A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms
Abstract
1. Introduction
2. Materials and Methods
2.1. Mathematical Model
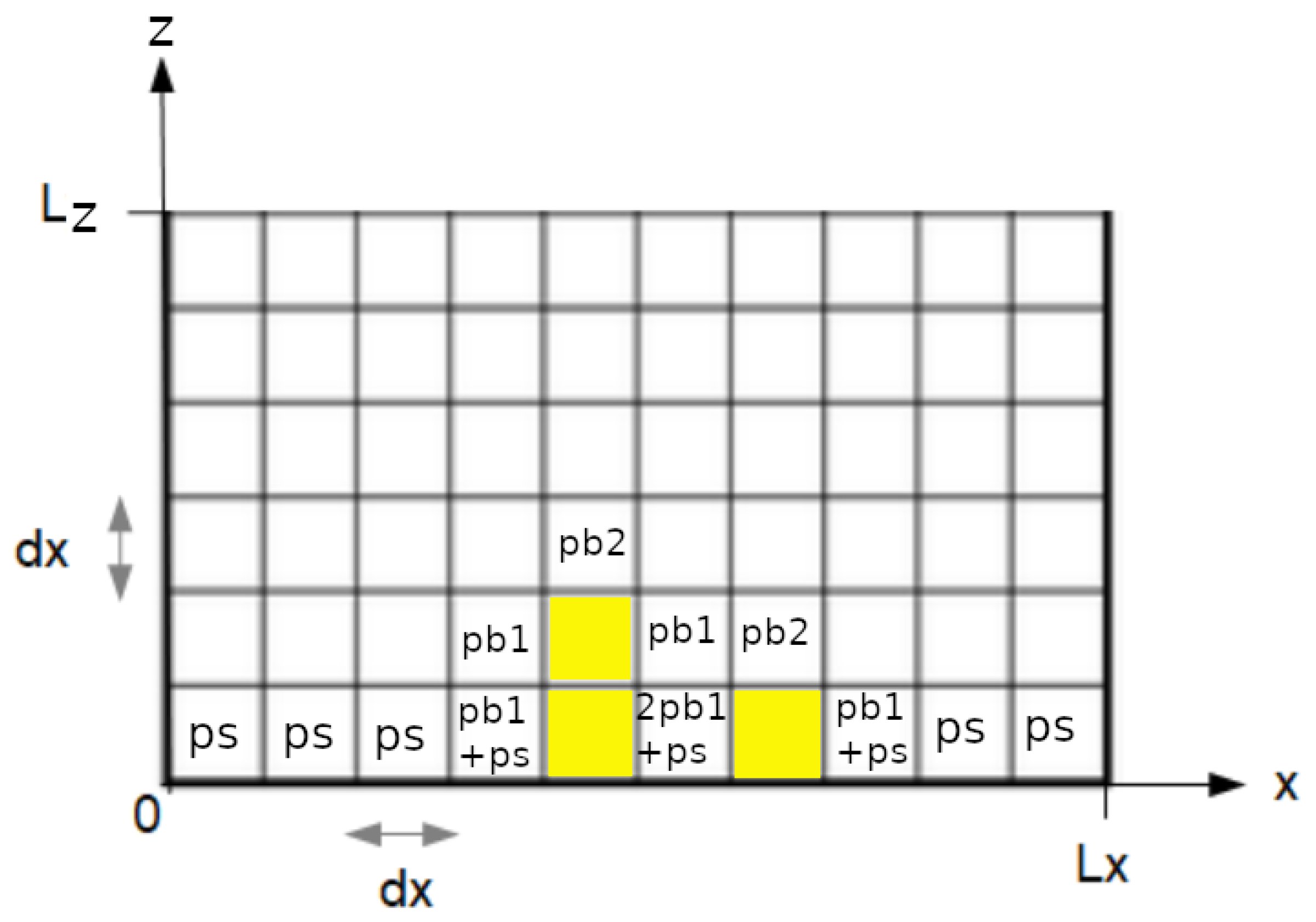
2.1.1. Attachment Algorithm
- : the attachment probability on the surface.
- : the probability of horizontal attachment on the side of an element occupied by bacteria.
- : the probability of vertical attachment below or on the top of an element occupied by bacteria.
| Algorithm 1: Attachment algorithm |
 |
- (i)
- To save computing time, the place of the first bacterium attached to the surface can be chosen randomly without random Bernouilly test. In the same way, to force the number of microcolonies, several places can be initialized with bacteria on the surface.
- (ii)
- Instead of random selection of the row, it is possible to choose the first available place from the bottom in the selected column. This choice leads to more compact microcolonies without holes.
- (iii)
- Instead of running the process until a given number of bacteria are attached, a number (depending on the concentration of the medium) of attachment tests by minute can be chosen. Then, the number of attached bacteria depends on the time of the initialization process.
2.1.2. Model Extensions
- Several species of bacteria. To simulate the attachment of k species of bacteria, more initial data are needed: the required number of attached bacteria, the proportion of the species s, the symmetrical matrix of size giving the probabilities of inter-bacterial attachment ( is the horizontal attachment probability of species s on species r, is for the vertical attachment), and the attachment probability on the substratum for the species s. There is an attachment probability matrix by species and can take on integer values from 0 to k. However, the algorithm is almost the same: the random choice of the species (with the constraint of respecting the given proportions) is added at the beginning of each iteration.
- 3D model of attachment. The domain is a 3D straight block of size and the size of the matrices are adapted: b with size , with size if k species are present. In the algorithm, the choice of the column j is made in a 2D grid instead of a discretized line, the periodic conditions are applied on the four side boundaries, and the update of the attachment probability matrix is a bit more complex because each cube of the mesh has six adjacent elements.
- Non-homogeneous surface. If the attachment surface is made with different materials, it is only necessary to define a value of by material and adapt the initialization of matrix accordingly.
- Non-flat surface. The rectangular domain (or the block) is defined as previously but b and are initialized to indicate the position of the surface: and if the element is filled with the material of the substratum.
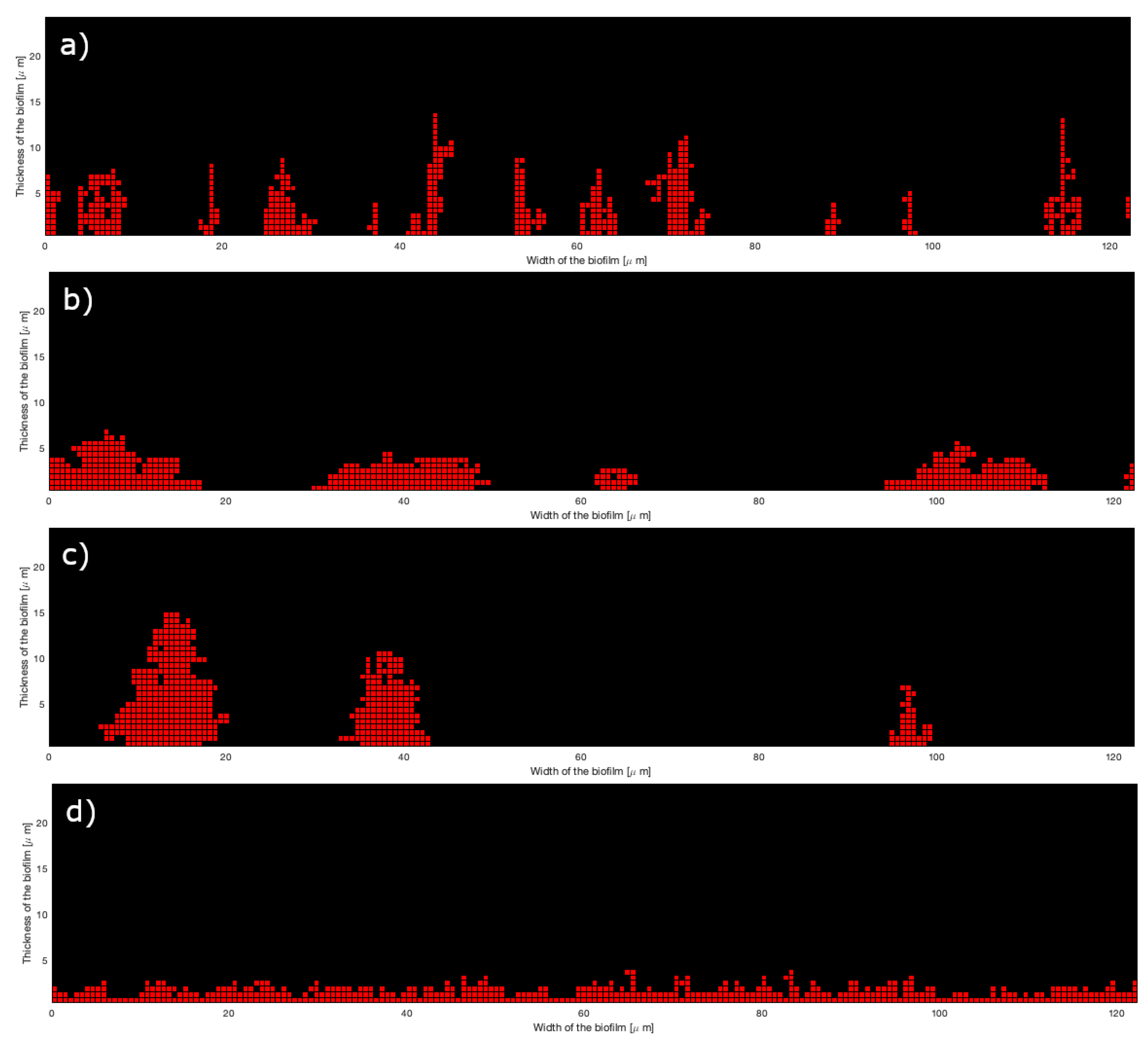
- Non-constant parameters of attachment. Specific shapes can be obtained by varying the value of parameters in time or depending on the number of attached bacteria in the process. For instance, a tall mushroom shape is obtained with a very low horizontal attachment probability replaced by a high value after half of the attachment process.
2.2. Experimental Initial Bacteria Microcolonies
2.2.1. Bacteria and Media
2.2.2. Bacterial Growth Conditions to Assess Initial Attachment
2.2.3. Confocal Laser Microscopy and Imaging
2.2.4. qPCR Quantification
2.2.5. Statistical Analysis
3. Results
3.1. Characteristics of the Attachment Model
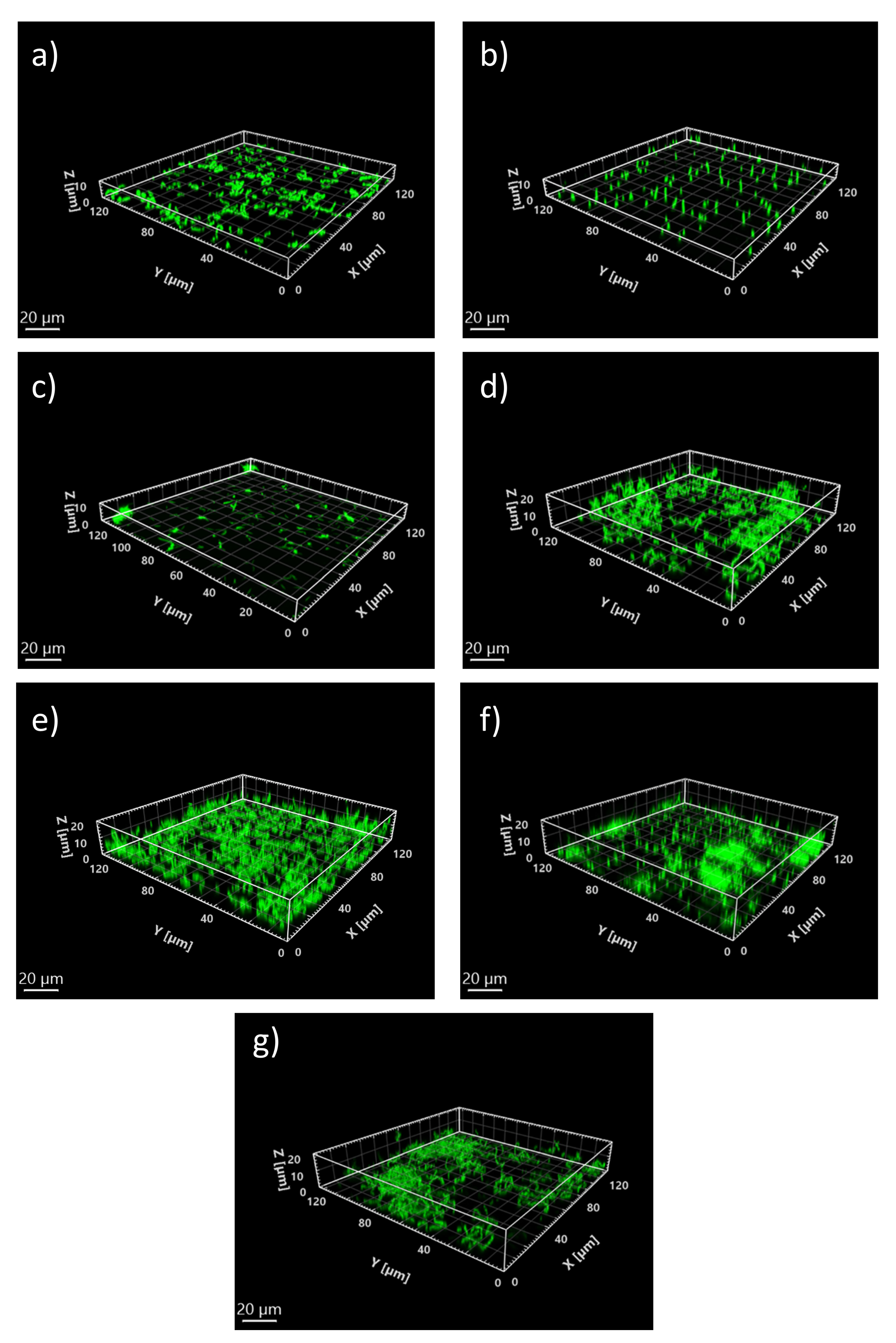
3.2. Biological Characterization of Biofilms Initiation by Oral Bacteria
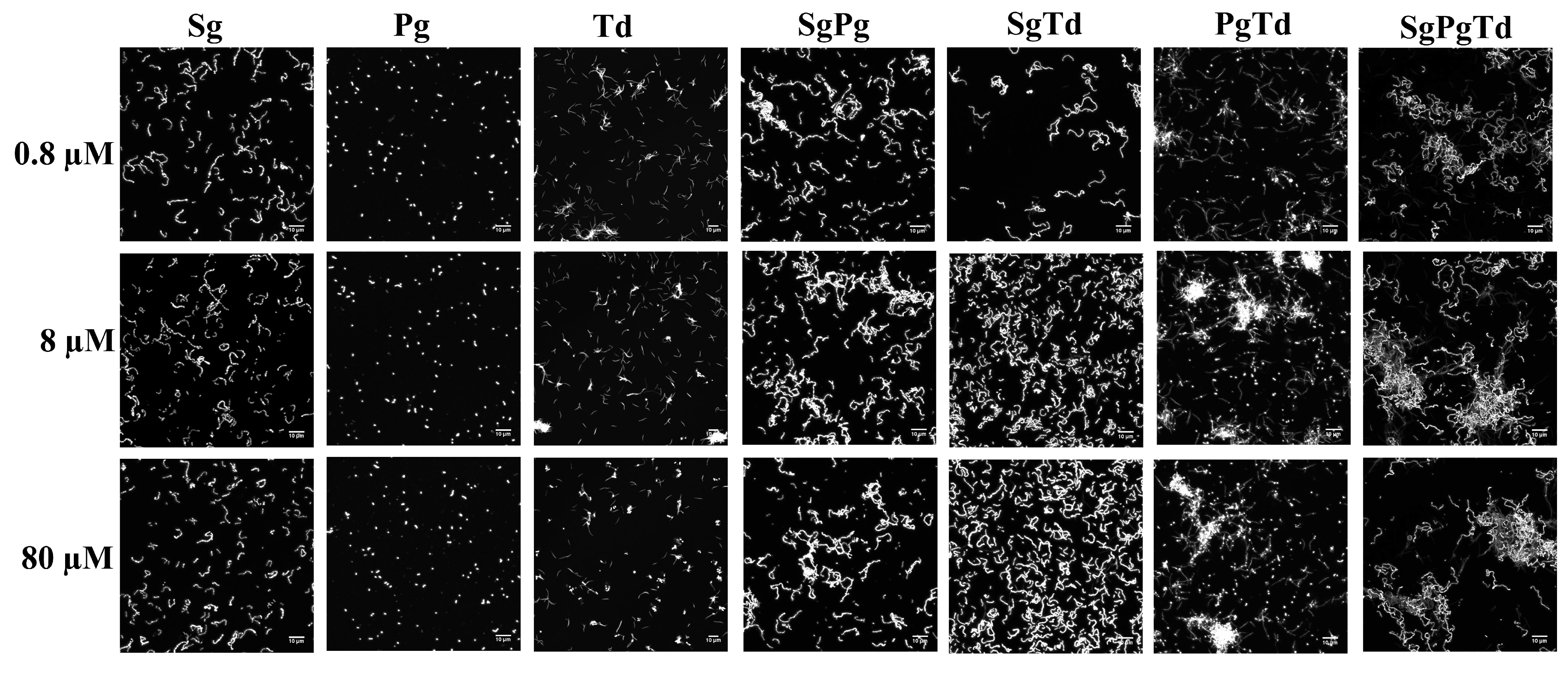
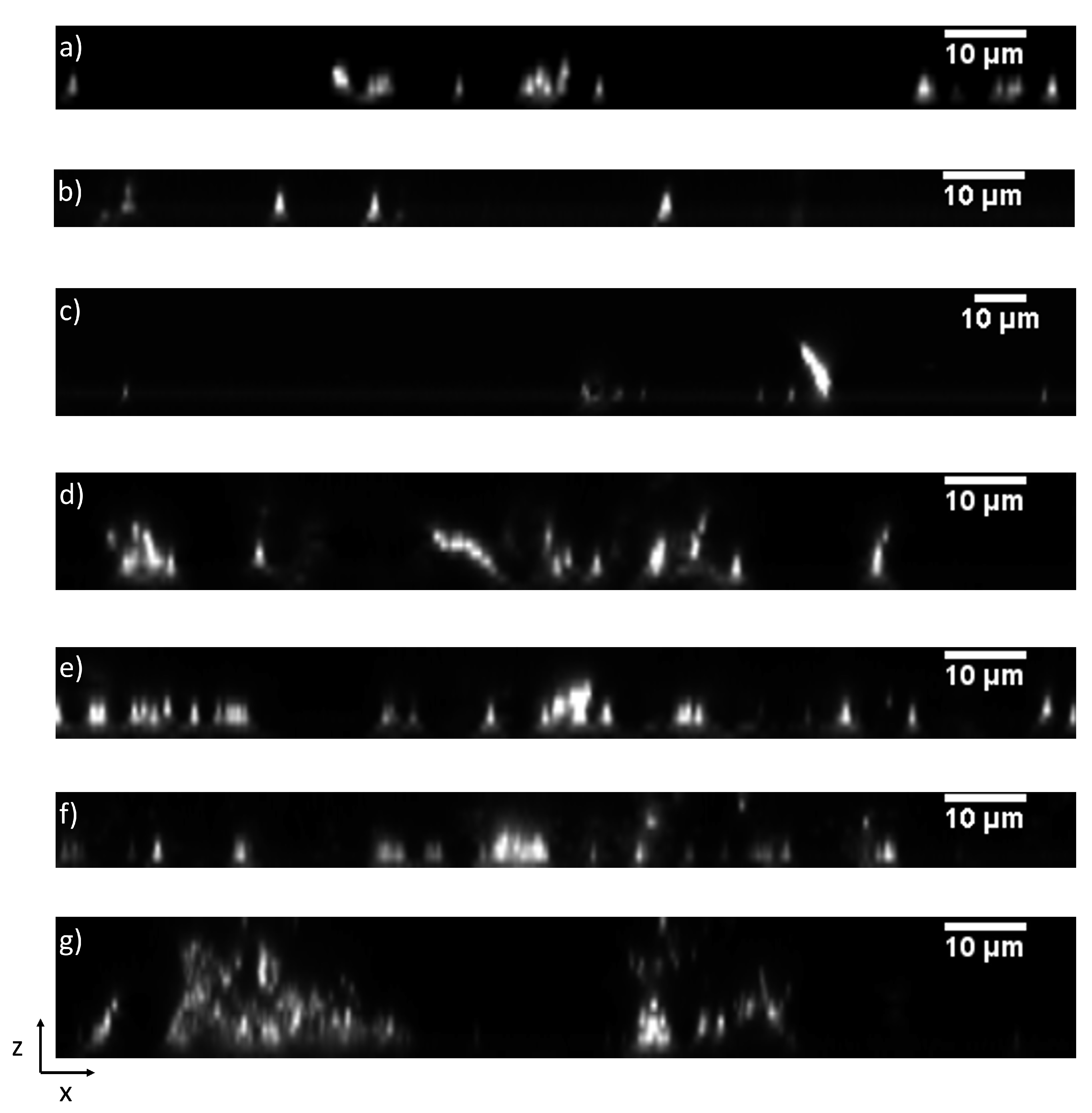
3.2.1. Effect of Iron Levels and Interspecies Associations on Initial Structure
3.2.2. Effect of Iron and Interspecies Association on Attachment of Each Species in Mono and Multi-Species Conditions
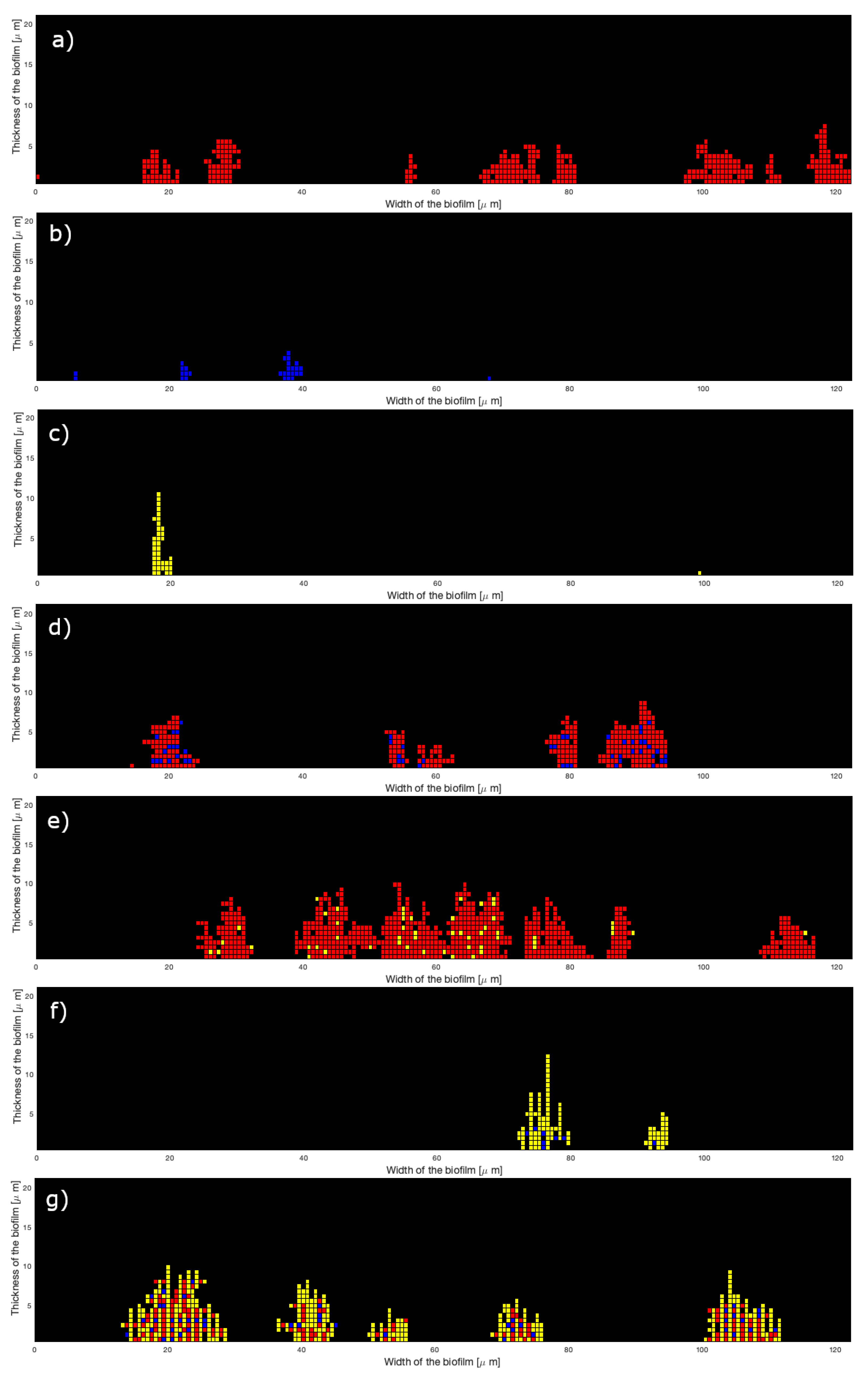
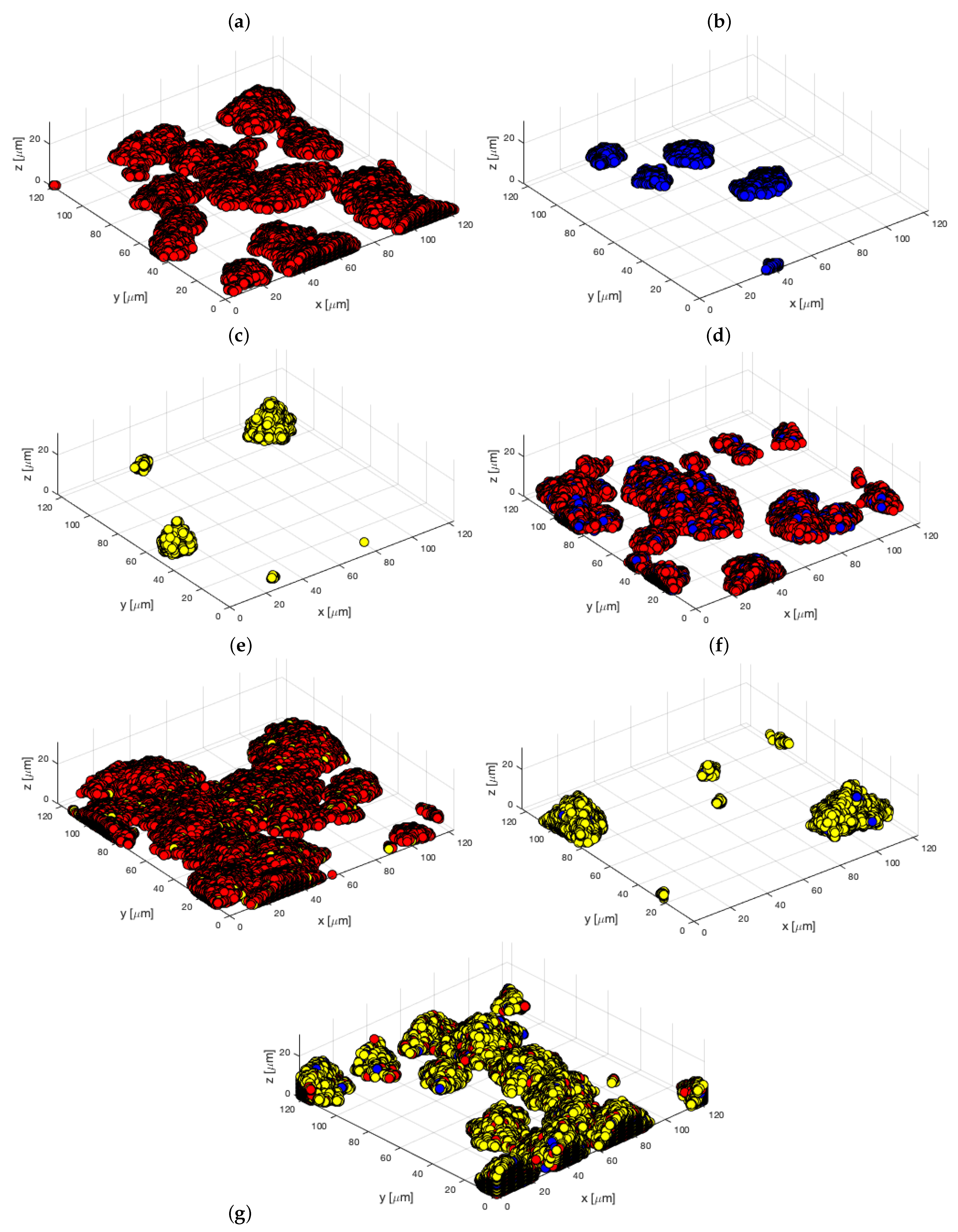
3.3. Simulation of Oral Bacterial Attachment
4. Discussion
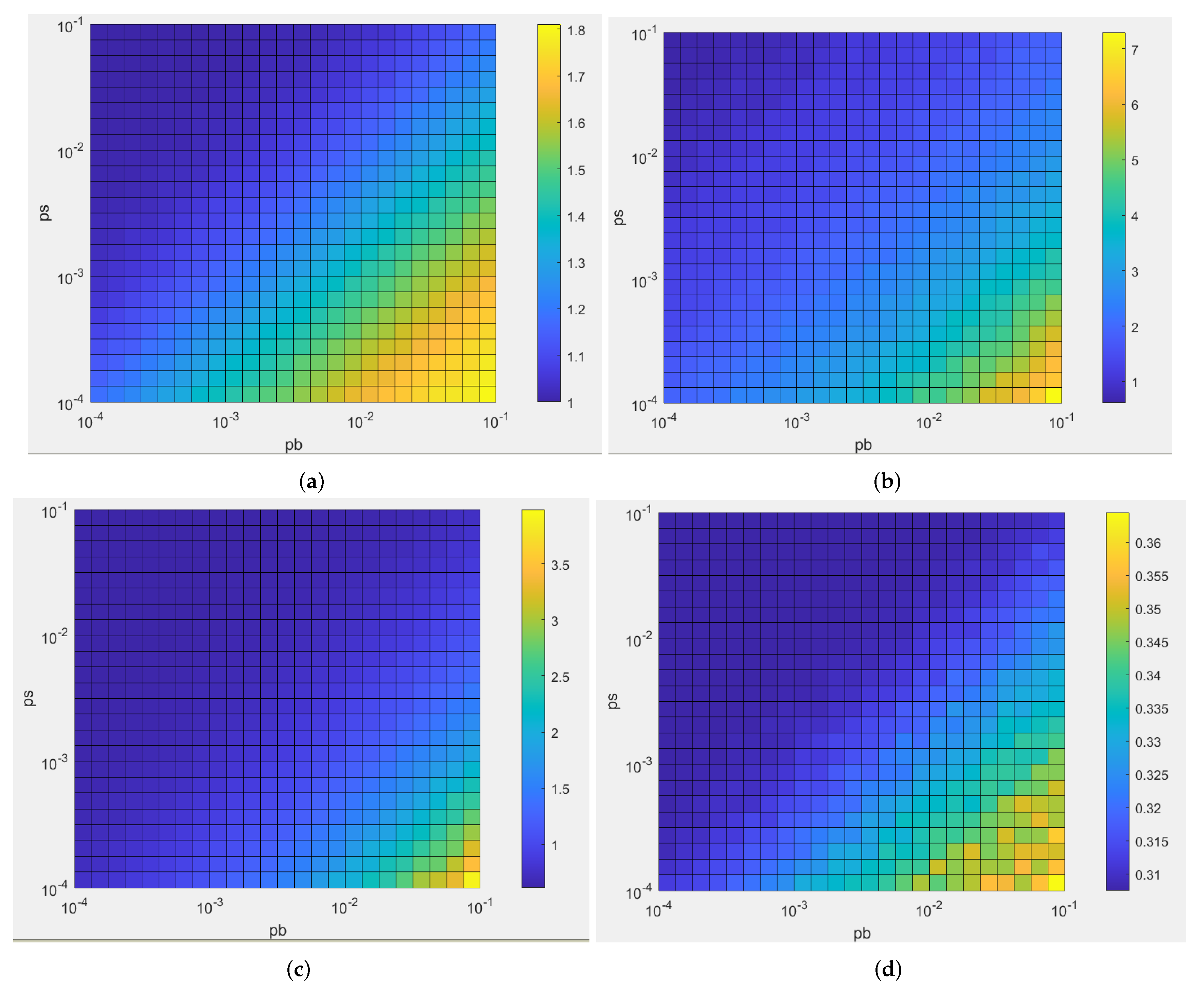
4.1. Dependence of the Characteristics of the Microcolonies on the Algorithm Parameters
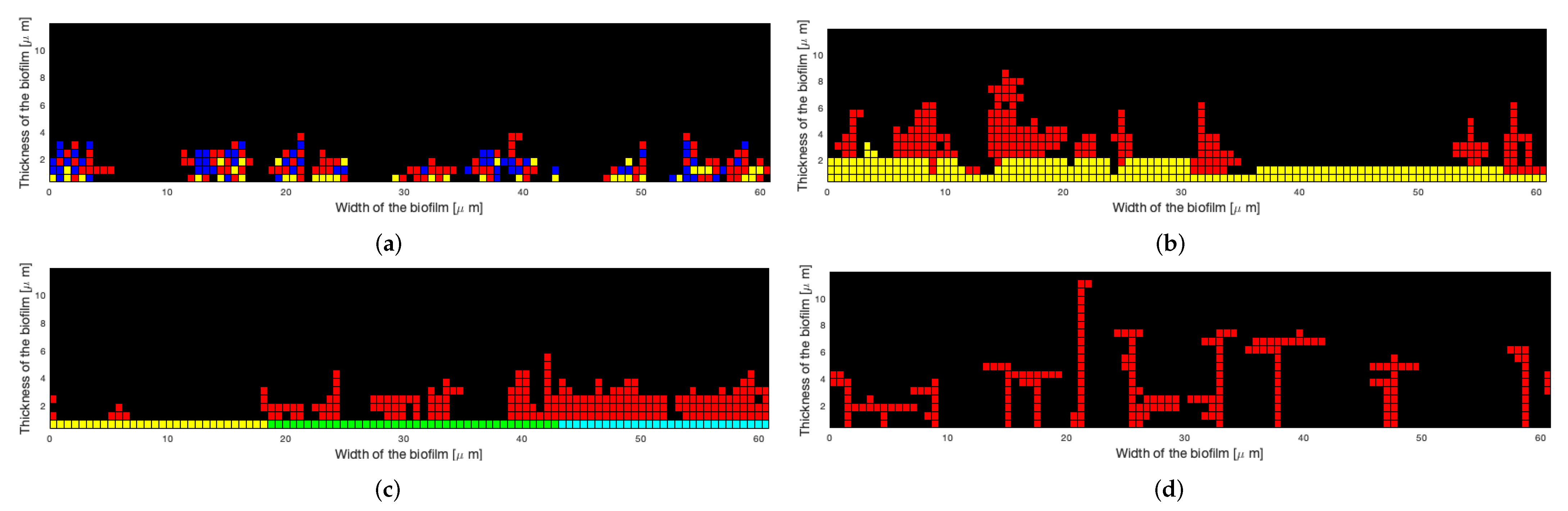
4.2. Ability of the Model to Fit the Experimental Oral Microcolonies
- For both T. denticola-containing inocula (SgTd and PgTd), the lowest errors were obtained by setting horizontal probabilities at the highest value, whereas the vertical probability was 250 times less. This would mean that bacteria attach predominantly next to other bacteria and less on top of them.
- The reverse was observed with SgPg-containing inocula, with a vertical probability higher than the horizontal one for best fitting. Overall, horizontal and vertical probabilities for this type of species interaction are higher, suggesting that S. gordonii and P. gingivalis would attach better together than the other types of species.
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Flemming, H.C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: An emergent form of bacterial life. Nat. Rev. Microbiol. 2016, 14, 563–575. [Google Scholar] [CrossRef]
- Muhammad, M.H.; Idris, A.L.; Fan, X.; Guo, Y.; Yu, Y.; Jin, X.; Qiu, J.; Guan, X.; Huang, T. Beyond Risk: Bacterial Biofilms and Their Regulating Approaches. Front. Microbiol. 2020, 11, 928. [Google Scholar] [CrossRef]
- Feng, Q.; Guo, W.; Wang, T.; Cristina Macias Alvarez, L.; Luo, M.; Ge, R.; Zhou, C.; Zhang, Q.; Luo, J. Iron coupling with carbon fiber to stimulate biofilms formation in aerobic biological film systems for improved decentralized wastewater treatment: Performance, mechanisms and implications. Bioresour. Technol. 2021, 319, 124151. [Google Scholar] [CrossRef]
- Berne, C.; Ellison, C.K.; Ducret, A.; Brun, Y.V. Bacterial adhesion at the single-cell level. Nat. Rev. Microbiol. 2018, 16, 616–627. [Google Scholar] [CrossRef]
- Zheng, S.; Bawazir, M.; Dhall, A.; Kim, H.E.; He, L.; Heo, J.; Hwang, G. Implication of Surface Properties, Bacterial Motility, and Hydrodynamic Conditions on Bacterial Surface Sensing and Their Initial Adhesion. Front. Bioeng. Biotechnol. 2021, 9, 82. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, T. Review of mathematical models for biofilms. Solid State Commun. 2010, 150, 1009–1022. [Google Scholar] [CrossRef]
- Klapper, I.; Dockery, J. Mathematical Description of Microbial Biofilms. SIAM Rev. 2010, 52, 221–265. [Google Scholar] [CrossRef]
- Kreft, J.U.; Picioreanu, C.; Wimpenny, J.W.; van Loosdrecht, M.C. Individual-based modelling of biofilms. Microbiology 2001, 147, 2897–2912. [Google Scholar] [CrossRef]
- Martin, B.; Tamanai-Shacoori, Z.; Bronsard, J.; Ginguené, F.; Meuric, V.; Mahé, F.; Bonnaure-Mallet, M. A new mathematical model of bacterial interactions in two-species oral biofilms. PLoS ONE 2017, 12, e0173153. [Google Scholar] [CrossRef]
- Picioreanu, C.; van Loosdrecht, M.C.; Heijnen, J.J. Mathematical Modeling of Biofilm Structure with a Hybrid Differential-Discrete Cellular Automaton Approach. Biotechnol. Bioeng. 1998, 58, 101–116. [Google Scholar] [CrossRef]
- Wimpenny, J.W.T.; Colasanti, R. A unifying hypothesis for the structure of microbial biofilms based on cellular automaton models. FEMS Microbiol. Ecol. 1997, 22, 1–16. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, H. Modeling multidimensional and multispecies biofilms in porous media. Biotechnol. Bioeng. 2017, 114, 1679–1687. [Google Scholar] [CrossRef]
- Zhang, X.; Zhou, X.; Xi, H.; Sun, J.; Liang, X.; Wei, J.; Xiao, X.; Liu, Z.; Li, S.; Liang, Z.; et al. Interpretation of adhesion behaviors between bacteria and modified basalt fiber by surface thermodynamics and extended DLVO theory. Colloids Surf. B Biointerfaces 2019, 177, 454–461. [Google Scholar] [CrossRef]
- Moraes, J.O.; Cruz, E.A.; Pinheiro, I.; Oliveira, T.C.M.; Alvarenga, V.; Sant’Ana, A.S.; Magnani, M. An ordinal logistic regression approach to predict the variability on biofilm formation stages by five Salmonella Enterica Strains on polypropylene and glass surfaces as affected by pH, temperature and NaCl. Food Microbiol. 2019, 83, 95–103. [Google Scholar] [CrossRef]
- Cerone, A.; Marsili, E. A Formal Model for the Simulation and Analysis of Early Biofilm Formation. In From Data to Models and Back; Bowles, J., Broccia, G., Nanni, M., Eds.; Series Title: Lecture Notes in Computer, Science; Springer International Publishing: Berlin/Heidelberg, Germany, 2021; Volume 12611, pp. 134–151. [Google Scholar] [CrossRef]
- Tamayo, L.; Melo, F.; Caballero, L.; Hamm, E.; Díaz, M.; Leal, M.; Guiliani, N.; Urzúa, M. Does Bacterial Elasticity Affect Adhesion to Polymer Fibers? ACS Appl. Mater. Interfaces 2020, 12, 14507–14517. [Google Scholar] [CrossRef]
- Zammuto, V.; Rizzo, M.G.; Spanò, A.; Spagnuolo, D.; Di Martino, A.; Morabito, M.; Manghisi, A.; Genovese, G.; Guglielmino, S.; Calabrese, G.; et al. Effects of crude polysaccharides from marine macroalgae on the adhesion and biofilm formation of Pseudomonas aeruginosa and Staphylococcus aureus. Algal Res. 2022, 63, 102646. [Google Scholar] [CrossRef]
- Lee, C.K.; Vachier, J.; de Anda, J.; Zhao, K.; Baker, A.E.; Bennett, R.R.; Armbruster, C.R.; Lewis, K.A.; Tarnopol, R.L.; Lomba, C.J.; et al. Social Cooperativity of Bacteria during Reversible Surface Attachment in Young Biofilms: A Quantitative Comparison of Pseudomonas Aeruginosa PA14 PAO1. mBio 2020, 11, e02644-19. [Google Scholar] [CrossRef]
- Rodriguez, D.; Einarsson, B.; Carpio, A. Biofilm growth on rugose surfaces. Phys. Rev. E 2012, 86, 061914. [Google Scholar] [CrossRef]
- O’Toole, G.; Kaplan, H.B.; Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 2000, 54, 49–79. [Google Scholar] [CrossRef]
- Kolenbrander, P.E.; Andersen, R.N.; Blehert, D.S.; Egland, P.G.; Foster, J.S.; Palmer, R.J. Communication among Oral Bacteria. Microbiol. Mol. Biol. Rev. 2002, 66, 486–505. [Google Scholar] [CrossRef]
- Sbordone, L.; Bortolaia, C. Oral microbial biofilms and plaque-related diseases: Microbial communities and their role in the shift from oral health to disease. Clin. Oral Investig. 2003, 7, 181–188. [Google Scholar] [CrossRef]
- Hojo, K.; Nagaoka, S.; Ohshima, T.; Maeda, N. Bacterial Interactions in Dental Biofilm Development. J. Dent. Res. 2009, 88, 982–990. [Google Scholar] [CrossRef]
- Byrne, S.J.; Dashper, S.G.; Darby, I.B.; Adams, G.G.; Hoffmann, B.; Reynolds, E.C. Progression of chronic periodontitis can be predicted by the levels of Porphyromonas gingivalis and Treponema denticola in Subgingival Plaque. Oral Microbiol. Immunol. 2009, 24, 469–477. [Google Scholar] [CrossRef]
- Boyer, E.; Le Gall-David, S.; Martin, B.; Fong, S.B.; Loréal, O.; Deugnier, Y.; Bonnaure-Mallet, M.; Meuric, V. Increased transferrin saturation is associated with subgingival microbiota dysbiosis and severe periodontitis in genetic haemochromatosis. Sci. Rep. 2018, 8, 15532. [Google Scholar] [CrossRef]
- Chathoth, K.N. Effect of Environmental Signals on the Development of Oral Biofilm. Ph.D. Thesis, University Rennes 1, Rennes, France, 2021. [Google Scholar]
- Chathoth, K.; Martin, B.; Cornelis, P.; Yvenou, S.; Bonnaure-Mallet, M.; Baysse, C. The events that may contribute to subgingival dysbiosis: A focus on the interplay between iron, sulfide and oxygen. FEMS Microbiol. Lett. 2020, 367, fnaa100. [Google Scholar] [CrossRef]
- Chen, C.C.; Teng, L.J.; Chang, T.C. Identification of Clinically Relevant Viridans Group Streptococci Sequence Analysis 16S-23S Ribosomal DNA Spacer Region. J. Clin. Microbiol. 2004, 42, 2651–2657. [Google Scholar] [CrossRef]
- Watanabe, T.; Maruyama, F.; Nozawa, T.; Aoki, A.; Okano, S.; Shibata, Y.; Oshima, K.; Kurokawa, K.; Hattori, M.; Nakagawa, I.; et al. Complete genome sequence of the bacterium Porphyromonas gingivalis TDC60, Which Causes Periodontal Disease. J. Bacteriol. 2011, 193, 4259–4260. [Google Scholar] [CrossRef]
- Chan, E.C.; Siboo, R.; Keng, T.; Psarra, N.; Hurley, R.; Cheng, S.L.; Iugovaz, I. Treponema denticola (ex Brumpt 1925) Sp. Nov., Nom. Rev., Identif. New Spirochete Isol. Periodontal Pockets. Int. J. Syst. Bacteriol. 1993, 43, 196–203. [Google Scholar] [CrossRef]
- Martin, B.; Chathoth, K.; Ouali, S.; Meuric, V.; Bonnaure-Mallet, M.; Baysse, C. New growth media for oral bacteria. J. Microbiol. Methods 2018, 153, 10–13. [Google Scholar] [CrossRef]
- Leschine, S.B.; Canale-Parola, E. Rifampin as a selective agent for isolation of oral spirochetes. J. Clin. Microbiol. 1980, 12, 792–795. [Google Scholar] [CrossRef]
- Heydorn, A.; Nielsen, A.T.; Hentzer, M.; Sternberg, C.; Givskov, M.; Ersboll, B.K.; Molin, S. Quantification of biofilm structures by the novel computer program COMSTAT. Microbiology 2000, 146, 2395–2407. [Google Scholar] [CrossRef]
- Ammann, T.W.; Bostanci, N.; Belibasakis, G.N.; Thurnheer, T. Validation of a quantitative real-time PCR assay and comparison with fluorescence microscopy and selective agar plate counting for species-specific quantification of an in vitro subgingival biofilm model. J. Periodontal Res. 2013, 48, 517–526. [Google Scholar] [CrossRef]



| Bacteria | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sg | 365 | 0.10 | 0.10 | 0.27 | 0.01 | 0.03 | 0.08 | 0.16 | |
| Pg | 29 | 0.06 | 0.10 | 0.31 | 0.01 | 0 | 0.03 | 0.21 | |
| Td | 99 | 0.01 | 0.10 | 0.02 | 0.02 | 0 | 0.05 | 0.12 | |
| SgPg | 240 | 0.05 | 0.20 | 0.28 | 0.02 | 0.01 | 0.02 | 0.18 | |
| SgTd | 126 | 0.001 | 0.25 | 0.12 | 0 | 0.07 | 0.40 | 0.45 | |
| PgTd | 75 | 0.25 | 0.001 | 0.28 | 0.01 | 0.03 | 0.35 | 0.02 | |
| SgPgTd | 500 | 0.23 | 0.06 | 0.22 | 0.70 | 0.37 |
| Bacteria | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sg | 360 | 0.10 | 0.10 | 0.28 | 0 | 0.02 | 0.03 | 0 | |
| Pg | 28 | 0.06 | 0.10 | 0.34 | 0 | 0 | 0.04 | 0.29 | |
| Td | 39 | 0.01 | 0.10 | 0 | 0.03 | 0 | 0.02 | 0.30 | |
| SgPg | 340 | 0.05 | 0.20 | 0.18 | 0.02 | 0 | 0 | 0.20 | |
| SgTd | 800 | 0.25 | 0.001 | 0.21 | 0.05 | 0.06 | 0.19 | 0.05 | |
| PgTd | 110 | 0.25 | 0.001 | 0.31 | 0.01 | 0.08 | 1.00 | 0.34 | |
| SgPgTd | 560 | 0.03 | 0.02 | 0.01 | 0.05 | 0.27 |
| Bacteria | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sg | 315 | 0.10 | 0.10 | 0.31 | 0 | 0.02 | 0.01 | 0.02 | |
| Pg | 29 | 0.06 | 0.10 | 0.36 | 0.02 | 0 | 0.08 | 0.24 | |
| Td | 16 | 0.01 | 0.10 | 0 | 0 | 0 | 0.17 | 0.31 | |
| SgPg | 400 | 0.001 | 0.25 | 0.11 | 0.02 | 0.04 | 0.05 | 0.13 | |
| SgTd | 970 | 0.25 | 0.001 | 0.18 | 0.06 | 0.07 | 0.20 | 0.02 | |
| PgTd | 110 | 0.25 | 0.001 | 0.28 | 0.01 | 0.09 | 1.43 | 0.57 | |
| SgPgTd | 545 | 0 | 0.01 | 0.03 | 0 | 0.21 |
| Bacteria | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sg | 78,000 | 0.20 | 0.03 | 0.22 | 0.01 | 0.08 | 0.10 | 0.12 | |
| Pg | 5900 | 0.20 | 0.03 | 0.30 | 0.01 | 0 | 0.05 | 0.07 | |
| Td | 7300 | 0.08 | 0.10 | 0.06 | 0.02 | 0 | 0.09 | 0.06 | |
| SgPg | 70,000 | 0.05 | 0.20 | 0.16 | 0 | 0 | 0.04 | 0.07 | |
| SgTd | 166,000 | 0.2 | 0.02 | 0.18 | 0.01 | 0.03 | 0.01 | 0.02 | |
| PgTd | 24,000 | 0.25 | 0.001 | 0.25 | 0.01 | 0.08 | 0.90 | 0.90 | |
| SgPgTd | 114,000 | 0.01 | 0.02 | 0.01 | 0.06 | 0.06 |
| Iron Concentration and Model | Bacteria | |||
|---|---|---|---|---|
| 0.8 M 2D model | Sg | 1 | ||
| Pg | ||||
| Td | ||||
| 8 M 2D model | Sg | 1 | ||
| Pg | ||||
| Td | ||||
| 8 M 3D model | Sg | |||
| Pg | ||||
| Td | ||||
| 80 M 2D model | Sg | 1 | ||
| Pg | ||||
| Td |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chathoth, K.; Fostier, L.; Martin, B.; Baysse, C.; Mahé, F. A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms. Microorganisms 2022, 10, 686. https://doi.org/10.3390/microorganisms10040686
Chathoth K, Fostier L, Martin B, Baysse C, Mahé F. A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms. Microorganisms. 2022; 10(4):686. https://doi.org/10.3390/microorganisms10040686
Chicago/Turabian StyleChathoth, Kanchana, Louis Fostier, Bénédicte Martin, Christine Baysse, and Fabrice Mahé. 2022. "A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms" Microorganisms 10, no. 4: 686. https://doi.org/10.3390/microorganisms10040686
APA StyleChathoth, K., Fostier, L., Martin, B., Baysse, C., & Mahé, F. (2022). A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms. Microorganisms, 10(4), 686. https://doi.org/10.3390/microorganisms10040686






