A Complex Dance: Measuring the Multidimensional Worlds of Influenza Virus Evolution and Anti-Influenza Immune Responses
Abstract
1. Introduction
2. Hemagglutinin (HA) and Its Antibodies
3. Complexity of Human Immune Responses against Influenza Virus
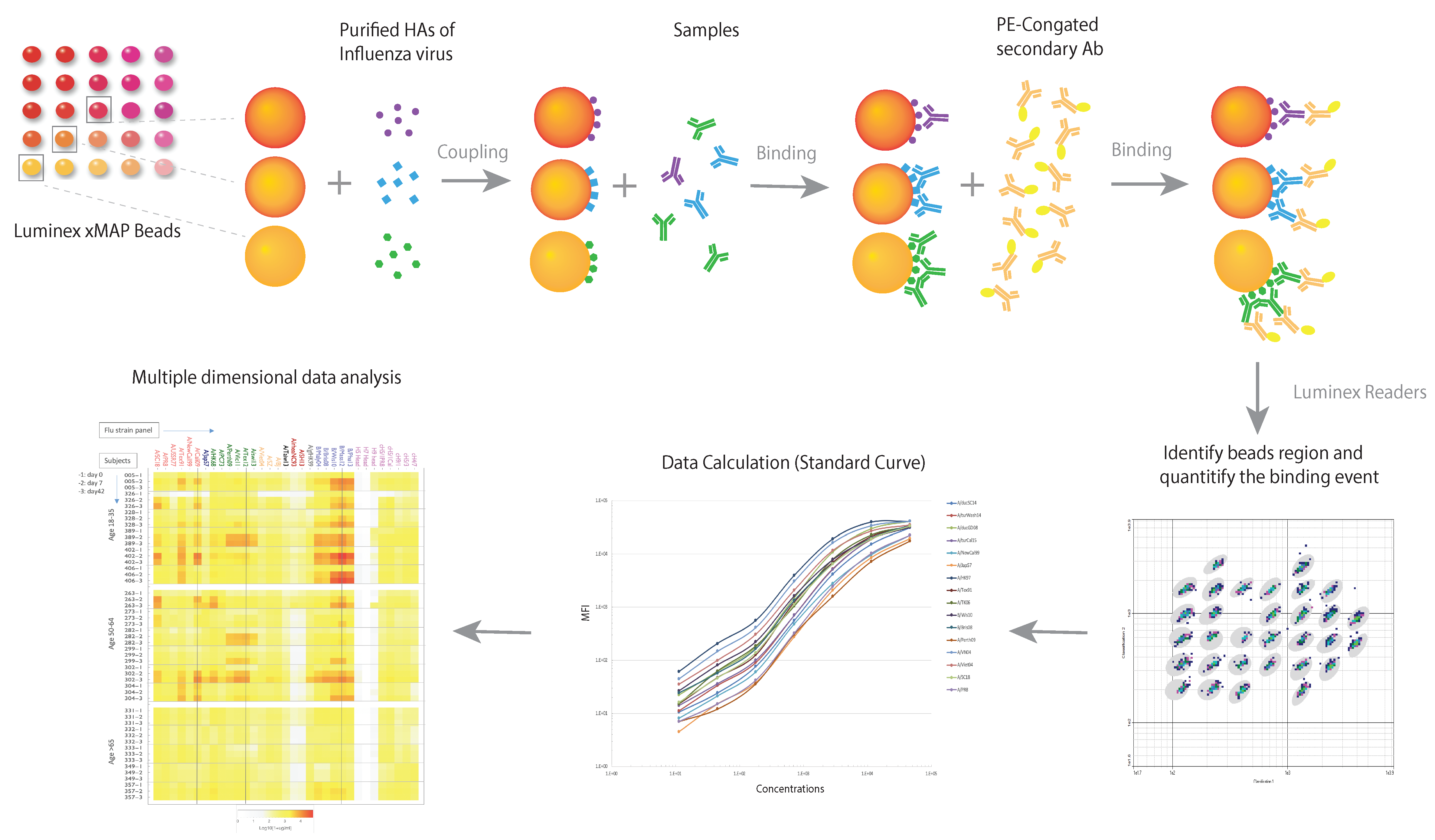
4. Multidimensional Assays (MDAs) for Anti-Influenza Antibodies
5. Current Applications of MDA
5.1. Determination of the Antigenicity of HA of Influenza Virus
5.2. Identify the Binding Profiles of Broad Cross-Reactive mAb
5.3. Detection of the Magnitude and Breadth of Serologic Responses to Influenza Infection or Vaccination
5.4. Detection of Antibodies in B Cell Culture Medium and Body Fluid
6. Future Applications of MDA
6.1. Population Studies with Micro-Sampling Techniques
6.2. Comprehensive Antigenic Study of HA Proteins
6.3. Detecting Cross-Reactive Antibodies against Other Viral Proteins of Influenza Virus
7. Limitations of MDAs
8. Summary
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| MDA | Multiple dimensional assay |
| mAb | monoclonal antibody |
| HA | hemagglutinin |
| NA | neuraminidase |
| MBC | memory B cell |
| HAI | hemagglutinin inhibition assay |
| MN | microneutralization assay |
| bcAb | broad cross-reactive antibody |
| bnAb | broad neutralizing antibody |
References
- Iuliano, A.D.; Roguski, K.M.; Chang, H.H.; Muscatello, D.J.; Palekar, R.; Tempia, S.; Cohen, C.; Gran, J.M.; Schanzer, D.; Cowling, B.J.; et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet 2018, 391, 1285–1300. [Google Scholar] [CrossRef]
- Koel, B.F.; Burke, D.F.; Bestebroer, T.M.; Van Der Vliet, S.; Zondag, G.C.M.; Vervaet, G.; Skepner, E.; Lewis, N.S.; Spronken, M.I.J.; Russell, C.A.; et al. Substitutions Near the Receptor Binding Site Determine Major Antigenic Change During Influenza Virus Evolution. Science 2013, 342, 976–979. [Google Scholar] [CrossRef] [PubMed]
- Hilleman, M.R. Realities and enigmas of human viral influenza: Pathogenesis, epidemiology and control. Vaccine 2002, 20, 3068–3087. [Google Scholar] [CrossRef]
- Carrat, F.; Flahault, A. Influenza vaccine: The challenge of antigenic drift. Vaccine 2007, 25, 6852–6862. [Google Scholar] [CrossRef]
- Lambert, L.C.; Fauci, A.S. Influenza Vaccines for the Future. New Engl. J. Med. 2010, 363, 2036–2044. [Google Scholar] [CrossRef] [PubMed]
- CDC (Centers for Disease Control and Prevention). Selecting Viruses for the Seasonal Influenza Vaccine; CDC: Atlanta, GA, USA, 2018.
- Erbelding, E.J.; Post, D.J.; Stemmy, E.J.; Roberts, P.C.; Augustine, A.D.; Ferguson, S.; Paules, C.I.; Graham, B.S.; Fauci, A.S. A Universal Influenza Vaccine: The Strategic Plan for the National Institute of Allergy and Infectious Diseases. J. Infect. Dis. 2018, 218, 347–354. [Google Scholar] [CrossRef]
- Zost, S.J.; Parkhouse, K.; Gumina, M.E.; Kim, K.; Diaz Perez, S.; Wilson, P.C.; Treanor, J.J.; Sant, A.J.; Cobey, S.; Hensley, S.E. Contemporary H3N2 influenza viruses have a glycosylation site that alters binding of antibodies elicited by egg-adapted vaccine strains. Proc. Natl. Acad. Sci. USA 2017, 114, 12578–12583. [Google Scholar] [CrossRef]
- Francis, T., Jr. On the Doctrine of Original Antigenic Sin. Proc. Am. Philos. Soc. 1960, 104, 572–578. [Google Scholar] [CrossRef]
- Gostic, K.M.; Ambrose, M.; Worobey, M.; Lloyd-Smith, J.O. Potent protection against H5N1 and H7N9 influenza via childhood hemagglutinin imprinting. Science 2016, 354, 722–726. [Google Scholar] [CrossRef]
- Schulman, J.L.; Kilbourne, E.D. Induction of Partial Specific Heterotypic Immunity in Mice by a Single Infection with Influenza a Virus. J. Bacteriol. 1965, 89, 170–174. [Google Scholar]
- Nguyen, H.H.; Zemlin, M.; Ivanov, I.; Andrasi, J.; Zemlin, C.; Vu, H.L.; Schelonka, R.; Schroeder, H.W., Jr.; Mestecky, J. Heterosubtypic immunity to influenza A virus infection requires a properly diversified antibody repertoire. J. Virol. 2007, 81, 9331–9338. [Google Scholar] [CrossRef] [PubMed]
- Hirst, G.K. The quantitative determination of influenza virus and antibodies by means of red cell agglutination. J. Exp. Med. 1942, 75, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Hirst, G.K. Adsorption of influenza hemagglutinins and virus by red blood cells. J. Exp. Med. 1942, 76, 195–209. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.L.; Puck, J.; Hughes, B.J.; Cate, T.R. Microneutralization test for influenza A and B and parainfluenza 1 and 2 viruses that uses continuous cell lines and fresh serum enhancement. J. Clin. Microbiol. 1980, 12, 426–432. [Google Scholar]
- Remarque, E.J.; de Bruijn, I.A.; Boersma, W.J.; Masurel, N.; Ligthart, G.J. Altered antibody response to influenza H1N1 vaccine in healthy elderly people as determined by HI, ELISA, and neutralization assay. J. Med. Virol. 1998, 55, 82–87. [Google Scholar] [CrossRef]
- Kilbourne, E.D. Taxonomy and Comparative Virology of the Influenza Viruses. In Influenza; Springer: Boston, MA, USA, 1987; pp. 25–32. [Google Scholar]
- Nachbagauer, R.; Choi, A.; Hirsh, A.; Margine, I.; Iida, S.; Barrera, A.; Ferres, M.; Albrecht, R.A.; García-Sastre, A.; Bouvier, N.M. Defining the antibody cross-reactome directed against the influenza virus surface glycoproteins. Nat. Immunol. 2017, 18, 464. [Google Scholar] [CrossRef]
- Yamashita, M.; Krystal, M.; Fitch, W.M.; Palese, P. Influenza B virus evolution: Co-circulating lineages and comparison of evolutionary pattern with those of influenza A and C viruses. Virology 1988, 163, 112–122. [Google Scholar] [CrossRef]
- Krammer, F.; Fouchier, R.A.M.; Eichelberger, M.C.; Webby, R.J.; Shaw-Saliba, K.; Wan, H.; Wilson, P.C.; Compans, R.W.; Skountzou, I.; Monto, A.S. NAction! How Can Neuraminidase-Based Immunity Contribute to Better Influenza Virus Vaccines? MBio 2018, 9. [Google Scholar] [CrossRef]
- Mair, C.M.; Ludwig, K.; Herrmann, A.; Sieben, C. Receptor binding and pH stability—How influenza A virus hemagglutinin affects host-specific virus infection. Biochim. Biophys. Acta 2014, 1838, 1153–1168. [Google Scholar] [CrossRef]
- Brandenburg, B.; Koudstaal, W.; Goudsmit, J.; Klaren, V.; Tang, C.; Bujny, M.V.; Korse, H.J.; Kwaks, T.; Otterstrom, J.J.; Juraszek, J. Mechanisms of hemagglutinin targeted influenza virus neutralization. PLoS ONE 2013, 8, e80034. [Google Scholar] [CrossRef]
- Coudeville, L.; Bailleux, F.; Riche, B.; Megas, F.; Andre, P.; Ecochard, R. Relationship between haemagglutination- inhibiting antibody titres and clinical protection against influenza: Development and application of a bayesian random-effects model. BMC Med. Res. Methodol. 2010, 10, 18. [Google Scholar] [CrossRef]
- Caton, A.J.; Brownlee, G.G.; Yewdell, J.W.; Gerhard, W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 1982, 31, 417–4127. [Google Scholar] [CrossRef]
- Wiley, D.C.; Wilson, I.A.; Skehel, J.J. Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 1981, 289, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Angeletti, D.; Gibbs, J.S.; Angel, M.; Kosik, I.; Hickman, H.D.; Frank, G.M.; Das, S.R.; Wheatley, A.K.; Prabhakaran, M.; Leggat, D.J.; et al. Defining B cell immunodominance to viruses. Nat. Immunol. 2017, 18, 456–463. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Wilson, I.A. Structural characterization of an early fusion intermediate of influenza virus hemagglutinin. J. Virol. 2011, 85, 5172–5182. [Google Scholar] [CrossRef]
- Yassine, H.M.; Boyington, J.C.; McTamney, P.M.; Wei, C.J.; Kanekiyo, M.; Kong, W.P.; Gallagher, J.R.; Wang, L.; Zhang, Y.; Joyce, M.G.; et al. Hemagglutinin-stem nanoparticles generate heterosubtypic influenza protection. Nat. Med. 2015, 21, 1065–1070. [Google Scholar] [CrossRef]
- Krammer, F.; Palese, P. Universal Influenza Virus Vaccines That Target the Conserved Hemagglutinin Stalk and Conserved Sites in the Head Domain. J. Infect. Dis. 2019, 219, S62–S67. [Google Scholar] [CrossRef]
- Ellebedy, A.H. Immunizing the Immune: Can We Overcome Influenza’s Most Formidable Challenge? Vaccines 2018, 6, 68. [Google Scholar] [CrossRef]
- Nogales, A.; Piepenbrink, M.S.; Wang, J.; Ortega, S.; Basu, M.; Fucile, C.F.; Treanor, J.J.; Rosenberg, A.F.; Zand, M.S.; Keefer, M.C.; et al. A Highly Potent and Broadly Neutralizing H1 Influenza-Specific Human Monoclonal Antibody. Sci. Rep. 2018, 8, 4374. [Google Scholar] [CrossRef]
- Hong, M.; Lee, P.S.; Hoffman, R.M.B.; Zhu, X.; Krause, J.C.; Laursen, N.S.; Yoon, S.I.; Song, L.; Tussey, L.; Crowe, J.E.; et al. Antibody Recognition of the Pandemic H1N1 Influenza Virus Hemagglutinin Receptor Binding Site. J. Virol. 2013, 87, 12471–12480. [Google Scholar] [CrossRef]
- Whittle, J.R.R.; Zhang, R.; Khurana, S.; King, L.R.; Manischewitz, J.; Golding, H.; Dormitzer, P.R.; Haynes, B.F.; Walter, E.B.; Moody, M.A.; et al. Broadly neutralizing human antibody that recognizes the receptor-binding pocket of influenza virus hemagglutinin. Proc. Natl. Acad. Sci. USA 2011, 108, 14216–14221. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Krause, J.C.; McBride, R.; Paulson, J.C.; Crowe, J.E.; Wilson, I.A. A recurring motif for antibody recognition of the receptor-binding site of influenza hemagglutinin. Nat. Struct. Mol. Biol. 2013, 20, 363–370. [Google Scholar] [CrossRef]
- Ekiert, D.C.; Kashyap, A.K.; Steel, J.; Rubrum, A.; Bhabha, G.; Khayat, R.; Lee, J.H.; Dillon, M.A.; O’Neil, R.E.; Faynboym, A.M.; et al. Cross-neutralization of influenza A viruses mediated by a single antibody loop. Nature 2012, 489, 526–532. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.S.; Yoshida, R.; Ekiert, D.C.; Sakai, N.; Suzuki, Y.; Takada, A.; Wilson, I.A. Heterosubtypic antibody recognition of the influenza virus hemagglutinin receptor binding site enhanced by avidity. Proc. Natl. Acad. Sci. USA 2012, 109, 17040–17045. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.S.; Ohshima, N.; Stanfield, R.L.; Yu, W.; Iba, Y.; Okuno, Y.; Kurosawa, Y.; Wilson, I.A. Receptor mimicry by antibody F045–092 facilitates universal binding to the H3 subtype of influenza virus. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef]
- Yoshida, R.; Igarashi, M.; Ozaki, H.; Kishida, N.; Tomabechi, D.; Kida, H.; Ito, K.; Takada, A. Cross-Protective Potential of a Novel Monoclonal Antibody Directed against Antigenic Site B of the Hemagglutinin of Influenza A Viruses. PLoS Pathog. 2009, 5, e1000350. [Google Scholar] [CrossRef]
- Lee, J.; Boutz, D.R.; Chromikova, V.; Joyce, M.G.; Vollmers, C.; Leung, K.; Horton, A.P.; DeKosky, B.J.; Lee, C.H.; Lavinder, J.J.; et al. Molecular-level analysis of the serum antibody repertoire in young adults before and after seasonal influenza vaccination. Nat. Med. 2016, 22, 1456–1464. [Google Scholar] [CrossRef]
- Ekiert, D.C.; Bhabha, G.; Elsliger, M.A.; Friesen, R.H.; Jongeneelen, M.; Throsby, M.; Goudsmit, J.; Wilson, I.A. Antibody recognition of a highly conserved influenza virus epitope. Science 2009, 324, 246–251. [Google Scholar] [CrossRef]
- Sui, J.; Hwang, W.C.; Perez, S.; Wei, G.; Aird, D.; Chen, L.M.; Santelli, E.; Stec, B.; Cadwell, G.; Ali, M.; et al. Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat. Struct. Mol. Biol. 2009, 16, 265–273. [Google Scholar] [CrossRef]
- Kashyap, A.K.; Steel, J.; Rubrum, A.; Estelles, A.; Briante, R.; Ilyushina, N.A.; Xu, L.; Swale, R.E.; Faynboym, A.M.; Foreman, P.K.; et al. Protection from the 2009 H1N1 Pandemic Influenza by an Antibody from Combinatorial Survivor-Based Libraries. PLoS Pathog. 2010, 6, e1000990. [Google Scholar] [CrossRef]
- Ekiert, D.C.; Friesen, R.H.; Bhabha, G.; Kwaks, T.; Jongeneelen, M.; Yu, W.; Ophorst, C.; Cox, F.; Korse, H.J.; Brandenburg, B.; et al. A highly conserved neutralizing epitope on Group 2 influenza A viruses. Science 2011, 333, 843–850. [Google Scholar] [CrossRef] [PubMed]
- Friesen, R.H.E.; Lee, P.S.; Stoop, E.J.M.; Hoffman, R.M.B.; Ekiert, D.C.; Bhabha, G.; Yu, W.; Juraszek, J.; Koudstaal, W.; Jongeneelen, M.; et al. A common solution to Group 2 influenza virus neutralization. Proc. Natl. Acad. Sci. USA 2014, 111, 445–450. [Google Scholar] [CrossRef] [PubMed]
- Kallewaard, N.L.; Corti, D.; Collins, P.J.; Neu, U.; McAuliffe, J.M.; Benjamin, E.; Wachter-Rosati, L.; Palmer-Hill, F.J.; Yuan, A.Q.; Walker, P.A.; et al. Structure and Function Analysis of an Antibody Recognizing All Influenza A Subtypes. Cell 2016, 166, 596–608. [Google Scholar] [CrossRef] [PubMed]
- Lang, S.; Xie, J.; Zhu, X.; Wu, N.C.; Lerner, R.A.; Wilson, I.A. Antibody 27F3 Broadly Targets Influenza A Group 1 and 2 Hemagglutinins through a Further Variation in V H 1-69 Antibody Orientation on the HA Stem. Cell Rep. 2017, 20, 2935–2943. [Google Scholar] [CrossRef]
- Corti, D.; Voss, J.; Gamblin, S.J.; Codoni, G.; Macagno, A.; Jarrossay, D.; Vachieri, S.G.; Pinna, D.; Minola, A.; Vanzetta, F.; et al. A Neutralizing Antibody Selected from Plasma Cells That Binds to Group 1 and Group 2 Influenza A Hemagglutinins. Science 2011, 333, 850–856. [Google Scholar] [CrossRef]
- Dreyfus, C.; Laursen, N.S.; Kwaks, T.; Zuijdgeest, D.; Khayat, R.; Ekiert, D.C.; Lee, J.H.; Metlagel, Z.; Bujny, M.V.; Jongeneelen, M.; et al. Highly Conserved Protective Epitopes on Influenza B Viruses. Science 2012, 337, 1343–1348. [Google Scholar] [CrossRef]
- Henry Dunand, C.J.; Leon, P.E.; Huang, M.; Choi, A.; Chromikova, V.; Ho, I.Y.; Tan, G.S.; Cruz, J.; Hirsh, A.; Zheng, N.Y.; et al. Both Neutralizing and Non-Neutralizing Human H7N9 Influenza Vaccine-Induced Monoclonal Antibodies Confer Protection. Cell Host Microbe 2016, 19, 800–813. [Google Scholar] [CrossRef]
- Mullarkey, C.E.; Bailey, M.J.; Golubeva, D.A.; Tan, G.S.; Nachbagauer, R.; He, W.; Novakowski, K.E.; Bowdish, D.M.; Miller, M.S.; Palese, P.; et al. Broadly Neutralizing Hemagglutinin Stalk-Specific Antibodies Induce Potent Phagocytosis of Immune Complexes by Neutrophils in an Fc-Dependent Manner. MBio 2016, 7. [Google Scholar] [CrossRef]
- Bangaru, S.; Lang, S.; Schotsaert, M.; Vanderven, H.A.; Zhu, X.; Kose, N.; Bombardi, R.; Finn, J.A.; Kent, S.J.; Gilchuk, P.; et al. A Site of Vulnerability on the Influenza Virus Hemagglutinin Head Domain Trimer Interface. Cell 2019, 177, 1136–1152.e18. [Google Scholar] [CrossRef]
- Xu, R.; Ekiert, D.C.; Krause, J.C.; Hai, R.; Crowe, J.E.; Wilson, I.A. Structural basis of preexisting immunity to the 2009 H1N1 pandemic influenza virus. Science 2010, 328, 357–360. [Google Scholar] [CrossRef]
- Dawood, F.S.; Iuliano, A.D.; Reed, C.; Meltzer, M.I.; Shay, D.K.; Cheng, P.Y.; Bandaranayake, D.; Breiman, R.F.; Brooks, W.A.; Buchy, P.; et al. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: A modelling study. Lancet Infect. Dis. 2012, 12, 687–695. [Google Scholar] [CrossRef]
- Nguyen, A.M.; Noymer, A. Influenza mortality in the United States, 2009 pandemic: Burden, timing and age distribution. PLoS ONE 2013, 8, e64198. [Google Scholar] [CrossRef] [PubMed]
- De St.Groth, S.F.; Webster, R. Disquisitions on original antigenic sin: I. Evidence in man. J. Exp. Med. 1966, 124, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Zarnitsyna, V.I.; Ellebedy, A.H.; Davis, C.; Jacob, J.; Ahmed, R.; Antia, R. Masking of antigenic epitopes by antibodies shapes the humoral immune response to influenza. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.S.; Gardner, T.J.; Krammer, F.; Aguado, L.C.; Tortorella, D.; Basler, C.F.; Palese, P. Neutralizing antibodies against previously encountered influenza virus strains increase over time: A longitudinal analysis. Sci. Transl. Med. 2013, 5, 198ra107. [Google Scholar] [CrossRef]
- Zhang, A.; Stacey, H.D.; Mullarkey, C.E.; Miller, M.S. Original Antigenic Sin: How First Exposure Shapes Lifelong Anti-Influenza Virus Immune Responses. J. Immunol. 2019, 202, 335–340. [Google Scholar] [CrossRef]
- Lessler, J.; Riley, S.; Read, J.M.; Wang, S.; Zhu, H.; Smith, G.J.D.; Guan, Y.; Jiang, C.Q.; Cummings, D.A.T. Evidence for Antigenic Seniority in Influenza A (H3N2) Antibody Responses in Southern China. PLoS Pathog. 2012, 8, e1002802. [Google Scholar] [CrossRef]
- Huijskens, E.G.; Reimerink, J.; Mulder, P.G.; van Beek, J.; Meijer, A.; de Bruin, E.; Friesema, I.; de Jong, M.D.; Rimmelzwaan, G.F.; Peeters, M.F.; et al. Profiling of humoral response to influenza A(H1N1)pdm09 infection and vaccination measured by a protein microarray in persons with and without history of seasonal vaccination. PLoS ONE 2013, 8, e54890. [Google Scholar] [CrossRef]
- Koopmans, M.; de Bruin, E.; Godeke, G.J.; Friesema, I.; van Gageldonk, R.; Schipper, M.; Meijer, A.; van Binnendijk, R.; Rimmelzwaan, G.F.; de Jong, M.D.; et al. Profiling of humoral immune responses to influenza viruses by using protein microarray. Clin. Microbiol. Infect. 2012, 18, 797–807. [Google Scholar] [CrossRef]
- Te Beest, D.; de Bruin, E.; Imholz, S.; Wallinga, J.; Teunis, P.; Koopmans, M.; van Boven, M. Discrimination of influenza infection (A/2009 H1N1) from prior exposure by antibody protein microarray analysis. PLoS ONE 2014, 9, e113021. [Google Scholar] [CrossRef]
- Wang, J.; Hilchey, S.P.; DeDiego, M.; Perry, S.; Hyrien, O.; Nogales, A.; Garigen, J.; Amanat, F.; Huertas, N.; Krammer, F.; et al. Broad cross-reactive IgG responses elicited by adjuvanted vaccination with recombinant influenza hemagglutinin (rHA) in ferrets and mice. PLoS ONE 2018, 13, e0193680. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Wang, J.; Garigen, J.; Treanor, J.J.; Zand, M.S. Continuous Readout versus Titer-Based Assays of Influenza Vaccine Trials: Sensitivity, Specificity, and False Discovery Rates. Comput. Math. Methods Med. 2019, 2019, 9287120. [Google Scholar] [CrossRef] [PubMed]
- Legutki, J.B.; Magee, D.M.; Stafford, P.; Johnston, S.A. A general method for characterization of humoral immunity induced by a vaccine or infection. Vaccine 2010, 28, 4529–4537. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, R.; Supnet, M.; Jasinskas, A.; Jain, A.; Taghavian, O.; Obiero, J.; Milton, D.K.; Chen, W.H.; Grantham, M.; Webby, R.; et al. Protein Microarray Analysis of the Specificity and Cross-Reactivity of Influenza Virus Hemagglutinin-Specific Antibodies. mSphere 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hilchey, S.P.; Hyrien, O.; Huertas, N.; Perry, S.; Ramanunninair, M.; Bucher, D.; Zand, M.S. Multi-Dimensional Measurement of Antibody-Mediated Heterosubtypic Immunity to Influenza. PLoS ONE 2015, 10, e0129858. [Google Scholar] [CrossRef]
- Luminex Instruments. Available online: https://www.luminexcorp.com/instruments/ (accessed on 5 May 2019).
- Watson, D.S.; Reddy, S.M.; Brahmakshatriya, V.; Lupiani, B. A multiplexed immunoassay for detection of antibodies against avian influenza virus. J. Immunol. Methods 2009, 340, 123–131. [Google Scholar] [CrossRef]
- Keynan, Y.; Bodnarchuk, T.; Wayne, S.; Li, Y.; Fowke, K.R. Evaluation of influenza-specific humoral response by microbead array analysis. Can. J. Infect. Dis. Med. Microbiol. 2011, 22, 25–29. [Google Scholar] [CrossRef]
- Zand, M.S.; Wang, J.; Hilchey, S. Graphical Representation of Proximity Measures for Multidimensional Data: Classical and Metric Multidimensional Scaling. Math. J. 2015, 17. [Google Scholar] [CrossRef][Green Version]
- Tesini, B.L.; Kanagaiah, P.; Wang, J.; Hahn, M.; Halliley, J.L.; Chaves, F.A.; Nguyen, P.Q.T.; Nogales, A.; DeDiego, M.L.; Anderson, C.S.; et al. Broad Hemagglutinin-Specific Memory B Cell Expansion by Seasonal Influenza Virus Infection Reflects Early-Life Imprinting and Adaptation to the Infecting Virus. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Clark, A.M.; DeDiego, M.L.; Anderson, C.S.; Wang, J.; Yang, H.; Nogales, A.; Martinez-Sobrido, L.; Zand, M.S.; Sangster, M.Y.; Topham, D.J. Antigenicity of the 2015-2016 seasonal H1N1 human influenza virus HA and NA proteins. PLoS ONE 2017, 12, e0188267. [Google Scholar] [CrossRef]
- Huang, J.; Hilchey, S.P.; Wang, J.; Gerigan, J.; Zand, M.S. IL-15 enhances cross-reactive antibody recall responses to seasonal H3 influenza viruses in vitro. F1000Research 2017, 6, 2015. [Google Scholar] [CrossRef] [PubMed]
- Järvinen, K.M.; Wang, J.; Seppo, A.E.; Zand, M. Novel multiplex assay for profiling influenza antibodies in breast milk and serum of mother-infant pairs. F1000Research 2019. [Google Scholar] [CrossRef]
- Li, D.; Wang, J.; Treanor, J.J.; Zand, M.S. Improved Specificity and False Discovery Rates for Multiplex Analysis of Changes in Strain-Specific Anti-Influenza IgG. Comput. Math. Methods Med. 2019, 2019, 3053869. [Google Scholar] [CrossRef] [PubMed]
- Germeraad, E.; Achterberg, R.; Venema, S.; Post, J.; de Leeuw, O.; Koch, G.; van der Wal, F.J.; Beerens, N. The development of a multiplex serological assay for avian influenza based on Luminex technology. Methods 2019, 158, 54–60. [Google Scholar] [CrossRef] [PubMed]
- De Bruin, E.; Loeber, J.G.; Meijer, A.; Castillo, G.M.; Cepeda, M.L.G.; Torres-Sepúlveda, M.R.; Borrajo, G.J.C.; Caggana, M.; Giguere, Y.; Meyer, M.; et al. Evolution of an influenza pandemic in 13 countries from 5 continents monitored by protein microarray from neonatal screening bloodspots. J. Clin. Virol. 2014, 61, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Freidl, G.S.; de Bruin, E.; van Beek, J.; Reimerink, J.; de Wit, S.; Koch, G.; Vervelde, L.; van den Ham, H.J.; Koopmans, M.P. Getting more out of less—A quantitative serological screening tool for simultaneous detection of multiple influenza A hemagglutinin-types in chickens. PLoS ONE 2014, 9, e108043. [Google Scholar] [CrossRef] [PubMed]
- Freidl, G.S.; van den Ham, H.J.; Boni, M.F.; de Bruin, E.; Koopmans, M.P. Changes in heterosubtypic antibody responses during the first year of the 2009 A(H1N1) influenza pandemic. Sci. Rep. 2016, 6, 20385. [Google Scholar] [CrossRef]
- Te Beest, D.E.; de Bruin, E.; Imholz, S.; Koopmans, M.; van Boven, M. Heterosubtypic cross-reactivity of HA1 antibodies to influenza A, with emphasis on nonhuman subtypes (H5N1, H7N7, H9N2). PLoS ONE 2017, 12, e0181093. [Google Scholar] [CrossRef]
- Desbien, A.L.; Van Hoeven, N.; Reed, S.J.; Casey, A.C.; Laurance, J.D.; Baldwin, S.L.; Duthie, M.S.; Reed, S.G.; Carter, D. Development of a high density hemagglutinin protein microarray to determine the breadth of influenza antibody responses. Biotechniques 2013, 54, 345–348. [Google Scholar] [CrossRef]
- Price, J.V.; Jarrell, J.A.; Furman, D.; Kattah, N.H.; Newell, E.; Dekker, C.L.; Davis, M.M.; Utz, P.J. Characterization of influenza vaccine immunogenicity using influenza antigen microarrays. PLoS ONE 2013, 8, e64555. [Google Scholar] [CrossRef]
- Meade, P.; Latorre-Margalef, N.; Stallknecht, D.E.; Krammer, F. Development of an influenza virus protein microarray to measure the humoral response to influenza virus infection in mallards. Emerg. Microbes Infect. 2017, 6, e110. [Google Scholar] [CrossRef] [PubMed]
- Freidl, G.S.; Binger, T.; Muller, M.A.; de Bruin, E.; van Beek, J.; Corman, V.M.; Rasche, A.; Drexler, J.F.; Sylverken, A.; Oppong, S.K.; et al. Serological evidence of influenza A viruses in frugivorous bats from Africa. PLoS ONE 2015, 10, e0127035. [Google Scholar] [CrossRef] [PubMed]
- Mace, C.R.; Topham, D.J.; Mosmann, T.R.; Quataert, S.A.; Treanor, J.J.; Miller, B.L. Label-free, arrayed sensing of immune response to influenza antigens. Talanta 2011, 83, 1000–1005. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bucukovski, J.; Latorre-Margalef, N.; Stallknecht, D.E.; Miller, B.L. A Multiplex Label-Free Approach to Avian Influenza Surveillance and Serology. PLoS ONE 2015, 10, e0134484. [Google Scholar] [CrossRef]
- Smith, D.J. Mapping the Antigenic and Genetic Evolution of Influenza Virus. Science 2004, 305, 371–376. [Google Scholar] [CrossRef]
- Bangaru, S.; Zhang, H.; Gilchuk, I.M.; Voss, T.G.; Irving, R.P.; Gilchuk, P.; Matta, P.; Zhu, X.; Lang, S.; Nieusma, T.; et al. A multifunctional human monoclonal neutralizing antibody that targets a unique conserved epitope on influenza HA. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef]
- Wrammert, J.; Smith, K.; Miller, J.; Langley, W.A.; Kokko, K.; Larsen, C.; Zheng, N.Y.; Mays, I.; Garman, L.; Helms, C.; et al. Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 2008, 453, 667–671. [Google Scholar] [CrossRef]
- Hill, D.L.; Pierson, W.; Bolland, D.J.; Mkindi, C.; Carr, E.J.; Wang, J.; Houard, S.; Wingett, S.W.; Audran, R.; Wallin, E.F.; et al. The adjuvant GLA-SE promotes human Tfh cell expansion and emergence of public TCRbeta clonotypes. J. Exp. Med. 2019. [Google Scholar] [CrossRef]
- Pinna, D.; Corti, D.; Jarrossay, D.; Sallusto, F.; Lanzavecchia, A. Clonal dissection of the human memory B-cell repertoire following infection and vaccination. Eur. J. Immunol. 2009, 39, 1260–1270. [Google Scholar] [CrossRef]
- Sangster, M.Y.; Baer, J.; Santiago, F.W.; Fitzgerald, T.; Ilyushina, N.A.; Sundararajan, A.; Henn, A.D.; Krammer, F.; Yang, H.; Luke, C.J.; et al. B cell response and hemagglutinin stalk-reactive antibody production in different age cohorts following 2009 H1N1 influenza virus vaccination. Clin. Vaccine Immunol. 2013, 20, 867–876. [Google Scholar] [CrossRef][Green Version]
- Kok, M.G.M.; Fillet, M. Volumetric absorptive microsampling: Current advances and applications. J. Pharm. Biomed. Anal. 2018, 147, 288–296. [Google Scholar] [CrossRef]
- Wang, J.; Li, D.; Wiltse, A.; Emo, J.; Hilchey, S.P.; Zand, M.S. Application of Volumetric Absorptive Micro Sampling to Measure Multidimensional Anti-Influenza Hemagglutinin Igg Antibodies by MPlex-Flu Assay. J. Clin. Transl. Sci. 2019, 3, 332–343. [Google Scholar] [CrossRef]
- Sebastian, S.; Lambe, T. Clinical Advances in Viral-Vectored Influenza Vaccines. Vaccines 2018, 6, 29. [Google Scholar] [CrossRef] [PubMed]
- Kreijtz, J.H.; Bodewes, R.; van Amerongen, G.; Kuiken, T.; Fouchier, R.A.; Osterhaus, A.D.; Rimmelzwaan, G.F. Primary influenza A virus infection induces cross-protective immunity against a lethal infection with a heterosubtypic virus strain in mice. Vaccine 2007, 25, 612–620. [Google Scholar] [CrossRef] [PubMed]
- Quinones-Parra, S.; Grant, E.; Loh, L.; Nguyen, T.H.; Campbell, K.A.; Tong, S.Y.; Miller, A.; Doherty, P.C.; Vijaykrishna, D.; Rossjohn, J.; et al. Preexisting CD8+ T-cell immunity to the H7N9 influenza A virus varies across ethnicities. Proc. Natl. Acad. Sci. USA 2014, 111, 1049–1054. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Jarvis, D.L. Protein N-glycosylation in the baculovirus-insect cell system. Curr. Drug Targets 2007, 8, 1116–1125. [Google Scholar] [CrossRef] [PubMed]
- York, I.A.; Stevens, J.; Alymova, I.V. Influenza virus N-linked glycosylation and innate immunity. Biosci. Rep. 2019, 39. [Google Scholar] [CrossRef]

| Methods | Target Antigen | Species | Isotype | Sample Type(s) | Reference |
|---|---|---|---|---|---|
| Luminex array | NP, M1 and NS1 proteins | Chicken, turkey | IgY | Serum | [69] |
| Whole HA of H1, H3, H5, Flu B | Human | IgA1, IgG1 | Serum | [70] | |
| Whole HA of H1, H3, Flu B | Ferret, mouse, human | IgG, IgA, IgM | Serum | [67] | |
| Whole HA of H1, H3, H5 | Human | IgG | Serum | [71] | |
| Whole HA of H1, H2, H3, H5, H7, H9, Flu B and chimeric HA | Human | IgG | Serum MBC culture | [63,72,73,74] | |
| Whole HA of H1, H2, H3, H5, H7, H9, Flu B | Human | IgG | Purified mAb | [31] | |
| Whole HA of H1, H2, H3, H5, H7, H9, Flu B and chimeric HA | Human | IgA, IgG | Breast milk Infant serum | [75] | |
| Whole HA of H1, H2, H3, H5, H7, H9, Flu B and chimeric HA | Human | IgG | Serum | [64,76] | |
| H1-16 whole HA, and N1-9 whole NA Avian flu | Chicken | IgY | Serum | [77] | |
| Microarray | Random sequence peptides | Human | IgG | Serum | [65] |
| Head domain of HA of H1, H2, H3, H5, H7, H9 | Human | IgG | Serum or dry blood spots | [60,61,62,78,79,80,81] | |
| H1-H16 and H18 whole HA protein and/or HA peptides | Human | IgG | Serum | [66,82,83] | |
| H1-H18 whole HA | Chicken, duck, bat | IgY, IgG | Serum | [79,84,85] | |
| Arrayed Imaging | H1, H3, H6, H5 | Human | IgG | Serum | [86] |
| Reflectometry (AIR) | H1-H12 and Flu B | Mallard duck | IgY | Serum | [87] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, J.; Wiltse, A.; Zand, M.S. A Complex Dance: Measuring the Multidimensional Worlds of Influenza Virus Evolution and Anti-Influenza Immune Responses. Pathogens 2019, 8, 238. https://doi.org/10.3390/pathogens8040238
Wang J, Wiltse A, Zand MS. A Complex Dance: Measuring the Multidimensional Worlds of Influenza Virus Evolution and Anti-Influenza Immune Responses. Pathogens. 2019; 8(4):238. https://doi.org/10.3390/pathogens8040238
Chicago/Turabian StyleWang, Jiong, Alexander Wiltse, and Martin S. Zand. 2019. "A Complex Dance: Measuring the Multidimensional Worlds of Influenza Virus Evolution and Anti-Influenza Immune Responses" Pathogens 8, no. 4: 238. https://doi.org/10.3390/pathogens8040238
APA StyleWang, J., Wiltse, A., & Zand, M. S. (2019). A Complex Dance: Measuring the Multidimensional Worlds of Influenza Virus Evolution and Anti-Influenza Immune Responses. Pathogens, 8(4), 238. https://doi.org/10.3390/pathogens8040238





