Characteristics of Listeria Monocytogenes Strains Persisting in a Meat Processing Facility over a 4-Year Period
Abstract
1. Introduction
2. Results
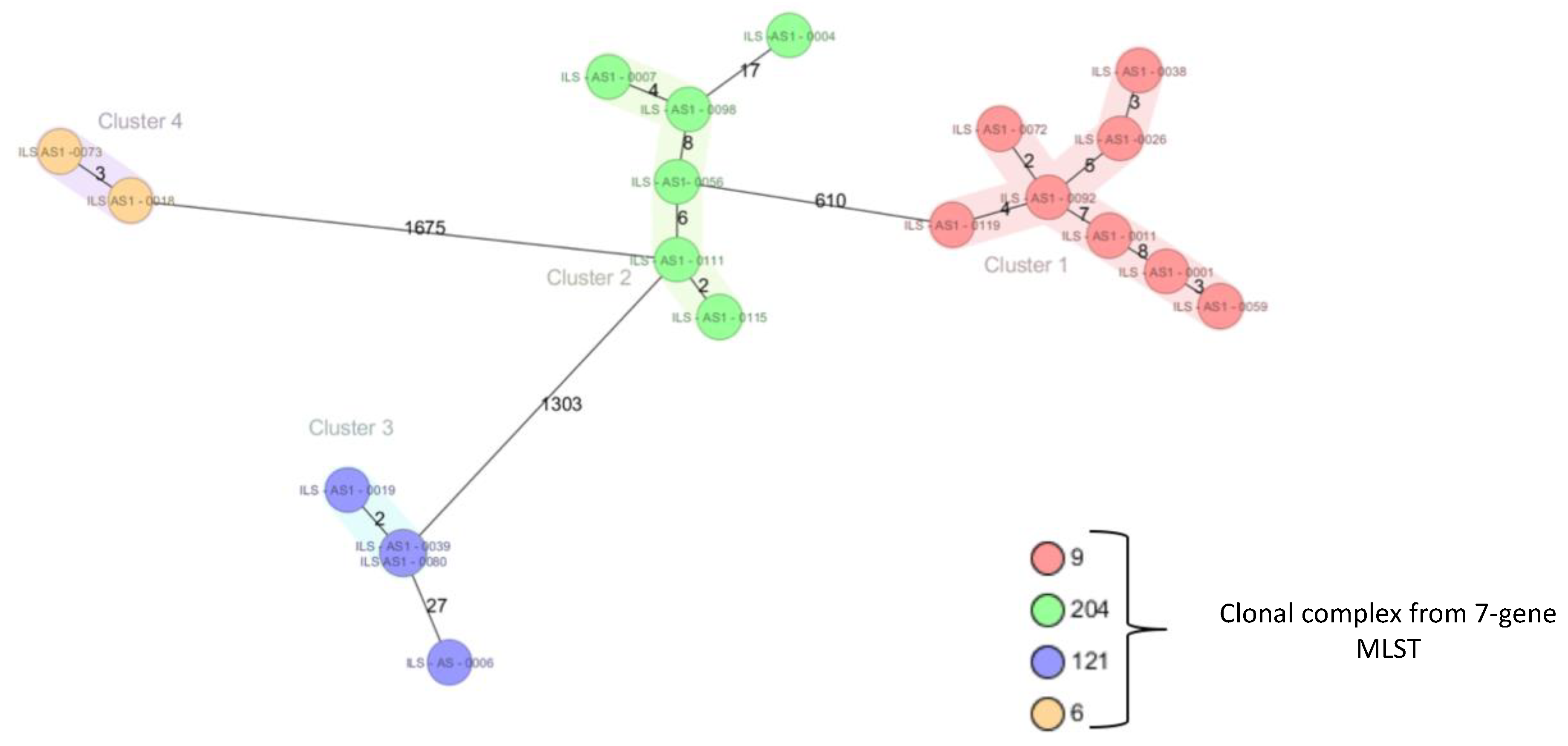
2.1. Subtyping of 124 Listeria Monocytogenes Isolated from a Swiss Meat Plant
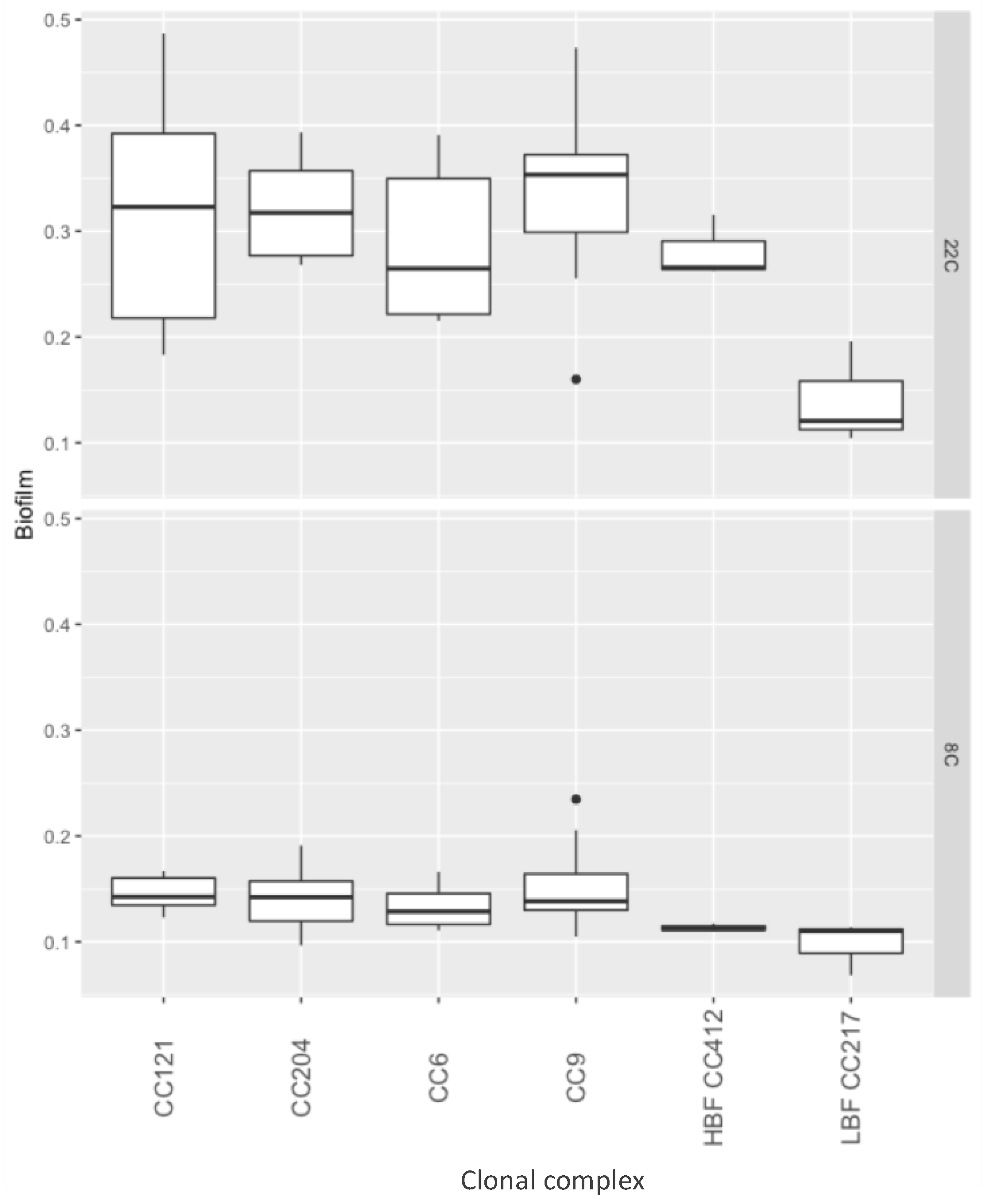
2.2. Biofilm Formation
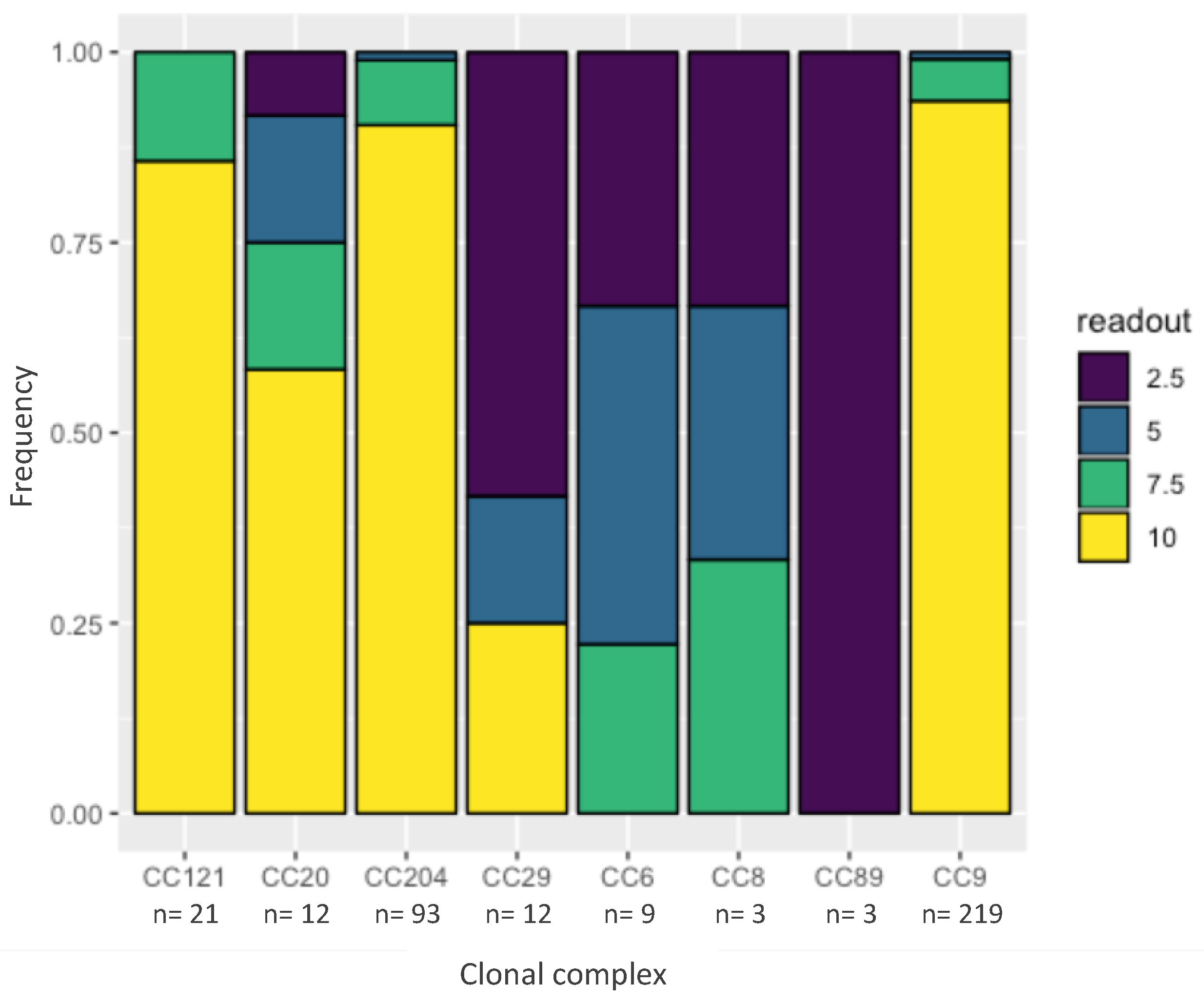
2.3. Tolerance to Benzalkonium Chloride
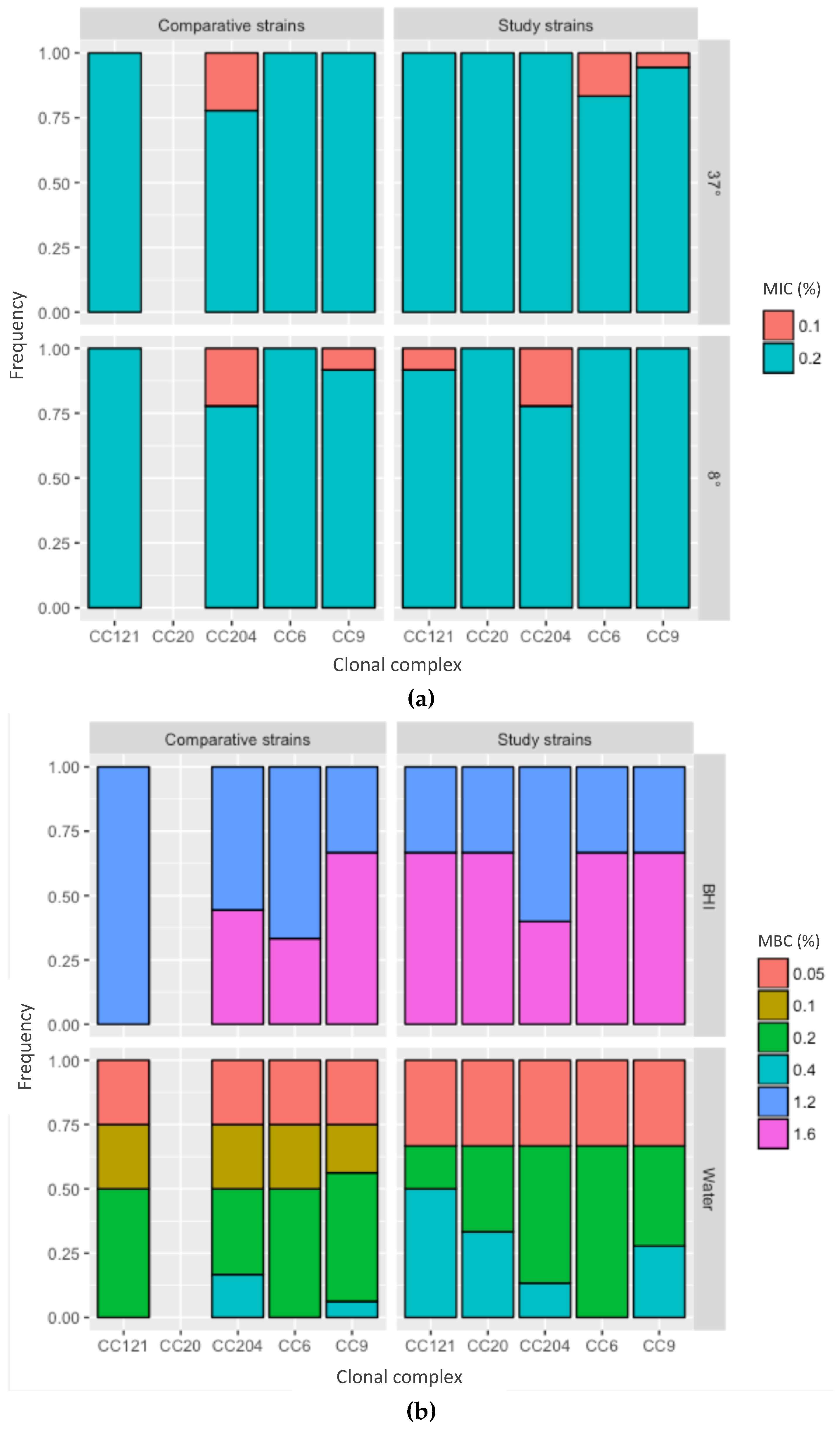
2.4. Minimal Inhibitory Concentration and Minimal Bactericidal Concentration of Peracetic Acid against L. monocytogenes
3. Discussion
4. Material and Methods
4.1. Bacterial Strains and Preparation of Bacterial Cultures
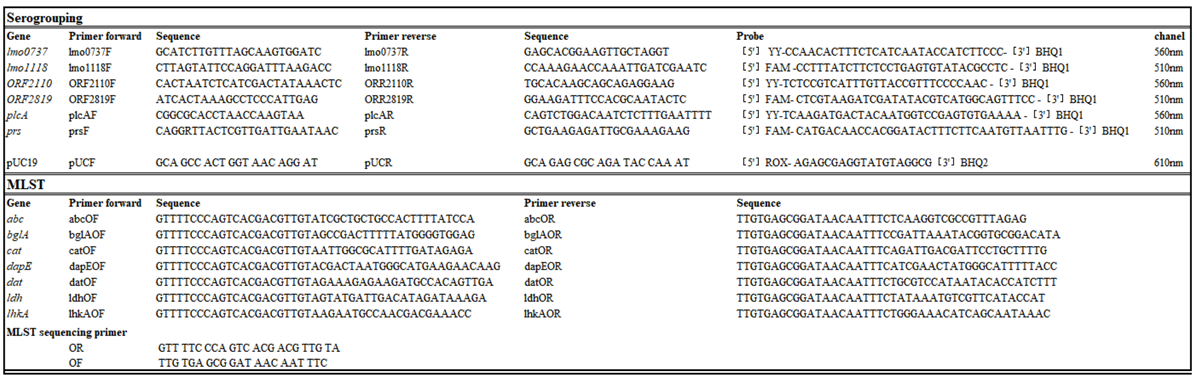
4.2. Serogrouping by qPCR
4.3. Multi Locus Sequence Typing (MLST)
4.4. Whole Genome Sequencing
4.5. Biofilm Formation
4.6. Tolerance to Benzalkonium Chloride
4.7. Minimal Inhibitory Concentration and Minimal Bactericidal Concentration of Peracetic Acid against L. monocytogenes
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barton Behravesh, C.; Jones, T.F.; Vugia, D.J.; Long, C.; Marcus, R.; Smith, K.; Thomas, S.; Zansky, S.; Fullerton, K.E.; Henao, O.L.; et al. Deaths associated with bacterial pathogens transmitted commonly through food: Foodborne diseases active surveillance network (FoodNet), 1996–2005. J. Infect. Dis. 2011, 204, 263–267. [Google Scholar] [CrossRef] [PubMed]
- de Valk, H.; Jacquet, C.; Goulet, V.; Vaillant, V.; Perra, A.; Simon, F.; Desenclos, J.C.; Martin, P. Surveillance of Listeria infections in Europe. Euro Surveill. 2005, 10, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Popovic, I.; Heron, B.; Covacin, C. Listeria: An Australian perspective (2001–2010). Foodborne Pathog. Dis. 2014, 11, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Werber, D.; Hille, K.; Frank, C.; Dehnert, M.; Altmann, D.; Müller-Nordhorn, J.; Koch, J.; Stark, K. Years of potential life lost for six major enteric pathogens, Germany, 2004–2008. Epidemiol. Infect. 2012, 141, 961–968. [Google Scholar] [CrossRef] [PubMed]
- Scallan, E.; Hoekstra, R.M.; Mahon, B.E.; Jones, T.F.; Griffin, P.M. An assessment of the human health impact of seven leading foodborne pathogens in the United States using disability adjusted life years. Epidemiol. Infect. 2015, 143, 2795–2804. [Google Scholar] [CrossRef] [PubMed]
- Olsen, S.J.; Patrick, M.; Hunter, S.B.; Reddy, V.; Kornstein, L.; MacKenzie, W.R.; Lane, K.; Bidol, S.; Stoltman, G.A.; Frye, D.M.; et al. Multistate outbreak of Listeria monocytogenes infection linked to delicatessen turkey meat. Clin. Infect. Dis. 2005, 40, 962–967. [Google Scholar] [CrossRef] [PubMed]
- Orsi, R.H.; Borowsky, M.L.; Lauer, P.; Young, S.K.; Nusbaum, C.; Galagan, J.E.; Birren, B.W.; Ivy, R.A.; Sun, Q.; Graves, L.M.; et al. Short-term genome evolution of Listeria monocytogenes in a non-controlled environment. BMC Genom. 2008, 9, 539. [Google Scholar] [CrossRef] [PubMed]
- Vongkamjan, K.; Roof, S.; Stasiewicz, M.J.; Wiedmann, M. Persistent Listeria monocytogenes subtypes isolated from a smoked fish processing facility included both phage susceptible and resistant isolates. Food Microbiol. 2013, 35, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Leong, D.; Alvarez-Ordóñez, A.; Jordan, K. Monitoring occurrence and persistence of Listeria monocytogenes in foods and food processing environments in the Republic of Ireland. Front. Microbiol. 2014, 5, 436. [Google Scholar] [CrossRef] [PubMed]
- Ruppitsch, W.; Pietzka, A.; Prior, K.; Bletz, S.; Fernandez, H.L.; Allerberger, F.; Harmsen, D.; Mellmann, A. Defining and Evaluating a Core Genome Multilocus Sequence Typing Scheme for Whole-Genome Sequence-Based Typing of Listeria monocytogenes. J. Clin. Microbol. 2015, 53, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Borucki, M.K.; Peppin, J.D.; White, D.; Loge, F.; Call, D.R. Variation in Biofilm Formation among Strains of Listeria monocytogenes. Appl. Environ. Microbiol. 2003, 69, 7336–7342. [Google Scholar] [CrossRef] [PubMed]
- Agresti, A. Categorical Data Analysis; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2002. [Google Scholar]
- Elhanafi, D.; Dutta, V.; Kathariou, S. Genetic characterization of plasmid-associated benzalkonium chloride resistance determinants in a Listeria monocytogenes strain from the 1998-1999 outbreak. Appl. Environ. Microbiol. 2010, 76, 8231–8238. [Google Scholar] [CrossRef] [PubMed]
- Muller, A.; Rychli, K.; Muhterem-Uyar, M.; Zaiser, A.; Stessl, B.; Guinane, C.M.; Cotter, P.D.; Wagner, M.; Schmitz-Esser, S. Tn6188—A novel transposon in Listeria monocytogenes responsible for tolerance to benzalkonium chloride. PLoS ONE 2013, 8, e76835. [Google Scholar] [CrossRef] [PubMed]
- Kovacevic, J.; Ziegler, J.; Walecka-Zacharska, E.; Reimer, A.; Kitts, D.D.; Gilmour, M.W. Tolerance of Listeria monocytogenes to Quaternary Ammonium Sanitizers Is Mediated by a Novel Efflux Pump Encoded by emrE. Appl. Environ. Microbiol. 2015, 82, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Maury, M.M.; Tsai, Y.-H.; Charlier, C.; Touchon, M.; Chenal-Francisque, V.; Leclercq, A.; Criscuolo, A.; Gaultier, C.; Roussel, S.; Brisabois, A.; et al. Uncovering Listeria monocytogenes hypervirulence by harnessing its biodiversity. Nat. Genet. 2016, 48, 308–313. [Google Scholar] [CrossRef] [PubMed]
- Martín, B.; Perich, A.; Gómez, D.; Yangüela, J.; Rodríguez, A.; Garriga, M.; Aymerich, T. Diversity and distribution of Listeria monocytogenes in meat processing plants. Food Microbiol. 2014, 44, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Ebner, R.; Stephan, R.; Althaus, D.; Brisse, S.; Maury, M.; Tasara, T. Phenotypic and genotypic characteristics of Listeria monocytogenes strains isolated during 2011–2014 from different food matrices in Switzerland. Food Control 2015, 321–326. [Google Scholar] [CrossRef]
- Nielsen, E.M.; Björkman, J.T.; Kiil, K.; Grant, K.; Dallman, T.; Painset, A.; Amar, C.; Roussel, S.; Guillier, L.; Félix, B.; et al. Closing gaps for performing a risk assessment on Listeria monocytogenes in ready-to-eat (RTE) foods: Activity 3, the comparison of isolates from different compartments along the food chain, and from humans using whole genome sequencing (WGS) analysis. EFSA Support. Publ. 2017, 14, 539. [Google Scholar] [CrossRef]
- Althaus, D.; Lehner, A.; Brisse, S.; Maury, M.; Tasara, T.; Stephan, R. Characterization of Listeria monocytogenes Strains Isolated During 2011–2013 from Human Infections in Switzerland. Foodborne Pathog. Dis. 2014, 11, 753–758. [Google Scholar] [CrossRef] [PubMed]
- Harvey, J.; Keenan, K.P.; Gilmour, A. Assessing biofilm formation by Listeria monocytogenes strains. Food Microbiol. 2007, 24, 380–392. [Google Scholar] [CrossRef] [PubMed]
- Heir, E.; Møretrø, T.; Simensen, A.; Langsrud, S. Listeria monocytogenes strains show large variations in competitive growth in mixed culture biofilms and suspensions with bacteria from food processing environments. Int. J. Food Microbiol. 2018, 275, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Lundén, J.M.; Miettinen, M.K.; Autio, T.J.; Korkeala, H.J. Persistent Listeria monocytogenes strains show enhanced adherence to food contact surface after short contact times. J. Food Prot. 2000, 63, 1204–1207. [Google Scholar] [CrossRef] [PubMed]
- Djordjevic, D.; Wiedmann, M.; McLandsborough, L.A. Microtiter plate assay for assessment of Listeria monocytogenes biofilm formation. Appl. Environ. Microbiol. 2002, 68, 2950–2958. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, V.; Wiedmann, M.; Teixeira, P.; Stasiewicz, M.J. Listeria monocytogenes persistence in food-associated environments: Epidemiology, strain characteristics, and implications for public health. J. Food Prot. 2014, 77, 150–170. [Google Scholar] [CrossRef] [PubMed]
- Meier, A.B.; Guldimann, C.; Markkula, A.; Pöntinen, A.; Korkeala, H.; Tasara, T. Comparative Phenotypic and Genotypic Analysis of Swiss and Finnish Listeria monocytogenes Isolates with Respect to Benzalkonium Chloride Resistance. Front. Microbiol. 2017, 8, 397. [Google Scholar] [CrossRef] [PubMed]
- Hegstad, K.; Langsrud, S.; Lunestad, B.T.; Scheie, A.A.; Sunde, M.; Yazdankhah, S.P. Does the wide use of quaternary ammonium compounds enhance the selection and spread of antimicrobial resistance and thus threaten our health? Microb. Drug Resist. 2010, 16, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Poimenidou, S.V.; Dalmasso, M.; Papadimitriou, K.; Fox, E.M.; Skandamis, P.N.; Jordan, K. Virulence Gene Sequencing Highlights Similarities and Differences in Sequences in Listeria monocytogenes Serotype 1/2a and 4b Strains of Clinical and Food Origin From 3 Different Geographic Locations. Front. Microbiol. 2018, 9, 1103. [Google Scholar] [CrossRef] [PubMed]
- Skowron, K.; Hulisz, K.; Gryń, G.; Olszewska, H.; Wiktorczyk, N.; Paluszak, Z. Comparison of selected disinfectants efficiency against Listeria monocytogenes biofilm formed on various surfaces. Int. Microbiol. 2018, 21, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Oxaran, V.; Dittmann, K.K.; Lee, S.H.I.; Chaul, L.T.; Fernandes de Oliveira, C.A.; Corassin, C.H.; Alves, V.F.; De Martinis, E.C.P.; Gram, L. Behavior of Foodborne Pathogens Listeria monocytogenes and Staphylococcus aureus in Mixed-Species Biofilms Exposed to Biocides. Appl. Environ. Microbiol. 2018, 84, 627. [Google Scholar] [CrossRef] [PubMed]
- Saá Ibusquiza, P.; Herrera, J.J.R.; Cabo, M.L. Resistance to benzalkonium chloride, peracetic acid and nisin during formation of mature biofilms by Listeria monocytogenes. Food Microbiol. 2011, 28, 418–425. [Google Scholar] [CrossRef] [PubMed]
- da Silva Fernandes, M.; Kabuki, D.Y.; Kuaye, A.Y. Behavior of Listeria monocytogenes in a multi-species biofilm with Enterococcus faecalis and Enterococcus faecium and control through sanitation procedures. Int. J. Food Microbiol. 2015, 200, 5–12. [Google Scholar] [CrossRef] [PubMed]
- Winniczuk, P.P.; Parish, M.E. Minimum inhibitory concentrations of antimicrobials against micro-organisms related to citrus juice. Food Microbiol. 1997, 14, 373–381. [Google Scholar] [CrossRef]
- Alasri, A.; Roques, C.; Michel, G.; Cabassud, C.; Aptel, P. Bactericidal properties of peracetic acid and hydrogen peroxide, alone and in combination, and chlorine and formaldehyde against bacterial water strains. Can. J. Microbiol. 1992, 38, 635–642. [Google Scholar] [CrossRef] [PubMed]
- Vieira, R.G.L.; da Silva Moraes, T.; de Oliveira Silva, L.; Bianchi, T.C.; Veneziani, R.C.S.; Ambrósio, S.R.; Bastos, J.K.; Pires, R.H.; Martins, C.H.G. In vitro studies of the antibacterial activity of Copaifera spp. oleoresins, sodium hypochlorite, and peracetic acid against clinical and environmental isolates recovered from a hemodialysis unit. Antimicrob. Resist. Infect. Control 2018, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Kastbjerg, V.G.; Gram, L. Industrial disinfectants do not select for resistance in Listeria monocytogenes following long term exposure. Int. J. Food Microbiol. 2012, 160, 11–15. [Google Scholar] [CrossRef] [PubMed]
- Vitullo, M.; Grant, K.A.; Sammarco, M.L.; Tamburro, M.; Ripabelli, G.; Amar, C.F.L. Real-time PCRs assay for serogrouping Listeria monocytogenes and differentiation from other Listeria spp. Mol. Cell. Probes 2013, 27, 68–70. [Google Scholar] [CrossRef] [PubMed]
- Yanisch-Perron, C.; Vieira, J.; Messing, J. Improved M13 phage cloning vectors and host strains: Nucleotide sequences of the M13mpl8 and pUC19 vectors. Gene 1985, 33, 103–119. [Google Scholar] [CrossRef]
- Ragon, M.; Wirth, T.; Hollandt, F.; Lavenir, R.; Lecuit, M.; Le, M.A.; Brisse, S. A new perspective on Listeria monocytogenes evolution. PLoS Pathog. 2008, 4, e1000146. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Langsrud, S.; Sidhu, M.S.; Heir, E.; Holck, A. Bacterial disinfectant resistance—A challenge for the food industry. Int. J. Biodeter. Biodegrad. 2003, 51, 283–290. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Jordan, K.; Fox, E.M.; Wagner, M. (Eds.) Listeria monocytogenes—Methods and Protocols; Springer: New York, NY, USA, 2014; Volume 1157. [Google Scholar]
- Wiegand, I.; Hilpert, K.; Hancock, R.E.W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, A.; Bruun Brockhoff, P.; Bojesen Christensen, H. lmerTest: Tests in Linear Mixed Effects Models. 2016. Available online: https://CRAN. R-project. org/package=lmerTest (accessed on 14 December 2018).
- Lenth, R.V. Least-Squares Means: The RPackage lsmeans. J. Stat. Softw. 2016, 69, 1–33. [Google Scholar] [CrossRef]
- Venable, W.N.; Ripley, B.D. Modern Applied Statistics with S, 4th ed.; Springer: Berlin/Heidelberg, Germany, 2002. [Google Scholar]
- Wickham, H. ggplot2: Elegant graphics for data analysis. In ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar]
| Strain | Isolation Date | Source | Purpose | Serogroup | Sequence Type | CC | WGS | brcABC | qacH | emrE | Lineage |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ILS AS1-0001 | 2015/10/15 | FPE | Study strain | 2 | 9 | 9 | 1 | absent | absent | absent | II |
| ILS AS1-0002 | 2015/10/19 | FP | Study strain | 1 | 121 | 121 | n/a | n/a | n/a | II | |
| ILS AS1-0003 | 2015/11/9 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0004 | 2015/11/14 | FPE | Study strain | 1 | 204 | 204 | 2* | present | absent | absent | II |
| ILS AS1-0005 | 2016/2/2 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0006 | 2016/3/9 | FPE | Study strain | 1 | 121 | 121 | 3* | absent | present | absent | II |
| ILS AS1-0007 | 2016/10/20 | FP | Study strain | 1 | 204 | 204 | 2 | present | absent | absent | II |
| ILS AS1-0008 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0009 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0010 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0011 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | 1 | present | absent | absent | II |
| ILS AS1-0012 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0013 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0014 | 2017/1/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0015 | 2017/6/26 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0016 | 2017/6/29 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0017 | 2017/7/7 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0018 | 2017/7/7 | FPE | Study strain | 4 | 6 | 6 | 4 | absent | absent | absent | I |
| ILS AS1-0019 | 2017/8/10 | FP | Study strain | 1 | 121 | 121 | 3 | absent | present | absent | II |
| ILS AS1-0020 | 2017/9/12 | FP | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0021 | 2017/9/13 | FP | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0022 | 2017/9/18 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0023 | 2017/9/18 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0024 | 2017/11/6 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0025 | 2017/11/6 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0026 | 2017/11/13 | FPE | Study strain | 1 | 204 | 204 | 1 | absent | absent | absent | II |
| ILS AS1-0027 | 2017/11/23 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0028 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0029 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0030 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0031 | 2017/12/5 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0032 | 2017/12/7 | FPE | Study strain | 1 | 20 | 20 | n/a | n/a | n/a | II | |
| ILS AS1-0033 | 2017/12/7 | FPE | Study strain | 1 | 20 | 20 | n/a | n/a | n/a | II | |
| ILS AS1-0034 | 2017/12/7 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0035 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0036 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0037 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0038 | 2017/12/5 | FPE | Study strain | 2 | 9 | 9 | 1 | absent | absent | absent | II |
| ILS AS1-0039 | 2017/12/14 | FPE | Study strain | 1 | 121 | 121 | 3 | absent | present | absent | II |
| ILS AS1-0040 | 2018/1/23 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0041 | 2018/1/23 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0042 | 2018/1/23 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0043 | 2018/1/23 | FPE | Study strain | 1 | 29 | 29 | n/a | n/a | n/a | II | |
| ILS AS1-0044 | 2018/1/23 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0045 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0046 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0047 | 2018/1/25 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0048 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0049 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0050 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0051 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0052 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0053 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0054 | 2018/1/25 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0055 | 2018/1/25 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0056 | 2018/1/25 | FPE | Study strain | 1 | 204 | 204 | 2 | present | absent | absent | II |
| ILS AS1-0057 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0058 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0059 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | 1 | present | absent | absent | II |
| ILS AS1-0060 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0061 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0062 | 2018/1/26 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0063 | 2018/1/26 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0064 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0065 | 2018/1/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0066 | 2018/1/25 | FPE | Study strain | 1 | 391 | 89 | n/a | n/a | n/a | II | |
| ILS AS1-0067 | 2018/2/6 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0068 | 2018/2/6 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0069 | 2018/2/7 | FPE | Study strain | 1 | 29 | 29 | n/a | n/a | n/a | II | |
| ILS AS1-0070 | 2018/2/6 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0071 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0072 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | 1 | present | absent | absent | II |
| ILS AS1-0073 | 2018/2/9 | FPE | Study strain | 4 | 6 | 6 | 4 | absent | absent | absent | I |
| ILS AS1-0074 | 2018/2/9 | FPE | Study strain | 1 | 29 | 29 | n/a | n/a | n/a | II | |
| ILS AS1-0075 | 2018/2/9 | FPE | Study strain | 1 | 8 | 8 | n/a | n/a | n/a | II | |
| ILS AS1-0076 | 2018/2/9 | FPE | Study strain | 1 | 20 | 20 | n/a | n/a | n/a | II | |
| ILS AS1-0077 | 2018/2/9 | FPE | Study strain | 1 | 29 | 29 | n/a | n/a | n/a | II | |
| ILS AS1-0078 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0079 | 2018/2/9 | FPE | Study strain | 1 | 121 | 121 | n/a | n/a | n/a | II | |
| ILS AS1-0080 | 2018/2/9 | FPE | Study strain | 1 | 121 | 121 | 3 | absent | present | absent | II |
| ILS AS1-0081 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0082 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0083 | 2018/2/9 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0084 | 2018/2/9 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0085 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0086 | 2018/2/9 | FPE | Study strain | 1 | 20 | 20 | n/a | n/a | n/a | II | |
| ILS AS1-0087 | 2018/2/9 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0088 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0089 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0090 | 2018/2/9 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0091 | 2018/2/10 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0092 | 2018/2/13 | FPE | Study strain | 2 | 9 | 9 | 1 | present | absent | absent | II |
| ILS AS1-0093 | 2018/3/21 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0094 | 2018/4/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0095 | 2018/4/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0096 | 2018/4/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0097 | 2018/4/5 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0098 | 2018/5/22 | FPE | Study strain | 1 | 204 | 204 | 2 | present | absent | absent | II |
| ILS AS1-0099 | 2018/5/22 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0100 | 2018/3/21 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0101 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0102 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0103 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0104 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | present | absent | absent | II | |
| ILS AS1-0105 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0106 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0107 | 2018/5/25 | FPE | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0108 | 2018/5/25 | FPE | Study strain | 4 | 6 | 6 | n/a | n/a | n/a | I | |
| ILS AS1-0109 | 2018/6/19 | FP | Study strain | 2 | 9 | 9 | n/a | n/a | n/a | II | |
| ILS AS1-0110 | 2018/6/25 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0111 | 2018/6/25 | FPE | Study strain | 1 | 204 | 204 | 2 | present | absent | absent | II |
| ILS AS1-0112 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0113 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0114 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0115 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | 2 | present | absent | absent | II |
| ILS AS1-0116 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0117 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0118 | 2018/6/26 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0119 | 2028/6/25 | FP | Study strain | 2 | 9 | 9 | 1 | present | absent | absent | II |
| ILS AS1-0120 | 2018/7/16 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0121 | 2018/7/16 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0122 | 2018/7/16 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS AS1-0123 | 2018/8/3 | FP | Study strain | 1 | 121 | 121 | n/a | n/a | n/a | II | |
| ILS AS1-0124 | 2018/8/9 | FPE | Study strain | 1 | 204 | 204 | n/a | n/a | n/a | II | |
| ILS-AS-R-001 | PAA reference | 204 | 204 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-002 | PAA reference | 204 | 204 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-003 | PAA reference | 204 | 204 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-004 | PAA reference | 9 | 9 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-005 | PAA reference | 9 | 9 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-006 | PAA reference | 9 | 9 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-007 | PAA reference | 9 | 9 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-008 | PAA reference | 121 | 121 | n/a | n/a | n/a | II | ||||
| ILS-AS-R-009 | PAA reference | 6 | 6 | n/a | n/a | n/a | I | ||||
| N586 | LBF | 3a° | 412 | 412 | n/a | n/a | n/a | II | |||
| N11-1850 | HBF | 4b° | 1290 | 217 | n/a | n/a | n/a | I |
 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stoller, A.; Stevens, M.J.A.; Stephan, R.; Guldimann, C. Characteristics of Listeria Monocytogenes Strains Persisting in a Meat Processing Facility over a 4-Year Period. Pathogens 2019, 8, 32. https://doi.org/10.3390/pathogens8010032
Stoller A, Stevens MJA, Stephan R, Guldimann C. Characteristics of Listeria Monocytogenes Strains Persisting in a Meat Processing Facility over a 4-Year Period. Pathogens. 2019; 8(1):32. https://doi.org/10.3390/pathogens8010032
Chicago/Turabian StyleStoller, Andrea, Marc J. A. Stevens, Roger Stephan, and Claudia Guldimann. 2019. "Characteristics of Listeria Monocytogenes Strains Persisting in a Meat Processing Facility over a 4-Year Period" Pathogens 8, no. 1: 32. https://doi.org/10.3390/pathogens8010032
APA StyleStoller, A., Stevens, M. J. A., Stephan, R., & Guldimann, C. (2019). Characteristics of Listeria Monocytogenes Strains Persisting in a Meat Processing Facility over a 4-Year Period. Pathogens, 8(1), 32. https://doi.org/10.3390/pathogens8010032





