Novel Coronavirus: Current Understanding of Clinical Features, Diagnosis, Pathogenesis, and Treatment Options
Abstract
1. Introduction
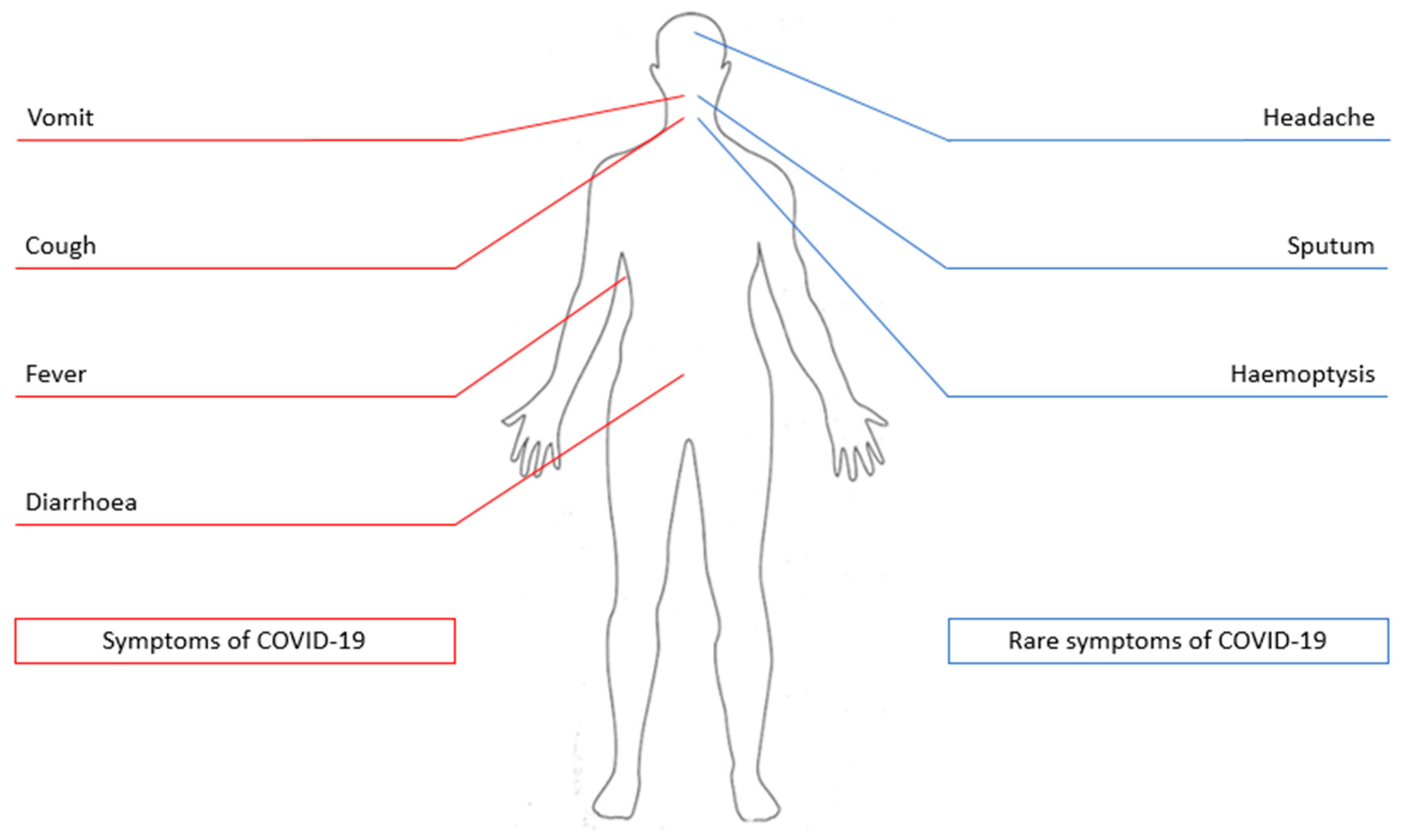
2. Clinical Aspects
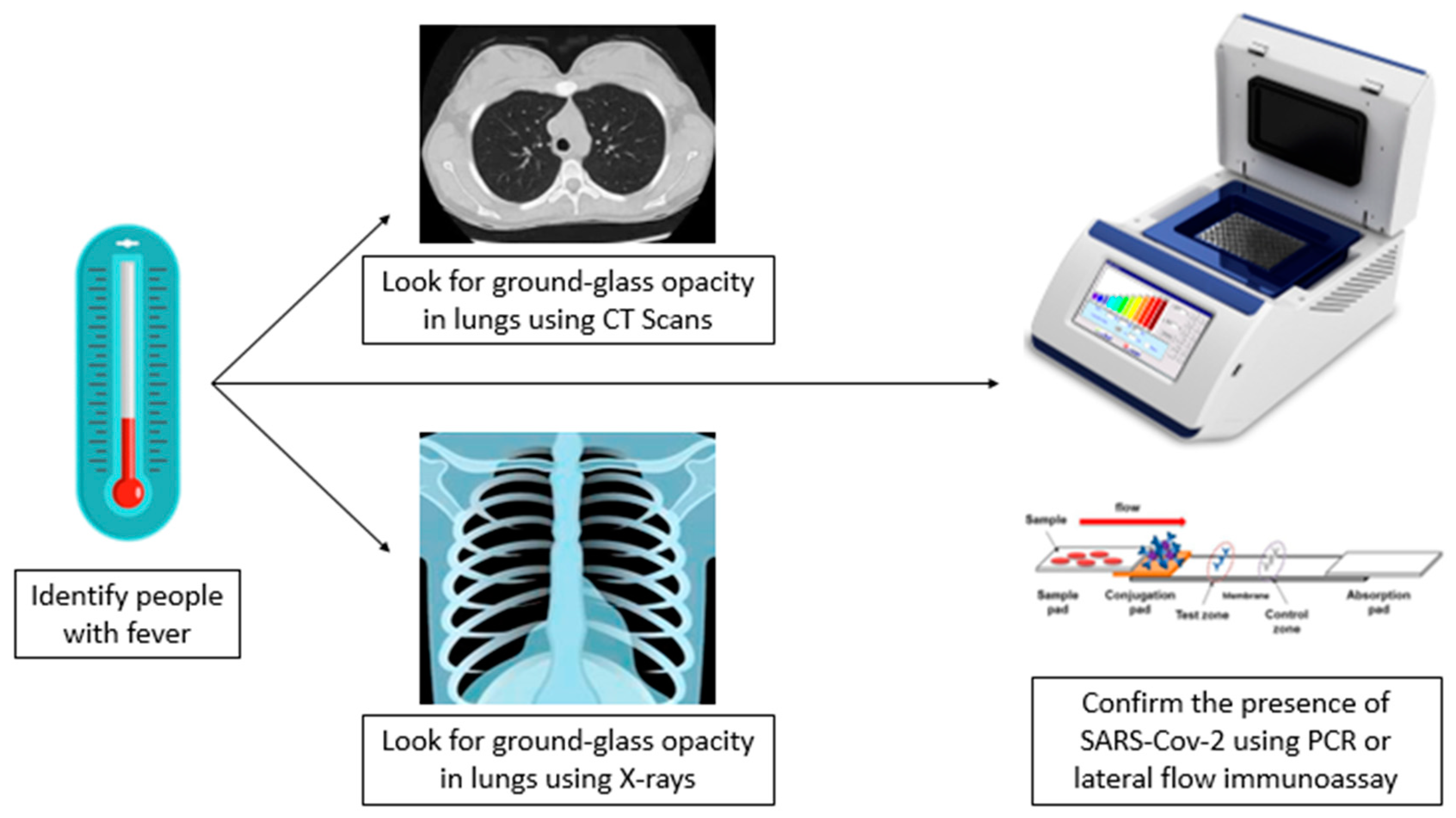
3. Diagnosis
4. Mode of Transmission
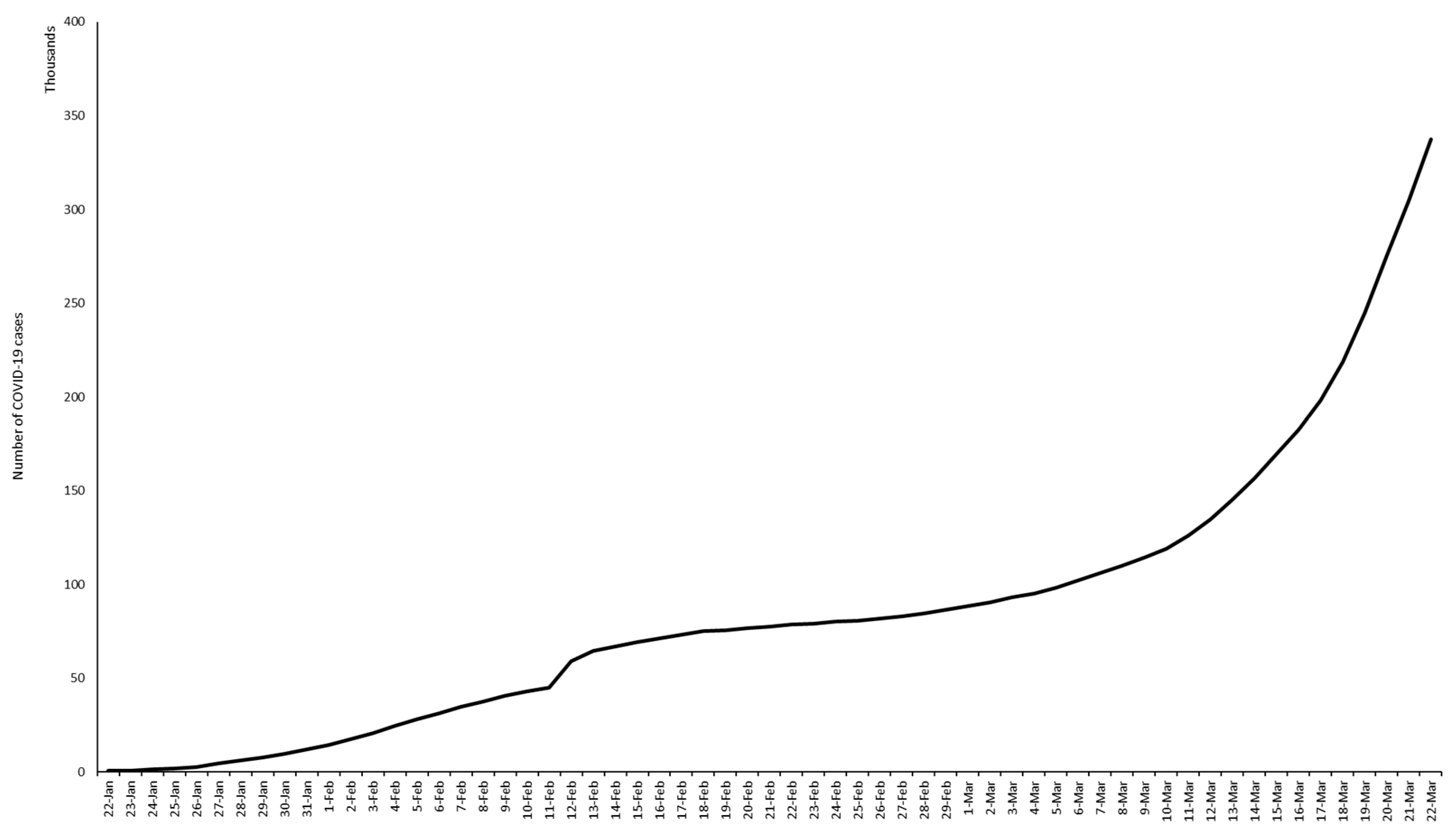
5. Epidemiology
6. Pathogenesis
7. Treatment
8. Developing a Vaccine
9. Developing Drugs
10. Challenges Faced due to SARS-CoV-2
11. Future Studies
12. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Wu, Z.; McGoogan, J.M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: Summary of a report of 72314 cases from the Chinese Center for Disease Control and Prevention. JAMA 2020, 323, 1239–1242. [Google Scholar] [CrossRef]
- WHO. WHO Director-General’s opening remarks at the media briefing on COVID-19—11 March 2020. World Health Organ. 2020. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020 (accessed on 24 March 2020).
- Chan, J.F.W.; Lau, S.K.P.; Woo, P.C.Y. The emerging novel Middle East respiratory syndrome coronavirus: The “knowns” and “unknowns”. J. Formos. Med. Assoc. 2013, 112, 372–381. [Google Scholar] [CrossRef]
- Weiss, S.R.; Leibowitz, J.L. Coronavirus pathogenesis. In Advances in Virus Research; Academic Press: Cambridge, MA, USA, 2011; Volume 81, pp. 85–164. [Google Scholar]
- Zumla, A.; Chan, J.F.; Azhar, E.I.; Hui, D.S.; Yuen, K.Y. Coronaviruses—Drug discovery and therapeutic options. Nat. Rev. Drug Discov. 2016, 15, 327. [Google Scholar] [CrossRef] [PubMed]
- Channappanavar, R.; Zhao, J.; Perlman, S. T cell-mediated immune response to respiratory coronaviruses. Immunol. Res. 2014, 59, 118–128. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.C.; Lau, S.K.; Woo, P.C.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus as an agent of emerging and reemerging infection. Clin. Microbiol. Rev. 2007, 20, 660–694. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Lau, S.K.; To, K.K.; Cheng, V.C.; Woo, P.C.; Yuen, K.Y. Middle East respiratory syndrome coronavirus: Another zoonotic betacoronavirus causing SARS-like disease. Clin. Microbiol. Rev. 2015, 28, 465–522. [Google Scholar] [CrossRef] [PubMed]
- Zaki, A.M.; Van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Drosten, C.; Günther, S.; Preiser, W.; Van Der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Gretebeck, L.M.; Subbarao, K. Animal models for SARS and MERS coronaviruses. Curr. Opin. Virol. 2015, 13, 123–129. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Cheng, Z.J.; Shan, J. 2019 Novel coronavirus: Where we are and what we know. Infection 2020, 48, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Lam, T.T.Y.; Shum, M.H.H.; Zhu, H.C.; Tong, Y.G.; Ni, X.B.; Liao, Y.S.; Wei, W.; Cheung, W.Y.M.; Li, W.J.; Li, L.F.; et al. Identification of 2019-nCoV related coronaviruses in Malayan pangolins in southern China. BioRxiv 2020. [Google Scholar] [CrossRef]
- Wu, D.; Wu, T.; Liu, Q.; Yang, Z. The SARS-CoV-2 outbreak: What we know. Int. J. Infect. Dis. 2020, 94, 44–48. [Google Scholar] [CrossRef] [PubMed]
- EWP (Epidemiology Working Group for NCIP Epidemic Response CCfDCaP). The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) in China. Chin. J. Epidemiol. 2020, 41, 145. [Google Scholar]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.; Lau, E.H.; Wong, J.Y.; et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Ou, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. MedRxiv 2020. [Google Scholar] [CrossRef]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus–infected pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Pan, Y.; Guan, H.; Zhou, S.; Wang, Y.; Li, Q.; Zhu, T.; Hu, Q.; Xia, L. Initial CT findings and temporal changes in patients with the novel coronavirus pneumonia (2019-nCoV): A study of 63 patients in Wuhan, China. Eur. Radiol. 2020. [Google Scholar] [CrossRef]
- Shi, H.; Han, X.; Jiang, N.; Cao, Y.; Alwalid, O.; Gu, J.; Fan, Y.; Zheng, C. Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: A descriptive study. Lancet Infect. Dis. 2020, 20, 425–434. [Google Scholar] [CrossRef]
- He, F.; Deng, Y.; Li, W. Coronavirus Disease 2019 (COVID-19): What we know? J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Yu, F.; Du, L.; Ojcius, D.M.; Pan, C.; Jiang, S. Measures for diagnosing and treating infections by a novel coronavirus responsible for a pneumonia outbreak originating in Wuhan, China. Microbes Infect. 2020, 22, 74–79. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef]
- Sheridan, C. Fast, Portable Tests Come Online to Curb Coronavirus Pandemic. 2020. Available online: https://www.nature.com/articles/d41587–020-00010-2?utm_source=facebook&utm_medium=social&utm_content=organic&utm_campaign=NGMT_USG_JC01_GL_NRJournals (accessed on 24 March 2020).
- Peeri, N.C.; Shrestha, N.; Rahman, M.S.; Zaki, R.; Tan, Z.; Bibi, S.; Baghbanzadeh, M.; Aghamohammadi, N.; Zhang, W.; Haque, U. The SARS, MERS and novel coronavirus (COVID-19) epidemics, the newest and biggest global health threats: What lessons have we learned? Int. J. Epidemiol. 2020. [Google Scholar] [CrossRef]
- Chan-Yeung, M.; Xu, R.H. SARS: Epidemiology. Respirology 2003, 8, S9–S14. [Google Scholar] [CrossRef]
- Phelan, A.L.; Katz, R.; Gostin, L.O. The novel coronavirus originating in Wuhan, China: Challenges for global health governance. JAMA 2020, 323, 709–710. [Google Scholar] [CrossRef]
- Shen, K.; Yang, Y.; Wang, T.; Zhao, D.; Jiang, Y.; Jin, R.; Zheng, Y.; Xu, B.; Xie, Z.; Lin, L.; et al. Diagnosis, treatment, and prevention of 2019 novel coronavirus infection in children: Experts’ consensus statement. World J. Pediatrics 2020. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First case of 2019 novel coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Zhu, H.; Wang, L.; Fang, C.; Peng, S.; Zhang, L.; Chang, G.; Xia, S.; Zhou, W. Clinical analysis of 10 neonates born to mothers with 2019-nCoV pneumonia. Transl. Pediatrics 2020, 9, 51. [Google Scholar] [CrossRef] [PubMed]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- WMHC. Report of Clustering Pneumonia of Unknown Etiology in Wuhan City. Wuhan Munic. Health Comm. 2019. Available online: http://wjw.wuhan.gov.cn/front/web/showDetail/2019123108989 (accessed on 24 March 2020).
- Worldometers. 2020. Available online: www.worldometers.info (accessed on 24 March 2020).
- He, Y.; Zhou, Y.; Liu, S.; Kou, Z.; Li, W.; Farzan, M.; Jiang, S. Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: Implication for developing subunit vaccine. Biochem. Biophys. Res. Commun. 2004, 324, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [PubMed]
- Spruth, M.; Kistner, O.; Savidis-Dacho, H.; Hitter, E.; Crowe, B.; Gerencer, M.; Brühl, P.; Grillberger, L.; Reiter, M.; Tauer, C.; et al. A double-inactivated whole virus candidate SARS coronavirus vaccine stimulates neutralising and protective antibody responses. Vaccine 2006, 24, 652–661. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.V.; Massare, M.J.; Barnard, D.L.; Kort, T.; Nathan, M.; Wang, L.; Smith, G. Chimeric severe acute respiratory syndrome coronavirus (SARS-CoV) S glycoprotein and influenza matrix 1 efficiently form virus-like particles (VLPs) that protect mice against challenge with SARS-CoV. Vaccine 2011, 29, 6606–6613. [Google Scholar] [CrossRef]
- Fett, C.; DeDiego, M.L.; Regla-Nava, J.A.; Enjuanes, L.; Perlman, S. Complete protection against severe acute respiratory syndrome coronavirus-mediated lethal respiratory disease in aged mice by immunization with a mouse-adapted virus lacking E protein. J. Virol. 2013, 87, 6551–6559. [Google Scholar] [CrossRef]
- He, Y.; Lu, H.; Siddiqui, P.; Zhou, Y.; Jiang, S. Receptor-binding domain of severe acute respiratory syndrome coronavirus spike protein contains multiple conformation-dependent epitopes that induce highly potent neutralizing antibodies. J. Immunol. 2005, 174, 4908–4915. [Google Scholar] [CrossRef]
- Chi, H.; Zheng, X.; Wang, X.; Wang, C.; Wang, H.; Gai, W.; Perlman, S.; Yang, S.; Zhao, J.; Xia, X. DNA vaccine encoding Middle East respiratory syndrome coronavirus S1 protein induces protective immune responses in mice. Vaccine 2017, 35, 2069–2075. [Google Scholar] [CrossRef]
- Martin, J.E.; Louder, M.K.; Holman, L.A.; Gordon, I.J.; Enama, M.E.; Larkin, B.D.; Andrews, C.A.; Vogel, L.; Koup, R.A.; Roederer, M.; et al. A SARS DNA vaccine induces neutralizing antibody and cellular immune responses in healthy adults in a Phase I clinical trial. Vaccine 2008, 26, 6338–6343. [Google Scholar] [CrossRef]
- Modjarrad, K.; Roberts, C.C.; Mills, K.T.; Castellano, A.R.; Paolino, K.; Muthumani, K.; Reuschel, E.L.; Robb, M.L.; Racine, T.; Oh, M.D.; et al. Safety and immunogenicity of an anti-Middle East respiratory syndrome coronavirus DNA vaccine: A phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect. Dis. 2019, 19, 1013–1022. [Google Scholar] [CrossRef]
- Sheahan, T.P.; Sims, A.C.; Leist, S.R.; Schäfer, A.; Won, J.; Brown, A.J.; Montgomery, S.A.; Hogg, A.; Babusis, D.; Clarke, M.O.; et al. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat. Commun. 2020, 11, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Xiao, G.; Chen, Y.; He, Y.; Niu, J.; Escalante, C.R.; Xiong, H.; Farmar, J.; Debnath, A.K.; Tien, P.; et al. Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: Implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet 2004, 363, 938–947. [Google Scholar] [CrossRef]
- Lu, L.; Liu, Q.; Zhu, Y.; Chan, K.H.; Qin, L.; Li, Y.; Wang, Q.; Chan, J.F.W.; Du, L.; Yu, F.; et al. Structure-based discovery of Middle East respiratory syndrome coronavirus fusion inhibitor. Nat. Commun. 2014, 5, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Traggiai, E.; Becker, S.; Subbarao, K.; Kolesnikova, L.; Uematsu, Y.; Gismondo, M.R.; Murphy, B.R.; Rappuoli, R.; Lanzavecchia, A. An efficient method to make human monoclonal antibodies from memory B cells: Potent neutralization of SARS coronavirus. Nat. Med. 2004, 10, 871–875. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Bao, L.; Chen, C.; Zou, T.; Xue, Y.; Li, F.; Lv, Q.; Gu, S.; Gao, X.; Cui, S.; et al. Human neutralizing monoclonal antibody inhibition of middle east respiratory syndrome coronavirus replication in the common marmoset. J. Infect. Dis. 2017, 215, 1807–1815. [Google Scholar] [CrossRef]
- Li, X.; Yu, J.; Zhang, Z.; Ren, J.; Peluffo, A.E.; Zhang, W.; Zhao, Y.; Yan, K.; Cohen, D.; Wang, W. Network Bioinformatics Analysis Provides Insight into Drug Repurposing for COVID-2019. Preprints 2020. [Google Scholar] [CrossRef][Green Version]
- Gordon, D.; Jang, G.; Bouhaddou, M.; Xu, J.; Obernier, K.; O’Meara, M.; Guo, J.; Swaney, D.; Tummino, T.; Huttenhain, R.; et al. A SARS-CoV-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug-Repurposing. bioRxiv 2020. [Google Scholar] [CrossRef]
- Branswell, H. WHO to Launch Multinational Trial to Jumpstart Search for Coronavirus Drugs. 2020. Available online: https://www.statnews.com/2020/03/18/who-to-launch-multinational-trial-to-jumpstart-search-for-coronavirus-drugs/ (accessed on 24 March 2020).
- Kupferschmidt, K.; Cohen, J. WHO Launches Global Megatrial of the Four Most Promising Coronavirus Treatments. 2020. Available online: https://www.sciencemag.org/news/2020/03/who-launches-global-megatrial-four-most-promising-coronavirus-treatments (accessed on 24 March 2020).
- Erol, A. High-dose intravenous vitamin C treatment for COVID-19. OSF Prepr. 2020. [Google Scholar] [CrossRef]
- Qiu, J. Hidden Coronavirus Infections Could Be Seeding New Outbreaks. 2020. Available online: https://www.nature.com/articles/d41586-020-00822-x?utm_source=fbk_nnc&utm_medium=social&utm_campaign=naturenews&sf231783776=1&fbclid=IwAR1aOwVDe-1W5t5oszR6zBxwnEFwdk_bb911DhuiQ78HZdfjR1Qq4-eG-fk (accessed on 24 March 2020).
- Wang, C.; Liu, L.; Hao, X.; Guo, H.; Wang, Q.; Huang, J.; He, N.; Yu, H.; Lin, X.; Pan, A.; et al. Evolving Epidemiology and Impact of Non-pharmaceutical Interventions on the Outbreak of Coronavirus Disease 2019 in Wuhan, China. medRxiv 2020. [Google Scholar] [CrossRef]
- Nishiura, H.; Kobayashi, T.; Suzuki, A.; Jung, S.; Hayashi, K.; Kinoshita, R.; Yang, Y.; Yuan, B.; Akhmetzhanov, A.; Linton, N.; et al. Estimation of the asymptomatic ratio of novel coronavirus infections (COVID-19). Int. J. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Woelfel, R.; Corman, V.; Guggemos, W.; Seilmaier, M.; Zange, S.; Mueller, M.; Niemeyer, D.; Vollmar, P.; Rothe, C.; Hoelscher, M.; et al. Clinical presentation and virological assessment of hospitalized cases of coronavirus disease 2019 in a travel-associated transmission cluster. medRxiv 2020. [Google Scholar] [CrossRef]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N. Engl. J. Med. 2020, 382, 1177–1179. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mungroo, M.R.; Khan, N.A.; Siddiqui, R. Novel Coronavirus: Current Understanding of Clinical Features, Diagnosis, Pathogenesis, and Treatment Options. Pathogens 2020, 9, 297. https://doi.org/10.3390/pathogens9040297
Mungroo MR, Khan NA, Siddiqui R. Novel Coronavirus: Current Understanding of Clinical Features, Diagnosis, Pathogenesis, and Treatment Options. Pathogens. 2020; 9(4):297. https://doi.org/10.3390/pathogens9040297
Chicago/Turabian StyleMungroo, Mohammad Ridwane, Naveed Ahmed Khan, and Ruqaiyyah Siddiqui. 2020. "Novel Coronavirus: Current Understanding of Clinical Features, Diagnosis, Pathogenesis, and Treatment Options" Pathogens 9, no. 4: 297. https://doi.org/10.3390/pathogens9040297
APA StyleMungroo, M. R., Khan, N. A., & Siddiqui, R. (2020). Novel Coronavirus: Current Understanding of Clinical Features, Diagnosis, Pathogenesis, and Treatment Options. Pathogens, 9(4), 297. https://doi.org/10.3390/pathogens9040297






