Reduction of Arcobacter at Two Conventional Wastewater Treatment Plants in Southern Arizona, USA
Abstract
1. Introduction
2. Results
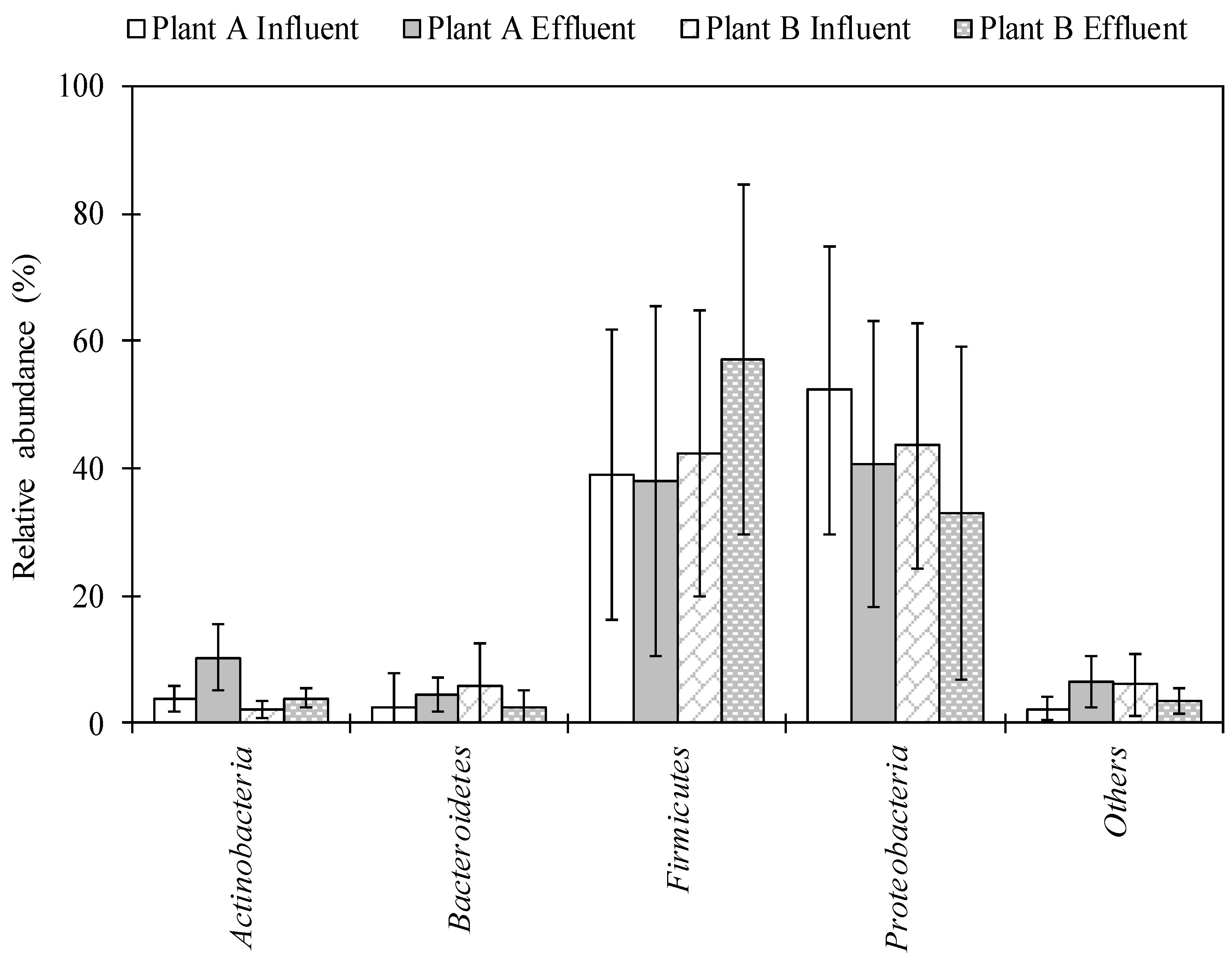
2.1. Characterization of Bacterial Community Using Next-Generation Sequencing (NGS) in Wastewater Samples
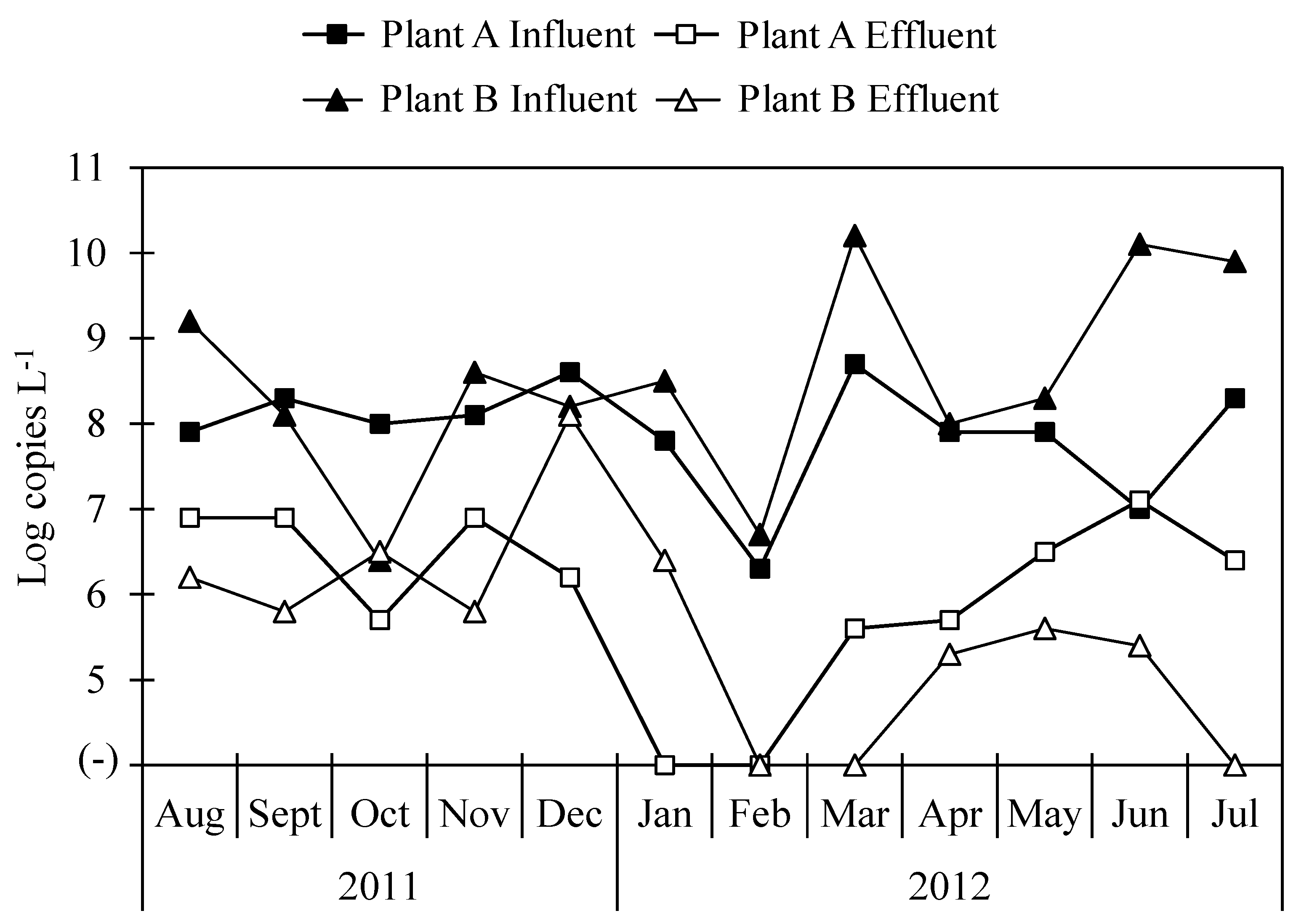
2.2. Occurrence of Total Bacteria, Arcobacter and Associated Virulence Genes in Wastewater Samples
2.3. Reduction Ratios of Arcobacter and Virulence Genes During Wastewater Treatment
3. Discussion
4. Materials and Methods
4.1. Collection of WWTP Samples
4.2. Bacterial DNA Extraction
4.3. NGS for Characterization of Bacterial Communities
4.4. qPCR of Total Bacteria and Arcobacter
4.5. Statistical Analysis
Author Contributions
Funding
Acknowledgements
Conflicts of Interest
References
- Norton-Brandão, D.; Scherrenberg, S.M.; van Lier, J.B. Reclamation of used urban waters for irrigation purposes–a review of treatment technologies. J. Environ. Manag. 2013, 122, 85–98. [Google Scholar] [CrossRef] [PubMed]
- Lazarova, V.; Levine, B.; Sack, J.; Cirelli, G.; Jeffrey, P.; Muntau, H.; Salgot, M.; Brissaud, F. Role of water reuse for enhancing integrated water management in Europe and Mediterranean countries. Water Sci. Technol. 2001, 43, 25–33. [Google Scholar] [CrossRef]
- Asano, T.; Cotruvo, J.A. Groundwater recharge with reclaimed municipal wastewater: Health and regulatory considerations. Water Res. 2004, 38, 1941–1951. [Google Scholar] [CrossRef] [PubMed]
- Angelakis, A.N.; Durham, B. Water recycling and reuse in EUREAU countries: Trends and challenges. Desalination 2008, 218, 3–12. [Google Scholar] [CrossRef]
- Bixio, D.; Thoeye, C.; Wintgens, T.; Hochstrat, R.; Melin, T.; Chikurel, H.; Aharoni, A.; Durham, B. Wastewater reclamation and reuse in the European Union and Israel: Status quo and future prospects. Int. Rev. Environ. Strateg. 2006, 6, 251–268. [Google Scholar]
- Rodriguez-Manzano, J.; Alonso, J.L.; Ferrús, M.A.; Moreno, Y.; Amorós, I.; Calgua, B.; Hundesa, A.; Guerrero-Latorre, L.; Carratala, A.; Rusiñol, M.; et al. Standard and new faecal indicators and pathogens in sewage treatment plants, microbiological parameters for improving the control of reclaimed water. Water Sci. Technol. 2012, 66, 2517–2523. [Google Scholar] [CrossRef]
- Vandamme, P.; Falsen, E.; Rossau, R.; Hoste, B.; Segers, P.; Tytgat, R.; De Ley, J. Revision of Campylobacter, Helicobacter, and Wolinella taxonomy: Emendation of generic descriptions and proposal of Arcobacter gen. nov. Int. J. Syst. Bacteriol. 1991, 41, 88–103. [Google Scholar] [CrossRef]
- Vandamme, P.; Vancanneyt, M.; Pot, B.; Mels, L.; Hoste, B.; Dewettinck, D.; Vlaes, L.; Van Den Borre, C.; Higgins, R.; Hommez, J.; et al. Polyphasic taxonomic study of the emended genus Arcobacter with Arcobacter butzleri comb. nov. and Arcobacter skirrowii sp. nov., an aerotolerant bacterium isolated from veterinary specimens. Int. J. Syst. Bacteriol. 1992, 42, 344–356. [Google Scholar] [CrossRef][Green Version]
- Banting, G.S.; Braithwaite, S.; Scott, C.; Kim, J.; Jeon, B.; Ashbolt, N.; Ruecker, N.; Tymensen, L.; Charest, J.; Pintar, K.; et al. Evaluation of various Campylobacter -specific qPCR assays for detection and enumeration of Campylobacteraceae in irrigation water and wastewater using a miniaturized MPN-qPCR assay. Appl. Environ. Microbiol. 2016, 82, 4743–4756. [Google Scholar] [CrossRef]
- Webb, A.L.; Taboada, E.N.; Selinger, L.B.; Boras, V.F.; Inglis, G.D. Efficacy of wastewater treatment on Arcobacter butzleri density and strain diversity. Water Res. 2016, 105, 291–296. [Google Scholar] [CrossRef]
- Gao, P.; Xu, W.; Sontag, P.; Li, X.; Xue, G. Correlating microbial community compositions with environmental factors in activated sludge from four full-scale municipal wastewater treatment plants in Shanghai, China. Appl. Microbiol. Biotechnol. 2016, 100, 4663–4673. [Google Scholar] [CrossRef] [PubMed]
- Jia, S.; Zhang, X.; Miao, Y.; Zhao, Y.; Ye, L.; Li, B.; Zhang, T. Fate of antibiotic resistance genes and their associations with bacterial community in livestock breeding wastewater and its receiving river water. Water Res. 2017, 124, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Ju, F.; Li, B.; Ma, L.; Wang, Y.; Huang, D.; Zhang, T. Antibiotic resistance genes and human bacterial pathogens: Co- occurrence, removal, and enrichment in municipal sewage sludge digesters. Water Res. 2016, 91, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Lu, P. Characterization of bacterial communities in sediments receiving various wastewater effluents with high-throughput sequencing analysis. Microb. Ecol. 2014, 67, 612–623. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, X.-X.; Wang, Z.; Huang, K.; Wang, Y.; Liang, W.; Tan, Y.; Liu, B.; Tang, J. Bacterial pathogens and community composition in advanced sewage treatment systems revealed by metagenomics analysis based on high-throughput sequencing. PLoS ONE 2015, 10, e0125549. [Google Scholar] [CrossRef]
- Snaidr, J.; Amann, R.; Huber, I.; Ludwig, W. Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl. Environ. Microbiol. 1997, 63, 2884–2896. [Google Scholar]
- Stampi, S.; De Luca, G.; Varoli, O.; Zanetti, F. Occurrence, removal and seasonal variation of thermophilic Campylobacters and Arcobacter in sewage sludge. Zentralblatt für Hygiene und Umweltmedizin 1999, 202, 19–27. [Google Scholar] [CrossRef]
- Al-jassim, N.; Ansari, M.I.; Harb, M.; Hong, P. Removal of bacterial contaminants and antibiotic resistance genes by conventional wastewater treatment processes in Saudi Arabia: Is the treated wastewater safe to reuse for agricultural irrigation? Water Res. 2015, 73, 277–290. [Google Scholar] [CrossRef]
- Diergaardt, S.M.; Venter, S.N.; Spreeth, A.; Theron, J.; Brözel, V.S. The occurrence of campylobacters in water sources in South Africa. Water Res. 2004, 38, 2589–2595. [Google Scholar] [CrossRef]
- Collado, L.; Inza, I.; Guarro, J.; Figueras, M.J. Presence of Arcobacter spp. in environmental waters correlates with high levels of fecal pollution. Environ. Microbiol. 2008, 10, 1635–1640. [Google Scholar] [CrossRef]
- Collado, L.; Kasimir, G.; Perez, U.; Bosch, A.; Pinto, R.; Saucedo, G.; Huguet, J.M.; Jose, M. Occurrence and diversity of Arcobacter spp. along the Llobregat River catchment, at sewage effluents and in a drinking water treatment plant. Water Res. 2010, 44, 3696–3702. [Google Scholar] [CrossRef] [PubMed]
- González, A.; Botella, S.; Montes, R.M.; Moreno, Y.; Ferrús, M.A. Direct detection and identification of Arcobacter species by multiplex PCR in chicken and wastewater samples from Spain. J. Food Prot. 2007, 70, 341–347. [Google Scholar] [CrossRef] [PubMed]
- Levican, A.; Collado, L.; Figueras, M.J. The use of two culturing methods in parallel reveals a high prevalence and diversity of Arcobacter spp. in a wastewater treatment plant. BioMed. Res. Int. 2016, 2016, 8132058. [Google Scholar] [CrossRef] [PubMed]
- Moreno, Y.; Botella, S.; Alonso, J.L.; Ferrus, M.A.; Hernandez, M.; Hernandez, J. Specific detection of Arcobacter and Campylobacter strains in water and sewage by PCR and Fluorescent In Situ Hybridization. Appl. Environ. Microbiol. 2003, 69, 1181–1186. [Google Scholar] [CrossRef]
- Akinglouchu, F. Isolation of Arcobacter Species from Different Water Sources and Characterization of Isolated Species by Molecular Techniques. Master’s Thesis, Graduate School of Engineering and Sciences of Izmir Institute of Technology, Izmir, Turkey, 2011. [Google Scholar]
- Merga, J.Y.; Royden, A.; Pandey, A.K.; Williams, N.J. Arcobacter spp. isolated from untreated domestic effluent. Lett. Appl. Microbiol. 2014, 59, 122–126. [Google Scholar] [CrossRef]
- Millar, J.A.; Raghavan, R. Accumulation and expression of multiple antibiotic resistance genes in Arcobacter cryaerophilus that thrives in sewage. PeerJ 2017, 5, e3269. [Google Scholar] [CrossRef]
- McLellan, S.L.; Huse, S.M.; Mueller-Spitz, S.R.; Andreishcheva, E.N.; Sogin, M.L. Diversity and population structure of sewage-derived microorganisms in wastewater treatment plant influent. Environ. Microbiol. 2010, 12, 378–392. [Google Scholar] [CrossRef]
- Sigala, J.; Unc, A. Pyrosequencing estimates of the diversity of antibiotic resistant bacteria in a wastewater system. Water Sci. Technol. 2013, 67, 1534–1543. [Google Scholar] [CrossRef]
- Fong, T.T.; Mansfield, L.S.; Wilson, D.L.; Schwab, D.J.; Molloy, S.L.; Rose, J.B. Massive microbiological groundwater contamination associated with a waterborne outbreak in Lake Erie, South Bass Island, Ohio. Environ. Health Perspect. 2007, 115, 856–864. [Google Scholar] [CrossRef]
- Ghaju Shrestha, R.; Tanaka, Y.; Malla, B.; Tandukar, S.; Bhandari, D.; Inoue, D.; Sei, K.; Sherchand, J.B.; Haramoto, E. Development of a quantitative PCR assay for Arcobacter spp. and its application to environmental water samples. Microbes Environ. 2018, 33, 309–316. [Google Scholar] [CrossRef]
- Zhang, T.; Shao, M.F.; Ye, L. 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J. 2012, 6, 1137–1147. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.Y.; Lauder, H.; Cruwys, H.; Falletta, P.; Beaudette, L.A. Development and application of an oligonucleotide microarray and real-time quantitative PCR for detection of wastewater bacterial pathogens. Sci. Total Environ. 2008, 398, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Iglewski, B.H. Pseudomonas. In Medical Microbiology; Baron, S., Ed.; The University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996; Chapter 27. [Google Scholar]
- Cai, L.; Ju, F.; Zhang, T. Tracking human sewage microbiome in a municipal wastewater treatment plant. Appl. Microbiol. Biotechnol. 2014, 98, 3317–3326. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Zhang, T. Detecting human bacterial pathogens in wastewater treatment plants by a high-throughput shotgun sequencing technique. Environ. Sci. Technol. 2013, 47, 5433–5441. [Google Scholar] [CrossRef] [PubMed]
- Varela, A.R.; Manaia, C.M. Human health implications of clinically relevant bacteria in wastewater habitats. Environ. Sci. Pollut. Res. 2013, 20, 3550–3569. [Google Scholar] [CrossRef] [PubMed]
- Hsu, T.T.D.; Lee, J. Global distribution and prevalence of Arcobacter in food and water. Zoonoses Public Health 2015, 62, 579–589. [Google Scholar] [CrossRef]
- Collado, L.; Figueras, M.J. Taxonomy, epidemiology, and clinical relevance of the genus Arcobacter. Clin. Microbiol. Rev. 2011, 24, 174–192. [Google Scholar] [CrossRef]
- Kitajima, M.; Haramoto, E.; Iker, B.C.; Gerba, C.P. Occurrence of Cryptosporidium, Giardia, and Cyclospora in influent and effluent water at wastewater treatment plants in Arizona. Sci. Total Environ. 2014, 484, 129–136. [Google Scholar] [CrossRef]
- Kitajima, M.; Iker, B.C.; Pepper, I.L.; Gerba, C.P. Relative abundance and treatment reduction of viruses during wastewater treatment processes—Identification of potential viral indicators. Sci. Total Environ. 2014, 488, 290–296. [Google Scholar] [CrossRef]
- Douidah, L.; De Zutter, L.; Baré, J.; De Vos, P.; Vandamme, P.; Vandenberg, O.; Van Den Abeele, A.M.; Houf, K. Occurrence of putative virulence genes in Arcobacter species isolated from humans and animals. J. Clin. Microbiol. 2012, 50, 735–741. [Google Scholar] [CrossRef]
- Tabatabaei, M.; Shirzad Aski, H.; Shayegh, H.; Khoshbakht, R. Occurrence of six virulence-associated genes in Arcobacter species isolated from various sources in Shiraz, Southern Iran. Microb. Pathog. 2014, 66, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Girbau, C.; Guerra, C.; Martínez-Malaxetxebarria, I.; Alonso, R.; Fernández-Astorga, A. Prevalence of ten putative virulence genes in the emerging foodborne pathogen Arcobacter isolated from food products. Food Microbiol. 2015, 52, 146–149. [Google Scholar] [CrossRef] [PubMed]
- Whiteduck-Léveillée, J.; Cloutier, M.; Topp, E.; Lapen, D.R.; Talbot, G.; Villemur, R.; Khan, I.U.H. Development and evaluation of multiplex PCR assays for rapid detection of virulence-associated genes in Arcobacter species. J. Microbiol. Methods 2016, 121, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Chu, B.T.T.; Petrovich, M.L.; Chaudhary, A.; Wright, D.; Murphy, B.; Wells, G.; Poretsky, R. Metagenomic analysis reveals the impact of wastewater treatment plants on the dispersal of microorganisms. Appl. Environ. Microbiol. 2018, 84, e02168-17. [Google Scholar]
- Xue, J.; Schmitz, B.W.; Caton, K.; Zhang, B.; Zabaleta, J.; Garai, J.; Taylor, C.M.; Romanchishina, T.; Gerba, C.P.; Pepper, I.L.; et al. Assessing the spatial and temporal variability of bacterial communities in two Bardenpho wastewater treatment systems via Illumina MiSeq sequencing. Sci. Total Environ. 2019, 657, 1543–1552. [Google Scholar] [CrossRef]
- Ghaju Shrestha, R.; Tanaka, Y.; Malla, B.; Bhandari, D.; Tandukar, S.; Inoue, D.; Sei, K.; Sherchand, J.B.; Haramoto, E. Next-generation sequencing identification of pathogenic bacterial genes and their relationship with fecal indicator bacteria in different water sources in the Kathmandu Valley, Nepal. Sci. Total Environ. 2017, 601, 278–284. [Google Scholar] [CrossRef]
- Baker, G.C.; Smith, J.J.; Cowan, D.A. Review and re-analysis of domain-specific 16S primers. J. Microbiol. Methods 2003, 55, 541–555. [Google Scholar] [CrossRef]
- Takai, K.; Horikoshi, K. Rapid detection and quantification of members of the archael community by quantitative PCR using fluorogenic probes. Appl. Environ. Microbiol. 2000, 66, 5066–5072. [Google Scholar] [CrossRef]
| Potential Pathogenic Genus | Plant A | Plant B | ||||
|---|---|---|---|---|---|---|
| Influent (Average ± s.d.) | Effluent (Average ± s.d.) | p-Value | Influent (Average ± s.d.) | Effluent (Average ± s.d.) | p-Value | |
| Achromobacter | 0.2 ± 0.4 | 0.1 ± 0.1 | 0.35 | 0.1 ± 0.3 | 0.0 ± 0.1 | 0.42 |
| Acidovorax | 0.9 ± 1.5 | 2.8 ± 2.9 | 0.06 | 2.0 ± 1.7 | 1.1 ± 1.5 | 0.15 |
| Acinetobacter | 6.2 ± 11.8 | 3.2 ± 8.0 | 0.48 | 1.7 ± 1.6 | 4.0 ± 11.8 | 0.52 |
| Actinobacillus | 0.1 ± 0.1 | 0.1 ± 0.2 | 0.40 | 1.3 ± 1.8 | 0.0 ± 0.1 | 0.04 |
| Arcobacter | 1.0 ± 1.0 | 0.5 ± 0.5 | 0.21 | 4.1 ± 5.7 | 0.9 ± 2.5 | 0.10 |
| Bacillus | 18.6 ± 15.4 | 25.8 ± 24.7 | 0.40 | 18.9 ± 18.7 | 33.2 ± 25.3 | 0.13 |
| Bacteroides | 0.1 ± 0.2 | 0.1 ± 0.2 | 0.58 | 1.6 ± 2.1 | 0.2 ± 0.4 | 0.04 |
| Bifidobacterium | 0.8 ± 0.5 | 0.7 ± 0.4 | 0.36 | 1.0 ± 0.6 | 1.3 ± 0.9 | 0.26 |
| Blautia | 2.4 ± 2.1 | 0.4 ± 0.4 | 0.01 | 2.5 ± 1.8 | 1.4 ± 3.1 | 0.34 |
| Brevundimonas | 0.2 ± 0.5 | 0.1 ± 0.1 | 0.22 | 0.1 ± 0.2 | 0.0 ± 0.1 | 0.57 |
| Chromobacterium | <0.01 | 0.5 ± 0.7 | 0.05 | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.46 |
| Chryseobacterium | 1.0 ± 2.0 | 1.1 ± 1.1 | 0.94 | 1.5 ± 1.6 | 0.5 ± 0.8 | 0.08 |
| Clostridium | 0.3 ± 0.2 | 0.4 ± 0.2 | 0.68 | 0.5 ± 0.3 | 0.4 ± 0.3 | 0.19 |
| Comamonas | 0.5 ± 0.4 | 0.2 ± 0.2 | 0.03 | 1.1 ± 0.8 | 0.4 ± 0.4 | 0.03 |
| Eikenella | 0.6 ± 0.5 | 0.1 ± 0.1 | 0.00 | 0.6 ± 0.5 | 0.2 ± 0.4 | 0.06 |
| Enterobacter | 1.6 ± 1.3 | 0.4 ± 0.9 | 0.03 | 0.6 ± 0.6 | 1.7 ± 3.8 | 0.37 |
| Enterococcus | 0.5 ± 0.4 | 0.1 ± 0.1 | 0.00 | 0.3 ± 0.4 | 0.2 ± 0.2 | 0.13 |
| Erwinia | 8.5 ± 12.0 | <0.01 | 0.03 | 3.3 ± 5.6 | 0.5 ± 0.9 | 0.10 |
| Flavobacterium | 0.1 ± 0.1 | 0.8 ± 0.8 | 0.00 | 0.4 ± 0.8 | 0.2 ± 0.3 | 0.41 |
| Klebsiella | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.69 | 0.3 ± 0.4 | 0.1 ± 0.3 | 0.38 |
| Gordonia | 0.1 ± 0.0 | 0.6 ± 1.6 | 0.32 | 0.0 ± 0.1 | 0.2 ± 0.2 | 0.01 |
| Lactococcus | 0.3 ± 0.6 | <0.01 | 0.13 | <0.01 | <0.01 | 0.14 |
| Leptotrichia | 0.2 ± 0.3 | 0.5 ± 0.5 | 0.11 | 0.8 ± 1.0 | 0.1 ± 0.2 | 0.03 |
| Megasphaera | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.85 | 0.4 ± 0.3 | 0.2 ± 0.3 | 0.17 |
| Microbacterium | 0.8 ± 0.8 | 0.3 ± 0.4 | 0.04 | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.86 |
| Mycobacterium | 0.2 ± 0.2 | 6.5 ± 4.0 | 0.00 | 0.1 ± 0.0 | 0.1 ± 0.1 | 0.06 |
| Neisseria | 0.9 ± 1.0 | 0.5 ± 0.7 | 0.15 | 2.3 ± 2.8 | 0.2 ± 0.5 | 0.03 |
| Paenibacillus | 0.4 ± 0.4 | 2.7 ± 3.1 | 0.02 | 0.5 ± 1.3 | 2.7 ± 3.2 | 0.04 |
| Parabacteroides | 0.0 ± 0.1 | <0.01 | 0.95 | 0.5 ± 0.6 | 0.0 ± 0.1 | 0.04 |
| Paracoccus | 0.6 ± 0.6 | 0.2 ± 0.4 | 0.09 | 0.3 ± 0.3 | 0.4 ± 0.7 | 0.65 |
| Plesiomonas | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.54 | 0.7 ± 1.1 | 0.2 ± 0.4 | 0.15 |
| Prevotella | <0.01 | <0.01 | 0.21 | 0.5 ± 0.7 | 0.0 ± 0.1 | 0.06 |
| Pseudomonas | 9.8 ± 24.3 | 0.1 ± 0.2 | 0.19 | 8.7 ± 22.8 | 5.1 ± 17.3 | 0.67 |
| Psychrobacter | 0.4 ± 0.5 | 0.0 ± 0.1 | 0.08 | <0.01 | 0.1 ± 0.1 | 0.62 |
| Rhodococcus | 0.7 ± 0.5 | <0.01 | 0.00 | 0.2 ± 0.2 | 0.1 ± 0.1 | 0.18 |
| Sebaldella | 0.1 ± 0.1 | 0.0 ± 0.0 | 0.05 | 0.7 ± 1.2 | 0.1 ± 0.3 | 0.13 |
| Sphingobacterium | 0.6 ± 1.9 | 0.1 ± 0.1 | 0.34 | <0.01 | 0.1 ± 0.1 | 0.31 |
| Stenotrophomonas | 1.7 ± 4.7 | 0.8 ± 1.9 | 0.56 | 0.3 ± 0.4 | 0.6 ± 1.3 | 0.41 |
| Streptococcus | 6.4 ± 4.1 | 1.1 ± 0.8 | 0.00 | 5.1 ± 5.0 | 2.4 ± 4.2 | 0.17 |
| Sutterella | <0.01 | 1.7 ± 1.8 | 0.01 | 0.0 ± 0.1 | <0.01 | 0.16 |
| Veillonella | 0.1 ± 0.1 | 0.1 ± 0.1 | 0.86 | 0.4 ± 0.5 | 0.2 ± 0.2 | 0.20 |
| Yersinia | 0.2 ± 0.5 | 0.2 ± 0.4 | 0.89 | 0.8 ± 2.1 | 1.0 ± 2.5 | 0.84 |
| Time of Sample Collection | Plant A (Log Copies L−1) | Plant B (Log Copies L−1) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| ciaB | pldA | ciaB | pldA | ||||||
| Influent | Effluent | Influent | Effluent | Influent | Effluent | Influent | Effluent | ||
| 2011 | August | 7.3 | 6.2 | 5.7 | 4.5 | 8.5 | 5.1 | 6.5 | n.d. |
| September | 7.8 | 5.8 | 5.8 | n.d. | 7.1 | 4.4 | 5.7 | n.d. | |
| October | 7.9 | 5.1 | 5.9 | n.d. | 5.4 | 6.0 | n.d. | n.d. | |
| November | 7.7 | 6.2 | 5.7 | n.d. | 8.1 | n.d. | n.d. | n.d. | |
| December | 8.0 | 5.6 | 5.8 | 4.8 | 7.9 | 7.7 | n.d. | 5.3 | |
| 2012 | January | 7.4 | n.d. | n.d. | n.d. | 8.1 | 6.3 | n.d. | 5.1 |
| February | 6.4 | n.d. | n.d. | n.d. | 5.7 | n.d. | n.d. | n.d. | |
| March | 7.4 | n.d. | 5.9 | n.d. | 8.6 | n.d. | 7.2 | n.d. | |
| April | n.d. | n.d. | n.d. | n.d. | 6.9 | 4.6 | n.d. | n.d. | |
| May | 6.9 | 5.2 | 5.9 | n.d. | 7.0 | n.d. | 6.0 | n.d. | |
| June | 5.7 | 6.2 | n.d. | 5.0 | 9.3 | 4.6 | 7.7 | n.d. | |
| July | 7.7 | 5.4 | 6.2 | n.d. | 9.1 | n.d. | 7.5 | n.d. | |
| Mean ± s.d. | 7.3 ± 0.7 | 5.7 ± 0.5 | 5.9 ± 0.2 | 4.8 ± 0.3 | 7.6 ± 1.2 | 5.5 ± 1.2 | 6.8 ± 0.8 | 5.2 ± 0.1 | |
| Log Reduction Tested | Log Reduction (mean ± s.d.) | |
|---|---|---|
| Plant A | Plant B | |
| 16S rRNA gene of total bacteria | 1.6 ± 0.4 (n = 12) | 1.5 ± 0.4 (n = 12) |
| 16S rRNA gene of Arcobacter | 1.7 ± 0.9 (n = 10) | 2.3 ± 1.5 (n = 9) |
| ciaB gene of Arcobacter | 1.7 ± 1.0 (n = 8) | 2.1 ± 1.8 (n = 7) |
| pldA gene of Arcobacter | 1.1 ± 0.1 (n = 2) | n.d. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghaju Shrestha, R.; Sherchan, S.P.; Kitajima, M.; Tanaka, Y.; Gerba, C.P.; Haramoto, E. Reduction of Arcobacter at Two Conventional Wastewater Treatment Plants in Southern Arizona, USA. Pathogens 2019, 8, 175. https://doi.org/10.3390/pathogens8040175
Ghaju Shrestha R, Sherchan SP, Kitajima M, Tanaka Y, Gerba CP, Haramoto E. Reduction of Arcobacter at Two Conventional Wastewater Treatment Plants in Southern Arizona, USA. Pathogens. 2019; 8(4):175. https://doi.org/10.3390/pathogens8040175
Chicago/Turabian StyleGhaju Shrestha, Rajani, Samendra P. Sherchan, Masaaki Kitajima, Yasuhiro Tanaka, Charles P. Gerba, and Eiji Haramoto. 2019. "Reduction of Arcobacter at Two Conventional Wastewater Treatment Plants in Southern Arizona, USA" Pathogens 8, no. 4: 175. https://doi.org/10.3390/pathogens8040175
APA StyleGhaju Shrestha, R., Sherchan, S. P., Kitajima, M., Tanaka, Y., Gerba, C. P., & Haramoto, E. (2019). Reduction of Arcobacter at Two Conventional Wastewater Treatment Plants in Southern Arizona, USA. Pathogens, 8(4), 175. https://doi.org/10.3390/pathogens8040175








