Endophytic Bacteria Associated with Origanum heracleoticum L. (Lamiaceae) Seeds
Abstract
1. Introduction
2. Materials and Methods
2.1. Light Microscopy and Transmission Electron Microscopy
2.2. Cultivable Endophytic Bacteria Isolation and Growth Conditions
2.3. Random Amplified Polymorphic DNA (RAPD) Analysis
2.4. 16S rRNA Gene Amplification and Sequencing and ARDRA Analysis
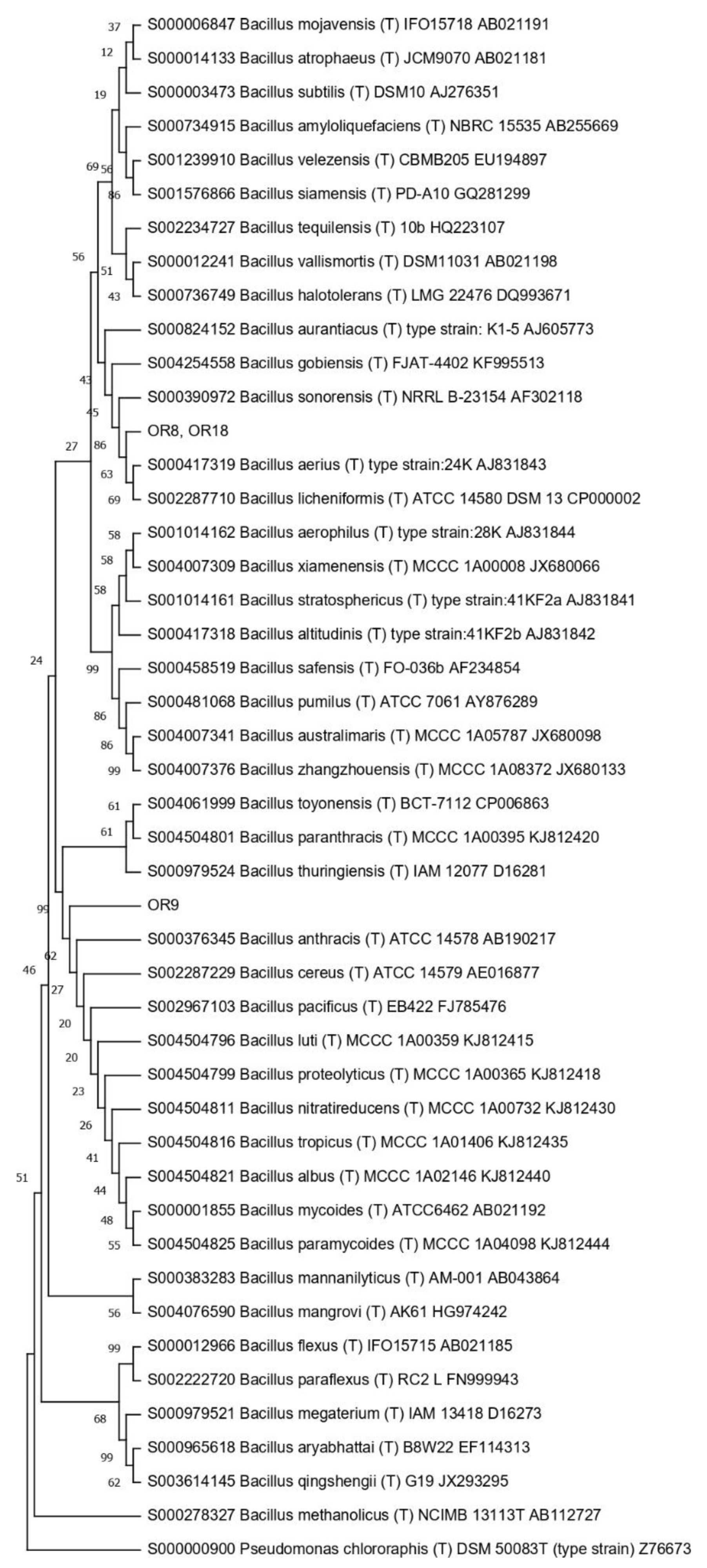
2.5. Phylogenetic Tree Construction
2.6. Antibiotic Resistance Tests
2.7. Cross-Streaking Test against Burkholderia cepacia Complex Strains
2.8. Genome Sequencing
2.9. Genome Assembly and Bioinformatic Analysis
3. Results
3.1. Electron Microscopy
3.2. Cultivable Endophytic Bacteria Isolation
3.3. Analysis of the Structure of the Seed-Associated Endophytic Community
3.4. Taxonomic Affiliation of the Endophytic Strains
3.5. Antibiotic Resistance Profiles
3.6. Cross-Streaking Test against Burkholderia cepacia Complex Strains
3.7. Genome Sequencing of Bacillus sp. OR9
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Compant, S.; Cambon, M.C.; Vacher, C.; Mitter, B.; Samad, A.; Sessitsch, A. The plant endosphere world—Bacterial life within plants. Environ. Microbiol. 2007, 23, 1812–1829. [Google Scholar] [CrossRef] [PubMed]
- Anyasi, R.O.; Atagana, H.I. Endophyte: Understanding the microbes and its applications. Pak. J. Biol. Sci. 2019, 22, 154–167. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.R.; Van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The Hidden World within Plants: Ecological and Evolutionary Considerations for Defining Functioning of Microbial Endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef] [PubMed]
- Compant, S.; Samad, A.; Faist, H.; Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 2019, 19, 29–37. [Google Scholar] [CrossRef]
- Taulé, C.; Vaz-Jauri, P.; Battistoni, F. Insights into the early stages of plant–endophytic bacteria interaction. World J. Microbiol. Biotechnol. 2021, 37, 13. [Google Scholar] [CrossRef]
- Nelson, E.B. The seed microbiome: Origins, interactions, and impacts. Plant Soil 2018, 422, 7–34. [Google Scholar] [CrossRef]
- Truyens, S.; Weyens, N.; Cuypers, A.; Vangronsveld, J. Bacterial seed endophytes: Genera, vertical transmission and interaction with plants. Environ. Microbiol. Rep. 2015, 7, 40–50. [Google Scholar] [CrossRef]
- Mano, H.; Tanaka, F.; Watanabe, A.; Kaga, H.; Okunishi, S.; Morisaki, H. Culturable Surface and Endophytic Bacterial Flora of the Maturing Seeds of Rice Plants (Oryza sativa) Cultivated in a Paddy Field. Microbes Environ. 2006, 21, 86–100. [Google Scholar] [CrossRef]
- Shade, A.; Jacques, M.A.; Barret, M. Ecological patterns of seed microbiome diversity, transmission, and assembly. Curr. Opin. Microbiol. 2017, 37, 15–22. [Google Scholar] [CrossRef]
- Escobar Rodríguez, C.; Antonielli, L.; Mitter, B.; Trognitz, F.; Sessitsch, A. Heritability and functional importance of the Setaria viridis bacterial seed microbiome. Phytobiomes J. 2020, 4, 40–52. [Google Scholar] [CrossRef]
- Khalaf, E.M.; Raizada, M.N. Taxonomic and functional diversity of cultured seed associated microbes of the cucurbit family. BMC Microbiol. 2016, 16, 131. [Google Scholar] [CrossRef]
- Khalaf, E.M.; Raizada, M.N. Bacterial seed endophytes of domesticated cucurbits antagonize fungal and oomycete pathogens including powdery mildew. Front. Microbiol. 2018, 9, 42. [Google Scholar] [CrossRef]
- Hardoim, P.R.; Hardoim, C.C.P.; Van Overbeek, L.S.; Van Elsas, J.D. Dynamics of seed-borne rice endophytes on early plant growth stages. PLoS ONE 2012, 7, e30438. [Google Scholar] [CrossRef]
- Johnston-Monje, D.; Raizada, M.N. Conservation and diversity of seed associated endophytes in Zea across boundaries of evolution, ethnography and ecology. PLoS ONE 2011, 6, e20396. [Google Scholar] [CrossRef]
- Mitter, B.; Pfaffenbichler, N.; Flavell, R.; Compant, S.; Antonielli, L.; Petric, A.; Berninger, T.; Naveed, M.; Sheibani-Tezerji, R.; von Maltzahn, G.; et al. A new approach to modify plant microbiomes and traits by introducing beneficial bacteria at flowering into progeny seeds. Front. Microbiol. 2017, 8, 11. [Google Scholar] [CrossRef]
- Alvin, A.; Miller, K.I.; Neilan, B.A. Exploring the potential of endophytes from medicinal plants as sources of antimycobacterial compounds. Microbiol. Res. 2014, 169, 483–495. [Google Scholar] [CrossRef]
- Del Giudice, L.; Massardo, D.R.; Pontieri, P.; Bertea, C.M.; Mombello, D.; Carata, E.; Tredici, S.M.; Talà, A.; Mucciarelli, M.; Groudeva, V.I.; et al. The microbial community of Vetiver root and its involvement into essential oil biogenesis. Environ. Microbiol. 2008, 10, 2824–2841. [Google Scholar] [CrossRef]
- Gouda, S.; Das, G.; Sen, S.K.; Shin, H.S.; Patra, J.K. Endophytes: A treasure house of bioactive compounds of medicinal importance. Front. Microbiol. 2018, 7, 1538. [Google Scholar] [CrossRef]
- Ruan, Q.; Patel, G.; Wang, J.; Luo, E.; Zhou, W.; Sieniawska, E.; Hao, X.; Kai, G. Current advances of endophytes as a platform for production of anti-cancer drug camptothecin. FCT 2021, 151, 112113. [Google Scholar] [CrossRef]
- Köberl, M.; Schmidt, R.; Ramadan, E.M.; Bauer, R.; Berg, G. The microbiome of medicinal plants: Diversity and importance for plant growth, quality, and health. Front. Microbiol. 2013, 4, 400. [Google Scholar] [CrossRef]
- Lombrea, A.; Antal, D.; Ardelean, F.; Avram, S.; Pavel, I.Z.; Vlaia, L.; Mut, A.M.; Diaconeasa, Z.; Dehelean, C.A.; Soica, C.; et al. A recent insight regarding the phytochemistry and bioactivity of Origanum vulgare L. Essential oil. Int. J. Mol. Sci. 2020, 21, 9653. [Google Scholar] [CrossRef]
- Chen, H.; Wu, H.; Yan, B.; Zhao, H.; Liu, F.; Zhang, H.; Sheng, Q.; Miao, F.; Liang, Z. Core microbiome of medicinal plant salvia miltiorrhiza seed: A rich reservoir of beneficial microbes for secondary metabolism? Int. J. Mol. Sci. 2018, 19, 672. [Google Scholar] [CrossRef]
- Govaerts, R.; Paton, A.; Harvey, Y.; Navarro, T.; Del Rosario García Peña, M. World Checklist of Lamiaceae; The Royal Botanic Gardens: Kew, UK, 2020; Available online: http://wcsp.science.kew.org (accessed on 26 September 2022).
- Napoli, E.; Giovino, A.; Carrubba, A.; How Yuen Siong, V.; Rinoldo, C.; Nina, O.; Ruberto, G. Variations of essential oil constituents in oregano (Origanum vulgare subsp. viridulum (O. heracleoticum) over cultivation cycles. Plants 2020, 9, 1174. [Google Scholar] [CrossRef]
- Spurr, A.R. A Low-Viscosity Epoxy Resin Embedding Medium for Electron Microscopy. J. Ultrastruct. Res. 1969, 26, 31–43. [Google Scholar] [CrossRef]
- Faddetta, T.; Abbate, L.; Alibrandi, P.; Arancio, W.; Siino, D.; Strati, F.; De Filippo, C.; Fatta del Bosco, S.; Carimi, F.; Puglia, A.M.; et al. The endophytic microbiota of Citrus limon is transmitted from seed to shoot highlighting differences of bacterial and fungal community structures. Sci. Rep. 2021, 11, 7078. [Google Scholar] [CrossRef]
- Castronovo, L.M.; Calonico, C.; Ascrizzi, R.; Del Duca, S.; Delfino, V.; Chioccioli, S.; Vassallo, A.; Strozza, I.; De Leo, M.; Biffi, S.; et al. The Cultivable Bacterial Microbiota Associated to the Medicinal Plant Origanum vulgare L.: From Antibiotic Resistance to Growth-Inhibitory Properties. Front. Microbiol. 2020, 11, 862. [Google Scholar] [CrossRef]
- Vaneechoutte, M.; Dijkshoorn, L.; Tjernberg, I.; Elaichouni, A.; De Vos, P.; Claeys, G.; Verschragen, G. Identification of Acinetobacter Genomic Species by Amplified Ribosomal DNA Restriction Analysis. J. Clin. Microbiol. 1995, 33, 11–15. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Cole, J.R.; Wang, Q.; Fish, J.A.; Chai, B.; McGarrell, D.M.; Sun, Y.; Brown, C.T.; Porras-Alfaro, A.; Kuske, C.R.; Tiedje, J.M. Ribosomal Database Project: Data and tools for high throughput rRNA analysis. Nucleic Acids Res. 2014, 42, 633–642. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Cardinale, M.; Viola, M.; Miceli, E.; Faddetta, T.; Puglia, A.M.; Maggini, V.; Tani, C.; Firenzuoli, F.; Schiff, S.; Bogani, P.; et al. The cypsela (achene) of Echinacea purpurea as a diffusion unit of a community of microorganisms. Appl. Microbiol. Biotechnol. 2021, 105, 2951–2965. [Google Scholar] [CrossRef] [PubMed]
- Mengoni, A.; Maida, I.; Chiellini, C.; Emiliani, G.; Mocali, S.; Fabiani, A.; Fondi, M.; Firenzuoli, F.; Fani, R. Antibiotic resistance differentiates Echinacea purpurea endophytic bacterial communities with respect to plant organs. Res. Microbiol. 2014, 165, 686–694. [Google Scholar] [CrossRef] [PubMed]
- Chiellini, C.; Maida, I.; Maggini, V.; Bosi, E.; Mocali, S.; Emiliani, G.; Perrin, E.; Firenzuoli, F.; Mangoni, A.; Fani, R. Preliminary data on antibacterial activity of Echinacea purpurea-associated bacterial communities against Burkholderia cepacia complex strains, opportunistic pathogens of Cystic Fibrosis patients. Microbiol. Res. 2017, 196, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Tabacchioni, S.; Ferri, L.; Manno, G.; Mentasti, M.; Cocchi, P.; Campana, S.; Ravenni, N.; Taccetti, G.; Dalmastri, C.; Chiarini, L.; et al. Use of the gyrB gene to discriminate among species of the Burkholderia cepacia complex. FEMS Microbiol. Lett. 2008, 281, 175–182. [Google Scholar] [CrossRef][Green Version]
- Maida, I.; Chiellini, C.; Mengoni, A.; Bosi, E.; Firenzuoli, F.; Fondi, M.; Fani, R. Antagonistic interactions between endophytic cultivable bacterial communities isolated from the medicinal plant Echinacea purpurea. Environ. Microbiol. 2016, 18, 2357–2365. [Google Scholar] [CrossRef]
- Semenzato, G.; Alonso- Vázquez, T.; Del Duca, S.; Vassallo, A.; Riccardi, C.; Zaccaroni, M.; Mucci, N.; Padula, A.; Emiliani, G.; Palumbo Piccionello, A.; et al. Genomic Analysis of Endophytic Bacillus-Related Strains Isolated from the Medicinal Plant Origanum vulgare L. Revealed the Presence of Metabolic Pathways Involved in the Biosynthesis of Bioactive Compounds. Microorganisms 2022, 10, 919. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35.37. [Google Scholar] [CrossRef]
- Chiellini, C.; Miceli, E.; Bacci, G.; Fagorzi, C.; Coppini, E.; Fibbi, D.; Bianconi, G.; Mengoni, A.; Canganella, F.; Fani, R. Spatial structuring of bacterial communities in epilithic biofilms in the Acquarossa river (Italy). FEMS Microbiol. Ecol. 2018, 94, fiy181. [Google Scholar] [CrossRef]
- Di Cello, F.; Fani, R. A molecular stategy for the study of natural bacterial communities by PCR-based techniques. Minerva Biotecnol. 1996, 8, 126–134. [Google Scholar]
- Grifoni, A.; Bazzicalupo, M.; Di Serio, C.; Fancelli, S.; Fani, R. Identification of Azospirillum Strains By Restriction Fragment Length Polymorphism of the 16S rDNA and of the Histidine Operon. FEMS Microbiol. Lett. 1995, 127, 85–91. [Google Scholar] [CrossRef]
- Schüßler, A.; Kluge, M. Geosiphon pyriforme, an Endocytosymbiosis Between Fungus and Cyanobacteria, and its Meaning as a Model System for Arbuscular Mycorrhizal Research. In The Mycota IX; Hock, B., Ed.; Springer: Berlin/Heidelberg, Germany, 2001; pp. 151–161. [Google Scholar]
- Falsini, S.; Tani, C.; Sambuco, G.; Papini, A.; Faraoni, P.; Campigli, S.; Ghelardini, L.; Bleve, G.; Rizzo, D.; Ricciolini, M.; et al. Anatomical and biochemical studies of Spartium junceum infected by Xylella fastidiosa subsp. multiplex ST 87. Protoplasma 2022, 259, 103–115. [Google Scholar] [CrossRef]
- Yamauchi, K.E.; Snel, J. Transmission electron microscopic demonstration of phagocytosis and intracellular processing of segmented filamentous bacteria by intestinal epithelial cells of the chick ileum. Infect. Immun. 2000, 68, 6496–6504. [Google Scholar] [CrossRef]
- Esposito-Polesi, N.P.; De Abreu-Tarazi, M.F.; De Almeida, C.V.; Tsai, S.M.; de Almeida, M. Investigation of Endophytic Bacterial Community in Supposedly Axenic Cultures of Pineapple and Orchids with Evidence on Abundant Intracellular Bacteria. Curr. Microbiol. 2017, 74, 103–113. [Google Scholar] [CrossRef]
- Compant, S.; Mitter, B.; Colli-Mull, J.G.; Gangl, H.; Sessitsch, A. Endophytes of Grapevine Flowers, Berries, and Seeds: Identification of Cultivable Bacteria, Comparison with Other Plant Parts, and Visualization of Niches of Colonization. Microb. Ecol. 2011, 62, 188–197. [Google Scholar] [CrossRef]
- Granér, G.; Persson, P.; Meijer, J.; Alström, S. A study on microbial diversity in different cultivars of Brassica napus in relation to its wilt pathogen, Verticillium longisporum. FEMS Microbiol. Lett. 2003, 224, 269–276. [Google Scholar] [CrossRef]
- Chiellini, C.; Maida, I.; Emiliani, G.; Mengoni, A.; Mocali, S.; Fabiani, A.; Biffi, S.; Maggini, V.; Gori, L.; Vannacci, A.; et al. Endophytic and rhizospheric bacterial communities isolated from the medicinal plants echinacea purpurea and echinacea angustifolia. Int. Microbiol. 2014, 17, 165–174. [Google Scholar] [CrossRef]
- Emiliani, G.; Mengoni, A.; Maida, I.; Perrini, E.; Chiellini, C.; Fondi, M.; Gallo, E.; Gori, L.; Maggini, V.; Vannacci, A.; et al. Linking bacterial endophytic communities to essential oils: Clues from Lavandula angustifolia Mill. Evid. Based Complement. Altern. Med. 2014, 2014, 650905. [Google Scholar] [CrossRef]
- Liu, Y.; Zuo, S.; Zou, Y.; Wang, J.; Song, W. Investigation on diversity and population succession dynamics of indigenous bacteria of the maize spermosphere. World J. Microbiol. Biotechnol. 2012, 28, 391–396. [Google Scholar] [CrossRef]
- Maggini, V.; Miceli, E.; Fagorzi, C.; Maida, I.; Fondi, M.; Perrin, E.; Mengoni, A.; Bogani, P.; Chiellini, C.; Mocali, S.; et al. Antagonism and antibiotic resistance drive a species-specific plant microbiota differentiation in Echinacea spp. FEMS Microbiol. Ecol. 2018, 94, fiy118. [Google Scholar] [CrossRef] [PubMed]
- Keswani, C.; Prakash, O.; Bharti, N.; Vílchez, I.; Sansinenea, E.; Lally, R.D.; Borriss, R.; Singh, S.P.; Gupta, V.K.; Fraceto, L.F.; et al. Re-addressing the biosafety issues of plant growth promoting rhizobacteria. Sci. Total Environ. 2019, 690, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Polito, G.; Semenzato, G.; Del Duca, S.; Castronovo, L.M.; Vassallo, A.; Chioccioli, S.; Borsetti, D.; Calabretta, V.; Puglia, A.M.; Fani, R.; et al. Endophytic Bacteria and Essential Oil from Origanum vulgare ssp. vulgare Share Some VOCs with an Antibacterial Activity. Microorganisms 2022, 10, 1424. [Google Scholar] [CrossRef] [PubMed]
- Cendrowski, S.; MacArthur, W.; Hanna, P. Bacillus anthracis requires siderophore biosynthesis for growth in macrophages and mouse virulence. Mol. Microbiol. 2004, 51, 407–417. [Google Scholar] [CrossRef]
- Koppisch, A.T.; Dhungana, S.; Hill, K.K.; Boukhalfa, H.; Heine, H.S.; Colip, L.A.; Romero, R.B.; Shou, Y.; Ticknor, L.O.; Marrone, B.L.; et al. Petrobactin is produced by both pathogenic and non-pathogenic isolates of the Bacillus cereus group of bacteria. BioMetals 2008, 21, 581–589. [Google Scholar] [CrossRef]
- Adeleke, B.S.; Ayangbenro, A.S.; Babalola, O.O. Genomic analysis of endophytic bacillus cereus T4S and its plant growth-promoting traits. Plants 2021, 10, 1776. [Google Scholar] [CrossRef]
- Liu, Y.; Ding, S.; Shen, J.; Zhu, K. Nonribosomal antibacterial peptides that target multidrug-resistant bacteria. Nat. Prod. Rep. 2019, 36, 573–592. [Google Scholar] [CrossRef]
- Hudson, G.A.; Mitchell, D.A. RiPP antibiotics: Biosynthesis and engineering potential. Curr. Opin. Microbiol. 2018, 45, 61–69. [Google Scholar] [CrossRef]










| Strain | Species | Origin |
|---|---|---|
| FCF3 | B. cepacia | CF patients |
| FCF23 | B. cenocepacia | |
| LMG16656 | B. cenocepacia | |
| LMG21462 | B. cenocepacia | |
| LMG24506 | B. cenocepacia | |
| K56–2 | B. cenocepacia | |
| LMG13010 | B. multivorans | |
| LMG19182 | B. ambifaria | Environment |
| LMG1222 | B. cepacia | |
| LMG19230 | B. cenocepacia | |
| LMG17588 | B. multivorans |
| ARDRA Profiles | RAPD Haplotypes | Isolates | Genus | Accession Number |
|---|---|---|---|---|
| 1 | 1 | OR1 | Pseudomonas | OP522403 |
| 2 | 2 | OR2 | Pantoea | OP522404 |
| 3 | OR11 | Pantoea | - | |
| 3 | 4 | OR3 | Erwinia | OP522405 |
| 5 | OR6; OR15 | Erwinia | - | |
| 4 | 6 | OR4 | Paenibacillus | OP522406 |
| 5 | 7 | OR5 | Pseudomonas | OP522407 |
| 8 | OR19 | Pseudomonas | - | |
| 9 | OR28 | Pseudomonas | - | |
| 10 | OR65 | Pseudomonas | - | |
| 6 | 11 | OR7 | Peribacillus | OP522408 |
| 7 | 12 | OR8 | Bacillus | OP522409 |
| 13 | OR18 | Bacillus | - | |
| 8 | 14 | OR9 | Bacillus | OP522410 |
| 9 | 15 | OR12 | Peribacillus | OP522411 |
| 10 | 16 | OR24 | N.A. a | N.A. |
| 11 | 17 | OR31 | Staphylococcus | OP522412 |
| 12 | 18 | OR36 | Kocuria | OP522413 |
| 13 | 19 | OR51 | Erwinia | OP522414 |
| 14 | 20 | OR56 | Pseudomonas | OP522415 |
| MIC (µg/mL) | |||||||
|---|---|---|---|---|---|---|---|
| Genus | Isolate | Str. | Tet. | Cip. | Kan. | Chl. | Rif |
| Bacillus | OR9 | 50 | 0.5 | 0.5 | 50 | 2.5 | 5 |
| Bacillus | OR8 | 50 | 1.25 | 0.5 | 5 | 25 | 5 |
| Bacillus | OR18 | 50 | 1.25 | 0.5 | 5 | 25 | 5 |
| N.A. a | OR24 | 50 | 1.25 | 5 | 5 | 5 | 5 |
| Erwinia | OR3 | 10 | 12.5 | 0.5 | 10 | 5 | 25 |
| Erwinia | OR6 | 10 | 5 | 0.5 | 10 | 5 | 10 |
| Erwinia | OR51 | 10 | 12.5 | 0.5 | 10 | 5 | 25 |
| Kocuria | OR36 | 10 | 12.5 | 5 | 50 | 5 | 10 |
| Paenibacillus | OR4 | n.a. a | n.a. a | n.a. a | n.a. a | n.a. a | n.a. a |
| Pantoea | OR2 | 10 | 5 | 0.5 | 50 | 2.5 | 10 |
| Pantoea | OR11 | 10 | 2.5 | 0.5 | 10 | 1 | 10 |
| Peribacillus | OR7 | 50 | 1.25 | 0.5 | 10 | 25 | 5 |
| Peribacillus | OR12 | 50 | 1.25 | 0.5 | 5 | 25 | 5 |
| Pseudomonas | OR1 | 10 | 2.5 | 0.5 | 2.5 | 25 | 5 |
| Pseudomonas | OR19 | 50 | 1.25 | 2.5 | 2.5 | 5 | 5 |
| Pseudomonas | OR5 | >50 | 25 | 0.5 | 10 | >50 | 25 |
| Pseudomonas | OR28 | >50 | 25 | 0.5 | 10 | >50 | 25 |
| Pseudomonas | OR56 | >50 | 25 | 0.5 | 10 | >50 | 25 |
| Pseudomonas | OR65 | >50 | >25 | 0.5 | 10 | >50 | 25 |
| Staphylococcus | OR31 | >50 | >25 | 0.5 | 5 | 2.5 | 5 |
| BCG type | From (nt) | To (nt) | Most Similar Known Cluster | % Similarity |
|---|---|---|---|---|
| Contig 1 | ||||
| Trans AT-PKS, NRPs | 495644 | 558461 | Bacillomycin D | (40%) |
| Terpene | 2943693 | 2964472 | Molybdenum cofactor | (17%) |
| NRPs | 3799358 | 3842588 | - | - |
| RiPP-like | 3864265 | 3873890 | - | - |
| RiPP-like | 3919957 | 3929781 | - | - |
| Betalactone | 3978104 | 4003065 | Fengycin | (40%) |
| NRPs | 4005346 | 4068767 | Puwainaphycin F/ MinutissamideA | (44%) |
| NRPs-like, NRPs | 4190374 | 4235399 | Bacillibactin | (38%) |
| Siderophore | 4609931 | 4624859 | Petrobactin | (100%) |
| NRPs | 4685992 | 4742532 | Octapeptin C4 | (17%) |
| RRE-containing, RiPP-like | 5304467 | 5324736 | - | - |
| Contig 2 | ||||
| RRE-containing | 150714 | 170974 | - | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Semenzato, G.; Faddetta, T.; Falsini, S.; Del Duca, S.; Esposito, A.; Padula, A.; Greco, C.; Mucci, N.; Zaccaroni, M.; Puglia, A.M.; et al. Endophytic Bacteria Associated with Origanum heracleoticum L. (Lamiaceae) Seeds. Microorganisms 2022, 10, 2086. https://doi.org/10.3390/microorganisms10102086
Semenzato G, Faddetta T, Falsini S, Del Duca S, Esposito A, Padula A, Greco C, Mucci N, Zaccaroni M, Puglia AM, et al. Endophytic Bacteria Associated with Origanum heracleoticum L. (Lamiaceae) Seeds. Microorganisms. 2022; 10(10):2086. https://doi.org/10.3390/microorganisms10102086
Chicago/Turabian StyleSemenzato, Giulia, Teresa Faddetta, Sara Falsini, Sara Del Duca, Antonia Esposito, Anna Padula, Claudia Greco, Nadia Mucci, Marco Zaccaroni, Anna Maria Puglia, and et al. 2022. "Endophytic Bacteria Associated with Origanum heracleoticum L. (Lamiaceae) Seeds" Microorganisms 10, no. 10: 2086. https://doi.org/10.3390/microorganisms10102086
APA StyleSemenzato, G., Faddetta, T., Falsini, S., Del Duca, S., Esposito, A., Padula, A., Greco, C., Mucci, N., Zaccaroni, M., Puglia, A. M., Papini, A., & Fani, R. (2022). Endophytic Bacteria Associated with Origanum heracleoticum L. (Lamiaceae) Seeds. Microorganisms, 10(10), 2086. https://doi.org/10.3390/microorganisms10102086











