Control Measures for SARS-CoV-2: A Review on Light-Based Inactivation of Single-Stranded RNA Viruses
Abstract
:1. Introduction
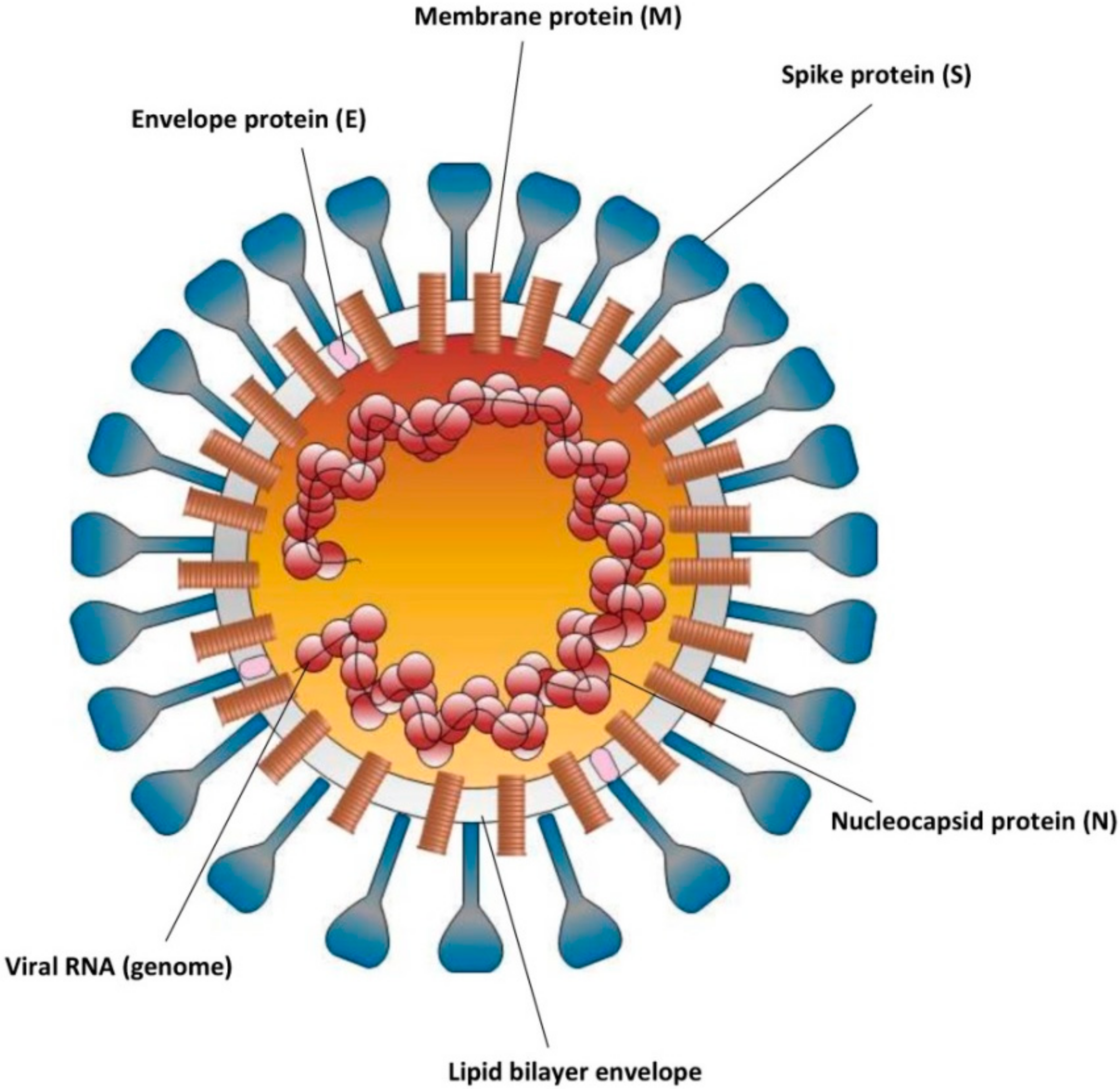
2. Virion Structure of SARS-CoV-2
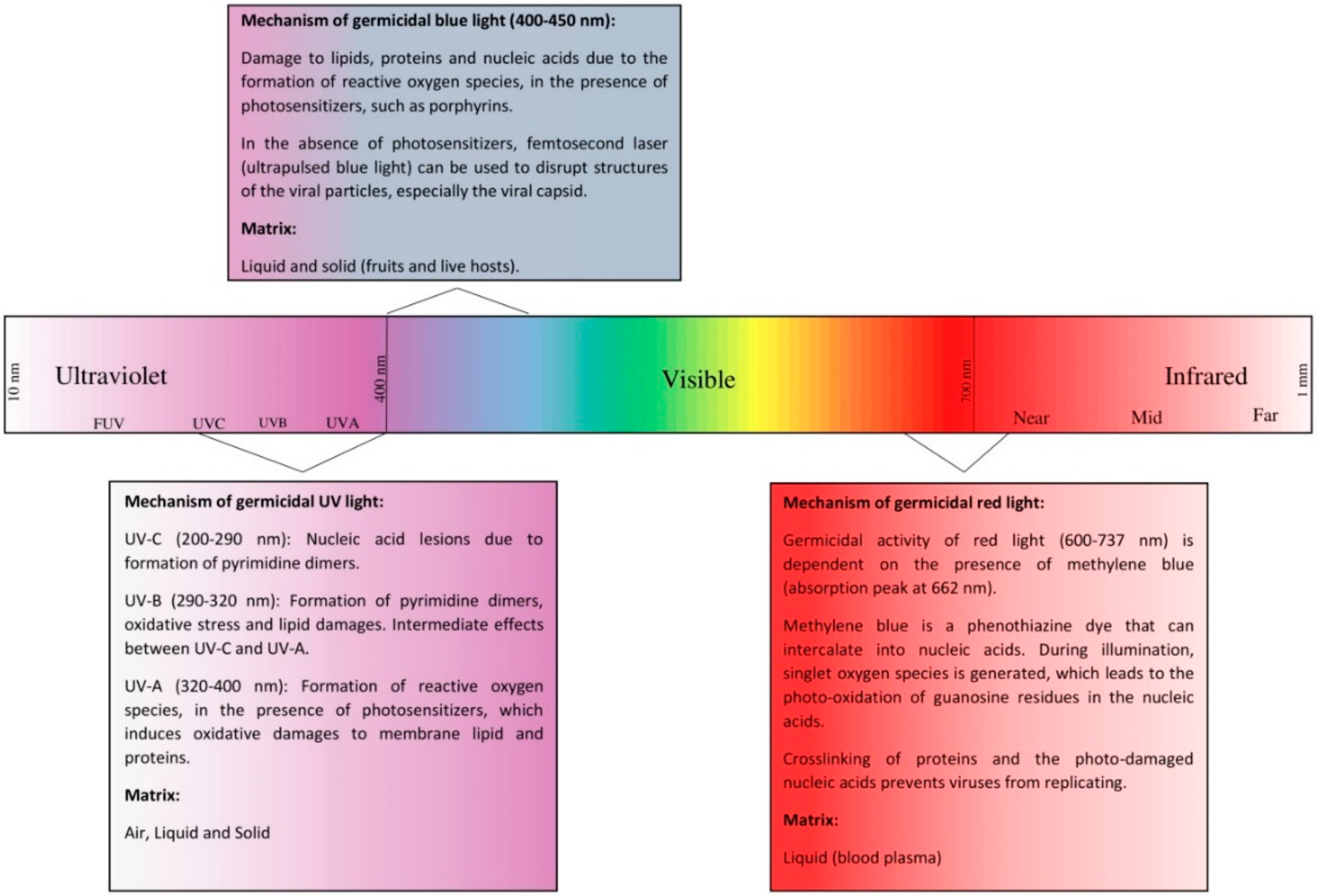
3. Viral Properties of Ultraviolet Light
3.1. Mechanism
3.2. Inactivation of Viruses in Air
3.3. Inactivation of Viruses in Liquid Matrix
3.4. Inactivation of Viruses on Solid Surfaces
3.5. Comparison of Light Dosage across Studies
4. Virucidal Properties of Blue Light
4.1. Mechanism
4.2. Viral Inactivation
5. Virucidal Properties of Red Light
6. Photosensitizing Agents for UV-A and UV-B
7. Hurdle Technology
8. Future Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Drosten, C.; Günther, S.; Preiser, W.; Van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.M.; et al. Identification of a Novel Coronavirus in Patients with Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef] [PubMed]
- Ksiazek, T.G.; Erdman, D.; Goldsmith, C.S.; Zaki, S.R.; Peret, T.; Emery, S.; Tong, S.; Urbani, C.; Comer, J.A.; Lim, W.; et al. A Novel Coronavirus Associated with Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2003, 348, 1953–1966. [Google Scholar] [CrossRef] [PubMed]
- Zaki, A.M.; Van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Isolation of a Novel Coronavirus from a Man with Pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical Features of Patients Infected with 2019 Novel Coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- CDC. Coronaviruses. Available online: https://www.cdc.gov/coronavirus/types.html (accessed on 2 May 2020).
- WHO. Middle East Respiratory Syndrome Coronavirus (MERS-CoV). Available online: https://www.who.int/emergencies/mers-cov/en/ (accessed on 2 May 2020).
- WHO. Summary of Probable SARS Cases with Onset of Illness from 1 November 2002 to 31 July 2003. Available online: https://www.who.int/csr/sars/country/table2004_04_21/en/ (accessed on 2 May 2020).
- WHO. Novel Coronavirus—China. Available online: https://www.who.int/csr/don/12-january-2020-novel-coronavirus-china/en/ (accessed on 2 May 2020).
- WHO. Modes of Transmission of Virus Causing COVID-19: Implications for IPC Precaution Recommendations. Available online: https://www.who.int/news-room/commentaries/detail/modes-of-transmission-of-virus-causing-covid-19-implications-for-ipc-precaution-recommendations (accessed on 20 May 2020).
- Bahl, P.; Doolan, C.; de Silva, C.; Chughtai, A.A.; Bourouiba, L.; MacIntyre, C.R. Airborne or Droplet Precautions for Health Workers Treating Coronavirus Disease 2019? J. Infect. Dis. 2020. [Google Scholar] [CrossRef] [Green Version]
- Bourouiba, L.; Dehandschoewercker, E.; Bush, J.W.M.; Bourouiba, L. Violent Expiratory Events: On Coughing and Sneezing. J. Fluid Mech. 2014, 745, 537–563. [Google Scholar] [CrossRef]
- Bourouiba, L. A Sneeze. N. Engl. J. Med. 2016, 375, e15. [Google Scholar] [CrossRef]
- Lee, J.; Yoo, D.; Ryu, S.; Ham, S.; Lee, K.; Yeo, M.; Min, K.; Yoon, C. Quantity, Size Distribution, and Characteristics of Cough-generated Aerosol Produced by Patients with an Upper Respiratory Tract Infection. Aerosol Air Qual. Res. 2019, 19, 840–853. [Google Scholar] [CrossRef]
- Parienta, D.; Morawska, L.; Johnson, G.; Ristovski, Z.; Hargreaves, M.; Mengersen, K.; Corbett, S.; Chao, C.; Li, Y.; Katoshevski, D. Theoretical Analysis of the Motion and Evaporation of Exhaled Respiratory Droplets of Mixed Composition. J. Aerosol Sci. 2010, 45, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Wei, J.; Li, Y. Enhanced Spread of Expiratory Droplets by Turbulence in a Cough Jet. Build. Environ. 2015, 93, 86–96. [Google Scholar] [CrossRef]
- Liu, L.; Wei, J.; Li, Y.; Ooi, A. Evaporation and Dispersion of Respiratory Droplets from Coughing. Indoor Air 2017, 27, 179–190. [Google Scholar] [CrossRef]
- Wei, J.; Li, Y. Human Cough as a Two-Stage Jet and Its Role in Particle Transport. PLoS ONE 2017, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, X.; Li, Y.; Chwang, A.T.Y.; Ho, P.L.; Seto, W.H. How Far Droplets Can Move in Indoor Environments—Revisiting the Wells Evaporation-Falling Curve. Indoor Air 2007, 17, 211–225. [Google Scholar] [CrossRef]
- Zhu, S.; Kato, S.; Yang, J.-H. Study on Transport Characteristics of Saliva Droplets Produced by Coughing in a Calm Indoor Environment. Build. Environ. 2006, 41, 1691–1702. [Google Scholar] [CrossRef]
- Guo, Z.-D.; Wang, Z.-Y.; Zhang, S.-F.; Li, X.; Li, L.; Li, C.; Cui, Y.; Fu, R.-B.; Dong, Y.-Z.; Chi, X.-Y.; et al. Aerosol and Surface Distribution of Severe Acute Respiratory Syndrome Coronavirus 2 in Hospital Wards, Wuhan, China, 2020. Emerg. Infect. Dis. 2020, 26, 1586–1591. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef]
- Fears, A.C.; Klimstra, W.B.; Duprex, P.; Hartman, A.; Weaver, S.C.; Plante, K.S.; Mirchandani, D.; Plante, J.A.; Aguilar, P.V.; Fernández, D.; et al. Persistence of Severe Acute Respiratory Syndrome Coronavirus 2 in Aerosol Suspensions. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef] [PubMed]
- Tang, A.; Tong, Z.D.; Wang, H.L.; Dai, Y.X.; Li, K.F.; Liu, J.N.; Wu, W.J.; Yuan, C.; Yu, M.L.; Li, P.; et al. Detection of Novel Coronavirus by RT-PCR in Stool Specimen from Asymptomatic Child, China. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA J. Am. Med. Assoc. 2020. [Google Scholar] [CrossRef] [Green Version]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Ahmed, W.; Angel, N.; Edson, J.; Bibby, K.; Bivins, A.; O’brien, J.W.; Choi, P.M.; Kitajima, M.; Simpson, S.L.; Li, J.; et al. Journal Pre-Proof First Confirmed Detection of SARS-CoV-2 in Untreated Wastewater in Australia: A Proof of Concept for the Wastewater Surveillance of COVID-19 in the Community. Sci. Total Environ. 2020, 728, 138764. [Google Scholar] [CrossRef]
- Medema, G.; Heijnen, L.; Elsinga, G.; Italiaander, R.; Brouwer, A. Presence of SARS-Coronavirus-2 RNA in Sewage and Correlation with Reported COVID-19 Prevalence in the Early Stage of the Epidemic in The Netherlands. Environ. Sci. Technol. Lett. 2020, 7, 511–514. [Google Scholar] [CrossRef]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial Undocumented Infection Facilitates the Rapid Dissemination of Novel Coronavirus (SARS-CoV2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef] [Green Version]
- Arons, M.M.; Hatfield, K.M.; Reddy, S.C.; Kimball, A.; James, A.; Jacobs, J.R.; Taylor, J.; Spicer, K.; Bardossy, A.C.; Oakley, L.P.; et al. Presymptomatic SARS-CoV-2 Infections and Transmission in a Skilled Nursing Facility. N. Engl. J. Med. 2020, 2081–2090. [Google Scholar] [CrossRef]
- Kimball, A.; Hatfield, K.M.; Arons, M.; James, A.; Taylor, J.; Spicer, K.; Bardossy, A.C.; Oakley, L.P.; Tanwar, S.; Chisty, Z.; et al. Asymptomatic and Presymptomatic SARS-CoV-2 Infections in Residents of a Long-Term Care Skilled Nursing Facility—King County, Washington, March 2020. Morb. Mortal. Wkly. Rep. Summ. CDC 2020, 69, 377–381. [Google Scholar] [CrossRef] [Green Version]
- Gandhi, M.; Yokoe, D.S.; Havlir, D.V. Asymptomatic Transmission, the Achilles’ Heel of Current Strategies to Control Covid-19. N. Engl. J. Med. 2020, 1–3. [Google Scholar] [CrossRef]
- Hijnen, D.; Marzano, A.V.; Eyerich, K.; GeurtsvanKessel, C.; Giménez-Arnau, A.M.; Joly, P.; Vestergaard, C.; Sticherling, M.; Schmidt, E. SARS-CoV-2 Transmission from Presymptomatic Meeting Attendee, Germany. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Aboubakr, H.A.; Sharafeldin, T.A.; Goyal, S.M. Stability of SARS-CoV-2 and Other Coronaviruses in the Environment and on Common Touch Surfaces and the Influence of Climatic Conditions: A review. Transbound. Emerg. Dis. 2020. [Google Scholar] [CrossRef]
- Gwynne, P.J.; Gallagher, M.P. Light as a Broad-Spectrum Antimicrobial. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Yin, R.; Dai, T.; Avci, P.; Jorge, A.E.S.; De Melo, W.C.M.A.; Vecchio, D.; Huang, Y.Y.; Gupta, A.; Hamblin, M.R. Light Based Anti-Infectives: Ultraviolet C Irradiation, Photodynamic Therapy, blue light, and beyond. Curr. Opin. Pharmacol. 2013, 13, 731–762. [Google Scholar] [CrossRef] [Green Version]
- Bintsis, T.; Litopoulou-Tzanetaki, E.; Robinson, R.K. Existing and Potential Applications of Ultraviolet Light in the Food Industry—A Critical Review. J. Sci. Food Agric. 2000, 80, 637–645. [Google Scholar] [CrossRef]
- Santin, G.C.; Oliveira, D.S.B.; Galo, R.; Borsatto, M.C.; Corona, S.A.M. Antimicrobial Photodynamic Therapy and Dental Plaque: A Systematic Review of the Literature. Sci. World J. 2014, 2014, 824538. [Google Scholar] [CrossRef] [Green Version]
- Gorbalenya, A.E.; Baker, S.C.; Baric, R.S.; de Groot, R.J.; Drosten, C.; Gulyaeva, A.A.; Haagmans, B.L.; Lauber, C.; Leontovich, A.M.; Neuman, B.W.; et al. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef] [Green Version]
- Yadav, P.; Potdar, V.; Cherian, S.S. Full-Genome Sequences of the First Two SARS-CoV-2 Viruses from India. Indian J. Med. Res. 2020, 151, 200–209. [Google Scholar]
- Sah, R.; Rodriguez-Morales, A.J.; Jha, R.; Chu, D.K.W.; Gu, H.; Peiris, M.; Bastola, A.; Lal, B.K.; Ojha, H.C.; Rabaan, A.A.; et al. Complete Genome Sequence of a 2019 Novel Coronavirus (SARS-CoV-2) Strain Isolated in Nepal. Microbiol. Resour. Announc. 2020, 9. [Google Scholar] [CrossRef] [Green Version]
- Khailany, R.; Safdar, M.; Ozaslan, M. Genomic Characterization of a Novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Woo, P.; Huang, Y.; Lau, S.; Yuen, K. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef] [Green Version]
- Kowalski, W. Genomic Model for the Prediction of Ultraviolet Inactivation Rate Constants for RNA and DNA Viruses. In Ultraviolet Germcidal Irradiation Handbook: UVGI Air and Surface; International Ultraviolet Association: Boston, MA, USA, 2009; pp. 4–10. [Google Scholar]
- Perlman, S.; Netland, J. Coronaviruses post-SARS: Update on replication and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 439–450. [Google Scholar] [CrossRef] [Green Version]
- He, Y.; Zhou, Y.; Siddiqui, P.; Jiang, S. Inactivated SARS-CoV vaccine elicits high titers of spike protein-specific antibodies that block receptor binding and virus entry. Biochem. Biophys. Res. Commun. 2004, 325, 445–452. [Google Scholar] [CrossRef]
- Schoeman, D.; Fielding, B.C. Coronavirus envelope protein: Current knowledge. Virol. J. 2019, 16, 1–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, D.X.; Yuan, Q.; Liao, Y. Coronavirus envelope protein: A small membrane protein with multiple functions. Cell. Mol. Life Sci. 2007, 64, 2043–2048. [Google Scholar] [CrossRef] [Green Version]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281.e6–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Tortorici, M.A.; Veesler, D. Structural Insights into Coronavirus Entry. Adv. Virus Res. 2019, 105, 93–116. [Google Scholar] [PubMed]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 2020, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, Y.J.; Walls, A.C.; Wang, Z.; Sauer, M.M.; Li, W.; Tortorici, M.A.; Bosch, B.J.; DiMaio, F.; Veesler, D. Structures of MERS-CoV spike glycoprotein in complex with sialoside attachment receptors. Nat. Struct. Mol. Biol. 2019, 26, 1151–1157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [Green Version]
- Wan, Y.; Shang, J.; Graham, R.; Baric, R.S.; Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: An Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020, 94, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Verdiá-Báguena, C.; Nieto-Torres, J.L.; Alcaraz, A.; DeDiego, M.L.; Torres, J.; Aguilella, V.M.; Enjuanes, L. Coronavirus E protein forms ion channels with functionally and structurally-involved membrane lipids. Virology 2012, 432, 485–494. [Google Scholar] [CrossRef] [Green Version]
- Nal, B.; Chan, C.; Kien, F.; Siu, L.; Tse, J.; Chu, K.; Kam, J.; Staropoli, I.; Crescenzo-Chaigne, B.; Escriou, N.; et al. Differential maturation and subcellular localization of severe acute respiratory syndrome coronavirus surface proteins S, M and E. J. Gen. Virol. 2005, 86, 1423–1434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ruch, T.R.; Machamer, C.E. The coronavirus E protein: Assembly and beyond. Viruses 2012, 4, 363–382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nieto-Torres, J.L.; DeDiego, M.L.; Álvarez, E.; Jiménez-Guardeño, J.M.; Regla-Nava, J.A.; Llorente, M.; Kremer, L.; Shuo, S.; Enjuanes, L. Subcellular location and topology of severe acute respiratory syndrome coronavirus envelope protein. Virology 2011, 415, 69–82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teoh, K.T.; Siu, Y.L.; Chan, W.L.; Schlüter, M.A.; Liu, C.J.; Peiris, J.S.M.; Bruzzone, R.; Margolis, B.; Nal, B. The SARS coronavirus E protein interacts with PALS1 and alters tight junction formation and epithelial morphogenesis. Mol. Biol. Cell 2010, 21, 3838–3852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cohen, J.R.; Lin, L.D.; Machamer, C.E. Identification of a Golgi Complex-Targeting Signal in the Cytoplasmic Tail of the Severe Acute Respiratory Syndrome Coronavirus Envelope Protein. J. Virol. 2011, 85, 5794–5803. [Google Scholar] [CrossRef] [Green Version]
- Ortego, J.; Escors, D.; Laude, H.; Enjuanes, L. Generation of a Replication-Competent, Propagation-Deficient Virus Vector Based on the Transmissible Gastroenteritis Coronavirus Genome. J. Virol. 2002, 76, 11518–11529. [Google Scholar] [CrossRef] [Green Version]
- Ortego, J.; Ceriani, J.E.; Patiño, C.; Plana, J.; Enjuanes, L. Absence of E protein arrests transmissible gastroenteritis coronavirus maturation in the secretory pathway. Virology 2007, 368, 296–308. [Google Scholar] [CrossRef] [Green Version]
- Curtis, K.M.; Yount, B.; Baric, R.S. Heterologous Gene Expression from Transmissible Gastroenteritis Virus Replicon Particles Downloaded from. J. Virol. 2002, 76, 1422–1434. [Google Scholar] [CrossRef] [Green Version]
- Dediego, M.L.; Álvarez, E.; Almazán, F.; Rejas, M.T.; Lamirande, E.; Roberts, A.; Shieh, W.-J.; Zaki, S.R.; Subbarao, K.; Enjuanes, L. A Severe Acute Respiratory Syndrome Coronavirus That Lacks the E Gene Is Attenuated In Vitro and In Vivo. J. Virol. 2007, 81, 1701–1713. [Google Scholar] [CrossRef] [Green Version]
- Hsieh, Y.C.; Li, H.C.; Chen, S.C.; Lo, S.Y. Interactions between M protein and other structural proteins of severe, acute respiratory syndrome-associated coronavirus. J. Biomed. Sci. 2008, 15, 707–717. [Google Scholar] [CrossRef] [Green Version]
- Neuman, B.W.; Kiss, G.; Kunding, A.H.; Bhella, D.; Baksh, M.F.; Connelly, S.; Droese, B.; Klaus, J.P.; Makino, S.; Sawicki, S.G.; et al. A structural analysis of M protein in coronavirus assembly and morphology. J. Struct. Biol. 2011, 174, 11–22. [Google Scholar] [CrossRef]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. In Coronaviruses: Methods and Protocols; Springer: New York, NY, USA, 2015; pp. 1–23. ISBN 9781493924387. [Google Scholar]
- Alsaadi, E.A.J.; Jones, I.M. Membrane binding proteins of coronaviruses. Future Virol. 2019, 14, 275–286. [Google Scholar] [CrossRef] [Green Version]
- Arndt, A.L.; Larson, B.J.; Hogue, B.G. A Conserved Domain in the Coronavirus Membrane Protein Tail Is Important for Virus Assembly. J. Virol. 2010, 84, 11418–11428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mcbride, R.; Van Zyl, M.; Fielding, B.C. The Coronavirus Nucleocapsid Is a Multifunctional Protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsin, W.C.; Chang, C.H.; Chang, C.Y.; Peng, W.H.; Chien, C.L.; Chang, M.F.; Chang, S.C. Nucleocapsid protein-dependent assembly of the RNA packaging signal of Middle East respiratory syndrome coronavirus. J. Biomed. Sci. 2018, 25, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, K.; Maeda, A.; Maeda, J.; Makino, S. Characterization of the Coronavirus M Protein and Nucleocapsid Interaction in Infected Cells. J. Virol. 2000, 74, 8127–8134. [Google Scholar] [CrossRef] [Green Version]
- Cui, L.; Wang, H.; Ji, Y.; Yang, J.; Xu, S.; Huang, X.; Wang, Z.; Qin, L.; Tien, P.; Zhou, X.; et al. The Nucleocapsid Protein of Coronaviruses Acts as a Viral Suppressor of RNA Silencing in Mammalian Cells. J. Virol. 2015, 89, 9029–9043. [Google Scholar] [CrossRef] [Green Version]
- Lu, X.; Pan, J.; Tao, J.; Guo, D. SARS-CoV nucleocapsid protein antagonizes IFN-β response by targeting initial step of IFN-β induction pathway, and its C-terminal region is critical for the antagonism. Virus Genes 2011, 42, 37–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozaki, Y.; Kawata, S. Introduction to FUV and DUV Spectrosocy. In Far-and Deep-Ultraviolet Spectroscopy; Springer: Tokyo, Japan, 2015; pp. 1–16. [Google Scholar]
- Tanabe, I.; Ozaki, Y. Far- and deep-ultraviolet spectroscopic investigations for titanium dioxide: Electronic absorption, Rayleigh scattering, and Raman spectroscopy. J. Mater. Chem. C 2016, 4, 7706–7717. [Google Scholar] [CrossRef]
- Kim, J.; Jang, J. Inactivation of airborne viruses using vacuum ultraviolet photocatalysis for a flow-through indoor air purifier with short irradiation time. Aerosol Sci. Technol. 2018, 52, 557–566. [Google Scholar] [CrossRef] [Green Version]
- Rutala, W.A.; Weber, D.J. Disinfection and Sterilization in Health Care Facilities: What Clinicians Need to Know. Clin. Infect. Dis. 2004, 39, 702–709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zimmerman, J.J. Ultraviolet irradiation and the mechanisms underlying its inactivation of infectious agents. Artic. Anim. Health Res. Rev. 2011, 12, 15–23. [Google Scholar] [CrossRef]
- Zaffina, S.; Camisa, V.; Lembo, M.; Vinci, M.R.; Tucci, M.G.; Borra, M.; Napolitano, A.; Cannatã, V. Accidental exposure to UV radiation produced by germicidal lamp: Case report and risk assessment. Photochem. Photobiol. 2012, 88, 1001–1004. [Google Scholar] [CrossRef] [PubMed]
- Dietz, L.; Horve, P.F.; Coil, D.A.; Fretz, M.; Eisen, J.A.; Van, K.; Wymelenberg, D. 2019 Novel Coronavirus (COVID-19) Pandemic: Built Environment Considerations To Reduce Transmission. Am. Soc. Microbiol. 2020, 5. [Google Scholar] [CrossRef] [Green Version]
- Haji Malayeri, A.; Mohseni, M.; Cairns, B.; Bolton, J.R. Fluence (UV Dose) Required to Achieve Incremental Log Inactivation of Bacteria, Protozoa, Viruses and Algae. IUVA News 2016, 18, 4–6. [Google Scholar]
- Welch, D.; Buonanno, M.; Grilj, V.; Shuryak, I.; Crickmore, C.; Bigelow, A.W.; Randers-Pehrson, G.; Johnson, G.W.; Brenner, D.J. Far-UVC light: A new tool to control the spread of airborne-mediated microbial diseases. Sci. Rep. 2018, 8, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Buonanno, M.; Welch, D.; Shuryak, I.; Brenner, D.J. Far-UVC light efficiently and safely inactivates airborne human coronaviruses. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef]
- Kaidzu, S.; Sugihara, K.; Sasaki, M.; Nishiaki, A.; Igarashi, T.; Tanito, M. Evaluation of acute corneal damage induced by 222-nm and 254-nm ultraviolet light in Sprague–Dawley rats. Free Radic. Res. 2019, 53, 611–617. [Google Scholar] [CrossRef]
- Narita, K.; Asano, K.; Morimoto, Y.; Igarashi, T.; Nakane, A. Chronic irradiation with 222-nm UVC light induces neither DNA damage nor epidermal lesions in mouse skin, even at high doses. PLoS ONE 2018, 13, e0201259. [Google Scholar] [CrossRef]
- Buonanno, M.; Ponnaiya, B.; Welch, D.; Stanislauskas, M.; Randers-Pehrson, G.; Smilenov, L.; Lowy, F.D.; Owens, D.M.; Brenner, D.J. Germicidal Efficacy and Mammalian Skin Safety of 222-nm UV Light. Radiat. Res. 2017, 187, 493–501. [Google Scholar] [CrossRef] [Green Version]
- Pfeifer, G.P. Formation and Processing of UV Photoproducts: Effetcs of DNA Sequence and Chromatin Environment. Photochem. Photobiol. 1997, 65, 270–283. [Google Scholar] [CrossRef] [PubMed]
- Kowalski, W. Ultraviolet Germicidal Irradiation Handbook: UVGI for Air and Surface Disinfection; Springer: Berlin/Heidelberg, Germany, 2009; ISBN 9783642019982. [Google Scholar]
- Wurtmann, E.J.; Wolin, S.L. RNA under attack: Cellular handling of RNA damage RNA under attack: Cellular handling of RNA damage E. J. Wurtmann et.al. Crit. Rev. Biochem. Mol. Biol. 2009, 44, 34–49. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wigginton, K.R.; Menin, L.; Sigstam, T.; Gannon, G.; Cascella, M.; Hamidane, H.B.; Tsybin, Y.O.; Waridel, P.; Kohn, T. UV Radiation Induces Genome-Mediated, Site-Specific Cleavage in Viral Proteins. ChemBioChem 2012, 13, 837–845. [Google Scholar] [CrossRef] [PubMed]
- Beck, S.E.; Rodriguez, R.A.; Hawkins, M.A.; Hargy, T.M.; Larason, T.C.; Linden, K.G. Comparison of UV-Induced Inactivation and RNA Damage in MS2 Phage across the Germicidal UV Spectrum. Am. Soc. Microbiol. 2016, 82, 1468–1474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beck, S.E.; Rodriguez, R.A.; Linden, K.G.; Hargy, T.M.; Larason, T.C.; Wright, H.B. Wavelength Dependent UV Inactivation and DNA Damage of Adenovirus as Measured by Cell Culture Infectivity and Long Range Quantitative PCR. Environ. Sci. Technol. 2013, 48, 591–598. [Google Scholar] [CrossRef]
- Araud, E.; Fuzawa, M.; Shisler, J.L.; Li, J.; Nguyen, T.H. UV inactivation of rotavirus and tulane virus targets different components of the virions. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef]
- Hull, N.M.; Linden, K.G. Synergy of MS2 disinfection by sequential exposure to tailored UV wavelengths. Water Res. 2018, 143, 292–300. [Google Scholar] [CrossRef]
- Tseng, C.C.; Li, C.S. Inactivation of virus-containing aerosols by ultraviolet germicidal irradiation. Aerosol Sci. Technol. 2005, 39, 1136–1142. [Google Scholar] [CrossRef]
- Matallana-Surget, S.; Meador, J.A.; Joux, F.; Douki, T. Effect of the GC Content of DNA on the Distribution of UVB-Induced Bipyrimidine Photoproducts. Photochem. Photobiol. Sci. 2008, 7, 794–801. [Google Scholar] [CrossRef]
- Liu, Y.; Ning, Z.; Chen, Y.; Guo, M.; Liu, Y.; Gali, N.K.; Sun, L.; Duan, Y.; Cai, J.; Westerdahl, D.; et al. Aerodynamic analysis of SARS-CoV-2 in two Wuhan hospitals. Nature 2020, 582, 557–560. [Google Scholar] [CrossRef]
- Jiang, F.-C.; Jiang, X.-L.; Wang, Z.-G.; Meng, Z.-H.; Shao, S.-F.; Anderson, B.D.; Ma, M.-J. Detection of Severe Acute Respiratory Syndrome Coronavirus 2 RNA on Surfaces in Quarantine Rooms. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef] [PubMed]
- Mphaphlele, M.; Dharmadhikari, A.S.; Jensen, P.A.; Rudnick, S.N.; Van Reenen, T.H.; Pagano, M.A.; Leuschner, W.; Sears, T.A.; Milonova, S.P.; Van Der Walt, M.; et al. Institutional tuberculosis transmission: Controlled trial of upper room ultraviolet air disinfection: A basis for new dosing guidelines. Am. J. Respir. Crit. Care Med. 2015, 192, 477–484. [Google Scholar] [CrossRef] [Green Version]
- Escombe, A.R.; Moore, D.A.J.; Gilman, R.H.; Navincopa, M.; Ticona, E.; Mitchell, B.; Noakes, C.; Martinez, C.; Sheen, P.; Ramirez, R.; et al. Upper-room ultraviolet light and negative air ionization to prevent tuberculosis transmissio. PLoS Med. 2009, 6, 0312–0323. [Google Scholar] [CrossRef] [Green Version]
- Xu, P.; Peccia, J.; Fabian, P.; Martyny, J.W.; Fennelly, K.P.; Hernandez, M.; Miller, S.L. Efficacy of Ultraviolet Germicidal Irradiation of Upper-Room Air in Inactivating Airborne Bacterial Spores and Mycobacteria in Full-Scale Studies. Atmos. Environ. 2003, 37, 405–417. [Google Scholar] [CrossRef]
- Nardell, E.A.; Bucher, S.J.; Brickner, P.W.; Wang, C.; Vincent, R.L.; Becan-McBride, K.; James, M.A.; Michael, M.; Wright, J.D. Safety of upper-room ultraviolet germicidal air disinfection for room occupants: Results from the Tuberculosis Ultraviolet Shelter Study. Public Health Rep. 2008, 123, 52–60. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, S.L.; Linnes, J.; Luongo, J. Ultraviolet germicidal irradiation: Future directions for air disinfection and building applications. Photochem. Photobiol. 2013, 89, 777–781. [Google Scholar] [CrossRef]
- Cutler, T.D.; Wang, C.; Hoff, S.J.; Zimmerman, J.J. Effect of temperature and relative humidity on ultraviolet (UV 254) inactivation of airborne porcine respiratory and reproductive syndrome virus. Vet. Microbiol. 2012, 159, 47–52. [Google Scholar] [CrossRef]
- Mcdevitt, J.J.; Rudnick, S.N.; Radonovich, L.J. Aerosol Susceptibility of Influenza Virus to UV-C Light. Am. Soc. Microbiol. 2012. [Google Scholar] [CrossRef] [Green Version]
- Kormuth, K.A.; Lin, K.; Prussin, A.J.; Vejerano, E.P.; Tiwari, A.J.; Cox, S.S.; Myerburg, M.M.; Lakdawala, S.S.; Marr, L.C. Influenza virus infectivity is retained in aerosols and droplets independent of relative humidity. J. Infect. Dis. 2018, 218, 739–747. [Google Scholar] [CrossRef]
- Kormuth, K.A.; Lin, K.; Qian, Z.; Myerburg, M.M.; Marr, L.C.; Lakdawala, S.S. Environmental Persistence of Influenza Viruses Is Dependent upon Virus Type and Host Origin. mSphere 2019, 4. [Google Scholar] [CrossRef] [Green Version]
- Yang, W.; Elankumaran, S.; Marr, L.C. Relationship between Humidity and Influenza A Viability in Droplets and Implications for Influenza’s Seasonality. PLoS ONE 2012, 7, e46789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naunovic, Z.; Lim, S.; Blatchley, E.R. Investigation of microbial inactivation efficiency of a UV disinfection system employing an excimer lamp. Water Res. 2008, 42, 4838–4846. [Google Scholar] [CrossRef] [PubMed]
- Sosnin, E.A.; Lavrent’eva, L.V.; Erofeev, M.V.; Masterova, Y.V.; Kuznetzova, E.N.; Tarasenko, V.F. New Bactericidal UV Light Sources: Excilamps. At. Mol. Pulsed Laser 2004, 5483, 317–322. [Google Scholar] [CrossRef]
- Buonanno, M.; Randers-Pehrson, G.; Bigelow, A.W.; Trivedi, S.; Lowy, F.D.; Spotnitz, H.M.; Hammer, S.M.; Brenner, D.J. 207-nm UV Light—A Promising Tool for Safe Low-Cost Reduction of Surgical Site Infections. I: In Vitro Studies. PLoS ONE 2013, 8, e76968. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oppenländer, T. Mercury-free sources of VUV/UV radiation: Application of modern excimer lamps (excilamps) for water and air treatment. J. Environ. Eng. Sci. 2007, 6, 253–264. [Google Scholar] [CrossRef]
- Walker, C.M.; Ko, G. Effect of Ultraviolet Germicidal Irradiation on Viral Aerosols. Environ. Sci. Technol. 2007, 41, 5460–5465. [Google Scholar] [CrossRef] [PubMed]
- Gundy, P.M.; Gerba, C.P.; Pepper, I.L. Survival of Coronaviruses in Water and Wastewater. Food Environ. Virol. 2009, 1, 10–14. [Google Scholar] [CrossRef] [Green Version]
- Van Doremalen, N.; Bushmaker, T.; Karesh, W.B.; Munster, V.J. Stability of middle east respiratory syndrome coronavirus in milk. Emerg. Infect. Dis. 2014, 20, 1263–1264. [Google Scholar] [CrossRef]
- Chang, L.; Yan, Y.; Wang, L. Coronavirus Disease 2019: Coronaviruses and Blood Safety. Transfus. Med. Rev. 2020, 34, 75–80. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Maintaining a Safe and Adequate Blood Supply during the Pandemic Outbreak of Coronavirus Disease (COVID-19). Available online: https://apps.who.int/iris/handle/10665/333182 (accessed on 5 August 2020).
- FDA Updated Information for Blood Establishments Regarding the Novel Coronavirus Outbreak. Available online: https://www.fda.gov/vaccines-blood-biologics/safety-availability-biologics/updated-information-blood-establishments-regarding-novel-coronavirus-covid-19-outbreak (accessed on 5 August 2020).
- Roberts, P.; Hope, A. Virus inactivation by high intensity broad spectrum pulsed light. J. Virol. Methods 2003, 110, 61–65. [Google Scholar] [CrossRef]
- Qiao, Z.; Ye, Y.; Chang, H.; Thirunarayanan, D.; Wigginton, K.R. Nucleic Acid Photolysis by UV 254 and the Impact of Virus Encapsidation. Environ. Sci. Technol. 2018, 52, 10408–10415. [Google Scholar] [CrossRef]
- Kim, D.K.; Kim, S.J.; Kang, D.H. Inactivation modeling of human enteric virus surrogates, MS2, Qβ, and ΦX174, in water using UVC-LEDs, a novel disinfecting system. Food Res. Int. 2017, 91, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Duan, S.M.; Zhao, X.S.; Wen, R.F.; Huang, J.J.; Pi, G.H.; Zhang, S.X.; Han, J.; Bi, S.L.; Ruan, L.; Dong, X.P. Stability of SARS Coronavirus in Human Specimens and Environment and Its Sensitivity to Heating and UV Irradiation. Biomed. Environ. Sci. 2003, 16, 246–255. [Google Scholar]
- Thurston-Enriquez, J.A.; Haas, C.N.; Jacangelo, J.; Riley, K.; Gerba, C.P. Inactivation of Feline Calicivirus and Adenovirus Type 40 by UV Radiation. Appl. Environ. Microbiol. 2003, 69, 577–582. [Google Scholar] [CrossRef] [Green Version]
- Darnell, M.E.R.; Subbarao, K.; Feinstone, S.M.; Taylor, D.R. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J. Virol. Methods 2004, 121, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Azar Daryany, M.K.; Hosseini, S.M.; Raie, M.; Fakharie, J.; Zareh, A. Study on continuous (254 nm) and pulsed UV (266 and 355 nm) lights on BVD virus inactivation and its effects on biological properties of fetal bovine serum. J. Photochem. Photobiol. B Biol. 2009, 94, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Jean, J.; Morales-Rayas, R.; Anoman, M.N.; Lamhoujeb, S. Inactivation of hepatitis A virus and norovirus surrogate in suspension and on food-contact surfaces using pulsed UV light (pulsed light inactivation of food-borne viruses). Food Microbiol. 2011, 28, 568–572. [Google Scholar] [CrossRef]
- Vimont, A.; Fliss, I.; Jean, J. Efficacy and Mechanisms of Murine Norovirus Inhibition by Pulsed-Light Technology. Am. Soc. Microbiol. 2015, 81, 2950–2957. [Google Scholar] [CrossRef] [Green Version]
- Huang, Y.; Ye, M.; Cao, X.; Chen, H. Pulsed light inactivation of murine norovirus, Tulane virus, Escherichia coli O157:H7 and Salmonella in suspension and on berry surfaces. Food Microbiol. 2016, 61, 1–4. [Google Scholar] [CrossRef] [Green Version]
- Rattanakul, S.; Oguma, K. Inactivation kinetics and efficiencies of UV-LEDs against Pseudomonas aeruginosa, Legionella pneumophila, and surrogate microorganisms. Water Res. 2018, 130, 31–37. [Google Scholar] [CrossRef]
- Song, K.; Taghipour, F.; Mohseni, M. Microorganisms inactivation by wavelength combinations of ultraviolet light-emitting diodes (UV-LEDs). Sci. Total Environ. 2019, 665, 1103–1110. [Google Scholar] [CrossRef] [PubMed]
- Steinmann, E.; Gravemann, U.; Friesland, M.; Doerrbecker, J.; Müller, T.H.; Pietschmann, T.; Seltsam, A. Two pathogen reduction technologies—Methylene blue plus light and shortwave ultraviolet light—Effectively inactivate hepatitis C virus in blood products. Transfusion 2013, 53, 1010–1018. [Google Scholar] [CrossRef] [PubMed]
- Eickmann, M.; Gravemann, U.; Handke, W.; Tolksdorf, F.; Reichenberg, S.; Müller, T.H.; Seltsam, A. Inactivation of Ebola virus and Middle East respiratory syndrome coronavirus in platelet concentrates and plasma by ultraviolet C light and methylene blue plus visible light, respectively. Transfusion 2018, 58, 2202–2207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eickmann, M.; Gravemann, U.; Handke, W.; Tolksdorf, F.; Reichenberg, S.; Müller, T.H.; Seltsam, A. Inactivation of three emerging viruses—Severe acute respiratory syndrome coronavirus, Crimean–Congo haemorrhagic fever virus and Nipah virus—In platelet concentrates by ultraviolet C light and in plasma by methylene blue plus visible light. Vox Sang. 2020, 115, 146–151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Inagaki, H.; Saito, A.; Sugiyama, H.; Okabayashi, T.; Fujimoto, S. Rapid inactivation of SARS-CoV-2 with Deep-UV LED irradiation. Emerg. Microbes Infect. 2020, 1–8. [Google Scholar] [CrossRef]
- Bianco, A.; Biasin, M.; Pareschi, G.; Cavalleri, A.; Cavatorta, C.; Fenizia, F.; Galli, P.; Lessio, L.; Lualdi, M.; Redaelli, E.; et al. UV-C Irradiation Is Highly Effective in Inactivating and Inhibiting SARS-CoV-2 Replication. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Blá zquez, E.I.; Rodríguez, C.; Ródenas, J.; Navarro, N.; Riquelme, C.; Rosell, R.; Campbell, J.; Crenshaw, J.I.; Segalé, J.I.; Pujols, J.; et al. Evaluation of the effectiveness of the SurePure Turbulator ultraviolet-C irradiation equipment on inactivation of different enveloped and non-enveloped viruses inoculated in commercially collected liquid animal plasma. PLoS ONE 2019, 14, e0212332. [Google Scholar] [CrossRef] [Green Version]
- Pfaender, S.; Brinkmann, J.; Todt, D.; Riebesehl, N.; Steinmann, J.; Steinmann, J.; Pietschmann, T.; Steinmann, E. Mechanisms of Methods for Hepatitis C Virus Inactivation. Am. Soc. Microbiol. 2015, 81, 1616–1621. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Templeton, M.R.; Andrews, R.C.; Hofmann, R. Inactivation of particle-associated viral surrogates by ultraviolet light. Water Res. 2005, 39, 3487–3500. [Google Scholar] [CrossRef]
- Templeton, M.R.; Hofmann, R.; Andrews, R.C. UV inactivation of humic-coated bacteriophages MS2 and T4 in water. J. Environ. Eng. Sci. 2006, 5, 537–543. [Google Scholar] [CrossRef]
- Williams, O.W.; Sharafkhaneh, A.; Kim, V.; Dickey, B.F.; Evans, C.M. Airway Mucus From Production to Secretion. Am. J. Respir. Cell. Mol. Biol. 2006, 34, 527–536. [Google Scholar] [CrossRef]
- Vejerano, E.P.; Marr, L.C. Physico-chemical characteristics of evaporating respiratory fluid droplets. J. R. Soc. Interface 2018, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bohrerova, Z.; Shemer, H.; Lantis, R.; Impellitteri, C.A.; Linden, K.G. Comparative disinfection efficiency of pulsed and continuous-wave UV irradiation technologies. Water Res. 2008, 42, 2975–2982. [Google Scholar] [CrossRef] [PubMed]
- Elmnasser, N.; Guillou, S.; Leroi, F.; Orange, N.; Bakhrouf, A.; Federighi, M. Pulsed-light system as a novel food decontamination technology: A review. Can. J. Microbiol. 2007, 53, 813–821. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seltsam, A.; Müller, T.H. UVC irradiation for pathogen reduction of platelet concentrates and plasma. Transfus. Med. Hemotherapy 2011, 38, 43–54. [Google Scholar] [CrossRef]
- Kneissl, M.; Seong, T.Y.; Han, J.; Amano, H. The emergence and prospects of deep-ultraviolet light-emitting diode technologies. Nat. Photonics 2019, 13, 233–244. [Google Scholar] [CrossRef]
- Taniyasu, Y.; Kasu, M.; Makimoto, T. An aluminium nitride light-emitting diode with a wavelength of 210 nanometres. Nature 2006, 441, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, H.; Fujikawa, S.; Kamata, N. Recent progress in AlGaN-based deep-UV LEDs. Electron. Commun. Jpn. 2015, 98, 1–8. [Google Scholar] [CrossRef]
- Hijnen, W.A.M.; Beerendonk, E.F.; Medema, G.J. Inactivation credit of UV radiation for viruses, bacteria and protozoan (oo)cysts in water: A review. Water Res. 2006, 40, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Mattle, M.J.; Kohn, T. Inactivation and Tailing during UV 254 Disinfection of Viruses: Contributions of Viral Aggregation, Light Shielding within Viral Aggregates, and Recombination. Environ. Sci. Technol. 2012, 46, 10022–10030. [Google Scholar] [CrossRef] [PubMed]
- Luria, S.E.; Dulbecco, R. Genetic Recombinations Leading to Production of Active Bacteriophage. Genetics 1949, 34, 93–125. [Google Scholar] [PubMed]
- Center for Disease Control and Prevention Decontamination and Reuse of Filtering Facepiece Respirators: COVID-19. Available online: https://www.cdc.gov/coronavirus/2019-ncov/hcp/ppe-strategy/decontamination-reuse-respirators.html (accessed on 29 June 2020).
- Coulliette, A.D.; Perry, K.A.; Fisher, E.M.; Edwards, J.R.; Shaffer, R.E.; Noble-Wang, J. MS2 Coliphage as a Surrogate for 2009 Pandemic Influenza A (H1N1) Virus (pH1N1) in Surface Survival Studies on N95 Filtering Facepiece Respirators. J. Int. Soc. Respir. Prot. 2014, 21, 14–22. [Google Scholar] [PubMed]
- Coulliette, A.D.; Perry, K.A.; Edwards, J.R.; Noble-Wang, J.A. Persistence of the 2009 pandemic influenza a (H1N1) virus on N95 respirators. Appl. Environ. Microbiol. 2013, 79, 2148–2155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lore, M.B.; Heimbuch, B.K.; Brown, T.L.; Wander, J.D.; Hinrichs, S.H. Effectiveness of three decontamination treatments against influenza virus applied to filtering facepiece respirators. Ann. Occup. Hyg. 2012, 56, 92–101. [Google Scholar] [CrossRef] [Green Version]
- Heimbuch, B.K.; Wallace, W.H.; Kinney, K.; Lumley, A.E.; Wu, C.Y.; Woo, M.H.; Wander, J.D. A pandemic influenza preparedness study: Use of energetic methods to decontaminate filtering facepiece respirators contaminated with H1N1 aerosols and droplets. Am. J. Infect. Control 2011, 39. [Google Scholar] [CrossRef]
- Vo, E.; Rengasamy, S.; Shaffer, R. Development of a test system to evaluate procedures for decontamination of respirators containing viral droplets. Appl. Environ. Microbiol. 2009, 75, 7303–7309. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simmons, S.; Carrion, R.; Alfson, K.; Staples, H.; Jinadatha, C.; Jarvis, W.; Sampathkumar, P.; Chemaly, R.F.; Khawaja, F.; Povroznik, M.; et al. Deactivation of SARS-CoV-2 with Pulsed Xenon Ultraviolet: Implications for environmental COVID-19 control. Infect. Control Hosp. Epidemiol. 2020, 1–19. [Google Scholar] [CrossRef]
- Smith, J.S.; Hanseler, H.; Welle, J.; Rattray, R.; Campbell, M.; Brotherton, T.; Moudgil, T.; Pack, T.F.; Wegmann, K.; Jensen, S.; et al. Effect of various decontamination procedures on disposable N95 mask integrity and SARS-CoV-2 infectivity. J. Clin. Transl. Sci. 2020, 1–14. [Google Scholar] [CrossRef]
- Kariwa, H.; Fujii, N.; Takashima, I. Inactivation of SARS Coronavirus by Means of Povidone-Iodine, Physical Conditions, and Chemical Reagents. Jpn. J. Vet. Sci. 2004, 52, 105–112. [Google Scholar] [CrossRef] [Green Version]
- Sagripanti, J.L.; Lytle, C.D. Sensitivity to ultraviolet radiation of Lassa, vaccinia, and Ebola viruses dried on surfaces. Springer 2011, 156, 489–494. [Google Scholar] [CrossRef]
- Fino, V.R.; Kniel, K.E. UV light inactivation of hepatitis A virus, aichi virus, and feline calicivirus on strawberries, green onions, and lettuce. J. Food Prot. 2008, 71, 908–913. [Google Scholar] [CrossRef]
- Belliot, G.; Loutreul, J.; Estienney, M.; Cazeaux, C.; Nicorescu, I.; Aho, S.; Gervais, P.; Orange, N.; Pothier, P.; Morin, T. Potential of Pulsed Light to Inactivate Bacteriophage MS2 in Simple Liquid Medium and on Complex Foodstuffs. Food Environ. Virol. 2013, 5, 176–179. [Google Scholar] [CrossRef] [PubMed]
- Bedell, K.; Buchaklian, A.H.; Perlman, S. Efficacy of an automated multiple emitter whole-room Ultraviolet-C disinfection system against coronaviruses MHV and MERS-CoV. Infect. Control Hosp. Epidemiol. 2016, 37, 598–599. [Google Scholar] [CrossRef] [Green Version]
- Tseng, C.C.; Li, C.S. Inactivation of viruses on surfaces by ultraviolet germicidal irradiation. J. Occup. Environ. Hyg. 2007, 4, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Lytle, D.; Sagripanti, J.-L. Predicted Inactivation of Viruses of Relevance to Biodefense by Solar Radiation. J. Virol. 2005, 79, 14244–14252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nuanualsuwan, S.; Mariam, T.; Himathongkham, S.; Cliver, D.O. Ultraviolet Inactivation of Feline Calicivirus, Human EntericViruses and Coliphages¶. Photochem. Photobiol. 2002, 76, 406–410. [Google Scholar] [CrossRef]
- Maria, A.; Husman, R.; Bijkerk, P.; Lodder, W.; Van Den Berg, H.; Pribil, W.; Cabaj, A.; Gehringer, P.; Sommer, R.; Duizer, E. Calicivirus Inactivation by Nonionizing (253.7-Nanometer-Wavelength [UV]) and Ionizing (Gamma) Radiation. Appl. Environ. Microbiol. 2004, 70, 5089–5093. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Mauser, A.; Chao, S.F.; Remington, K.; Treckmann, R.; Kaiser, K.; Pifat, D.; Hotta, J.A. Virus inactivation and protein recovery in a novel ultraviolet-C reactor. Vox Sang. 2004, 86, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Simonet, J.; Gantzer, C. Inactivation of Poliovirus 1 and F-Specific RNA Phages and Degradation of Their Genomes by UV Irradiation at 254 Nanometers Downloaded from. Appl. Environ. Microbiol. 2006, 72, 7671–7677. [Google Scholar] [CrossRef] [Green Version]
- Butkus, M.A.; Labare, M.P.; Starke, J.A.; Moon, K.; Talbot, M. Use of Aqueous Silver To Enhance Inactivation of Coliphage MS-2 by UV Disinfection. Appl. Environ. Microbiol. 2004, 70, 2848–2853. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sommer, R.; Pribil, W.; Appelt, S.; Gehringer, P.; Eschweiler, H.; Leth, H.; Cabaj, A.; Haider, T. Inactivation of bacteriophages in water by means of non-ionizing (UV-253.7nm) and ionizing (gamma) radiation: A comparative approach. Water Res. 2001, 35, 3109–3116. [Google Scholar] [CrossRef]
- Lee, J.; Zoh, K.; Ko, G. Inactivation and UV Disinfection of Murine Norovirus with TiO 2 under Various Environmental Conditions. Appl. Environ. Microbiol. 2008, 74, 2111–2117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nishisaka-Nonaka, R.; Mawatari, K.; Yamamoto, T.; Kojima, M.; Shimohata, T.; Uebanso, T.; Nakahashi, M.; Emoto, T.; Akutagawa, M.; Kinouchi, Y.; et al. Irradiation by ultraviolet light-emitting diodes inactivates influenza a viruses by inhibiting replication and transcription of viral RNA in host cells. J. Photochem. Photobiol. B Biol. 2018, 189, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Josewin, S.W.; Kim, M.J.; Yuk, H.G. Inactivation of Listeria monocytogenes and Salmonella spp. on cantaloupe rinds by blue light emitting diodes (LEDs). Food Microbiol. 2018, 76, 219–225. [Google Scholar] [CrossRef]
- Li, X.; Kim, M.J.; Yuk, H.G. Influence of 405 nm light-emitting diode illumination on the inactivation of Listeria monocytogenes and Salmonella spp. on ready-to-eat fresh salmon surface at chilling storage for 8 h and their susceptibility to simulated gastric fluid. Food Control 2018, 88, 61–68. [Google Scholar] [CrossRef]
- Endarko, E.; Maclean, M.; Timoshkin, I.V.; MacGregor, S.J.; Anderson, J.G. High-intensity 405 nm light inactivation of Listeria monocytogenes. Photochem. Photobiol. 2012, 88, 1280–1286. [Google Scholar] [CrossRef]
- Paskeviciute, E.; Zudyte, B.; Luksiene, Z. Innovative nonthermal technologies: Chlorophyllin and visible light significantly reduce microbial load on Basil. Food Technol. Biotechnol. 2019, 57, 126–132. [Google Scholar] [CrossRef] [Green Version]
- Stephen Guffey, J.; Stephen, J.; Professor, A. The Use of 405nm and 464nm Blue Light to Inhibit Listeria monocytogenes in Ready-to-Eat (RTE) Meat. Eur. J. Acad. Essays ISSN 2016, 3, 76–80. [Google Scholar]
- Aurum, F.S.; Nguyen, L.T. Efficacy of photoactivated curcumin to decontaminate food surfaces under blue light emitting diode. J. Food Process. Eng. 2019, 42. [Google Scholar] [CrossRef]
- Srimagal, A.; Ramesh, T.; Sahu, J.K. Effect of light emitting diode treatment on inactivation of Escherichia coli in milk. LWT Food Sci. Technol. 2016, 71, 378–385. [Google Scholar] [CrossRef]
- MacLean, M.; Murdoch, L.E.; MacGregor, S.J.; Anderson, J.G. Sporicidal effects of high-intensity 405 nm visible light on endospore-forming bacteria. Photochem. Photobiol. 2013, 89, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Dai, T.; Gupta, A.; Murray, C.K.; Vrahas, M.S.; Tegos, G.P.; Hamblin, M.R. Blue light for infectious diseases: Propionibacterium acnes, Helicobacter pylori, and beyond? Drug Resist. Updat. 2012, 15, 223–236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bumah, V.V.; Aboualizadeh, E.; Masson-Meyers, D.S.; Eells, J.T.; Enwemeka, C.S.; Hirschmugl, C.J. Spectrally resolved infrared microscopy and chemometric tools to reveal the interaction between blue light (470 nm) and methicillin-resistant Staphylococcus aureus. J. Photochem. Photobiol. B Biol. 2017, 167, 150–157. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.J.; Mikš-Krajnik, M.; Kumar, A.; Yuk, H.G. Inactivation by 405 ± 5 nm light emitting diode on Escherichia coli O157: H7, Salmonella Typhimurium, and Shigella sonnei under refrigerated condition might be due to the loss of membrane integrity. Food Control 2016, 59, 99–107. [Google Scholar] [CrossRef]
- Wu, J.; Chu, Z.; Ruan, Z.; Wang, X.; Dai, T.; Hu, X. Changes of Intracellular Porphyrin, Reactive Oxygen Species, and Fatty Acids Profiles During Inactivation of Methicillin-Resistant Staphylococcus aureus by Antimicrobial Blue Light. Front. Physiol. 2018, 9. [Google Scholar] [CrossRef]
- Kim, M.J.; Bang, W.S.; Yuk, H.G. 405 ± 5 nm light emitting diode illumination causes photodynamic inactivation of Salmonella spp. on fresh-cut papaya without deterioration. Food Microbiol. 2017, 62, 124–132. [Google Scholar] [CrossRef]
- Maclean, M.; McKenzie, K.; Anderson, J.G.; Gettinby, G.; MacGregor, S.J. 405 Nm Light Technology for the Inactivation of Pathogens and Its Potential Role for Environmental Disinfection and Infection Control. J. Hosp. Infect. 2014, 88, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Wainwright, M. Photoinactivation of viruses. Photochem. Photobiol. Sci. 2004, 3, 406–411. [Google Scholar] [CrossRef]
- Luksiene, Z.; Zukauskas, A. Prospects of photosensitization in control of pathogenic and harmful micro-organisms. J. Appl. Microbiol. 2009, 107, 1415–1424. [Google Scholar] [CrossRef]
- Tomb, R.M.; White, T.A.; Coia, J.E.; Anderson, J.G.; MacGregor, S.J.; Maclean, M. Review of the Comparative Susceptibility of Microbial Species to Photoinactivation Using 380–480 nm Violet-Blue Light. Photochem. Photobiol. 2018, 94, 445–458. [Google Scholar] [CrossRef] [Green Version]
- Bumah, V.V.; Masson-Meyers, D.S.; Tong, W.; Castel, C.; Enwemeka, C.S. Optimizing the bactericidal effect of pulsed blue light on Propionibacterium acnes—A correlative fluorescence spectroscopy study. J. Photochem. Photobiol. B Biol. 2020, 202. [Google Scholar] [CrossRef] [PubMed]
- Enwemeka, C.; Bumah, V.; Masson-Meyers, D.; Castel, D.; Castel, C. Optimizing the antimicrobial efficacy of pulsed 450-nm light on Propionibacterium acnes through correlation with fluorescence spectroscopy. Photonics Dermatol. Plastic Surg. 2019, 12. [Google Scholar] [CrossRef]
- Gillespie, J.B.; Maclean, M.; Given, M.J.; Wilson, M.P.; Judd, M.D.; Timoshkin, I.V.; Macgregor, S.J. Efficacy of Pulsed 405-nm Light-Emitting Diodes for Antimicrobial Photodynamic Inactivation: Effects of Intensity, Frequency, and Duty Cycle. Photomed. Laser Surg. 2017, 35, 150–156. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tomb, R.M.; Maclean, M.; Coia, J.E.; Graham, E.; McDonald, M.; Atreya, C.D.; MacGregor, S.J.; Anderson, J.G. New Proof-of-Concept in Viral Inactivation: Virucidal Efficacy of 405 nm Light Against Feline Calicivirus as a Model for Norovirus Decontamination. Food Environ. Virol. 2017, 9, 159–167. [Google Scholar] [CrossRef] [Green Version]
- Tomb, R.M.; Maclean, M.; Herron, P.R.; Hoskisson, P.A.; MacGregor, S.J.; Anderson, J.G. Inactivation of Streptomyces phage ϕC31 by 405 nm light. Bacteriophage 2014, 4, e32129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ho, T.; Kim, A.; Kim, N.; Jin Roh, H.; Tho Ho, D.; Chun, W.-K.; Lee, Y.; Kim, D.-H. Effect of blue light emitting diode on viral hemorrhagic septicemia in olive flounder (Paralichthys olivaceus). Aquaculture 2020, 521, 735019. [Google Scholar] [CrossRef]
- Wu, J.; Hou, W.; Cao, B.; Zuo, T.; Xue, C.; Leung, A.W.; Xu, C.; Tang, Q.J. Virucidal efficacy of treatment with photodynamically activated curcumin on murine norovirus bio-accumulated in oysters. Photodiagnosis Photodyn. Ther. 2015, 12, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Richardson, T.B.; Porter, C.D. Inactivation of murine leukaemia virus by exposure to visible light. Virology 2005, 341, 321–329. [Google Scholar] [CrossRef] [Green Version]
- Tsen, K.T.; Tsen, S.W.D.; Chang, C.L.; Hung, C.F.; Wu, T.C.; Kiang, J.G. Inactivation of viruses by coherent excitations with a low power visible femtosecond laser. Virol. J. 2007, 4, 50. [Google Scholar] [CrossRef] [Green Version]
- Tsen, S.W.D.; Kingsley, D.H.; Poweleit, C.; Achilefu, S.; Soroka, D.S.; Wu, T.C.; Tsen, K.T. Studies of inactivation mechanism of non-enveloped icosahedral virus by a visible ultrashort pulsed laser. Virol. J. 2014, 11, 20. [Google Scholar] [CrossRef]
- Tsen, K.T.; Tsen, S.-W.D.; Fu, Q.; Lindsay, S.M.; Li, Z.; Cope, S.; Vaiana, S.; Kiang, J.G. Studies of inactivation of encephalomyocarditis virus, M13 bacteriophage, and Salmonella typhimurium by using a visible femtosecond laser: Insight into the possible inactivation mechanisms. J. Biomed. Opt. 2011, 16, 078003. [Google Scholar] [CrossRef] [PubMed]
- Berchtikou, A.; Greschner, A.A.; Tijssen, P.; Gauthier, M.A.; Ozaki, T. Accelerated inactivation of M13 bacteriophage using millijoule femtosecond lasers. J. Biophotonics 2020, 13. [Google Scholar] [CrossRef] [PubMed]
- Tsen, S.-W.D.; Chapa, T.; Beatty, W.; Tsen, K.-T.; Yu, D.; Achilefu, S. Inactivation of enveloped virus by laser-driven protein aggregation. J. Biomed. Opt. 2012, 17, 128002. [Google Scholar] [CrossRef] [PubMed]
- Kingsley, D.H.; Perez-Perez, R.E.; Boyd, G.; Sites, J.; Niemira, B.A. Evaluation of 405-nm monochromatic light for inactivation of Tulane virus on blueberry surfaces. J. Appl. Microbiol. 2018, 124, 1017–1022. [Google Scholar] [CrossRef] [PubMed]
- Idil, O.; Darcan, C.; Ozen, T.; Ozkanca, R. The effect of UV-A and various visible light wavelengths radiations on expression level of Escherichia coli oxidative enzymes in seawater. Jundishapur J. Microbiol. 2013, 6, 230–236. [Google Scholar] [CrossRef] [Green Version]
- Lian, C.; Piksa, M.; Yoshida, K.; Persheyev, S.; Pawlik, K.J.; Matczyszyn, K.; Samuel, I.D.W. Flexible organic light-emitting diodes for antimicrobial photodynamic therapy. npj Flex. Electron. 2019, 3, 1–6. [Google Scholar] [CrossRef]
- Angarano, V.; Smet, C.; Akkermans, S.; Watt, C.; Chieffi, A.; Van Impe, J.F.M. Visible light as an antimicrobial strategy for inactivation of pseudomonas fluorescens and staphylococcus epidermidis biofilms. Antibiotics 2020, 9, 171. [Google Scholar] [CrossRef] [Green Version]
- Floyd, R.A.; Schneider, J.E.; Dittmer, D.P. Methylene blue photoinactivation of RNA viruses. Antiviral Res. 2004, 61, 141–151. [Google Scholar] [CrossRef]
- Mohr, H. Virus Inactivation of Plasma By Methylene Blue/Light Exposure. Pediatr. Res. 1999, 45, 946. [Google Scholar] [CrossRef] [Green Version]
- Elikaei, A.; Sharifi, Z.; Hosseini, S.M.; Latifi, H.; Musavi Hosseini, M.K. Inactivation of model viruses suspended in fresh frozen plasma using novel methylene blue based device. Iran. J. Microbiol. 2014, 6, 41–45. [Google Scholar]
- Lee, D.H.; Blajchman, M.A. Platelet substitutes and novel methods of platelet preservation. Platelets 2007, 1297–1309. [Google Scholar] [CrossRef]
- Lin, L.; Hanson, C.V.; Alter, H.J.; Jauvin, V.; Bernard, K.A.; Murthy, K.K.; Metzel, P.; Corash, L. Inactivation of viruses in platelet concentrates by photochemical treatment with amotosalen and long-wavelength ultraviolet light. Transfusion 2005, 45, 580–590. [Google Scholar] [CrossRef]
- Hashem, A.M.; Hassan, A.M.; Tolah, A.M.; Alsaadi, M.A.; Abunada, Q.; Damanhouri, G.A.; El-Kafrawy, S.A.; Picard-Maureau, M.; Azhar, E.I.; Hindawi, S.I. Amotosalen and ultraviolet A light efficiently inactivate MERS-coronavirus in human platelet concentrates. Transfus. Med. 2019, 29, 434–441. [Google Scholar] [CrossRef] [Green Version]
- Hindawi, S.I.; Hashem, A.M.; Damanhouri, G.A.; El-Kafrawy, S.A.; Tolah, A.M.; Hassan, A.M.; Azhar, E.I. Inactivation of Middle East respiratory syndrome-coronavirus in human plasma using amotosalen and ultraviolet A light. Transfusion 2018, 58, 52–59. [Google Scholar] [CrossRef]
- Tonnetti, L.; Proctor, M.C.; Reddy, H.L.; Goodrich, R.P.; Leiby, D.A. Evaluation of the Mirasol platelet reduction technology system against Babesia microti in apheresis platelets and plasma. Transfusion 2010, 50, 1019–1027. [Google Scholar] [CrossRef]
- Keil, S.D.; Bowen, R.; Marschner, S. Inactivation of Middle East respiratory syndrome coronavirus (MERS-CoV) in plasma products using a riboflavin-based and ultraviolet light-based photochemical treatment. Transfusion 2016, 56, 2948–2952. [Google Scholar] [CrossRef] [Green Version]
- Keil, S.D.; Ragan, I.; Yonemura, S.; Hartson, L.; Dart, N.K.; Bowen, R. Inactivation of severe acute respiratory syndrome coronavirus 2 in plasma and platelet products using a riboflavin and ultraviolet light-based photochemical treatment. Vox Sang. 2020. [Google Scholar] [CrossRef]
- Ragan, I.; Hartson, L.; Pidcoke, H.; Bowen, R.; Goodrich, R. Pathogen reduction of SARS-CoV-2 virus in plasma and whole blood using riboflavin and UV light. PLoS ONE 2020, 15, e0233947. [Google Scholar] [CrossRef]
- Wiehe, A.; O’brien, J.M.; Senge, M.O. Trends and Targets in Antiviral Phototherapy. Photochem. Photobiol. Sci. 2018, 17. [Google Scholar] [CrossRef]
- Terrier, O.; Essere, B.; Yver, M.; Barthélémy, M.; Bouscambert-Duchamp, M.; Kurtz, P.; Vanmechelen, D.; Morfin, F.; Billaud, G.; Ferraris, O.; et al. ARTICLE IN PRESS G Model Cold oxygen plasma technology efficiency against different airborne respiratory viruses. J. Clin. Virol. 2009, 45, 119–124. [Google Scholar] [CrossRef]
- Li, D.; Baert, L.; De Jonghe, M.; Van Coillie, E.; Ryckeboer, J.; Devlieghere, F.; Uyttendaele, M. Inactivation of Murine Norovirus 1, Coliphage X174, and Bacillus fragilis Phage B40-8 on Surfaces and Fresh-Cut Iceberg Lettuce by Hydrogen Peroxide and UV Light. Appl. Environ. Microbiol. 2011, 77, 1399–1404. [Google Scholar] [CrossRef] [Green Version]
- Xie, Y.; Hajdok, C.; Mittal, G.S.; Warriner, K. Inactivation of MS2 F(+) coliphage on lettuce by a combination of UV light and hydrogen peroxide. J. Food Prot. 2008, 71, 903–907. [Google Scholar] [CrossRef]
- Kim, S.H.; Shahbaz, H.M.; Park, D.; Chun, S.; Lee, W.; Oh, J.W.; Lee, D.U.; Park, J. A combined treatment of UV-assisted TiO2 photocatalysis and high hydrostatic pressure to inactivate internalized murine norovirus. Innov. Food Sci. Emerg. Technol. 2017, 39, 188–196. [Google Scholar] [CrossRef]
- Akgün, M.P.; Ünlütürk, S. Effects of ultraviolet light emitting diodes (LEDs) on microbial and enzyme inactivation of apple juice. Int. J. Food Microbiol. 2017, 260, 65–74. [Google Scholar] [CrossRef] [PubMed]

| Virus | Initial Titre | Reduction | Light Dosage a | Light Source | Reference |
|---|---|---|---|---|---|
| MS2 bacteriophage * | 2 × 108 to 7 × 108 plaque-forming units (PFU)/mL | 1 log 2 log | 0.34–0.42 mJ/cm2 0.80–0.91 mJ/cm2 | low-pressure, mercury-discharge UV-C lamps (254 nm; 60–253.7 µW/cm2) | [97] |
| MS2 bacteriophage * MHV 1,** | 104 PFU/mL | <1 log <1 log | 2.6 mJ/cm2 0.6 mJ/cm2 | UV-C lamps (254 nm; 36 W) | [115] |
| PRRSV 2,*** | ca. 3.33 × 106 TCID50/mL | 3 log | 1.21 mJ/cm2 | low-pressure, mercury-vapor discharge UV-C lamps (254 nm; 110 V) | [106] |
| Influenza A *** | 5.62 × 103 focus forming units (FFU)/L air | 1.4 log | ca. 1.48 mJ/cm2 | low-pressure, mercury UV-C lamps (254 nm; 36 W) | [107] |
| Influenza A *** | 108 FFU/ml | 1.3 log | 2 mJ/cm2 | KrCl lamps (222 nm; 120 µW/cm2) | [84] |
| HCoV 229E ** HCoV OC43 ** | 107 to 108 TCID50/mL | 3 log 4 log | 1.70 mJ/cm2 1.56 mJ/cm2 | KrCl excimer lamps (222 nm; 0.5–2 mJ/cm2) | [85] |
| Virus | Initial Titre | Reduction | Light Dosage a | Light Source | Reference |
|---|---|---|---|---|---|
| Sindbis *** HAV 1,* EMCV 2,* | not available (NA) | 2.7 log to >7.2 log (no protein); 1.1 log to 5 log (protein) 3.8 log to >5.7 log (no protein); 1.3 log to >5.6 log (protein) 3.1 log to >5.9 log (no protein); 1.2 log to >6.1 log (protein) | 0.25–2 J/cm2 | Xenon lamp—pulsed white light (200–1100 nm; 300 µs) | [121] § |
| SARS-CoV ** | 7 × 106 PFU/mL | 6.85 log | 0.32 J/cm2 | UV-C (260 nm; >90 µW/cm2; 80 cm; 60 min) | [124] |
| MS2 bacteriophage * FCV 3,* | NA | 4 log | 0.12 J/cm2 0.04 J/cm2 | low-pressure, mercury-vapour lamp (254 nm; 8 W) | [125] |
| SARS-CoV ** | 106 TCID50/mL | 4 log | 1.45 J/cm2 | UV-C (254 nm; 4016 µm/cm2; 3 cm; 6 min) | [126] |
| BVDV 4,*** | 104.3 TCID50/mL | Complete inactivation (below the limit of detection of 1 log TCID50/mL) | 1.6 J/cm2 (no protein); 3.2 J/cm2 (protein) 127 J/cm2 (no protein); 190.5 J/cm2 (protein) | low-pressure, mercury lamp (254 nm; 40 W) nanosecond laser (266 nm; 8 ns per pulse) | [127] |
| HAV 1,* Murine NoV-1 5,* | 106 PFU/mL | 2.8 log to 4.8 log (no protein); 2.1 to 2.4 log (protein) 5 log (no protein); 2.8 to 4.1 log (protein) | 0.05 to 0.15 J/cm2 | xenon lamp—pulsed white light (200 to 1100 nm; 2 s) | [128] § |
| Murine NoV-1 5,* | 105 PFU/mL | 3.3 log | 2.07 J/cm2 | pulsed white light (200 to 1000 nm; 2 s) | [129] § |
| Murine NoV-1 5,* Tulane virus * | 106 PFU/mL 106 PFU/mL | 1.3 log to 5.8 log 1 log to 6 log | 0.31 to 4.94 J/cm2 | pulsed broad white light (180–1100 nm; 40% UV; 6 s) | [130] |
| Qβ bacteriophage * | NA | 3 log | 0.03 to 0.5 J/cm2 | UV LED (265, 280 and 300 nm; 0.99 and 1.01 mW/cm2) | [131] |
| MS2 bacteriophage * | 106 PFU/mL | ca. 1.5 log | 0.02 J/cm2 | UV LED (265 nm; 10 mW) | [132] |
| HCV 6,*** BVDV 4,*** | 105.61 TCID50/mL 106.25 TCID50/mL | 5 log 2.7 log | 0.2 J/cm2 0.2 J/cm2 | THERAFLEX UV-Platelets system (254 nm) | [133] |
| MERS-CoV ** Ebola virus *** | 106.37 to 106.43 TCID50/mL 106.84 to 106.96 TCID50/mL | 3.7 log to 3.8 log 4.5 log to 4.6 log | 0.15 J/cm2 0.15 J/cm2 | THERAFLEX UV-Platelets system (254 nm) | [134] |
| SARS-CoV ** CCHFV 7,*** Nipah virus *** | 105.8 to 106 TCID50/mL 104.6 to 105.2 TCID50/mL 106.2 to 106.5 TCID50/mL | 3.4 log to 3.6 log 2.2 log to 2.9 log 4.3 log to 4.6 log | 0.15 J/cm2 0.15 J/cm20.15 J/cm2 | THERAFLEX UV-Platelets system (254 nm) | [135] |
| SARS-CoV-2 ** | 3.7 × 104 PFU/mL | 3.33 log | 75 mJ/cm2 | Deep UV LED (280 nm) | [136] |
| SARS-CoV-2 ** | NA | ca. 5.7 log | 16 mJ/cm2 | low-pressure, mercury lamp (254 nm; 1.08 mW/cm2) | [137] |
| Virus | Initial Titre | Reduction | Light Dosage a | Light Source | Reference |
|---|---|---|---|---|---|
| SARS-CoV ** | 2.65 × 107 PFU/mL | 4 log | 0.12 J/cm2 | UV-C (254 nm; 134 µW/cm2) | [161] |
| MS2 bacteriophage * | 108 PFU/mL | 1 log | 3.20 mJ/cm2 | UV-C lamp (254 nm; 60–240 µW/cm2; 3 s to 6 min) | [166] |
| HAV 1,* FCV 2,* Aichi virus * | 6.93 × 106 to 6.93 × 108 PFU/mL | 2.44 log to 5.42 log 2.12 log to 4.46 log 1.71 log to 4.43 log | 0.24 J/cm2 | low pressure lamp (254 nm) | [163] |
| MS2 bacteriophage * | 107 PFU/mL | 3 log to 4 log (below detection limit of 1000 PFU/mL) | 4.32 to 7.20 J/cm2 | low-pressure, mercury-arc lamp (254 nm; 40 W) | [158] |
| HAV 1,* Murine NoV-1 3,* | 105 PFU/mL | 5 log 5 log (no protein); 2.3 log to 3.6 log (protein) | 0.09 J/cm2 0.06 J/cm2 | xenon lamp—pulsed white light (200 to 1100 nm; 2–3 s) | [128] § |
| Influenza A virus (H1N1) *** | 6.98 × 107 PFU/mL | 4.08 log to 5.75 log (PD 4 = 15 µm); 4.83 log to 5.08 log (PD 4 = 0.8 µm) | 1.8 J/cm2 | UV-C lamps (254 nm; 80 W) | [157] |
| Influenza A virus (H5N1) *** | 2.20 × 105 PFU/mL | 4.5 log | 1.8 J/cm2 | UV-C lamps (254 nm; 15 W) | [156] |
| MS2 bacteriophage * | 6.31 × 107 PFU/mL | 4.87 log (glass beads); 0.64 log (powdered black pepper); 0.12 log (garlic); 0.68 log (chopped mint) | 9.6 J/cm2 | pulsed white light (200–1100 nm; 0.96 J/cm2 per pulse; 1 s) | [164] § |
| Murine NoV-1 3,* | 105 PFU/mL | 3 log (no protein); 2.6 log to 3 log (protein) | 3.45 J/cm2 (no protein); 7.60 J/cm2 (protein) | pulsed white light (200 to 1000 nm; 2 to 6 s) | [129] |
| Virus | D90 (mJ/cm2) | Genome Size (kb) | Matrix | Reference |
|---|---|---|---|---|
| MS2 bacteriophage | 0.28 | 3.57 | Airborne | [97] |
| Influenza A (H1N1) | 1.04 | 13.50 | Airborne | [107] |
| Influenza A (H1N1) | 1.28 (222 nm) | 13.50 | Airborne | [84] |
| Human coronavirus 229E | 0.56 (222 nm) | 27.32 | Airborne | [85] |
| Human coronavirus OC43 | 0.39 (222 nm) | 30.74 | Airborne | [85] |
| Feline calicivirus | 6 | 7.58 | Liquid | [125] |
| Feline calicivirus | 47.85 | 7.68 | Liquid | [168] |
| Feline calicivirus | 5 | 7.68 | Liquid | [169] |
| Hepatitis A virus | 36.50 | 7.48 | Liquid | [168] |
| Hepatitis A virus | 4.5 | 7.48 | Liquid | [170] |
| Sindbis | 10 | 11.7 | Liquid | [170] |
| Poliovirus-1 | 24.10 | 7.44 | Liquid | [168] |
| Poliovirus-1 | 9.04 | 7.44 | Liquid | [171] |
| MS2 bacteriophage | 22.22 | 3.57 | Liquid | [169] |
| MS2 bacteriophage | 24.39 | 3.57 | Liquid | [172] |
| MS2 bacteriophage | 23 | 3.57 | Liquid | [125] |
| MS2 bacteriophage | 23.04 | 3.57 | Liquid | [168] |
| MS2 bacteriophage | 25.71 | 3.57 | Liquid | [171] |
| MS2 bacteriophage | 18.60 | 3.57 | Liquid | [173] |
| Qβ bacteriophage | 16.81 | 4.16 | Liquid | [171] |
| Qβ bacteriophage | 11.75 | 4.16 | Liquid | [131] |
| Qβ bacteriophage | 10.20 (265 nm) | 4.16 | Liquid | [131] |
| Qβ bacteriophage | 17.86 (280 nm) | 4.16 | Liquid | [131] |
| Qβ bacteriophage | 166.67 (300 nm) | 4.16 | Liquid | [131] |
| Murine norovirus-1 | 10 | 7.38 | Liquid | [174] |
| Influenza A (H1N1) | 6400 (365 nm) | 13.50 | Liquid | [175] |
| Influenza A (H1N1) | 260 (310 nm) | 13.50 | Liquid | [175] |
| Influenza A (H1N1) | 28 (280 nm) | 13.50 | Liquid | [175] |
| MS2 bacteriophage | 3.20 | 3.57 | Semi-solid (gelatin-based medium) | [166] |
| Virus | Initial Titre | Reduction | Light Dosage a | Light Source | Matrix | Reference |
|---|---|---|---|---|---|---|
| Bacteriophage ϕC31 α,* | 103, 105, 107 PFU/mL | 0.3 log (M); 2.7 to 7.1 log (NR); 3 log (PR) | 306.2 J/cm2 (M); 306.2 to 1400 J/cm2 (NR); 612.4 J/cm2 (PR) | blue light LED (405 nm; 2 cm 56.7 mW/cm2) | minimum (M), nutrient-rich (NR) and porphyrin-rich (PR) media | [197] |
| Murine NoV-1 4,* | ca. 108.5 PFU/mL ca. 104 PFU/mL | 1.32 log (5 µm curcumin); >3 log (20 µm curcumin) 0.75 log (10 µm curcumin); 1.15 log (20 µm curcumin) | 3.6 J/cm2 | blue LED (470 nm; 0.06 W/cm2; 60s) | culture medium (5 µm, 10 µm, and 20 µm curcumin) oyster (5 µm, 10 µm, and 20 µm curcumin) | [199] |
| FCV 2,* | 2 × 105 PFU/mL | 3.9 log (minimum); 4.5 to 5.1 log (ORM) | 2804 J/cm2 (M); 421 to 1400 J/cm2 (ORM) | blue light LED (405; 4 cm; 155.8 mW/cm2) | minimum (M) and organically-rich media (ORM) | [196] |
| Tulane virus * | 4 × 107 PFU/mL | 0.06 log (no PS 3); 0.51 to 1.01 log (PS 3) | 7.56 J/cm2 | blue light LED (405 nm; 7.5 cm; 4.2 mW/cm2) | blueberries | [206] |
| VHSV 1,*** | 105 (L15); 105.8 (mucus) PFU/mL | Complete inactivation | 715 J/cm2 | blue light LED (405 nm) | culture medium (L15); Flounder skin mucus | [198] |
| M13 bacteriophage β,* | 103 PFU/mL | Complete inactivation | ≥50 MW/cm2 | femtosecond laser; 425 nm; 40 mW; 10 h | water | [201] |
| MCMV 5,α,*** | 2 × 107 PFU/mL | 5 log | 810 J | femtosecond laser; 425 nm; 150 mW; 1.5 h | culture medium | [205] |
| Murine NoV-1 4,* | 5 × 107 PFU/mL | 3 log | ≥80 MW/cm2 | femtosecond laser; 425 nm; 120 mWs; 2 h | culture medium | [202] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hadi, J.; Dunowska, M.; Wu, S.; Brightwell, G. Control Measures for SARS-CoV-2: A Review on Light-Based Inactivation of Single-Stranded RNA Viruses. Pathogens 2020, 9, 737. https://doi.org/10.3390/pathogens9090737
Hadi J, Dunowska M, Wu S, Brightwell G. Control Measures for SARS-CoV-2: A Review on Light-Based Inactivation of Single-Stranded RNA Viruses. Pathogens. 2020; 9(9):737. https://doi.org/10.3390/pathogens9090737
Chicago/Turabian StyleHadi, Joshua, Magdalena Dunowska, Shuyan Wu, and Gale Brightwell. 2020. "Control Measures for SARS-CoV-2: A Review on Light-Based Inactivation of Single-Stranded RNA Viruses" Pathogens 9, no. 9: 737. https://doi.org/10.3390/pathogens9090737
APA StyleHadi, J., Dunowska, M., Wu, S., & Brightwell, G. (2020). Control Measures for SARS-CoV-2: A Review on Light-Based Inactivation of Single-Stranded RNA Viruses. Pathogens, 9(9), 737. https://doi.org/10.3390/pathogens9090737







