The StkSR Two-Component System Influences Colistin Resistance in Acinetobacter baumannii
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Media
2.2. Eukaryotic Cell Adherence Assay
2.3. Minimum Inhibitory Concentration and Disk Diffusion Susceptibility
2.4. Deletion Replacement Mutant Construction by Homologous Recombination
2.5. Cell Surface Hydrophobicity Tests
2.6. Transmission Electron Microscopy
2.7. SDS-PAGE and Lipooligosaccharide Silver Staining
2.8. RNA Isolation
2.9. Quantitative Reverse Transcription PCR
2.10. DNA Extraction
2.11. Library Quality Control and Bioinformatics
2.12. Statistical Analysis
2.13. Accession Numbers
3. Results
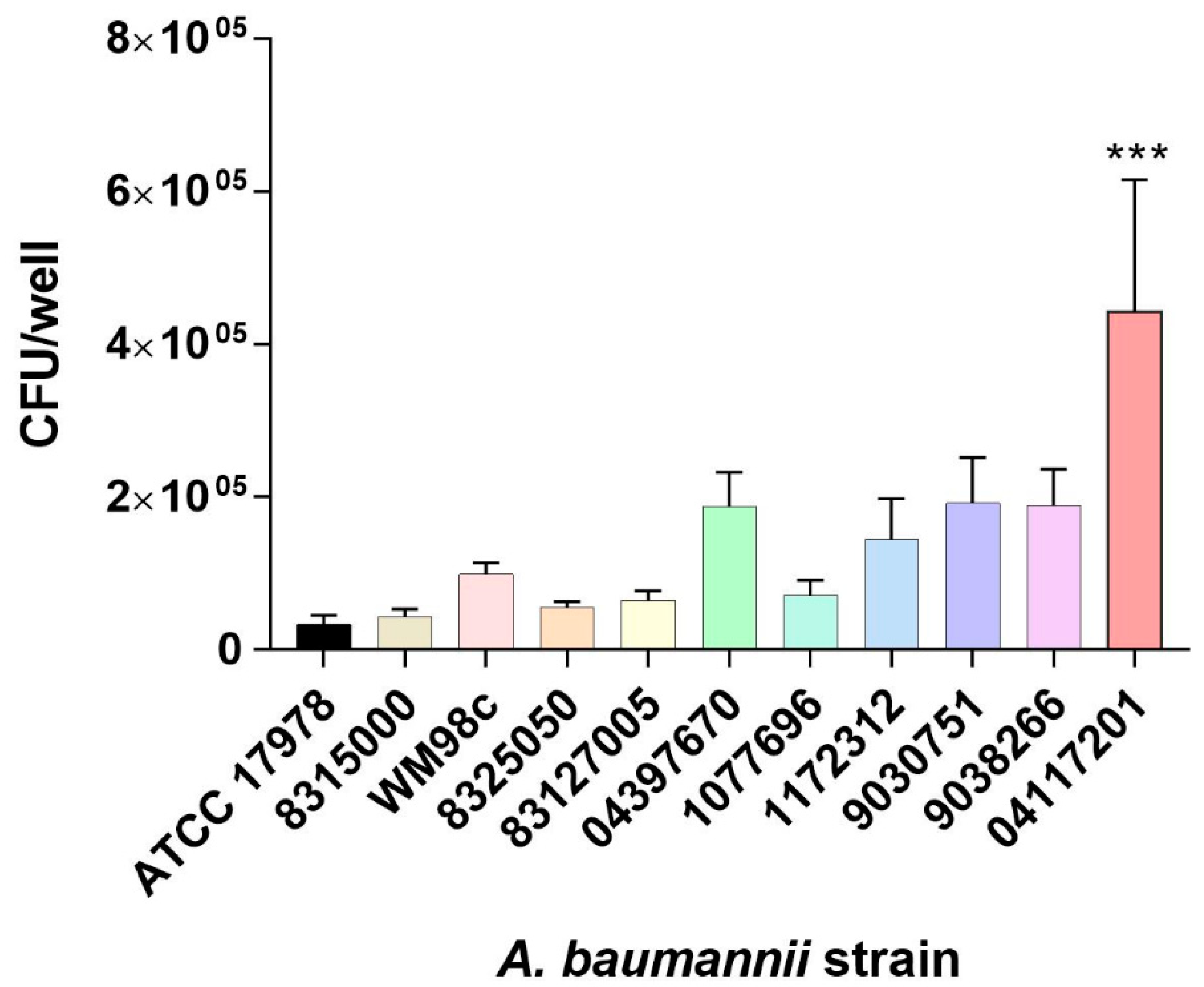
3.1. Selection of A. baumannii Isolate
3.2. Construction of an A. baumannii 04117201ΔstkR Derivative
3.3. Assessment of the A. baumannii ΔstkR Mutant Strain Antibiogram
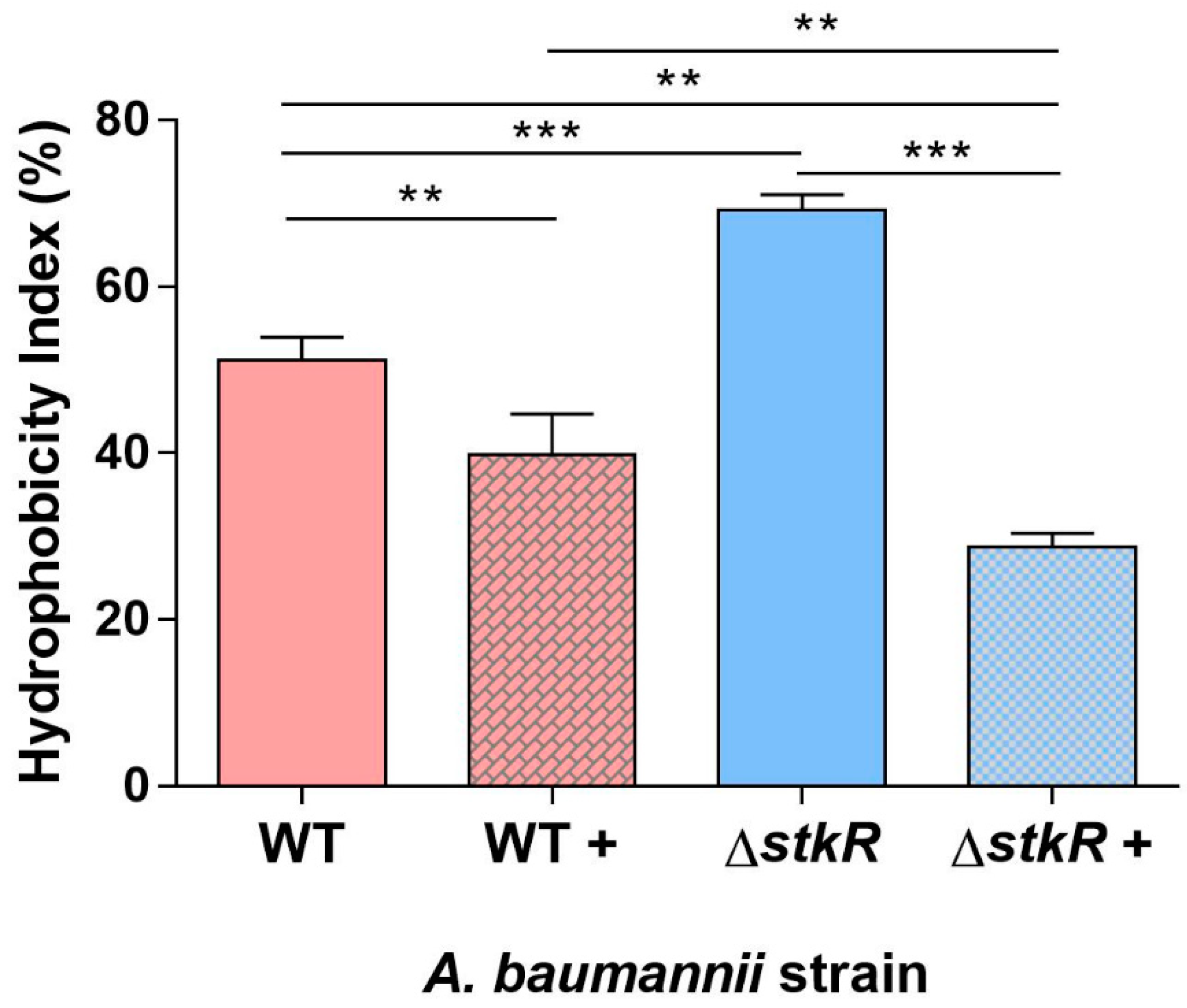
3.4. Examination of Cell Surface Hydrophobicity of the A. baumannii ΔstkR Mutant Strain When Exposed to Sub-MIC Levels of Colistin
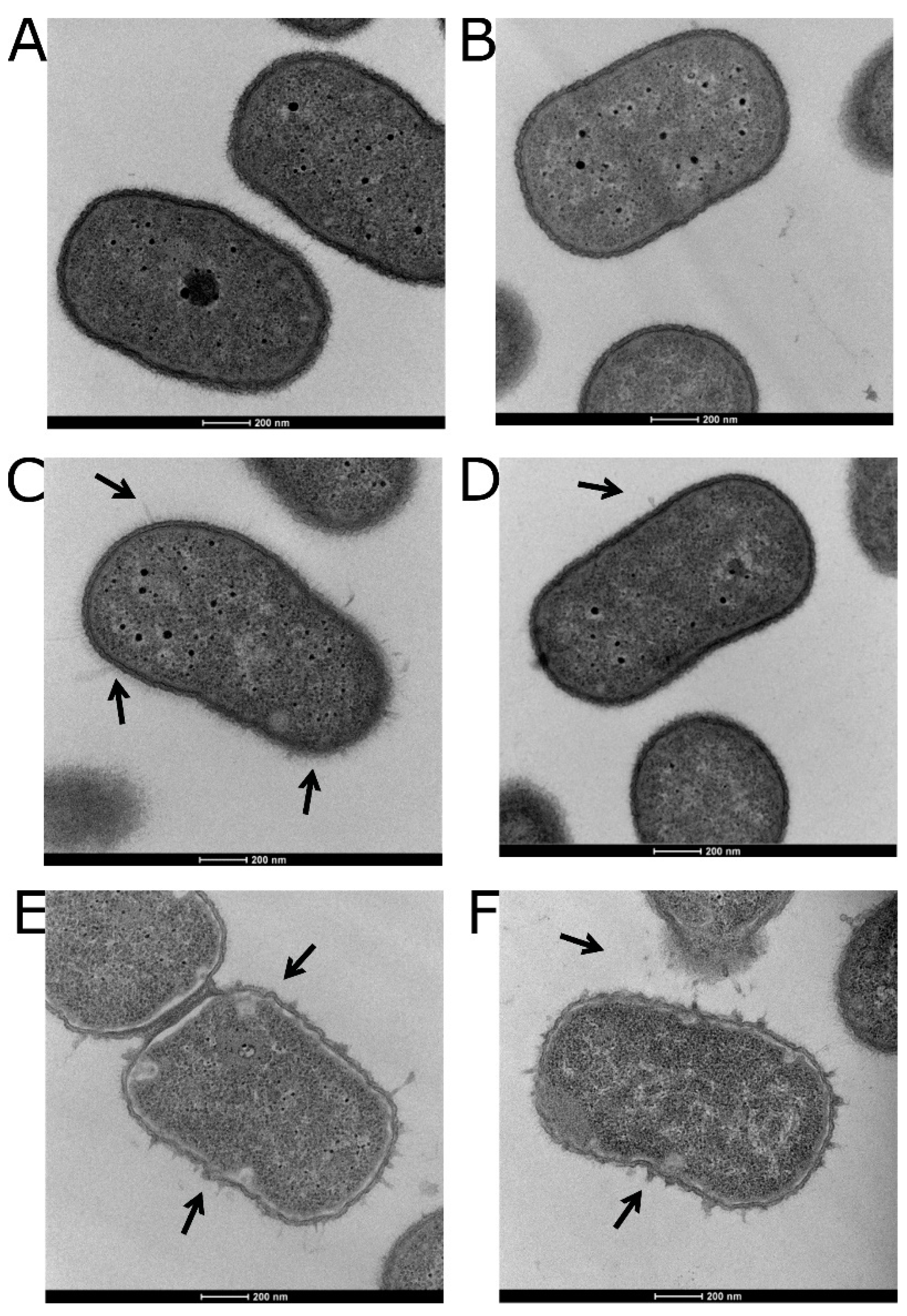
3.5. Visualizing the Effects of Colistin Stress on the Bacterial Cell Envelope
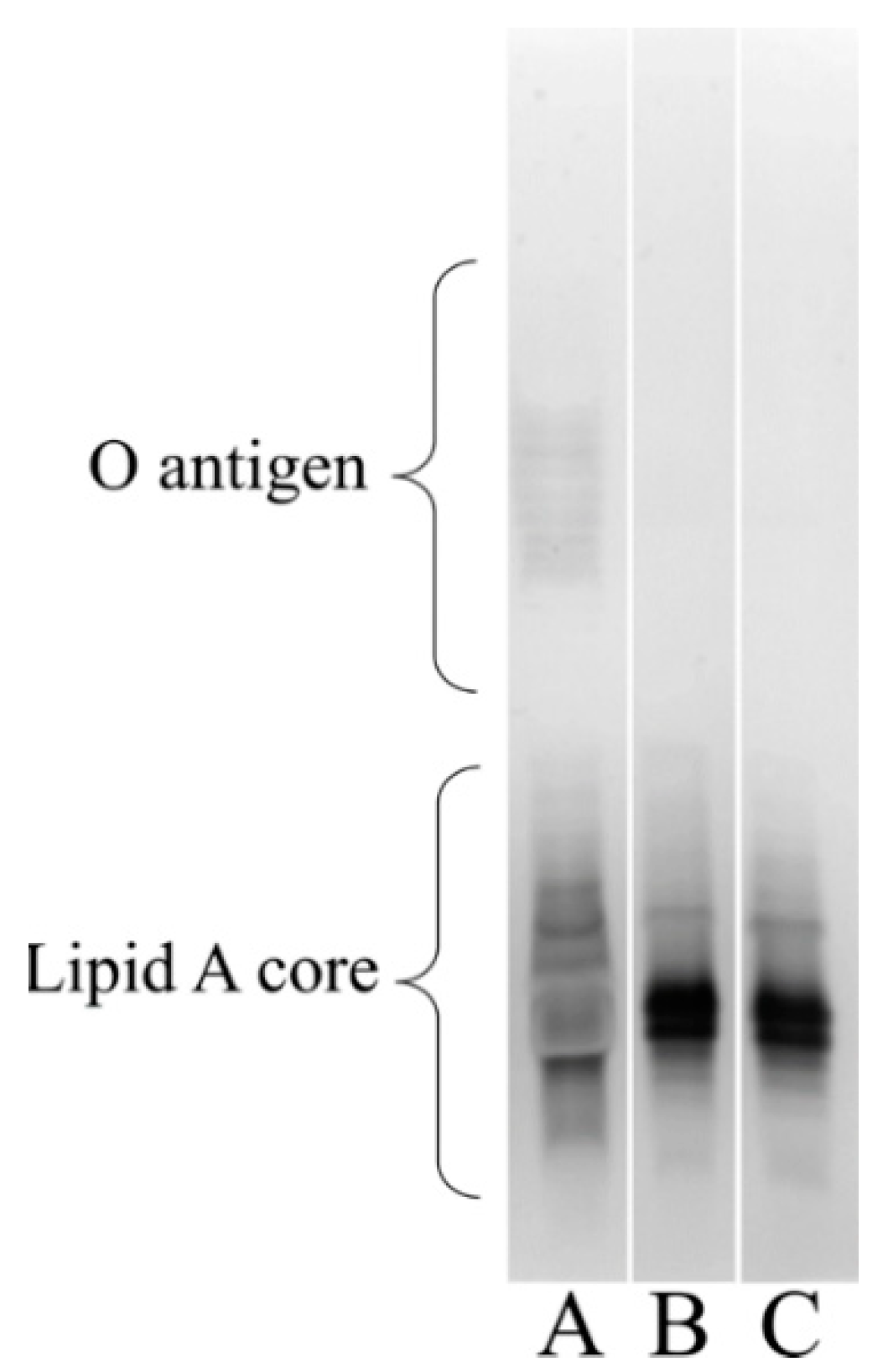
3.6. Visualizing the Lipooligosaccharide Composition of the A. baumannii ΔstkR Mutant and WT Strain
3.7. Transcriptional Profiling of the A. baumannii ΔstkR Mutant and WT Strain Treated with Sub-MIC Concentrations of Colistin
3.8. Comparative Genomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- WHO. Essential Medicines and Health Products: Global Priority List of Antibiotic-Resistant Bacteria to Guide Researcher, Discovery, and Development of New Antibiotics. 2017. Available online: www.who.int/medicines/publications/global-priority-list-antibiotic-resistant-bacteria/en/ (accessed on 27 February 2017).
- Sabnis, A.; Hagart, K.L.; Klockner, A.; Becce, M.; Evans, L.E.; Furniss, R.C.D.; Mavridou, D.A.; Murphy, R.; Stevens, M.M.; Davies, J.C.; et al. Colistin kills bacteria by targeting lipopolysaccharide in the cytoplasmic membrane. elife 2021, 10, e65836. [Google Scholar] [CrossRef] [PubMed]
- Erdem, F.; Abulaila, A.; Aktas, Z.; Oncul, O. In vitro evaluation of double carbapenem and colistin combinations against OXA-48, NDM carbapenemase-producing colistin-resistant Klebsiella pneumoniae strains. Antimicrob. Resis. Infect. Control 2020, 9, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.L.; Tomlinson, B.R.; Casella, L.G.; Shaw, L.N. Regulatory networks important for survival of Acinetobacter baumannii within the host. Curr. Opin. Microbiol. 2020, 55, 74–80. [Google Scholar] [CrossRef] [PubMed]
- De Silva, P.M.; Kumar, A. Signal transduction proteins in Acinetobacter baumannii: Role in antibiotic resistance, virulence, and potential as drug targets. Front. Microbiol. 2019, 10, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leung, L.M.; McElheny, C.L.; Gardner, F.M.; Chandler, C.E.; Bowler, S.L.; Mettus, R.T.; Spychala, C.N.; Fowler, E.L.; Opene, B.N.A.; Myers, R.A.; et al. A prospective study of Acinetobacter baumannii complex isolates and colistin susceptibility monitoring by mass spectrometry of microbial membrane glycolipids. J. Clin. Microbiol. 2019, 57, e01100-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trimble, M.J.; Mlynarcik, P.; Kolar, M.; Hancock, R.E. Polymyxin: Alternative mechanisms of action and resistance. Cold Spring Harbor Perspect. Med. 2016, 6, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Kyriakidis, I.; Vasileiou, E.; Pana, Z.D.; Tragiannidis, A. Acinetobacter baumannii antibiotic resistance mechanisms. Pathogens 2021, 10, 373. [Google Scholar] [CrossRef]
- Da Silva, G.J.; Domingues, S. Interplay between colistin resistance, virulence and fitness in Acinetobacter baumannii. Antibiotics 2017, 6, 28. [Google Scholar] [CrossRef] [Green Version]
- Bakthavatchalam, Y.D.; Pragasam, A.K.; Biswas, I.; Veeraraghavan, B. Polymyxin susceptibility testing, interpretative breakpoints and resistance mechanisms: An update. J. Global Antimicrob. Res. 2018, 12, 124–136. [Google Scholar] [CrossRef]
- Falagas, M.E.; Rafailidis, P.I.; Ioannidou, E.; Alexiou, V.G.; Matthaiou, D.K.; Karageorgopoulos, D.E.; Kapaskelis, A.; Nikita, D.; Michalopoulos, A. Colistin therapy for microbiologically documented multidrug-resistant Gram-negative bacterial infections: A retrospective cohort study of 258 patients. Internat. J. Antimicrob. Agents 2010, 35, 194–199. [Google Scholar] [CrossRef] [Green Version]
- Hancock, R.E. Peptide antibiotics. Lancet 1997, 349, 418–422. [Google Scholar] [CrossRef]
- Fiester, S.E.; Nwugo, C.C.; Penwell, W.F.; Neary, J.M.; Beckett, A.C.; Arivett, B.A.; Schmidt, R.E.; Geiger, S.C.; Connerly, P.L.; Menke, S.M.; et al. Role of the carboxy terminus of secA in iron acquisition, protein translocation, and virulence of the bacterial pathogen Acinetobacter Baumannii. Infect. Immun. 2015, 83, 1354–1365. [Google Scholar] [CrossRef] [Green Version]
- Srinivasan, V.B.; Vaidyanathan, V.; Rajamohan, G. AbuO, a TolC-like outer membrane protein of Acinetobacter baumannii, is involved in antimicrobial and oxidative stress resistance. Antimicrob. Agents Chemother. 2015, 59, 1236–1245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaddy, J.A.; Tomaras, A.P.; Actis, L.A. The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and in the interaction of this pathogen with eukaryotic cells. Infect. Immun. 2009, 77, 3150–3160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Howell, A.; Dubrac, S.; Noone, D.; Varughese, K.I.; Devine, K. Interactions between the YycFG and PhoPR two-component systems in Bacillus subtilis: The PhoR kinase phosphorylates the non-cognate YycF response regulator upon phosphate limitation. Molec. Microbiol. 2006, 59, 1199–1215. [Google Scholar] [CrossRef] [PubMed]
- Rowland, M.A.; Deeds, E.J. Crosstalk and the evolution of specificity in two-component signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 5550–5555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Capra, E.J.; Laub, M.T. Evolution of two-component signal transduction systems. Ann. Rev. Microbiol. 2012, 66, 325–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rajput, A.; Seif, Y.; Choudhary, K.S.; Dalldorf, C.; Poudel, S.; Monk, J.M.; Palsson, B.O. Pangenome analytics reveal two-component systems as conserved targets in ESKAPEE pathogens. mSystems 2021, 6, e00981-20. [Google Scholar] [CrossRef]
- Cerqueira, G.M.; Kostoulias, X.; Khoo, C.; Aibinu, I.; Qu, Y.; Traven, A.; Peleg, A.Y. A global virulence regulator in Acinetobacter baumannii and its control of the phenylacetic acid catabolic pathway. J. Infect. Dis. 2014, 210, 46–55. [Google Scholar] [CrossRef] [Green Version]
- Quesada, A.; Porrero, M.C.; Tellez, S.; Palomo, G.; Garcia, M.; Dominguez, L. Polymorphism of genes encoding PmrAB in colistin-resistant strains of Escherichia coli and Salmonella enterica isolated from poultry and swine. J. Antimicrob. Chemother. 2015, 70, 71–74. [Google Scholar] [CrossRef] [Green Version]
- Park, Y.K.; Choi, J.Y.; Shin, D.; Ko, K.S. Correlation between overexpression and amino acid substitution of the PmrAB locus and colistin resistance in Acinetobacter Baumannii. Int. J. Antimicrob. Agents 2011, 37, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Girardello, R.; Visconde, M.; Cayo, R.; Figueiredo, R.C.; Mori, M.A.; Lincopan, N.; Gales, A.C. Diversity of polymyxin resistance mechanisms among Acinetobacter baumannii clinical isolates. Diagnost.Microbiol. Infect. Dis. 2017, 87, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Adams, F.G.; Stroeher, U.H.; Hassan, K.A.; Marri, S.; Brown, M.H. Resistance to pentamidine is mediated by AdeAB, regulated by AdeRS, and influenced by growth conditions in Acinetobacter baumannii ATCC 17978. PLoS ONE 2018, 13, e0197412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richmond, G.E.; Evans, L.P.; Anderson, M.J.; Wand, M.E.; Bonney, L.C.; Ivens, A.; Chua, K.L.; Webber, M.A.; Sutton, J.M.; Peterson, M.L.; et al. The Acinetobacter baumannii two-component system AdeRS regulates genes required for multidrug efflux, biofilm formation, and virulence in a strain-specific manner. mBio 2016, 7, e00430-16. [Google Scholar] [CrossRef] [Green Version]
- Cheng, H.Y.; Chen, Y.F.; Peng, H.L. Molecular characterization of the PhoPQ-PmrD-PmrAB mediated pathway regulating polymyxin B resistance in Klebsiella pneumoniae CG43. J. Biomed. Sci. 2010, 17, 60–76. [Google Scholar] [CrossRef] [Green Version]
- Jaidane, N.; Naas, T.; Mansour, W.; Radhia, B.B.; Jerbi, S.; Boujaafar, N.; Bouallegue, O.; Bonnin, R.A. Genomic analysis of in vivo acquired resistance to colistin and rifampicin in Acinetobacter Baumannii. Internat. J. Antimicrob. Agents 2018, 51, 266–269. [Google Scholar] [CrossRef]
- Boinett, C.J.; Cain, A.K.; Hawkey, J.; Do Hoang, N.T.; Khanh, N.N.T.; Thanh, D.P.; Dordel, J.; Campbell, J.I.; Lan, N.P.H.; Mayho, M.; et al. Clinical and laboratory-induced colistin-resistance mechanisms in Acinetobacter Baumannii. Micro. Genom. 2019, 5, 1–10. [Google Scholar] [CrossRef]
- Beceiro, A.; Llobet, E.; Aranda, J.; Bengoechea, J.A.; Doumith, M.; Hornsey, M.; Dhanji, H.; Chart, H.; Bou, G.; Livermore, D.D.; et al. Phosphoethanolamine modification of lipid A in colistin-resistant variants of Acinetobacter baumannii mediated by the pmrAB two-component regulatory system. Antimicrob. Agents Chemother. 2011, 55, 3370–3379. [Google Scholar] [CrossRef] [Green Version]
- Jones, C.L.; Singh, S.S.; Alamneh, Y.; Casella, L.G.; Ernst, R.K.; Lesho, E.P.; Waterman, P.E.; Zurawski, D.V. In vivo fitness adaptations of colistin-resistant Acinetobacter baumannii isolates to oxidative stress. Antimicrob. Agents Chemother. 2017, 61, e00598-16. [Google Scholar] [CrossRef] [Green Version]
- Trebosc, V.; Gartenmann, S.; Totzl, M.; Lucchini, V.; Schellhorn, B.; Pieren, M.; Lociuro, S.; Gitzinger, M.; Tigges, M.; Bumann, D.; et al. Dissecting colistin resistance mechanisms in extensively drug-resistant Acinetobacter baumannii clinical isolates. mBio 2019, 10, e01083-19. [Google Scholar] [CrossRef] [Green Version]
- Deveson Lucas, D.; Crane, B.; Wright, A.; Han, M.L.; Moffatt, J.; Bulach, D.; Gladman, S.L.; Powell, D.; Aranda, J.; Seemann, T.; et al. Emergence of high-level colistin resistance in an Acinetobacter baumannii clinical isolate mediated by inactivation of the global regulator H-NS. Antimicrob. Agents Chemother. 2018, 62, e02442-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eijkelkamp, B.A.; Hassan, K.A.; Paulsen, I.T.; Brown, M.H. Investigation of the human pathogen Acinetobacter baumannii under iron limiting conditions. BMC Genom. 2011, 12, 126–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giles, S.K.; Stroeher, U.H.; Eijkelkamp, B.A.; Brown, M.H. Identification of genes essential for pellicle formation in Acinetobacter Baumannii. BMC Microbiol. 2015, 15, 116–129. [Google Scholar] [CrossRef] [Green Version]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Molec. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Simon, R.; Priefer, U.; Puhler, A. A broad host range mobilization system for in vivo genetic-engineering: Transposon mutagenesis in Gram-negative bacteria. Biotechnol. Appl. Microbiol. 1983, 1, 784–791. [Google Scholar] [CrossRef]
- Yanisch-Perron, C.; Vieira, J.; Messing, J. Improved M13 phage cloning vectors and host strains: Nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 1985, 33, 103–119. [Google Scholar] [CrossRef]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Disk Susceptibility Testing, Seventeenth Information Supplement; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2007; Volume 27, p. M100-S17. [Google Scholar]
- CLSI. Clinical Laboratory Standards Institute evaluation protocols. MLO-Med. Lab. Obs. 2006, 38, 40–41. [Google Scholar]
- Eijkelkamp, B.A.; Stroeher, U.H.; Hassan, K.A.; Elbourne, L.D.; Paulsen, I.T.; Brown, M.H. H-NS plays a role in expression of Acinetobacter baumannii virulence features. Infect. Immun. 2013, 81, 2574–2583. [Google Scholar] [CrossRef] [Green Version]
- Gui, S.; Li, R.; Feng, Y.; Wang, S. Transmission electron microscopic morphological study and flow cytometric viability assessment of Acinetobacter baumannii susceptible to Musca domestica cecropin. Sci. World J. 2014, 2014, 1–6. [Google Scholar]
- Tsai, C.M.; Frasch, C.E. A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal. Biochem. 1982, 119, 115–119. [Google Scholar] [CrossRef]
- Fomsgaard, A.; Freudenberg, M.A.; Galanos, C. Modification of the silver staining technique to detect lipopolysaccharide in polyacrylamide gels. J. Clin. Microbiol. 1990, 28, 2627–2631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brazma, A.; Vilo, J. Gene expression data analysis. Microbes Infect. 2001, 3, 823–829. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comp. Biol. 2017, 13, e1005595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Samuels, M.L.; Witmer, J.A.; Schaffner, A.A. Statistics for the Life Sciences, 4th ed.; Pearson Education: Boston, MA, USA, 2012. [Google Scholar]
- Brossard, K.A.; Campagnari, A.A. The Acinetobacter baumannii biofilm-associated protein plays a role in adherence to human epithelial cells. Infect. Immun. 2012, 80, 228–233. [Google Scholar] [CrossRef] [Green Version]
- Eijkelkamp, B.A.; Stroeher, U.H.; Hassan, K.A.; Papadimitrious, M.S.; Paulsen, I.T.; Brown, M.H. Adherence and motility characteristics of clinical Acinetobacter baumannii isolates. FEMS Microbiol. Lett. 2011, 323, 44–51. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Palmer, K.L.; Aye, L.M.; Whiteley, M. Nutritional cues control Pseudomonas aeruginosa multicellular behavior in cystic fibrosis sputum. J. Bacteriol. 2007, 189, 8079–8087. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marchand, I.; Damier-Piolle, L.; Courvalin, P.; Lambert, T. Expression of the RND-type efflux pump AdeABC in Acinetobacter baumannii is regulated by the AdeRS two-component system. Antimicrob. Agents Chemother. 2004, 48, 3298–3304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adams, M.D.; Goglin, K.; Molyneaux, N.; Hujer, K.M.; Lavender, H.; Jamison, J.J.; MacDonald, I.J.; Martin, K.M.; Russo, T.; Campagnari, A.A.; et al. Comparative genome sequence analysis of multidrug-resistant Acinetobacter Baumannii. J. Bacteriol. 2008, 190, 8053–8064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, M.F.; Lin, Y.Y.; Lan, C.Y. The role of the two-component system BaeSR in disposing chemicals through regulating transporter systems in Acinetobacter baumannii. PLoS ONE 2015, 10, e0132843. [Google Scholar] [CrossRef] [PubMed]
- Heermann, R.; Jung, K. The complexity of the ‘simple’ two-component system KdpD/KdpE in Escherichia coli. FEMS Microbiol. Lett. 2010, 304, 97–106. [Google Scholar] [CrossRef] [Green Version]
- Bhagirath, A.Y.; Li, Y.; Patidar, R.; Yerex, K.; Ma, X.; Kumar, A.; Duan, K. Two component regulatory systems and antibiotic resistance in Gram-negative pathogens. Internat. J. Molec. Sci. 2019, 20, 1781. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Xue, M.; Yu, L.; Qi, K.; Ni, J.; Chen, X.; Deng, R.; Shang, F.; Xue, T. QseBC is involved in the biofilm formation and antibiotic resistance in Escherichia coli isolated from bovine mastitis. Peer J. 2020, 8, e8833. [Google Scholar] [CrossRef] [Green Version]
- Meng, X.; Ahator, S.D.; Zhang, L.H. Molecular mechanisms of phosphate stress activation of Pseudomonas aeruginosa quorum sensing systems. mSphere 2020, 5, e00119-20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tomaras, A.P.; Flagler, M.J.; Dorsey, C.W.; Gaddy, J.A.; Actis, L.A. Characterization of a two-component regulatory system from Acinetobacter baumannii that controls biofilm formation and cellular morphology. Microbiology 2008, 154, 3398–3409. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, K.; Shen, D.; Yang, N.; Chou, S.H.; Gomelsky, M.; Qian, G. Coordinated control of the type IV pili and c-di-GMP-dependent antifungal antibiotic production in Lysobacter by the response regulator PilR. Molec. Plant Pathol. 2021, 22, 602–617. [Google Scholar] [CrossRef] [PubMed]
- Leech, A.J.; Sprinkle, A.; Wood, L.; Wozniak, D.J.; Ohman, D.E. The NtrC family regulator AlgB, which controls alginate biosynthesis in mucoid Pseudomonas aeruginosa, binds directly to the algD promoter. J. Bacteriol. 2008, 190, 581–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Delumeau, O.; Dutta, S.; Brigulla, M.; Kuhnke, G.; Hardwick, S.W.; Volker, U.; Yudkin, M.D.; Lewis, R.J. Functional and structural characterization of RsbU, a stress signaling protein phosphatase 2C. J. Biol. Chem. 2004, 279, 40927–40937. [Google Scholar] [CrossRef] [Green Version]
- Yoon, E.J.; Kim, H.S.; Woo, H.; Choi, Y.J.; Won, D.; Choi, J.R.; Kim, Y.A.; Jeong, S.H. Trajectory of genetic alterations associated with colistin resistance in Acinetobacter baumannii during an in-hospital outbreak of infection. J. Antimicrob. Chemother. 2021, 77, 69–73. [Google Scholar] [CrossRef]
- Moffatt, J.H.; Harper, M.; Harrison, P.; Hale, J.D.; Vinogradov, E.; Seemann, T.; Henry, R.; Crane, B.; St Michael, F.; Cox, A.D.; et al. Colistin resistance in Acinetobacter baumannii is mediated by complete loss of lipopolysaccharide production. Antimicrob. Agents Chemother. 2010, 54, 4971–4977. [Google Scholar] [CrossRef] [Green Version]
- Rosenberg, E.; Gottlieb, A.; Rosenberg, M. Inhibition of bacterial adherence to hydrocarbons and epithelial cells by emulsan. Infect. Immun. 1983, 39, 1024–1028. [Google Scholar] [CrossRef] [Green Version]
- Sun, B.; Liu, J.; Jiang, Y.; Shao, L.; Yang, S.; Chen, D. New mutations involved in colistin resistance in Acinetobacter Baumannii. Msphere 2020, 5, e00895-19. [Google Scholar] [CrossRef] [Green Version]
- Alves, C.S.; Melo, M.N.; Franquelim, H.G.; Ferre, R.; Planas, M.; Feliu, L.; Bardaji, E.; Kowalczyk, W.; Andreu, D.; Santos, N.C.; et al. Escherichia coli cell surface perturbation and disruption induced by antimicrobial peptides BP100 and pepR. J. Biol. Chem. 2010, 285, 27536–27544. [Google Scholar] [CrossRef] [Green Version]
- Vrancianu, C.O.; Gheorghe, I.; Czobor, I.B.; Chifiriuc, M.C. Antibiotic resistance profiles, molecular mechanisms and innovative treatment strategies of Acinetobacter Baumannii. Microorganisms 2020, 8, 935. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, L.A.; Herrera, C.M.; Fernandez, L.; Hankins, J.V.; Trent, M.S.; Hancock, R.E. The pmrCAB operon mediates polymyxin resistance in Acinetobacter baumannii ATCC 17978 and clinical isolates through phosphoethanolamine modification of Lipid A. Antimicrob. Agents Chemother. 2011, 55, 3743–3751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riveros-Rosas, H.; Julian-Sanchez, A.; Moreno-Hagelsieb, G.; Munoz-Clares, R.A. Aldehyde dehydrogenase diversity in bacteria of the Pseudomonas genus. Chem.-Biologic. Interac. 2019, 304, 83–87. [Google Scholar] [CrossRef]
- Price-Whelan, A.; Dietrich, L.E.; Newman, D.K. Rethinking ‘secondary’ metabolism: Physiological roles for phenazine antibiotics. Nat. Chem. Biol 2006, 2, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, Y.F.; Abou-Shleib, H.M.; Khalil, A.M.; El-Guink, N.M.; El-Nakeeb, M.A. Membrane permeabilization of colistin toward pan-drug resistant Gram-negative isolates. Brazil. J. Microbiol. 2016, 47, 381–388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hameed, F.; Khan, M.A.; Muhammad, H.; Sarwar, T.; Bilal, H.; Rehman, T.U. Plasmid-mediated mcr-1 gene in Acinetobacter baumannii and Pseudomonas aeruginosa: First report from Pakistan. Rev. Soc. Bras. Med. Tropic. 2019, 52, e20190237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henry, R.; Vithanage, N.; Harrison, P.; Seemann, T.; Coutts, S.; Moffatt, J.H.; Nation, R.L.; Li, J.; Harper, M.; Adler, B.; et al. Colistin-resistant, lipopolysaccharide-deficient Acinetobacter baumannii responds to lipopolysaccharide loss through increased expression of genes involved in the synthesis and transport of lipoproteins, phospholipids, and poly-beta-1,6-N-acetylglucosamine. Antimicrob. Agents Chemother. 2012, 56, 59–69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Velkov, T.; Thompson, P.E.; Nation, R.L.; Li, J. Structure activity relationships of polymyxin antibiotics. J. Med. Chem. 2010, 53, 1898–1916. [Google Scholar] [CrossRef] [Green Version]
- Soon, R.L.; Li, J.; Boyce, J.D.; Harper, M.; Adler, B.; Larson, I.; Nation, R.L. Cell surface hydrophobicity of colistin-susceptible vs resistant Acinetobacter baumannii determined by contact angles: Methodological considerations and implications. J. Appl. Microbiol. 2012, 113, 940–951. [Google Scholar] [CrossRef] [Green Version]
- Coetzee, J.; Corcoran, C.; Prentice, E.; Moodley, M.; Mendelson, M.; Poirel, L.; Nordmann, P.; Brink, A.J. Emergence of plasmid-mediated colistin resistance (MCR-1) among Escherichia coli isolated from South African patients. S. Afr. Med. J. 2016, 106, 449–450. [Google Scholar] [CrossRef]
- Liu, Y.Y.; Wang, Y.; Walsh, T.R.; Yi, L.X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, H.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Stokes, J.M.; MacNair, C.R.; Ilyas, B.; French, S.; Cote, J.P.; Bouwman, C.; Farha, M.A.; Sieron, A.O.; Whitfield, C.; Coombes, B.K.; et al. Pentamidine sensitizes Gram-negative pathogens to antibiotics and overcomes acquired colistin resistance. Nat. Microbiol. 2017, 2, 1–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elnahriry, S.S.; Khalifa, H.O.; Soliman, A.M.; Ahmed, A.M.; Hussein, A.M.; Shimamoto, T.; Shimamoto, T. Emergence of plasmid-mediated colistin resistance gene mcr-1 in a clinical Escherichia coli isolate from Egypt. Antimicrob. Agents Chemother. 2016, 60, 3249–3250. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, C.Y.; Ang, G.Y.; Chin, P.S.; Ngeow, Y.F.; Yin, W.F.; Chan, K.G. Emergence of mcr-1-mediated colistin resistance in Escherichia coli in Malaysia. Internat. J. Antimicrob. Agents 2016, 47, 504–505. [Google Scholar] [CrossRef]
- Vijayakumar, S.; Kanthan, K.; Veeraraghavan, B. Whole-genome shotgun sequences of seven colistin-resistant Acinetobacter baumannii isolates from bacteraemia. J. Antimicrob. Resis. 2018, 12, 155–156. [Google Scholar] [CrossRef]
- Moffatt, J.H.; Harper, M.; Adler, B.; Nation, R.L.; Li, J.; Boyce, J.D. Insertion sequence ISAba11 is involved in colistin resistance and loss of lipopolysaccharide in Acinetobacter Baumannii. Antimicrob. Agents Chemother. 2011, 55, 3022–3024. [Google Scholar] [CrossRef] [Green Version]
- Adams, M.D.; Nickel, G.C.; Bajaksouzian, S.; Lavender, H.; Murthy, A.R.; Jacobs, M.R.; Bonomo, R.A. Resistance to colistin in Acinetobacter baumannii associated with mutations in the PmrAB two-component system. Antimicrob. Agents Chemother. 2009, 53, 3628–3634. [Google Scholar] [CrossRef] [Green Version]
- Qureshi, Z.A.; Hittle, L.E.; O’Hara, J.A.; Rivera, J.A.; Syed, A.; Shields, R.K.; Pasculle, A.W.; Ernst, R.K.; Doi, Y. Colistin-resistant Acinetobacter baumannii: Beyond carbapenem resistance. Clin. Infect. Dis. 2015, 60, 1295–1303. [Google Scholar] [CrossRef] [Green Version]
- Lopez-Rojas, R.; Dominguez-Herrera, J.; McConnell, M.J.; Docobo-Perez, F.; Smani, Y.; Fernandez-Reyes, M.; Rivas, L.; Pachon, J. Impaired virulence and in vivo fitness of colistin-resistant Acinetobacter Baumannii. J. Infect. Dis. 2011, 203, 545–548. [Google Scholar] [CrossRef] [Green Version]
| A. baumannii Strains | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Compound ab | ATCC 17978 | 04117201 | 1172312 | WM98 c | 08315000 | 04397670 | 9030751 | 1077697 | 9038266 | 08325050 | 083217005 |
| KAN c | R | R | R | S | S | R | R | S | R | S | R |
| ERY | S | S | S | S | R | S | S | I | S | I | R |
| TET | S | R | R | I | R | I | R | I | R | R | R |
| AMP | 250 | >500 | >500 | 250 | >500 | >500 | >500 | >500 | >500 | >500 | >500 |
| CIP | 3.25 | 30 | 30 | 3.25 | >240 | 60 | 60 | 60 | 60 | >240 | >240 |
| CST | 4 | 8 | 8 | 16 | 8 | 8 | 32 | 32 | 16 | 4 | 4 |
| TEL | 0.93 | 1.87 | 1.87 | 1.87 | 3.75 | 3.75 | 1.87 | 1.87 | 3.75 | 3.75 | 3.75 |
| GEN | 32 | 500 | 500 | 16 | 125 | 250 | 125 | 125 | 250 | 125 | 125 |
| PMB | 8 | 16 | 16 | 16 | 8 | 16 | 32 | 32 | 8 | 8 | 8 |
| RIF | 8 | 8 | 8 | 8 | >64 | 4 | 4 | 4 | 4 | 4 | 4 |
| H2O2 d | 125 | 250 | 250 | 500 | 250 | 125 | 250 | 250 | 250 | 125 | 250 |
| Peg # in 04117201 | Location in 04117201 | A1S + # in ATCC 17978 | Location in ATCC 17978 | Present in SDF | Response Regulator a | Description |
|---|---|---|---|---|---|---|
| 25 | 18192–18935 | A1S_1753 | 2038890–2044193 | N | AdeR | TCS, known as AdeRS, involved in antibiotic resistance and AdeABC efflux pump [58] |
| 95 | 40498–39824 | A1S_2751 | 3193938- 3189264 | Y | PmrA | TCS, known as PmrAB involved in lipid A modification [59] |
| 226 | 180539–179853 | A1S_2883 | 3334601–3333044 | Y | BaeR | TCS known as BaeRS involved in chemical transport and regulation+ of AdeABC and AdeIJK pumps [60] |
| 519 | 97888–98604 | A1S_2137 | 2497340–2492622 | Y | KdpE | TCS, known as KdpED involved in potassium transport [61] |
| 838 | 118775–122230 | A1S_1394 | 1636507–1641464 | N | StkR | TCS known as GerE (renamed here as StkSR) |
| 1104 | 21883–21119 | A1S_3229 | 3720752–3725517 | Y | OmpR | TCS, EnvZ-OmpR, involved in osmotic stress [62] |
| 1270 | 84347–85009 | A1S_2288 | 2655031–2650369 | Y | QseB | TCS, known as QseBC involved in biofilm formation [63] |
| 1649 | 134448–135158 | A1S_3375 | 3881425–3886134 | Y | PhoB | TCS, known as PhoRB involved in phosphate stress and quorum sensing [64] |
| 1825 | 90896–91486 | A1S_2006 | 2319938–2324522 | N | NasT | Orphan response regulator known as NasT; no histidine kinase identifiable up or down stream |
| 1962 | 98859–99575 | A1S_0748 | 887026–892617 | Y | RstA | TCS, known as BfmR involved in biofilm formation [65] |
| 2189 | 37589–36954 | A1S_0233 | 259526–264163 | N | PilR | TCS, known as PilR involved in Type 4 fimbriae expression [66] |
| 2213 | 62365–63105 | A1S_0261 | 284993–289733 | Y | AlgB | TCS, known as AlgBZ involved in alginate biosynthesis [67] |
| 2465 | 1290–2258 | A1S_0621 | 670111–674210 | N | RsbU | Orphan response regulator known as RsbU; no histidine kinase identifiable up or down stream [68] |
| Acinetobacter baumannii Strain | |||
|---|---|---|---|
| Compound | WT (µg/mL) | ΔstkR (µg/mL) | Antibiotic Family Group |
| Novobiocin | 31 | 31 | Aminocoumaria |
| Amikacin | 10 | 10 | Aminoglycoside |
| Gentamicin | 500 | 500 | Aminoglycoside |
| Kanamycin | 3000 | >3000 | Aminoglycoside |
| Streptomycin | >300 | >300 | Aminoglycoside |
| Chloramphenicol; | 5 | 5–10 | Amphenicol |
| Rifampicin | 4 | 4–8 | Ansamycins |
| Ampicillin | >500 | >500 | Beta-lactam |
| Ciprofloxacin | 30 | 30 | Carboxy fluoroquinoline |
| Chlorhexidine | 7.5–15 | 15 | Chlorobenzenes |
| Triclosan | 0.15–0.65 | 0.31–0.65 | Diphenyl ethers |
| Tellurite | 1.87 | 1.87 | Metal |
| Triton X100 | 64 | 64 | Nonionic surfactant |
| Pentamidine | 125 | 125 | Phenol ether |
| Colistin | 8 | 16 | Polymyxin |
| Polymyxin | 16 | 32 | Polymyxin |
| Nalidixic acid | 1250 | 1250 | Quinolone |
| SNP Position a | Codon in WT | Codon in ΔstkR | Amino Acid in WT | Amino Acid in ΔstkR | Annotation of Region b |
|---|---|---|---|---|---|
| 176528 | - | - | - | - | Non-coding region |
| 538323 | GAG | GGG | Glu | Gly | Hypothetical protein (Repeat region) |
| 1001055 | - | - | - | - | Non-coding region |
| 1058011 | - | - | - | - | Non-coding region |
| 1098529 | - | - | - | - | Non-coding region |
| 1150782 | TAA | CAA | Stop | Gln | Aldehyde dehydrogenase |
| 1590889 | GCT | GCC | Ala | Ala | Hypothetical protein |
| 2730447 | TTT | TTC | Phe | Phe | Hypothetical protein |
| 2796261 | - | - | - | - | Non-coding region |
| 3132365 | TTC | TCC | Phe | Ser | Phenazine biosynthesis protein (PhzF) |
| 3201037 | CCC | TCC | Pro | Ser | Repeat region |
| 3361708 | - | - | - | - | Non-coding region |
| 3370031 | - | - | - | - | Non-coding region |
| 3445432 | AAA | GAA | Lys | Glu | Alcyl-CoA Dehydrogenase |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giles, S.K.; Stroeher, U.H.; Papudeshi, B.; Edwards, R.A.; Carlson-Jones, J.A.; Roach, M.; Brown, M.H. The StkSR Two-Component System Influences Colistin Resistance in Acinetobacter baumannii. Microorganisms 2022, 10, 985. https://doi.org/10.3390/microorganisms10050985
Giles SK, Stroeher UH, Papudeshi B, Edwards RA, Carlson-Jones JA, Roach M, Brown MH. The StkSR Two-Component System Influences Colistin Resistance in Acinetobacter baumannii. Microorganisms. 2022; 10(5):985. https://doi.org/10.3390/microorganisms10050985
Chicago/Turabian StyleGiles, Sarah K., Uwe H. Stroeher, Bhavya Papudeshi, Robert A. Edwards, Jessica AP. Carlson-Jones, Michael Roach, and Melissa H. Brown. 2022. "The StkSR Two-Component System Influences Colistin Resistance in Acinetobacter baumannii" Microorganisms 10, no. 5: 985. https://doi.org/10.3390/microorganisms10050985
APA StyleGiles, S. K., Stroeher, U. H., Papudeshi, B., Edwards, R. A., Carlson-Jones, J. A., Roach, M., & Brown, M. H. (2022). The StkSR Two-Component System Influences Colistin Resistance in Acinetobacter baumannii. Microorganisms, 10(5), 985. https://doi.org/10.3390/microorganisms10050985





