Diversity of Bacterial Biota in Capnodis tenebrionis (Coleoptera: Buprestidae) Larvae
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
Author Contributions
Funding
Conflicts of Interest
References
- Chornesky, E.A.; Bartuska, A.M.; Aplet, G.H.; Britton, K.O.; Cummings-Carlson, J.; Davis, F.W.; Eskow, J.; Gordon, D.R.; Gottschalk, K.W.; Haack, R.A.; et al. Science priorities for reducing the threat of invasive species to sustainable forestry. AIBS Bull. 2005, 55, 335–348. [Google Scholar] [CrossRef]
- Vasanthakumar, A.; Handelsman, J.O.; Schloss, P.D.; Bauer, L.S.; Raffa, K.F. Gut microbiota of an invasive subcortical beetle, Agrilus planipennis Fairmaire, across various life stages. Environ. Entomol. 2008, 37, 1344–1353. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Delalibera, I., Jr.; Handelsman, J.O.; Raffa, K.F. Bacteria associated with the guts of two wood boring beetles: Anoplophora glabripennis and Saperda vestita (Cerambycidae). Environ. Entomol. 2006, 35, 625–629. [Google Scholar] [CrossRef]
- Vasanthakumar, A.; Delalibera, I., Jr.; Handelsman, J.; Klepzig, K.D.; Schloss, P.D.; Raffa, K.F. Characterization of gut-associated bacteria in larvae and adults of the southern pine beetle, Dendroctonus frontalis Zimmermann. Environ. Entomol. 2006, 35, 1710–1717. [Google Scholar] [CrossRef]
- Douglas, A.E. The microbial dimension in insect nutritional ecology. Funct. Ecol. 2009, 23, 38–47. [Google Scholar] [CrossRef]
- Engel, P.; Moran, N.A. The gut microbiota of insects—Diversity in structure and function. FEMS Microbiol. Rev. 2016, 37, 699–735. [Google Scholar] [CrossRef]
- Welte, C.U.; de Graaf, R.M.; van den Bosch, T.J.; Op den Camp, H.J.; van Dam, N.M.; Jetten, M.S. Plasmids from the gut microbiome of cabbage root fly larvae encode SaxA that catalyzes the conversion of the plant toxin 2-phenylethyl isothiocyanate. Environ. Microbiol. 2016, 18, 1379–1390. [Google Scholar] [CrossRef]
- Hammer, T.J.; Bowers, M.D. Gut microbes may facilitate insect herbivory of chemically defended plants. Oecologia 2015, 179, 1–14. [Google Scholar] [CrossRef]
- Berasategui, A.; Salem, H.; Paetz, C.; Santoro, M.; Gershenzon, J.; Kaltenpoth, M.; Schmidt, A. Gut microbiota of the pine weevil degrades conifer diterpenes and increases insect fitness. Mol. Ecol. 2017, 26, 4099–4110. [Google Scholar] [CrossRef]
- Ben-Yehuda, S.; Assael, F.; Mendel, Z. Improved chemical control of Capnodis tenebrionis L. and C. carbonaria Klug (Coleoptera: Buprestidae) in stone-fruit plantations in Israel. Phytoparasitica 2000, 28, 27–41. [Google Scholar] [CrossRef]
- Gindin, G.; Mendel, Z.; Levitin, B.; Kumar, P.; Levi, T.; Shahi, P.; Khasdan, V.; Weinthal, D.; Kuznetsova, T.; Einav, M.; et al. The basis for rootstock resilient to Capnodis species: Screening for genes encoding δ-endotoxins from Bacillus thuringiensis. Pest Manag. Sci. 2014, 70, 1283–1290. [Google Scholar] [CrossRef] [PubMed]
- Strano, C.P.; Malacrinò, A.; Campolo, O.; Palmeri, V. Influence of host plant on Thaumetopoea pityocampa gut bacterial community. Microb. Ecol. 2018, 75, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.E.; Russell, J.A.; Moreau, C.S.; Kautz, S.; Sullam, K.E.; Hu, Y.I.; Basinger, U.; Mott, B.M.; Buck, N.; Wheeler, D.E. Highly similar microbial communities are shared among related and trophically similar ant species. Mol. Ecol. 2012, 21, 2282–2296. [Google Scholar] [CrossRef] [PubMed]
- Gómez Expósito, R.; Postma, J.; Raaijmakers, J.M.; De Bruijn, I. Diversity and activity of Lysobacter species from disease suppressive soils. Front. Microbiol. 2015, 6, 1243. [Google Scholar] [CrossRef]
- Ogura, J.; Toyoda, A.; Kurosawa, T.; Chong, A.L.; Chohnan, S.; Masaki, T. Purification, characterization, and gene analysis of cellulase (Cel8A) from Lysobacter sp. IB-9374. Biosci. Biotechnol. Biochem. 2006, 70, 2420–2428. [Google Scholar] [CrossRef]
- Puopolo, G.; Tomada, S.; Sonego, P.; Moretto, M.; Engelen, K.; Perazzolli, M.; Pertot, I. The Lysobacter capsici AZ78 genome has a gene pool enabling it to interact successfully with phytopathogenic microorganisms and environmental factors. Front. Microbiol. 2016, 7, 96. [Google Scholar] [CrossRef]
- Palumbo, J.D.; Sullivan, R.F.; Kobayashi, D.Y. Molecular characterization and expression in Escherichia coli of three β-1,3-glucanase genes from Lysobacter enzymogenes strain N4-7. J. Bacteriol. 2003, 185, 4362–4370. [Google Scholar] [CrossRef]
- Chen, J.; Moore, W.H.; Yuen, G.Y.; Kobayashi, D.; Caswell-Chen, E.P. Influence of Lysobacter enzymogenes strain C3 on nematodes. J. Nematol. 2006, 38, 233–239. [Google Scholar]
- Zhang, Z.; Yuen, G.Y. Biological control of Bipolaris sorakiniana on tall fescue by Stenotrophomonas maltophilia strain C3. Phytopathology 1999, 89, 817–822. [Google Scholar] [CrossRef]
- Yuen, G.Y.; Steadman, J.R.; Lindgren, D.T.; Schaff, D.; Jochum, C. Bean rust biological control using bacterial agents. Crop Prot. 2001, 20, 395–402. [Google Scholar] [CrossRef]
- Jochum, C.C.; Osborne, L.E.; Yuen, G.Y. Fusarium head blight biological control with Lysobacter enzymogenes. Biol. Control 2006, 39, 336–344. [Google Scholar] [CrossRef]
- Postma, J.; Schilder, M.T.; Bloem, J.; Van Leeuwen-Haagsma, W.K. Soil suppressiveness and functional diversity of the soil microflora in organic farming systems. Soil Biol. Biochem. 2008, 40, 2394–2406. [Google Scholar] [CrossRef]
- Brucker, R.M.; Baylor, C.M.; Walters, R.L.; Lauer, A.; Harris, R.N.; Minbiole, K.P.C. The identification of 2,4-diacetylphloroglucinol as an antifungal metabolite produced by cutaneous bacteria of the salamander Plethodon cinereus. J. Chem. Ecol. 2008, 34, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.P.; McDonald, I.R.; Wood, A.P. The family Methylobacteriaceae. In The Prokaryotes; Springer: Berlin/Heidelberg, Germany, 2014; pp. 313–340. [Google Scholar]
- Liu, Y.H.; Guo, J.W.; Salam, N.; Li, L.; Zhang, Y.G.; Han, J.; Mohamad, O.A.; Li, W.J. Culturable endophytic bacteria associated with medicinal plant Ferula songorica: Molecular phylogeny, distribution and screening for industrially important traits. 3 Biotech 2016, 6, 209. [Google Scholar] [CrossRef] [PubMed]
- Msaddak, A.; Duran, D.; Rejili, M.; Mars, M.; Argueso, T.R.; Imperial, J.; Palacios, J.; Rey, L. Diverse bacteria affiliated with the genera Microvirga, Phyllobacterium, and Bradyrhizobium nodulate Lupinus micranthus growing in soils of Northern Tunisia. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Zhou, N.; Sun, Y.T.; Chen, D.W.; Du, W.; Yang, H.; Liu, S.J. Harnessing microfluidic streak plate technique to investigate the gut microbiome of Reticulitermes chinensis. MicrobiologyOpen 2018, e00654. [Google Scholar] [CrossRef]
- Bignell, D.E. Morphology, physiology, biochemistry and functional design of the termite gut: An evolutionary wonderland. In Biology of Termites: A Modern Synthesis; Bignell, D.E., Roisin, Y., Lo, N., Eds.; Springer: Dordrecht, The Netherlands, 2011; pp. 375–412. [Google Scholar]
- Setiawan, R.; Sumerta, I.N.; Anita, S.; Narakusumo, R.P. Isolation and Enzymatic Activities of Bacteria Associated to Weevils (Coleoptera: Curculionidae). In Proceedings of the 4th International Symposium on Innovative Bioproduction, Bogor, Indonesia, 27 September 2017; pp. 39–47. [Google Scholar]
- Huang, S.; Sheng, P.; Zhang, H. Isolation and identification of cellulolytic bacteria from the gut of Holotrichia parallela larvae (Coleoptera: Scarabaeidae). Int. J. Mol. Sci. 2012, 13, 2563–2577. [Google Scholar] [CrossRef]
- Liang, X.; Fu, Y.; Tong, L.; Liu, H. Microbial shifts of the silkworm larval gut in response to lettuce leaf feeding. Appl. Microbiol. Biotechnol. 2014, 98, 3769–3776. [Google Scholar] [CrossRef]
- Danismazoglu, M.; Demir, İ.; Sevim, A.; Demirbag, Z.; Nalcacioglu, R. An investigation on the bacterial flora of Agriotes lineatus (Coleoptera: Elateridae) and pathogenicity of the flora members. Crop Prot. 2012, 40, 1–7. [Google Scholar] [CrossRef]
- Ince, I.A.; Demirbağ, Z.; Kati, H. Arthrobacter pityocampae sp. nov., isolated from Thaumetopoea pityocampa (Lep., Thaumetopoeidae). Int. J. Syst. Evol. Microbiol. 2014, 64, 3384–3389. [Google Scholar] [CrossRef]
- Latella, L.; Castioni, A.; Bignotto, L.; Salvetti, E.; Torriani, S.; Felis, G.E. Exploring gut microbiota composition of the cave beetles Neobathyscia pasai Ruffo, 1950 and Neobathyscia mancinii Jeannel, 1924 (Leiodidae; Cholevinae). Boll. Mus. Civ. Stor. Nat. Verona 2017, 41, 3–24. [Google Scholar]
- Thayer, D.W. Facultative wood-digesting bacteria from the hind-gut of the termite Reticulitermes hesperus. J. Gen. Microbiol. 1976, 95, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Normand, P.; Daffonchio, D.; Gtari, M. The Family Geodermatophilaceae. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014. [Google Scholar]
- Dickey, A.M.; Trease, A.J.; Jara-Cavieres, A.; Kumar, V.; Christenson, M.K.; Potluri, L.P.; Morgan, J.K.; Shatters, R.G., Jr.; Mckenzie, C.L.; Davis, P.H.; et al. Estimating bacterial diversity in Scirtothrips dorsalis (Thysanoptera: Thripidae) via next generation sequencing. Fla. Entomol. 2014, 97, 362–366. [Google Scholar] [CrossRef] [PubMed]
- Reyes, R.D.; Cafaro, M.J. Paratrechina longicornis ants in a tropical dry forest harbor specific Actinobacteria diversity. J. Basic Microbiol. 2015, 55, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Warnecke, F.; Luginbühl, P.; Ivanova, N.; Ghassemian, M.; Richardson, T.H.; Stege, J.T.; Cayouette, M.; McHardy, A.C.; Djordjevic, G.; Aboushadi, N.; et al. Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 2007, 450, 560–565. [Google Scholar] [CrossRef]
- Kikuchi, Y.; Hayatsu, M.; Hosokawa, T.; Nagayama, A.; Tago, K.; Fukatsu, T. Symbiont-mediated insecticide resistance. Proc. Natl. Acad. Sci. USA 2012, 109, 8618–8622. [Google Scholar] [CrossRef] [PubMed]
- Ping, L.; Büchler, R.; Mithöfer, A.; Svatoš, A.; Spiteller, D.; Dettner, K.; Gmeiner, S.; Piel, J.; Schlott, B.; Boland, W. A novel Dps-type protein from insect gut bacteria catalyses hydrolysis and synthesis of N-acyl amino acids. Environ. Microbiol. 2007, 9, 1572–1583. [Google Scholar] [CrossRef] [PubMed]
- Broderick, N.A.; Robinson, C.J.; McMahon, M.D.; Holt, J.; Handelsman, J.; Raffa, K.F. Contributions of gut bacteria to Bacillus thuringiensis-induced mortality vary across a range of Lepidoptera. BMC Biol. 2009, 7, 11. [Google Scholar] [CrossRef]
- Gindin, G.; Kuznetsowa, T.; Protasov, A.; Ben-Yehuda, S.; Mendel, Z. Artificial diet for two flat headed borers, Capnodis spp. (Coleoptera: Buprestidae). Eur. J. Entomol. 2009, 106, 573–581. [Google Scholar] [CrossRef]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
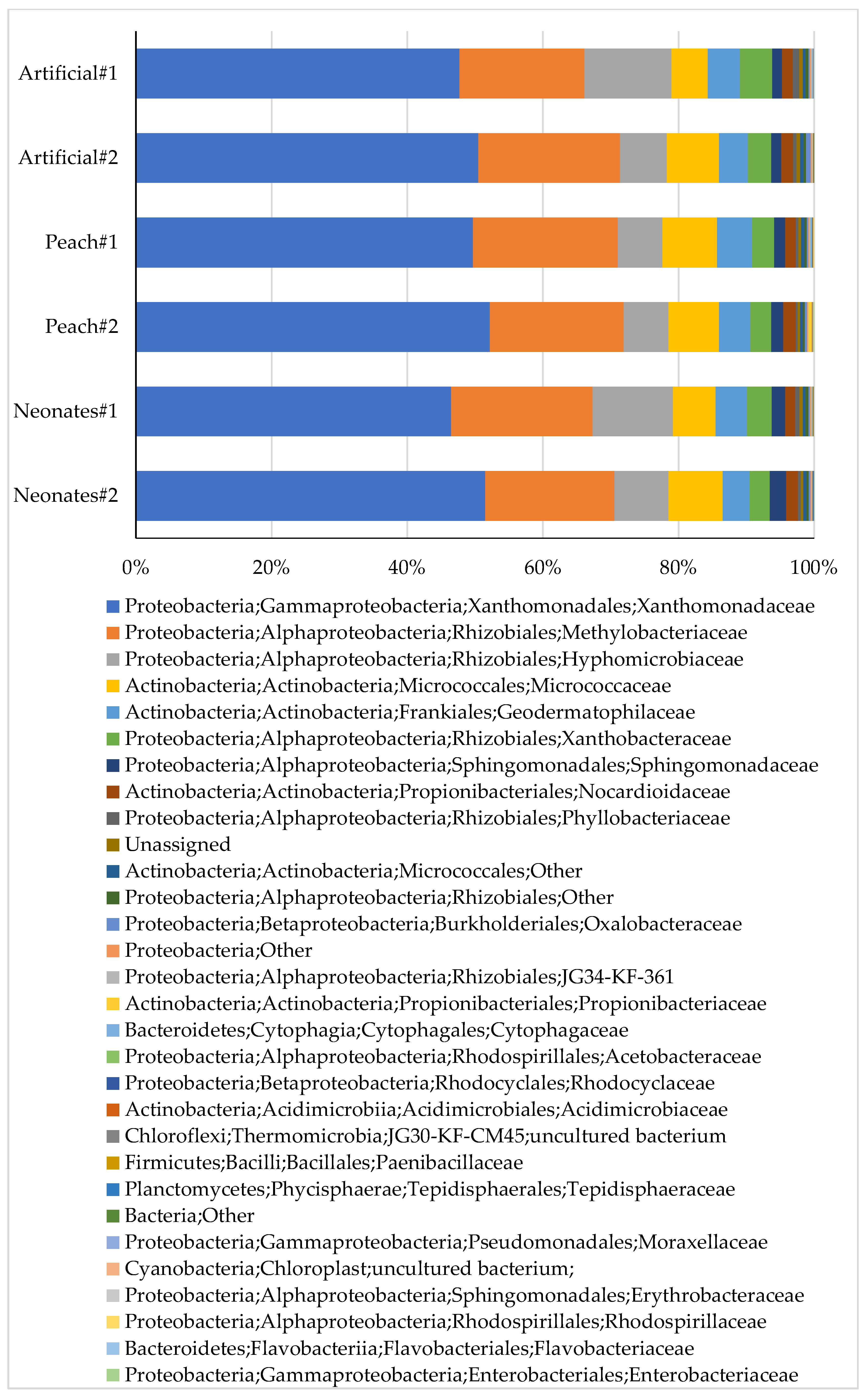
| Phylum | Artificial#1 | Artificial#2 | Neonates#1 | Neonates#2 | Peach#1 | Peach#2 |
|---|---|---|---|---|---|---|
| Proteobacteria | 87.22% | 85.12% | 86.60% | 85.21% | 83.86% | 84.69% |
| Actinobacteria | 12.09% | 14.25% | 12.83% | 14.23% | 15.38% | 14.85% |
| Unassigned | 0.54% | 0.56% | 0.49% | 0.37% | 0.42% | 0.35% |
| Others 1 | 0.15% | 0.07% | 0.08% | 0.18% | 0.34% | 0.11% |
| Class | Artificial#1 | Artificial#2 | Neonates#1 | Neonates#2 | Peach#1 | Peach#2 |
|---|---|---|---|---|---|---|
| Gammaproteobacteria | 47.69% | 50.47% | 46.49% | 51.49% | 49.67% | 52.31% |
| Alphaproteobacteria | 39.13% | 33.80% | 39.65% | 33.28% | 33.79% | 31.88% |
| Actinobacteria | 12.07% | 14.23% | 12.79% | 14.21% | 15.35% | 14.83% |
| Unassigned | 0.54% | 0.56% | 0.49% | 0.37% | 0.42% | 0.35% |
| Others 1 | 0.58% | 0.94% | 0.57% | 0.64% | 0.77% | 0.63% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barak, H.; Kumar, P.; Zaritsky, A.; Mendel, Z.; Ment, D.; Kushmaro, A.; Ben-Dov, E. Diversity of Bacterial Biota in Capnodis tenebrionis (Coleoptera: Buprestidae) Larvae. Pathogens 2019, 8, 4. https://doi.org/10.3390/pathogens8010004
Barak H, Kumar P, Zaritsky A, Mendel Z, Ment D, Kushmaro A, Ben-Dov E. Diversity of Bacterial Biota in Capnodis tenebrionis (Coleoptera: Buprestidae) Larvae. Pathogens. 2019; 8(1):4. https://doi.org/10.3390/pathogens8010004
Chicago/Turabian StyleBarak, Hana, Pradeep Kumar, Arieh Zaritsky, Zvi Mendel, Dana Ment, Ariel Kushmaro, and Eitan Ben-Dov. 2019. "Diversity of Bacterial Biota in Capnodis tenebrionis (Coleoptera: Buprestidae) Larvae" Pathogens 8, no. 1: 4. https://doi.org/10.3390/pathogens8010004
APA StyleBarak, H., Kumar, P., Zaritsky, A., Mendel, Z., Ment, D., Kushmaro, A., & Ben-Dov, E. (2019). Diversity of Bacterial Biota in Capnodis tenebrionis (Coleoptera: Buprestidae) Larvae. Pathogens, 8(1), 4. https://doi.org/10.3390/pathogens8010004








