.L.; Wijma, R.; Caixeta, L.S.; Al-Abri, M.A.; Giordano, J.O. Use of rumination and activity monitoring for the identification of dairy cows with health disorders: Part I. Metabolic and digestive disorders. J. Dairy Sci. 2016, 99, 7395–7410. [Google Scholar] [CrossRef] [PubMed]
- Stangaferro, M.L.; Wijma, R.; Caixeta, L.S.; Al-Abri, M.A.; Giordano, J.O. Use of rumination and activity monitoring for the identification of dairy cows with health disorders: Part III. Metritis. J. Dairy Sci. 2016, 99, 7422–7433. [Google Scholar] [CrossRef]
- Penner, G.B.; Beauchemin, K.A.; Mutsvangwa, T. An evaluation of the accuracy and precision of a stand-alone submersible continuous ruminal pH measurement system. J. Dairy Sci. 2006, 89, 2132–2140. [Google Scholar] [CrossRef]
- Zebeli, Q.; Tafaj, M.; Steingass, H.; Metzler, B.; Drochner, W. Effects of physically effective fiber on digestive processes and milk fat content in early lactating dairy cows fed total mixed rations. J. Dairy Sci. 2006, 89, 651–668. [Google Scholar] [CrossRef]
- Gharechahi, J.; Vahidi, M.F.; Sharifi, G.; Ariaeenejad, S.; Ding, X.Z.; Han, J.L.; Salekdeh, G.H. Lignocellulose degradation by rumen bacterial communities: New insights from metagenome analyses. Environ. Res. 2023, 229, 115925. [Google Scholar] [CrossRef]
- Sharma, V.; Vashishtha, A.; Jos, A.L.M.; Khosla, A.; Basu, N.; Yadav, R.; Bhatt, A.; Gulani, A.; Singh, P.; Lakhera, S.; et al. Phylogenomics of the Phylum Proteobacteria: Resolving the Complex Relationships. Curr. Microbiol. 2022, 79, 224. [Google Scholar] [CrossRef]
- Zhang, Y.K.; Zhang, X.X.; Li, F.D.; Li, C.; Li, G.Z.; Zhang, D.Y.; Song, Q.Z.; Li, X.L.; Zhao, Y.; Wang, W.M. Characterization of the rumen microbiota and its relationship with residual feed intake in sheep. Animal 2021, 15, 100161. [Google Scholar] [CrossRef] [PubMed]
- Ransom-Jones, E.; Jones, D.L.; McCarthy, A.J.; McDonald, J.E. The Fibrobacteres: An important phylum of cellulose-degrading bacteria. Microb. Ecol. 2012, 63, 267–281. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Wang, Y.; Xu, L.; Liu, J.; Zhu, W.; Mao, S. Microbiome-host co-oscillation patterns in remodeling of colonic homeostasis during adaptation to a high-grain diet in a sheep model. Anim. Microbiome 2020, 2, 22. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.; Borges, A.; Flament-Simon, S.C.; Simões, M. Quorum sensing architecture network in Escherichia coli virulence and pathogenesis. FEMS Microbiol. Rev. 2023, 47, fuad031. [Google Scholar] [CrossRef] [PubMed]
- Garner, M.R.; Flint, J.F.; Russell, J.B. Allisonella histaminiformans gen. nov., sp. nov. A novel bacterium that produces histamine, utilizes histidine as its sole energy source, and could play a role in bovine and equine laminitis. Syst. Appl. Microbiol. 2002, 25, 498–506. [Google Scholar] [CrossRef]
- Huang, H.T.; Guo, J.J.; Huang, Y.H.; Fu, Y.S. Histamine-induced changes in rat tracheal goblet cell mucin store and mucosal edema. Histochem. Cell Biol. 2013, 139, 717–726. [Google Scholar] [CrossRef]
- Walters, M.N.; Wolman, M.; Robertson, T.A.; Papadimitriou, J.M. The inhibitory effects of high molecular levan on transport across the vascular wall stimulated by histamine. J. Pathol. 1978, 124, 195–203. [Google Scholar] [CrossRef]
- Li, R.; Li, L.; Hong, P.; Lang, W.; Hui, J.; Yang, Y.; Zheng, X. β-Carotene prevents weaning-induced intestinal inflammation by modulating gut microbiota in piglets. Anim. Biosci. 2021, 34, 1221–1234. [Google Scholar] [CrossRef]
- Vartoukian, S.R.; Palmer, R.M.; Wade, W.G. The division “Synergistes”. Anaerobe 2007, 13, 99–106. [Google Scholar] [CrossRef]
- Vianna, M.E.; Conrads, G.; Gomes, B.P.; Horz, H.P. Quantification and characterization of Synergistes in endodontic infections. Oral Microbiol. Immunol. 2007, 22, 260–265. [Google Scholar] [CrossRef] [PubMed]
- Siqueira, J.F., Jr.; Rôças, I.N. Uncultivated phylotypes and newly named species associated with primary and persistent endodontic infections. J. Clin. Microbiol. 2005, 43, 3314–3319. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Xiong, J.; Yang, W.; Zhao, L.; Wang, T.; Qian, W.; Hu, S.; Wang, Q.; Aleem, M.T.; Miao, W.; et al. Metagenome of Gut Microbiota Provides a Novel Insight into the Pathogenicity of Balantioides coli in Weaned Piglets. Int. J. Mol. Sci. 2023, 24, 10791. [Google Scholar] [CrossRef]
- Zhao, Y.; Yu, S.; Li, L.; Zhao, H.; Li, Y.; Jiang, L.; Liu, M. Feeding citrus flavonoid extracts decreases bacterial endotoxin and systemic inflammation and improves immunometabolic status by modulating hindgut microbiome and metabolome in lactating dairy cows. Anim. Nutr. 2023, 13, 386–400. [Google Scholar] [CrossRef] [PubMed]
- Callahan, S.M.; Hancock, T.J.; Doster, R.S.; Parker, C.B.; Wakim, M.E.; Gaddy, J.A.; Johnson, J.G. A secreted sirtuin from Campylobacter jejuni contributes to neutrophil activation and intestinal inflammation during infection. Sci. Adv. 2023, 9, eade2693. [Google Scholar] [CrossRef] [PubMed]
- Ponziani, F.R.; Nesci, A.; Caputo, C.; Salvatore, L.; Picca, A.; Del Chierico, F.; Paroni Sterbini, F.; Marzetti, E.; Di Giorgio, A.; Santoro, L.; et al. High prevalence of lower limb atherosclerosis is linked with the gut-liver axis in patients with primary biliary cholangitis. Liver Int. 2023, 43, 370–380. [Google Scholar] [CrossRef] [PubMed]
- Bernard-Vailhe, M.A.; Besle, J.M.; Dore, J. Transformation of (sup14)C-Lignin-Labeled Cell Walls of Wheat by Syntrophococcus sucromutans, Eubacterium oxidoreducens, and Neocallimastix frontalis. Appl. Environ. Microbiol. 1995, 61, 379–381. [Google Scholar] [CrossRef]
- Hao, Y.; Guo, C.; Gong, Y.; Sun, X.; Wang, W.; Wang, Y.; Yang, H.; Cao, Z.; Li, S. Rumen Fermentation, Digestive Enzyme Activity, and Bacteria Composition between Pre-Weaning and Post-Weaning Dairy Calves. Animals 2021, 11, 2527. [Google Scholar] [CrossRef]
- Wang, T.; Sternes, P.R.; Guo, X.K.; Zhao, H.; Xu, C.; Xu, H. Autoimmune diseases exhibit shared alterations in the gut microbiota. Rheumatology 2023, 19, kead364. [Google Scholar] [CrossRef]
- Parfrey, L.W.; Moreau, C.S.; Russell, J.A. Introduction: The host-associated microbiome: Pattern, process and function. Mol. Ecol. 2018, 27, 1749–1765. [Google Scholar] [CrossRef]
- Chang, C.J.; Winn, B.J. Perturbations of the ocular surface microbiome and their effect on host immune function. Curr. Opin. Ophthalmol. 2023, 34, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Rice, R.V.; Stahmann, M.A.; Alberty, R.A. The interaction of lysine polypeptides and bovine plasma albumin. J. Biol. Chem. 1954, 209, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Thomas, B.L.; Guadagnin, A.R.; Fehlberg, L.K.; Sugimoto, Y.; Shinzato, I.; Drackley, J.K.; Cardoso, F.C. Feeding rumen-protected lysine to dairy cows prepartum improves performance and health of their calves. J. Dairy Sci. 2022, 105, 2256–2274. [Google Scholar] [CrossRef]
- Giallongo, F.; Harper, M.T.; Oh, J.; Lopes, J.C.; Lapierre, H.; Patton, R.A.; Parys, C.; Shinzato, I.; Hristov, A.N. Effects of rumen-protected methionine, lysine, and histidine on lactation performance of dairy cows. J. Dairy Sci. 2016, 99, 4437–4452. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, F.; Kong, F.; Cao, Z.; Wang, W.; Yang, H.; Wang, Y.; Bi, Y.; Li, S. Effect of Supplementing Different Levels of L-Glutamine on Holstein Calves during Weaning. Antioxidants 2022, 11, 542. [Google Scholar] [CrossRef] [PubMed]




| Items | Disease | Health | p-Value |
|---|---|---|---|
| Age (years) | 5.2 ± 0.58 | 4.9 ± 0.81 | 0.390 |
| DIM 1 (days) | 81 ± 2.12 | 80 ± 2.36 | 0.366 |
| Parity | 3.25 ± 0.89 | 2.90 ± 0.57 | 0.324 |
| Ruminal pH | 6.13 ± 0.4 | 6.22 ± 0.3 | 0.592 |
| Duration of rumination (min/d) | 324 ± 95 | 561 ± 62 | <0.05 |
| Milk yield (Kg/d) | 32.7 ± 6.2 | 47.1 ± 7.7 | <0.05 |
| Firmness 2 (Left rumen) | Yes | No | |
| Dullness 3 (Rumen) | Yes | No | |
| Rumen contraction strength 4 | 1 | 2.5 | <0.05 |
| Rumen contraction frequency (No./min) | 1.21 ± 0.7 | 2.34 ± 0.3 | <0.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Kong, F.; Liu, J.; Xia, J.; Du, W.; Li, S.; Wang, W. Comparative Analysis of Rumen Microbiota Composition in Dairy Cows with Simple Indigestion and Healthy Cows. Microorganisms 2023, 11, 2673. https://doi.org/10.3390/microorganisms11112673
Wang S, Kong F, Liu J, Xia J, Du W, Li S, Wang W. Comparative Analysis of Rumen Microbiota Composition in Dairy Cows with Simple Indigestion and Healthy Cows. Microorganisms. 2023; 11(11):2673. https://doi.org/10.3390/microorganisms11112673
Chicago/Turabian StyleWang, Shuo, Fanlin Kong, Jingjing Liu, Jianmin Xia, Wen Du, Shengli Li, and Wei Wang. 2023. "Comparative Analysis of Rumen Microbiota Composition in Dairy Cows with Simple Indigestion and Healthy Cows" Microorganisms 11, no. 11: 2673. https://doi.org/10.3390/microorganisms11112673
APA StyleWang, S., Kong, F., Liu, J., Xia, J., Du, W., Li, S., & Wang, W. (2023). Comparative Analysis of Rumen Microbiota Composition in Dairy Cows with Simple Indigestion and Healthy Cows. Microorganisms, 11(11), 2673. https://doi.org/10.3390/microorganisms11112673






