Establishment of the Bacterial Microbiota in a Lab-Reared Model Teleost Fish, the Medaka Oryzias latipes
Abstract
1. Introduction
2. Materials and Methods
2.1. Production of Life Stages
2.2. DNA Extraction and Sequencing
2.3. Sequence Analysis
3. Results
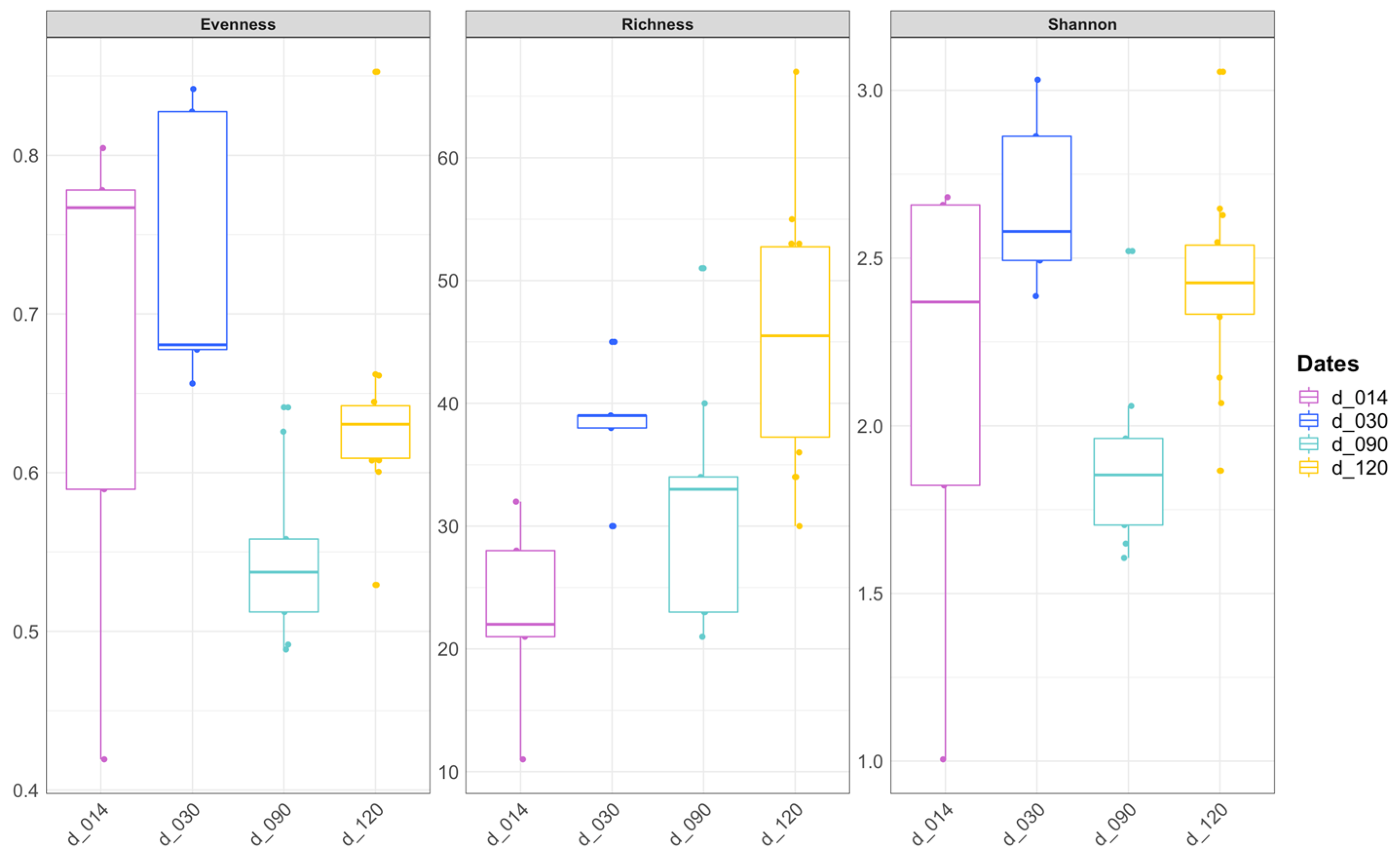
3.1. Diversity of Bacterial Communities
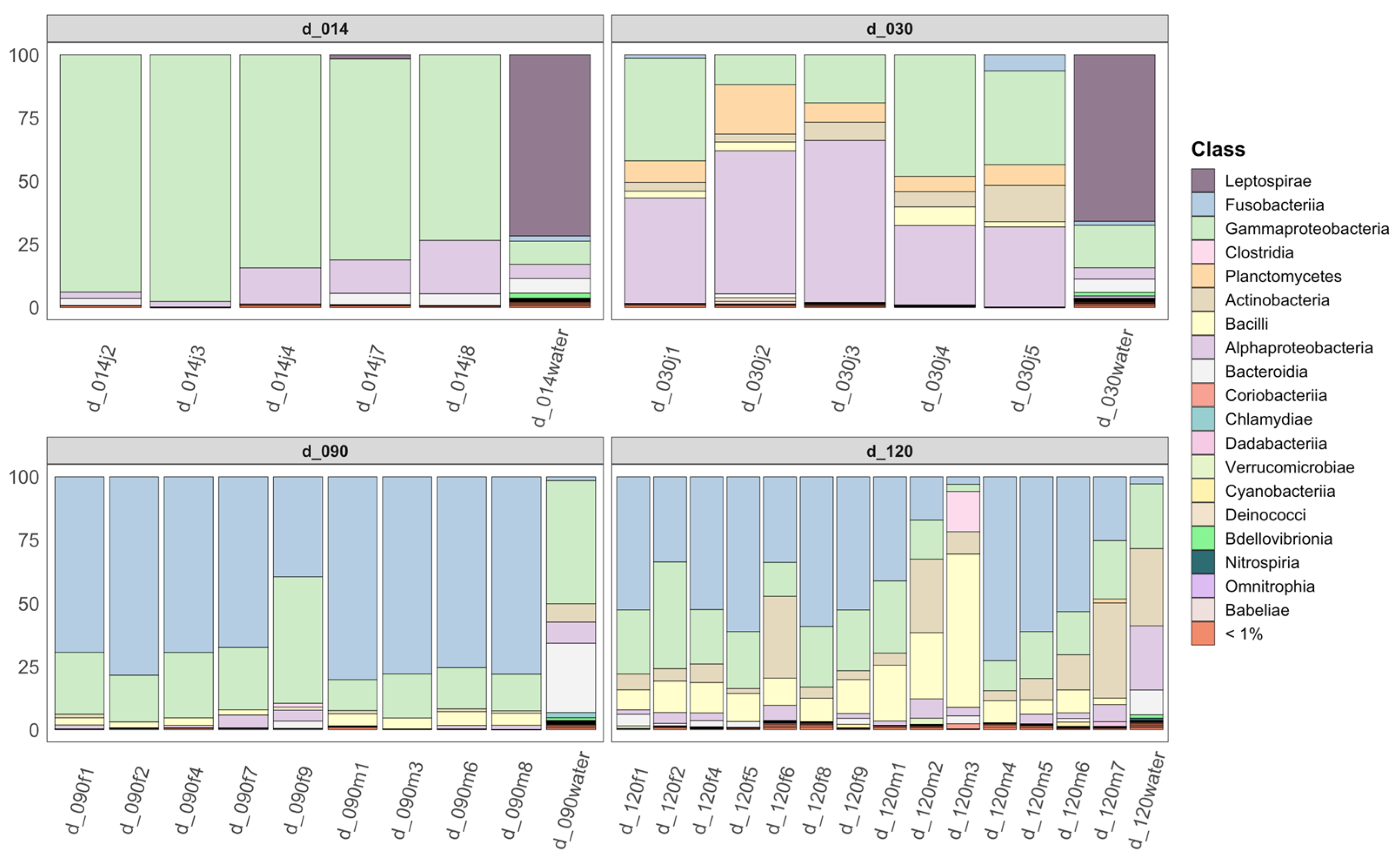
3.2. Community Composition as a Function of Life Stage
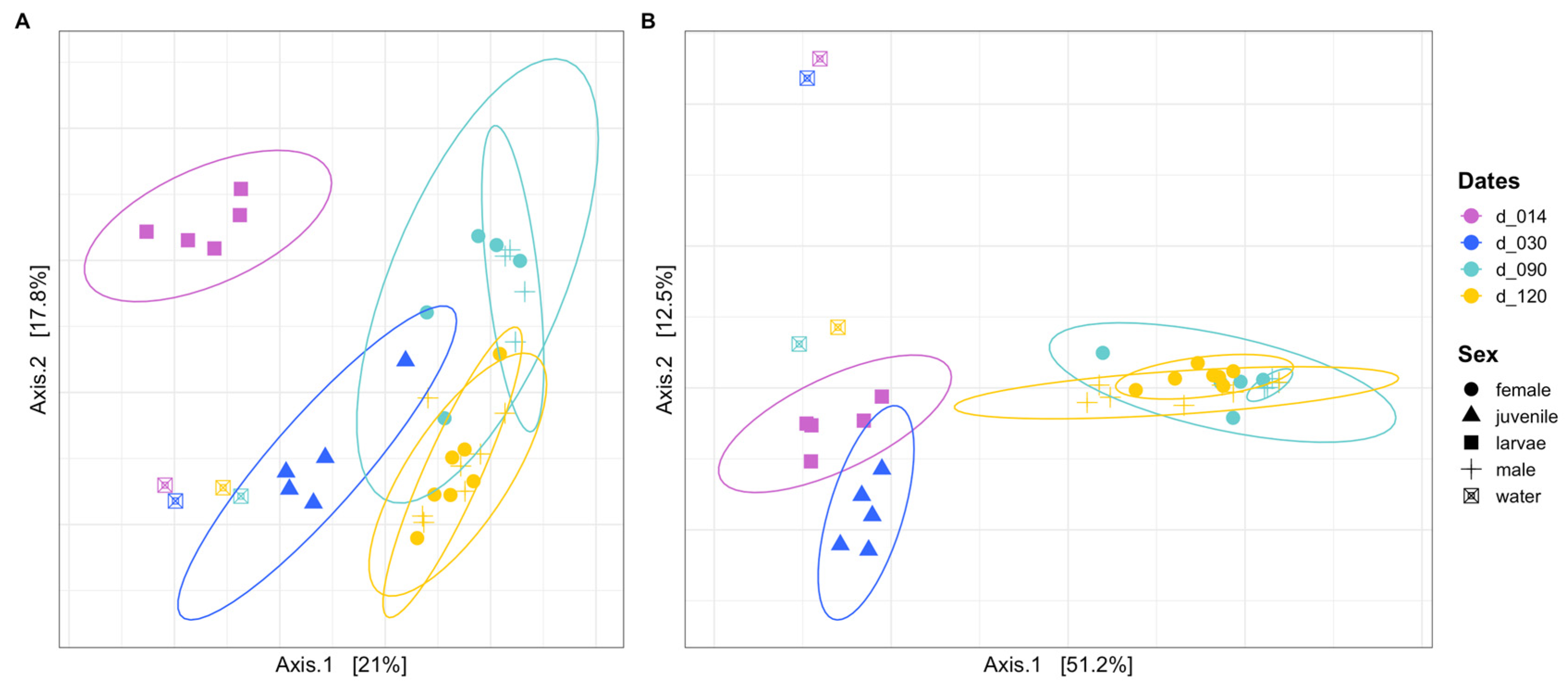
3.3. Community Structure and Dominant ASVs
4. Discussion
4.1. Bacteria Become Detectable Only after Mouth Opening
4.2. Community Shifts Occur throughout Development
4.3. No Evidence for Differences between Males and Females
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- McFall-Ngai, M.; Hadfield, M.G.; Bosch, T.C.G.; Carey, H.V.; Domazet-Loso, T.; Douglas, A.E.; Dubilier, N.; Eberl, G.; Fukami, T.; Gilbert, S.F.; et al. Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl. Acad. Sci. USA 2013, 110, 3229–3236. [Google Scholar] [CrossRef]
- Cryan, J.F.; Dinan, T.G. Talking about a microbiome revolution. Nat. Microbiol. 2019, 4, 552–553. [Google Scholar] [CrossRef] [PubMed]
- Adamovsky, O.; Buerger, A.N.; Wormington, A.M.; Ector, N.; Griffitt, R.J.; Bisesi, J.H.; Martyniuk, C.J. The gut microbiome and aquatic toxicology: An emerging concept for environmental health. Environ. Toxicol. Chem. 2018, 37, 2758–2775. [Google Scholar] [CrossRef] [PubMed]
- Duperron, S.; Halary, S.; Gallet, A.; Marie, B. Microbiome-Aware Ecotoxicology of Organisms: Relevance, Pitfalls, and Challenges. Front. Public Health 2020, 8, 407. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Wu, S.; Zeng, Z.; Fu, Z. Effects of environmental pollutants on gut microbiota. Environ. Pollut. 2017, 222, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeld, C.S. Gut Dysbiosis in Animals Due to Environmental Chemical Exposures. Front. Cell. Infect. Microbiol. 2017, 7, 396. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Zhao, A.; Xie, G.; Chi, Y.; Zhao, L.; Li, H.; Wang, C.; Bao, Y.; Jia, W.; Luther, M.; et al. Melamine-induced renal toxicity is mediated by the gut microbiota. Sci. Transl. Med. 2013, 5, 172ra22. [Google Scholar] [CrossRef]
- Claus, S.P.; Guillou, H.; Ellero-Simatos, S. The gut microbiota: A major player in the toxicity of environmental pollutants? NPJ Biofilms Microbiomes 2016, 2, 16003. [Google Scholar] [CrossRef] [PubMed]
- Wittbrodt, J.; Shima, A.; Schartl, M. Medaka—A model organism from the far East. Nat. Rev. Genet. 2002, 3, 53–64. [Google Scholar] [CrossRef]
- Qiao, Q.; Le Manach, S.; Huet, H.; Duvernois-Berthet, E.; Chaouch, S.; Duval, C.; Sotton, B.; Ponger, L.; Marie, A.; Mathéron, L.; et al. An integrated omic analysis of hepatic alteration in medaka fish chronically exposed to cyanotoxins with possible mechanisms of reproductive toxicity. Environ. Pollut. 2016, 219, 119–131. [Google Scholar] [CrossRef]
- Le Manach, S.; Sotton, B.; Huet, H.; Duval, C.; Paris, A.; Marie, A.; Yépremian, C.; Catherine, A.; Mathéron, L.; Vinh, J.; et al. Physiological effects caused by microcystin-producing and non-microcystin producing Microcystis aeruginosa on medaka fish: A proteomic and metabolomic study on liver. Environ. Pollut. 2018, 234, 523–537. [Google Scholar] [CrossRef]
- Yoon, J.B.; Hwang, S.; Yang, J.H.; Lee, S.; Bang, W.Y.; Moon, K.H. Dynamics of the Gut Microbiome and Transcriptome in Korea Native Ricefish (Oryzias latipes) during Chronic Antibiotic Exposure. Genes 2022, 13, 1243. [Google Scholar] [CrossRef]
- Kim, B.-M.; Kim, J.; Choi, I.-Y.; Raisuddin, S.; Au, D.W.T.; Leung, K.M.Y.; Wu, R.S.S.; Rhee, J.-S.; Lee, J.-S. Omics of the marine medaka (Oryzias melastigma) and its relevance to marine environmental research. Mar. Environ. Res. 2016, 113, 141–152. [Google Scholar] [CrossRef]
- Kang, H.-M.; Byeon, E.; Jeong, H.; Kim, M.-S.; Chen, Q.; Lee, J.-S. Different effects of nano- and microplastics on oxidative status and gut microbiota in the marine medaka Oryzias melastigma. J. Hazard. Mater. 2021, 405, 124207. [Google Scholar] [CrossRef] [PubMed]
- Ye, R.R.; Peterson, D.R.; Seemann, F.; Kitamura, S.-I.; Lee, J.S.; Lau, T.C.K.; Tsui, S.K.W.; Au, D.W.T. Immune competence assessment in marine medaka (Orzyias melastigma)-a holistic approach for immunotoxicology. Environ. Sci. Pollut. Res. Int. 2017, 24, 27687–27701. [Google Scholar] [CrossRef]
- Duperron, S.; Halary, S.; Habiballah, M.; Gallet, A.; Huet, H.; Duval, C.; Bernard, C.; Marie, B. Response of Fish Gut Microbiota to Toxin-Containing Cyanobacterial Extracts: A Microcosm Study on the Medaka (Oryzias latipes). Environ. Sci. Technol. Lett. 2019, 6, 341–347. [Google Scholar] [CrossRef]
- Gallet, A.; Halary, S.; Duval, C.; Huet, H.; Duperron, S.; Marie, B. Disruption of fish gut microbiota composition and holobiont’s metabolome by cyanobacterial blooms. bioRxiv 2021. [CrossRef]
- Foucault, P.; Gallet, A.; Duval, C.; Marie, B.; Duperron, S. Gut microbiota and holobiont metabolome composition of the Medaka fish (Oryzias latipes) are affected by a short exposure to the cyanobacterium Microcystis aeruginosa. Aquat. Toxicol. 2022, 253, 106329. [Google Scholar] [CrossRef]
- Hird, S.M. Evolutionary Biology Needs Wild Microbiomes. Front. Microbiol. 2017, 8, 725. [Google Scholar] [CrossRef]
- Alessandri, G.; Milani, C.; Mancabelli, L.; Mangifesta, M.; Lugli, G.A.; Viappiani, A.; Duranti, S.; Turroni, F.; Ossiprandi, M.C.; van Sinderen, D.; et al. The impact of human-facilitated selection on the gut microbiota of domesticated mammals. FEMS Microbiol. Ecol. 2019, 95, fiz121. [Google Scholar] [CrossRef] [PubMed]
- Stephens, W.Z.; Burns, A.R.; Stagaman, K.; Wong, S.; Rawls, J.F.; Guillemin, K.; Bohannan, B.J.M. The composition of the zebrafish intestinal microbial community varies across development. ISME J. 2016, 10, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; de Bruijn, I.; Jack, A.L.; Drynan, K.; van den Berg, A.H.; Thoen, E.; Sandoval-Sierra, V.; Skaar, I.; van West, P.; Diéguez-Uribeondo, J.; et al. Deciphering microbial landscapes of fish eggs to mitigate emerging diseases. ISME J. 2014, 8, 2002–2014. [Google Scholar] [CrossRef] [PubMed]
- Iwamatsu, T. Stages of normal development in the medaka Oryzias latipes. Mech. Dev. 2004, 121, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-H.; Kim, C.-C.; Koh, S.-J.; Shin, L.-S.; Cho, J.-K.; Han, K.-H. Egg Development and Morphology of Larva and Juvenile of the Oryzias latipes. Dev. Reprod. 2014, 18, 173–178. [Google Scholar] [CrossRef]
- Parada, A.E.; Needham, D.M.; Fuhrman, J.A. Every base matters: Assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ. Microbiol. 2016, 18, 1403–1414. [Google Scholar] [CrossRef]
- Fadeev, E.; Cardozo-Mino, M.G.; Rapp, J.Z.; Bienhold, C.; Salter, I.; Salman-Carvalho, V.; Molari, M.; Tegetmeyer, H.E.; Buttigieg, P.L.; Boetius, A. Comparison of Two 16S rRNA Primers (V3–V4 and V4–V5) for Studies of Arctic Microbial Communities. Front. Microbiol. 2021, 12, 637526. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Weiss, S.; Xu, Z.Z.; Peddada, S.; Amir, A.; Bittinger, K.; Gonzalez, A.; Lozupone, C.; Zaneveld, J.R.; Vázquez-Baeza, Y.; Birmingham, A.; et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 2017, 5, 27. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P. Vegan: Community Ecology Package; R Package Version 2.5–7. 2020; R Core Team: Vienna, Austria, 2022. [Google Scholar]
- Arbizu, P.M. pairwiseAdonis; R Core Team: Vienna, Austria, 2022. [Google Scholar]
- Bordenstein, S.R.; Theis, K.R. Host Biology in Light of the Microbiome: Ten Principles of Holobionts and Hologenomes. PLoS Biol. 2015, 13, e1002226. [Google Scholar] [CrossRef]
- Roeselers, G.; Mittge, E.K.; Stephens, W.Z.; Parichy, D.M.; Cavanaugh, C.M.; Guillemin, K.; Rawls, J.F. Evidence for a core gut microbiota in the zebrafish. ISME J. 2011, 5, 1595–1608. [Google Scholar] [CrossRef]
- Nyholm, L.; Odriozola, I.; Bideguren, G.M.; Aizpurua, O.; Alberdi, A. Gut microbiota differences between paired intestinal wall and digesta samples in three small species of fish. PeerJ 2022, 10, e12992. [Google Scholar] [CrossRef]
- Wilkins, L.G.E.; Rogivue, A.; Schütz, F.; Fumagalli, L.; Wedekind, C. Increased diversity of egg-associated bacteria on brown trout (Salmo trutta) at elevated temperatures. Sci. Rep. 2015, 5, 17084. [Google Scholar] [CrossRef]
- Bai, S.; Hou, G. Microbial communities on fish eggs from Acanthopagrus schlegelii and Halichoeres nigrescens at the XuWen coral reef in the Gulf of Tonkin. PeerJ 2020, 8, e8517. [Google Scholar] [CrossRef] [PubMed]
- Najafpour, B.; Pinto, P.I.S.; Moutou, K.A.; Canario, A.V.M.; Power, D.M. Factors Driving Bacterial Microbiota of Eggs from Commercial Hatcheries of European Seabass and Gilthead Seabream. Microorganisms 2021, 9, 2275. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.-H.; Wu, Z.-H.; Jian, J.-C.; Lu, Y.-S.; Tang, J.-F. Characterization of pathogenic Aeromonas veronii bv. veronii associated with ulcerative syndrome from chinese longsnout catfish (Leiocassis longirostris Günther). Braz. J. Microbiol. 2012, 43, 382–388. [Google Scholar] [CrossRef]
- Scheifler, M.; Sanchez-Brosseau, S.; Magnanou, E.; Desdevises, Y. Diversity and structure of sparids external microbiota (Teleostei) and its link with monogenean ectoparasites. Anim. Microbiome 2022, 4, 27. [Google Scholar] [CrossRef] [PubMed]
- Escalas, A.; Troussellier, M.; Yuan, T.; Bouvier, T.; Bouvier, C.; Mouchet, M.A.; Hernandez, D.F.; Miranda, J.R.; Zhou, J.; Mouillot, D. Functional diversity and redundancy across fish gut, sediment and water bacterial communities. Environ. Microbiol. 2017, 19, 3268–3282. [Google Scholar] [CrossRef]
- Tsuchiya, C.; Sakata, T.; Sugita, H. Novel ecological niche of Cetobacterium somerae, an anaerobic bacterium in the intestinal tracts of freshwater fish. Lett. Appl. Microbiol. 2008, 46, 43–48. [Google Scholar] [CrossRef]
- Llewellyn, M.S.; Boutin, S.; Hoseinifar, S.H.; Derome, N. Teleost microbiomes: The state of the art in their characterization, manipulation and importance in aquaculture and fisheries. Front. Microbiol. 2014, 5, 207. [Google Scholar] [CrossRef] [PubMed]
- Egerton, S.; Culloty, S.; Whooley, J.; Stanton, C.; Ross, R.P. The Gut Microbiota of Marine Fish. Front. Microbiol. 2018, 9, 873. [Google Scholar] [CrossRef] [PubMed]
- Bolnick, D.I.; Snowberg, L.K.; Hirsch, P.E.; Lauber, C.L.; Org, E.; Parks, B.; Lusis, A.J.; Knight, R.; Caporaso, J.G.; Svanbäck, R. Individual diet has sex-dependent effects on vertebrate gut microbiota. Nat. Commun. 2014, 5, 4500. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Rangel-Zúñiga, O.A.; Alcalá-Díaz, J.F.; Gómez-Delgado, F.; Pérez-Martínez, P.; Delgado-Lista, J.; Quintana-Navarro, G.M.; Landa, B.B.; Navas-Cortés, J.A.; Tena-Sempere, M.; et al. Intestinal Microbiota Is Influenced by Gender and Body Mass Index. PLoS ONE 2016, 11, e0154090. [Google Scholar] [CrossRef]
- Ma, Y.; Song, L.; Lei, Y.; Jia, P.; Lu, C.; Wu, J.; Xi, C.; Strauss, P.R.; Pei, D.-S. Sex dependent effects of silver nanoparticles on the zebrafish gut microbiota. Environ. Sci. Nano 2018, 5, 740–751. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duval, C.; Marie, B.; Foucault, P.; Duperron, S. Establishment of the Bacterial Microbiota in a Lab-Reared Model Teleost Fish, the Medaka Oryzias latipes. Microorganisms 2022, 10, 2280. https://doi.org/10.3390/microorganisms10112280
Duval C, Marie B, Foucault P, Duperron S. Establishment of the Bacterial Microbiota in a Lab-Reared Model Teleost Fish, the Medaka Oryzias latipes. Microorganisms. 2022; 10(11):2280. https://doi.org/10.3390/microorganisms10112280
Chicago/Turabian StyleDuval, Charlotte, Benjamin Marie, Pierre Foucault, and Sébastien Duperron. 2022. "Establishment of the Bacterial Microbiota in a Lab-Reared Model Teleost Fish, the Medaka Oryzias latipes" Microorganisms 10, no. 11: 2280. https://doi.org/10.3390/microorganisms10112280
APA StyleDuval, C., Marie, B., Foucault, P., & Duperron, S. (2022). Establishment of the Bacterial Microbiota in a Lab-Reared Model Teleost Fish, the Medaka Oryzias latipes. Microorganisms, 10(11), 2280. https://doi.org/10.3390/microorganisms10112280





