Characterization of a Novel Peptide from Pathogenic Leptospira and Its Cytotoxic Effect
Abstract
1. Introduction
2. Results
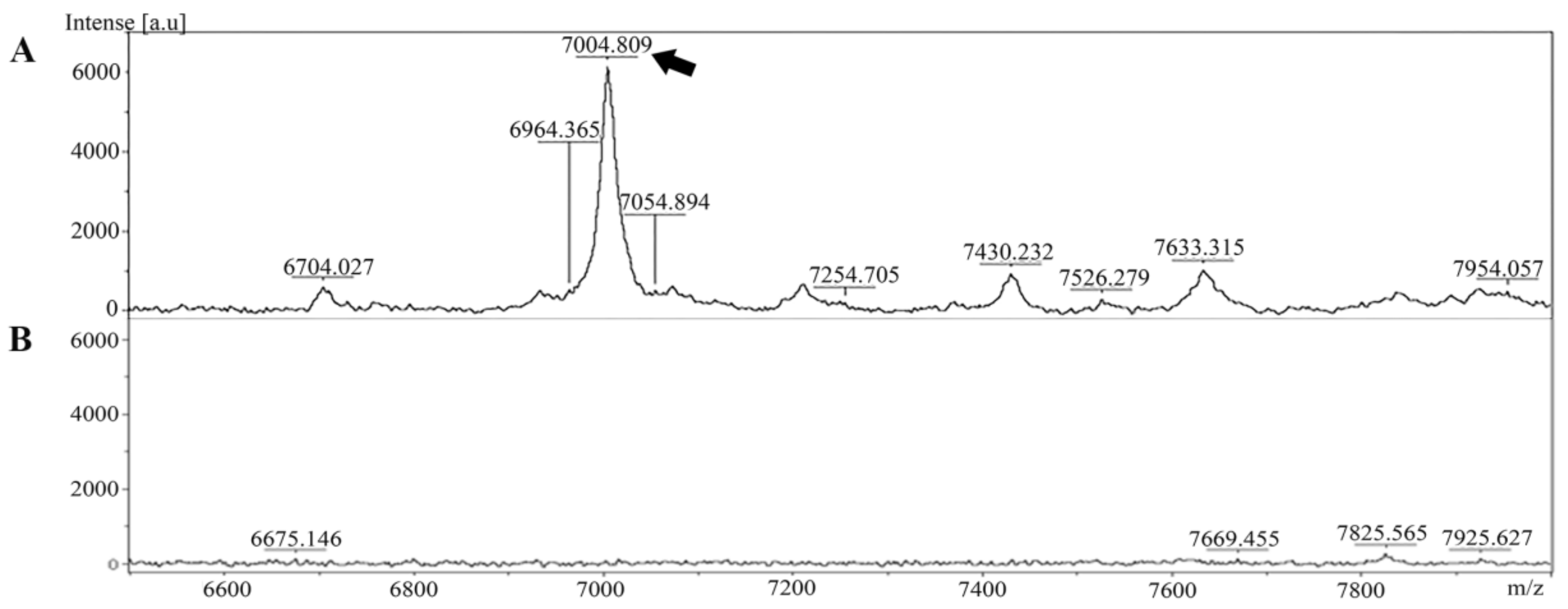
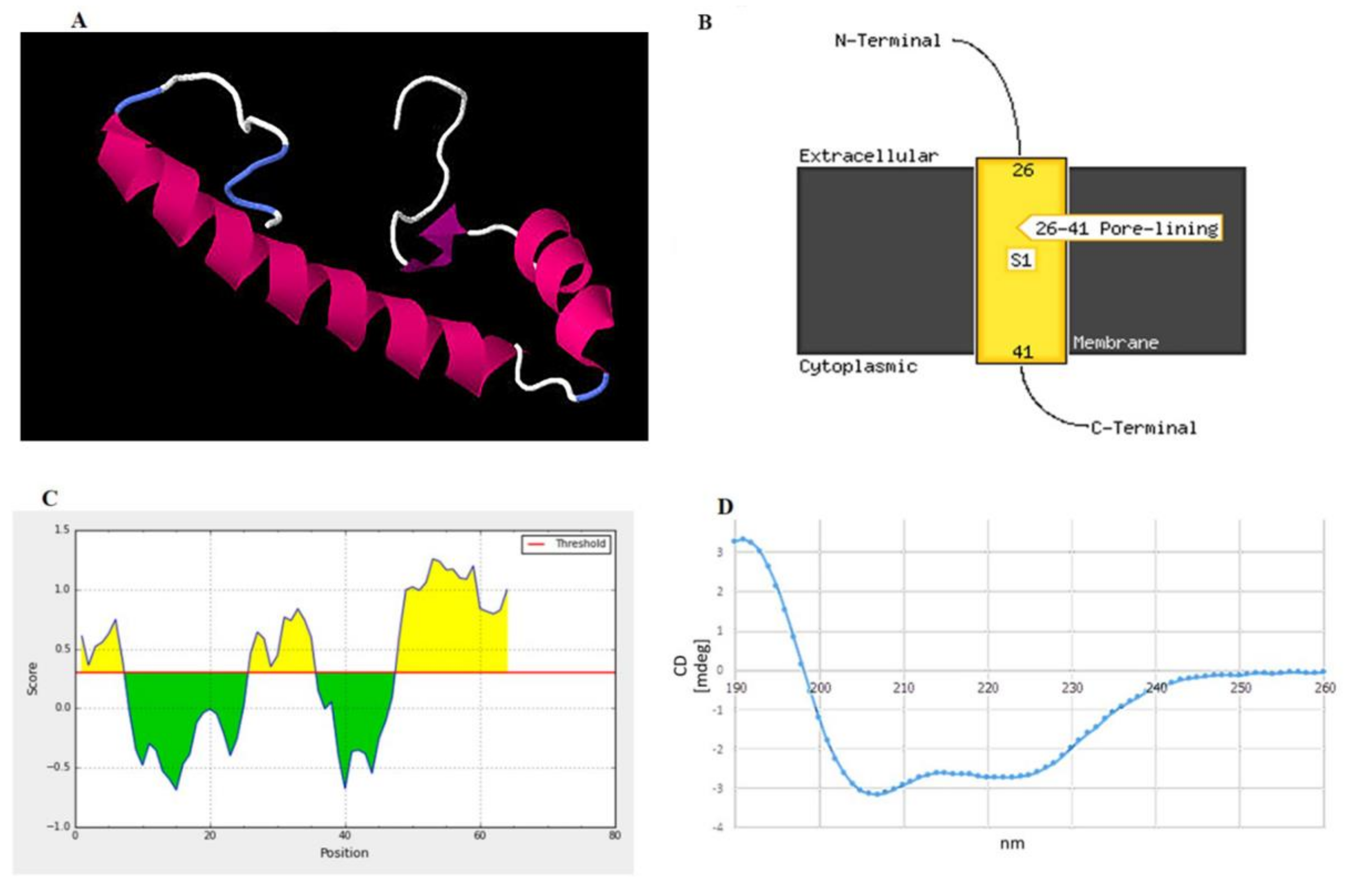
2.1. Identification and Characterization of the Specific Peptide
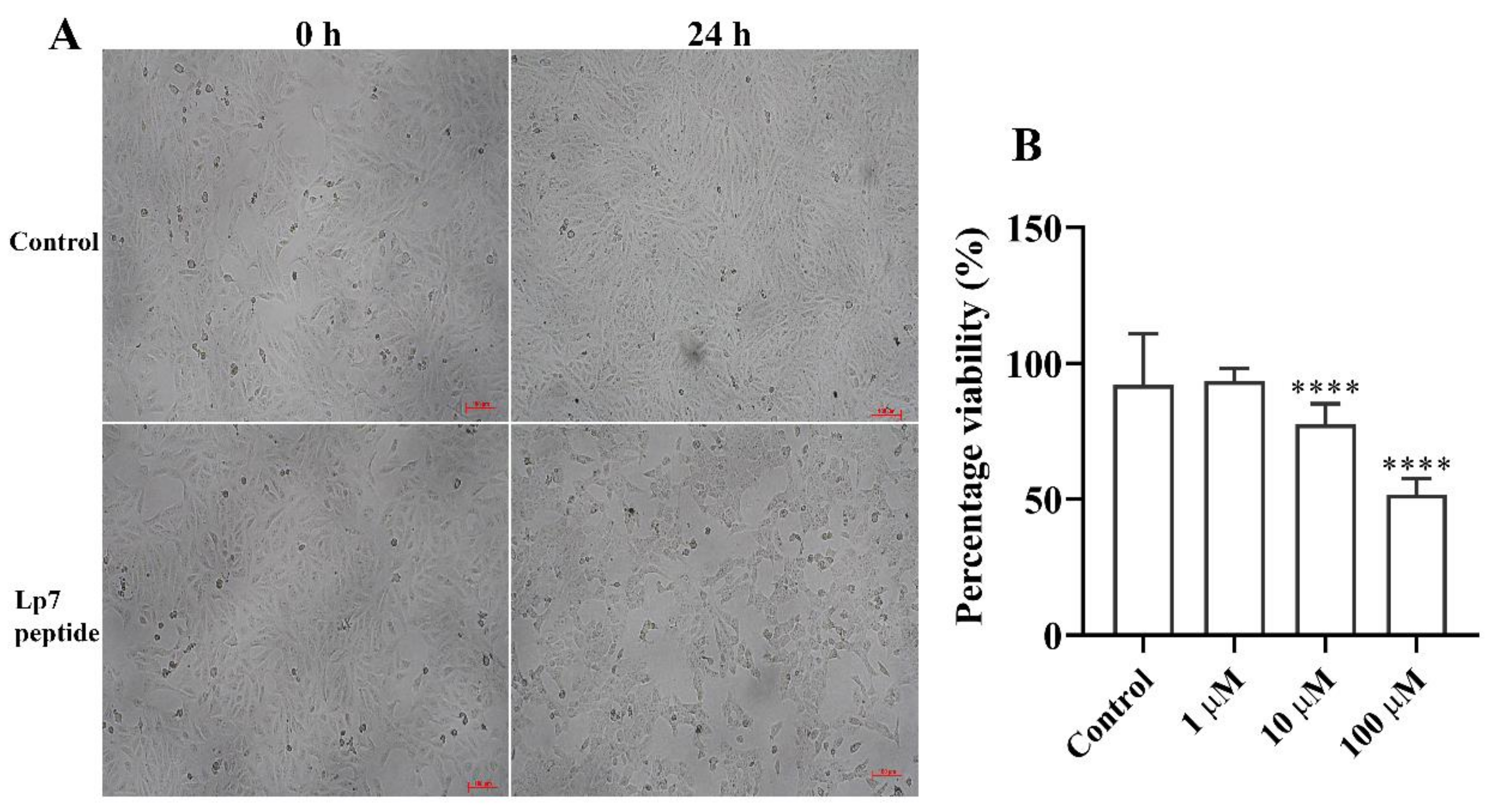
2.2. Effect of the Synthetic Peptide on the Viability of Vero Cells
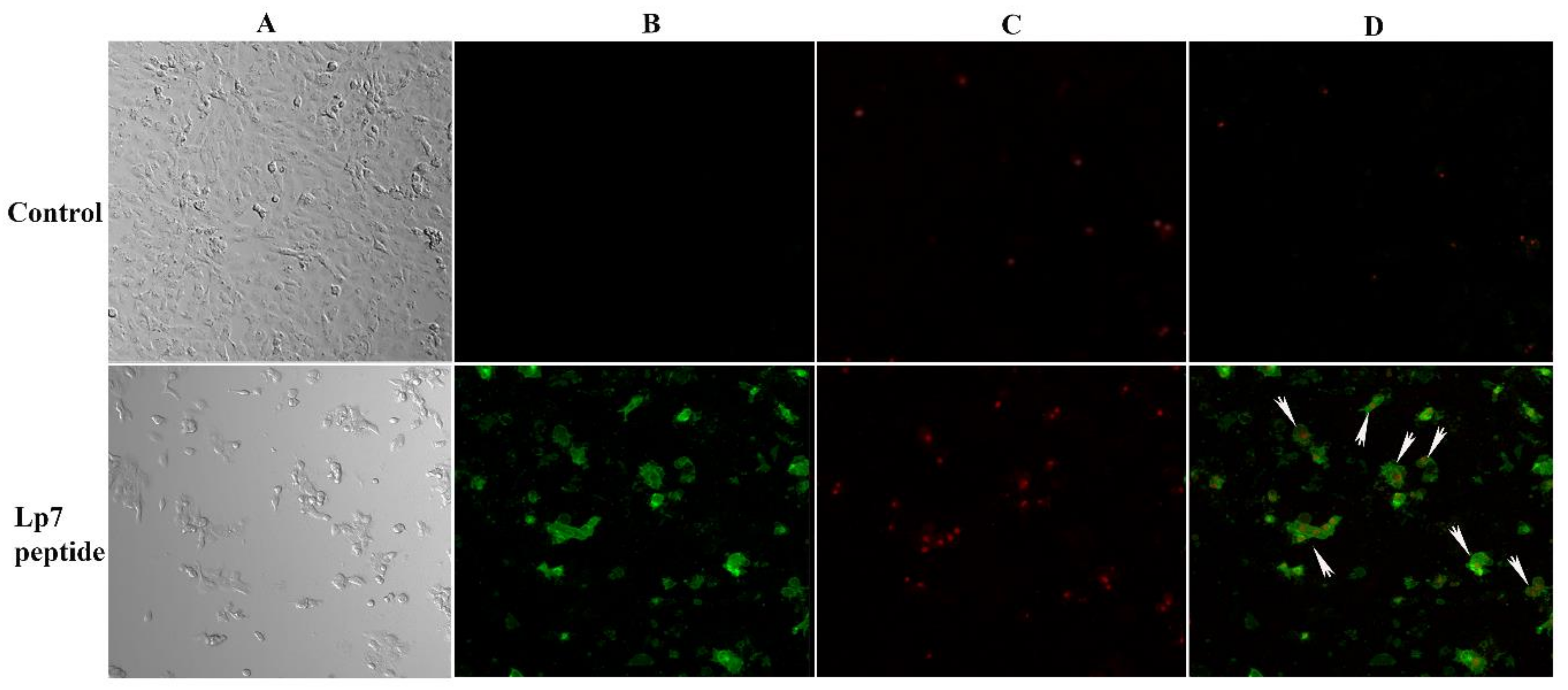
2.3. Induction of Apoptotic Programmed Cell Death by the Synthetic Peptide in Vero Cells
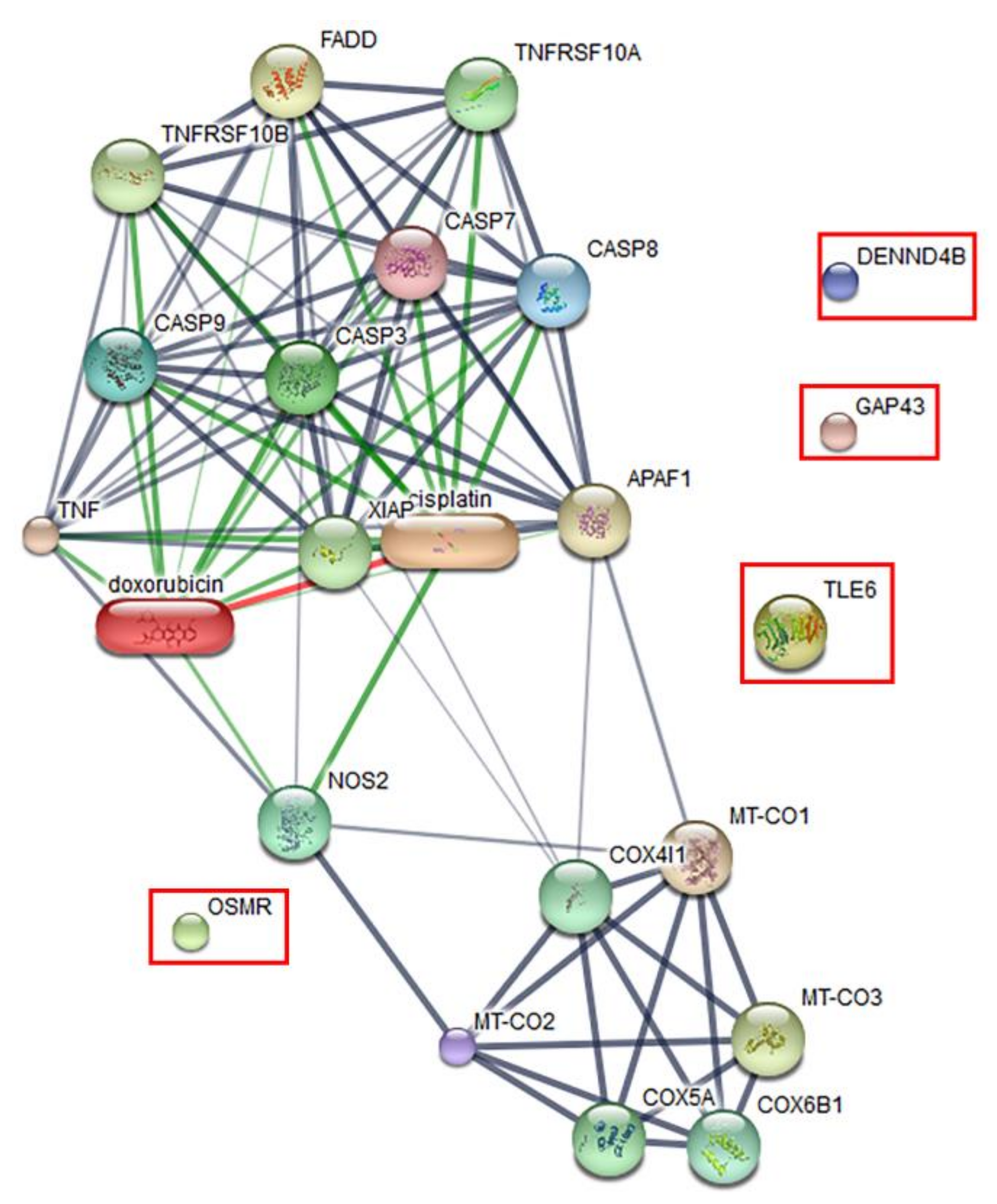
2.4. Proteomics Profile of Vero Cells Treated With the Synthetic Peptide
3. Discussion
4. Materials and Methods
4.1. Leptospira Cultivation
4.2. Extraction and Purification of the Peptide
4.3. Protein Identification by Liquid Chromatography–Tandem Mass Spectrometry (LC–MS/MS)
4.4. Characterization of the Novel Peptide Sequence
4.5. Cytotoxicity Testing on Vero Cells
4.6. Comparative Proteomics Study of the 7-kDa-Peptide-Treated Vero Cells
Author Contributions
Funding
Conflicts of Interest
References
- Costa, F.; Hagan, J.E.; Calcagno, J.; Kane, M.; Torgerson, P.; Martinez-Silveira, M.S.; Stein, C.; Abela-Ridder, B.; Ko, A.I. Global morbidity and mortality of leptospirosis: A Systematic Review. PLoS Negl. Trop. Dis. 2015, 9, e0003898. [Google Scholar] [CrossRef]
- Picardeau, M. Virulence of the zoonotic agent of leptospirosis: Still terra incognita? Nat. Rev. Microbiol. 2017, 15, 297–307. [Google Scholar] [CrossRef]
- Martinez-Lopez, D.G.; Fahey, M.; Coburn, J. Responses of human endothelial cells to pathogenic and non-pathogenic Leptospira species. PLoS Negl. Trop. Dis. 2010, 4, e918. [Google Scholar] [CrossRef]
- Lata, K.S.; Kumar, S.; Vaghasia, V.; Sharma, P.; Bhairappanvar, S.B.; Soni, S.; Das, J. Exploring leptospiral proteomes to identify potential candidates for vaccine design against leptospirosis using an immunoinformatics approach. Sci. Rep. 2018, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Murray, G. The molecular basis of leptospiral pathogenesis. Curr. Top. Microbiol. Immunol. 2014, 387, 139–185. [Google Scholar] [CrossRef]
- Yang, C.-W.; Wu, M.-S.; Pan, M.-J.; Hsieh, W.-J.; Vandewalle, A.; Huang, C.-C. The Leptospira outer membrane protein LipL32 induces tubulointerstitial nephritis-mediated gene expression in mouse proximal tubule cells. J. Am. Soc. Nephrol. 2002, 13, 2037–2045. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.-W.; Hung, C.-C.; Wu, M.-S.; Tian, Y.-C.; Chang, C.-T.; Pan, M.-J.; Vandewalle, A. Toll-like receptor 2 mediates early inflammation by leptospiral outer membrane proteins in proximal tubule cells. Kidney Int. 2006, 69, 815–822. [Google Scholar] [CrossRef]
- Sonthayanon, P.; Jaresitthikunchai, J.; Mangmee, S.; Thiangtrongjit, T.; Wuthiekanun, V.; Amornchai, P.; Newton, P.; Phetsouvanh, R.; Day, N.P.; Roytrakul, S. Whole cell matrix assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) for identification of Leptospira spp. in Thailand and Lao PDR. PLoS Negl. Trop. Dis. 2019, 13, e0007232. [Google Scholar] [CrossRef]
- Ren, S.-X.; Fu, G.; Jiang, X.-G.; Zeng, R.; Miao, Y.-G.; Xu, H.; Zhang, Y.-X.; Xiong, H.; Lu, G.; Lu, L.-F.; et al. Unique physiological and pathogenic features of Leptospira interrogans revealed by whole-genome sequencing. Nature 2003, 422, 888–893. [Google Scholar] [CrossRef]
- Eshghi, A.; Pappalardo, E.; Hester, S.; Thomas, B.; Pretre, G.; Picardeau, M. Pathogenic Leptospira interrogans exoproteins are primarily involved in heterotrophic processes. Infect. Immun. 2015, 83, 3061–3073. [Google Scholar] [CrossRef]
- Matsunaga, J.; Werneid, K.; Zuerner, R.L.; Frank, A.; Haake, D.A. LipL46 is a novel surface-exposed lipoprotein expressed during leptospiral dissemination in the mammalian host. Microbiology 2006, 152, 3777–3786. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Santos, J.V.; Pereira, P.R.; Fernandes, L.G.; Siqueira, G.H.; De Souza, G.O.; Filho, A.S.; Vasconcellos, S.A.; Heinemann, M.B.; Chapola, E.G.; Nascimento, A.L. Binding of human plasminogen by the lipoprotein LipL46 of Leptospira interrogans. Mol. Cell. Probes 2018, 37, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Ristow, P.; Bourhy, P.; da Cruz McBride, F.W.; Figueira, C.P.; Huerre, M.; Ave, P.; Girons, I.S.; I Ko, A.; Picardeau, M. The OmpA-like protein Loa22 is essential for leptospiral virulence. PLoS Pathog. 2007, 3, e97. [Google Scholar] [CrossRef]
- Ratet, G.; Santecchia, I.; D’Andon, M.F.; Vernel-Pauillac, F.; Wheeler, R.; Lenormand, P.; Fischer, F.; Lechat, P.; Haake, D.A.; Picardeau, M.; et al. LipL21 lipoprotein binding to peptidoglycan enables Leptospira interrogans to escape NOD1 and NOD2 recognition. PLoS Pathog. 2017, 13, e1006725. [Google Scholar] [CrossRef] [PubMed]
- Vieira, M.L.; Teixeira, A.F.; Pidde, G.; Ching, A.T.C.; Tambourgi, D.V.; Nascimento, A.L.T.O.; Herwald, H. Leptospira interrogans outer membrane protein LipL21 is a potent inhibitor of neutrophil myeloperoxidase. Virulence 2018, 9, 414–425. [Google Scholar] [CrossRef] [PubMed]
- Chasteen, T.G.; Fuentes, D.E.; Tantaleán, J.C.; Vásquez, C.C. Tellurite: History, oxidative stress, and molecular mechanisms of resistance. FEMS Microbiol. Rev. 2009, 33, 820–832. [Google Scholar] [CrossRef]
- Turkovicova, L.; Smidak, R.; Jung, G.; Turňa, J.; Lubec, G.; Aradska, J. Proteomic analysis of the TerC interactome: Novel links to tellurite resistance and pathogenicity. J. Proteom. 2016, 136, 167–173. [Google Scholar] [CrossRef]
- Taylor, D.E. Bacterial tellurite resistance. Trends Microbiol. 1999, 7, 111–115. [Google Scholar] [CrossRef]
- Merien, F.; Baranton, G.; Perolat, P. Invasion of Vero cells and induction of apoptosis in macrophages by pathogenic Leptospira interrogans are correlated with virulence. Infect. Immun. 1997, 65, 729–738. [Google Scholar] [CrossRef]
- Artiushin, S.; Timoney, J.F.; Nally, J.; Verma, A. Host-inducible immunogenic sphingomyelinase-like protein, Lk73.5, of Leptospira interrogans. Infect. Immun. 2004, 72, 742–749. [Google Scholar] [CrossRef]
- Lee, S.H.; Kim, S.; Park, S.C.; Kim, M.J. Cytotoxic activities of Leptospira interrogans hemolysin SphH as a pore-forming protein on mammalian cells. Infect. Immun. 2002, 70, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Matsui, M.; Roche, L.; Geroult, S.; Soupé-Gilbert, M.-E.; Monchy, D.; Huerre, M.; Goarant, C. Cytokine and chemokine expression in kidneys during chronic leptospirosis in reservoir and susceptible animal models. PLoS ONE 2016, 11, e0156084. [Google Scholar] [CrossRef] [PubMed]
- Ghandehari, F.; Behbahani, M.; Pourazar, A.; Noormohammadi, Z. In silico and in vitro studies of cytotoxic activity of different peptides derived from vesicular stomatitis virus G protein. Iran J. Basic Med. Sci. 2015, 18, 47–52. [Google Scholar]
- Barragán-Cárdenas, A.C.; Urrea-Pelayo, M.; Niño-Ramírez, V.A.; Umaña, A.; Vernot, J.P.; Parra-Giraldo, C.M.; Fierro-Medina, R.; Monroy, Z.J.R.; Castañeda, J.E.G. Selective cytotoxic effect against the MDA-MB-468 breast cancer cell line of the antibacterial palindromic peptide derived from bovine lactoferricin. RSC Adv. 2020, 10, 17593–17601. [Google Scholar] [CrossRef]
- Kerr, J.F.R.; Wyllie, A.H.; Currie, A.R. Apoptosis: A basic biological phenomenon with wide ranging implications in tissue kinetics. Br. J. Cancer 1972, 26, 239–257. [Google Scholar] [CrossRef] [PubMed]
- Ravindran, N.; García-Gareta, E. Kerr’s coining of ‘Apoptosis’ and its relevance in skin wound healing and fibrosis. Exp. Dermatol. 2015, 24, 99–100. [Google Scholar] [CrossRef]
- Gschwind, M.; Huber, G. Detection of Apoptotic or Necrotic Death in Neuronal Cells by Morphological, Biochemical, and Molecular Analysis. In Apoptosis Techniques and Protocols; Poirier, J., Ed.; Springer Science and Business Media LLC: Berlin, Germany, 2003; Volume 29, pp. 13–32. [Google Scholar]
- Ingh, T.V.D.; Hartman, E. Pathology of acute Leptospira interrogans serotype icterohaemorrhagiae infection in the Syrian hamster. Vet. Microbiol. 1986, 12, 367–376. [Google Scholar] [CrossRef]
- Du, P.; Li, S.-J.; Ojcius, D.M.; Li, K.-X.; Hu, W.-L.; Lin, X.; Sun, A.-H.; Yan, J. A novel Fas-binding outer membrane protein and lipopolysaccharide of Leptospira interrogans induce macrophage apoptosis through the Fas/FasL-caspase-8/-3 pathway. Emerg. Microbes Infect. 2018, 7, 1–17. [Google Scholar] [CrossRef]
- Zhang, Y.-X.; Geng, Y.; Yang, J.-W.; Guo, X.-K.; Zhao, G.-P. Cytotoxic activity and probable apoptotic effect of Sph2, a sphigomyelinase hemolysin from Leptospira interrogans strain Lai. BMB Rep. 2008, 41, 119–125. [Google Scholar] [CrossRef]
- Che, R.; Ding, S.; Zhang, Q.; Yang, W.; Yan, J.; Lin, X. Haemolysin Sph2 of Leptospira interrogans induces cell apoptosis via intracellular reactive oxygen species elevation and mitochondrial membrane injury. Cell. Microbiol. 2018, 21, e12959. [Google Scholar] [CrossRef]
- Yoshimura, S.-I.; Gerondopoulos, A.; Linford, A.; Rigden, D.J.; Barr, F.A. Family-wide characterization of the DENN domain Rab GDP-GTP exchange factors. J. Cell Biol. 2010, 191, 367–381. [Google Scholar] [CrossRef]
- I Benowitz, L.; Routtenberg, A. GAP-43: An intrinsic determinant of neuronal development and plasticity. Trends Neurosci. 1997, 20, 84–91. [Google Scholar] [CrossRef]
- Zhu, K.; Yan, L.; Zhang, X.; Lu, X.; Wang, T.; Yan, J.; Liu, X.; Qiao, J.; Li, L. Identification of a human subcortical maternal complex. Mol. Hum. Reprod. 2014, 21, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Mosley, B.; De Imus, C.; Friend, D.; Boiani, N.; Thoma, B.; Park, L.S.; Cosman, D. Dual oncostatin M (OSM) receptors. Cloning and characterization of an alternative signaling subunit conferring OSM-specific receptor activation. J. Biol. Chem. 1996, 271, 32635–32643. [Google Scholar] [CrossRef]
- Haider, S.R.; Reid, H.J.; Sharp, B.L. Tricine-SDS-PAGE. In Protein Electrophoresis. Methods in Molecular Biology (Methods and Protocols); Kurien, B., Scofield, R., Eds.; Humana Press: Totowa, NJ, USA, 2019; Volume 869, pp. 151–160. [Google Scholar]
- Shevchenko, A.; Tomas, H.; Havlis, J.; Olsen, J.V.; Mann, M.J. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 2006, 1, 2856–2860. [Google Scholar] [CrossRef]
- Louis-Jeune, C.; Andrade-Navarro, M.A.; Perez-Iratxeta, C. Prediction of protein secondary structure from circular dichroism using theoretically derived spectra. Proteins Struct. Funct. Bioinform. 2012, 80, 374–381. [Google Scholar] [CrossRef]
- Källberg, M.; Wang, H.; Wang, S.; Peng, J.; Wang, Z.; Lu, H.; Xu, J. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 2012, 7, 1511–1522. [Google Scholar] [CrossRef]
- Wang, L.; Li, W.; Liu, S.; Xu, J. RaptorX-Property: A web server for protein structure property prediction. Nucleic Acids Res. 2016, 44, W430–W435. [Google Scholar] [CrossRef]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2010, 27, 343–350. [Google Scholar] [CrossRef]
- Benkert, P.; Tosatto, S.C.E.; Schomburg, D. QMEAN: A comprehensive scoring function for model quality assessment. Proteins Struct. Funct. Bioinform. 2008, 71, 261–277. [Google Scholar] [CrossRef]
- Lovell, S.C.; Davis, I.W.; Arendall, W.B.; de Bakker, P.I.W.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure validation by Calpha geometry: Phi,psi and Cbeta deviation. Proteins 2003, 50, 437–450. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, H. Predicting Secretory Proteins with SignalP. In Protein Function Prediction. Methods in Molecular Biology; Kihara, D., Ed.; Humana Press: New York, NY, USA, 2017; Volume 1611, pp. 59–73. [Google Scholar]
- Käll, L.; Krogh, A.; Sonnhammer, E.L. A Combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 2004, 338, 1027–1036. [Google Scholar] [CrossRef] [PubMed]
- Käll, L.; Krogh, A.; Sonnhammer, E.L. Advantages of combined transmembrane topology and signal peptide prediction--the Phobius web server. Nucleic Acids Res. 2007, 35, W429–W432. [Google Scholar] [CrossRef] [PubMed]
- Bjellqvist, B.; Basse, B.; Olsen, E.; Celis, J.E. Reference points for comparisons of two-dimensional maps of proteins from different human cell types defined in a pH scale where isoelectric points correlate with polypeptide compositions. Electrophoresis 1994, 15, 529–539. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.E.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Jespersen, M.C.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017, 45, W24–W29. [Google Scholar] [CrossRef]
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paratsaphan, S.; Moonsom, S.; Reamtong, O.; Roytrakul, S.; Wuthiekanun, V.; Day, N.P.J.; Sonthayanon, P. Characterization of a Novel Peptide from Pathogenic Leptospira and Its Cytotoxic Effect. Pathogens 2020, 9, 906. https://doi.org/10.3390/pathogens9110906
Paratsaphan S, Moonsom S, Reamtong O, Roytrakul S, Wuthiekanun V, Day NPJ, Sonthayanon P. Characterization of a Novel Peptide from Pathogenic Leptospira and Its Cytotoxic Effect. Pathogens. 2020; 9(11):906. https://doi.org/10.3390/pathogens9110906
Chicago/Turabian StyleParatsaphan, Saksakon, Saengduen Moonsom, Onrapak Reamtong, Sittiruk Roytrakul, Vanaporn Wuthiekanun, Nicholas P. J. Day, and Piengchan Sonthayanon. 2020. "Characterization of a Novel Peptide from Pathogenic Leptospira and Its Cytotoxic Effect" Pathogens 9, no. 11: 906. https://doi.org/10.3390/pathogens9110906
APA StyleParatsaphan, S., Moonsom, S., Reamtong, O., Roytrakul, S., Wuthiekanun, V., Day, N. P. J., & Sonthayanon, P. (2020). Characterization of a Novel Peptide from Pathogenic Leptospira and Its Cytotoxic Effect. Pathogens, 9(11), 906. https://doi.org/10.3390/pathogens9110906





