In silico Identification of Novel Toxin Homologs and Associated Mobile Genetic Elements in Clostridium perfringens
Abstract
:1. Introduction
2. Results
2.1. Identification of Toxin Homologs in C. perfringens Isolates
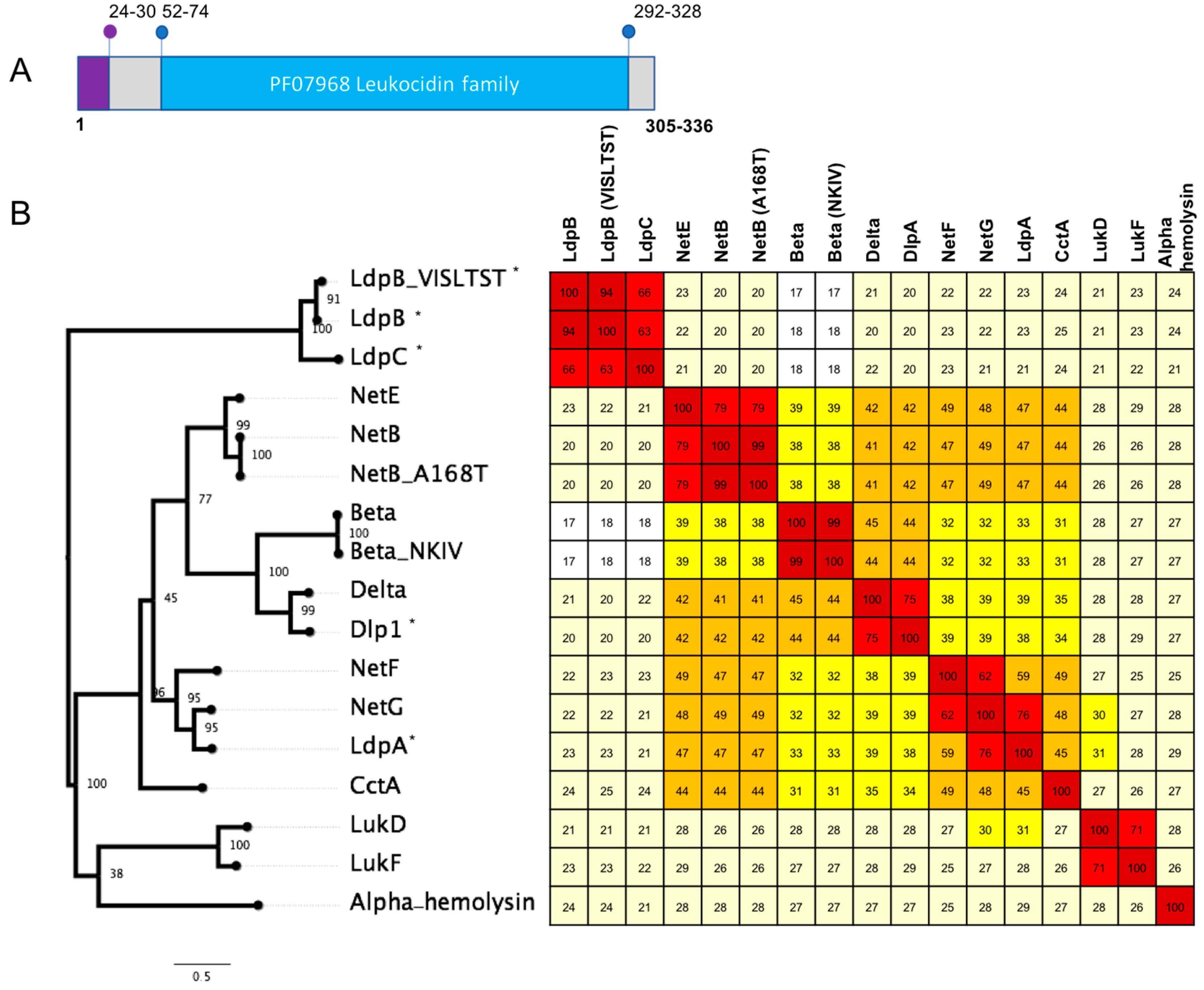
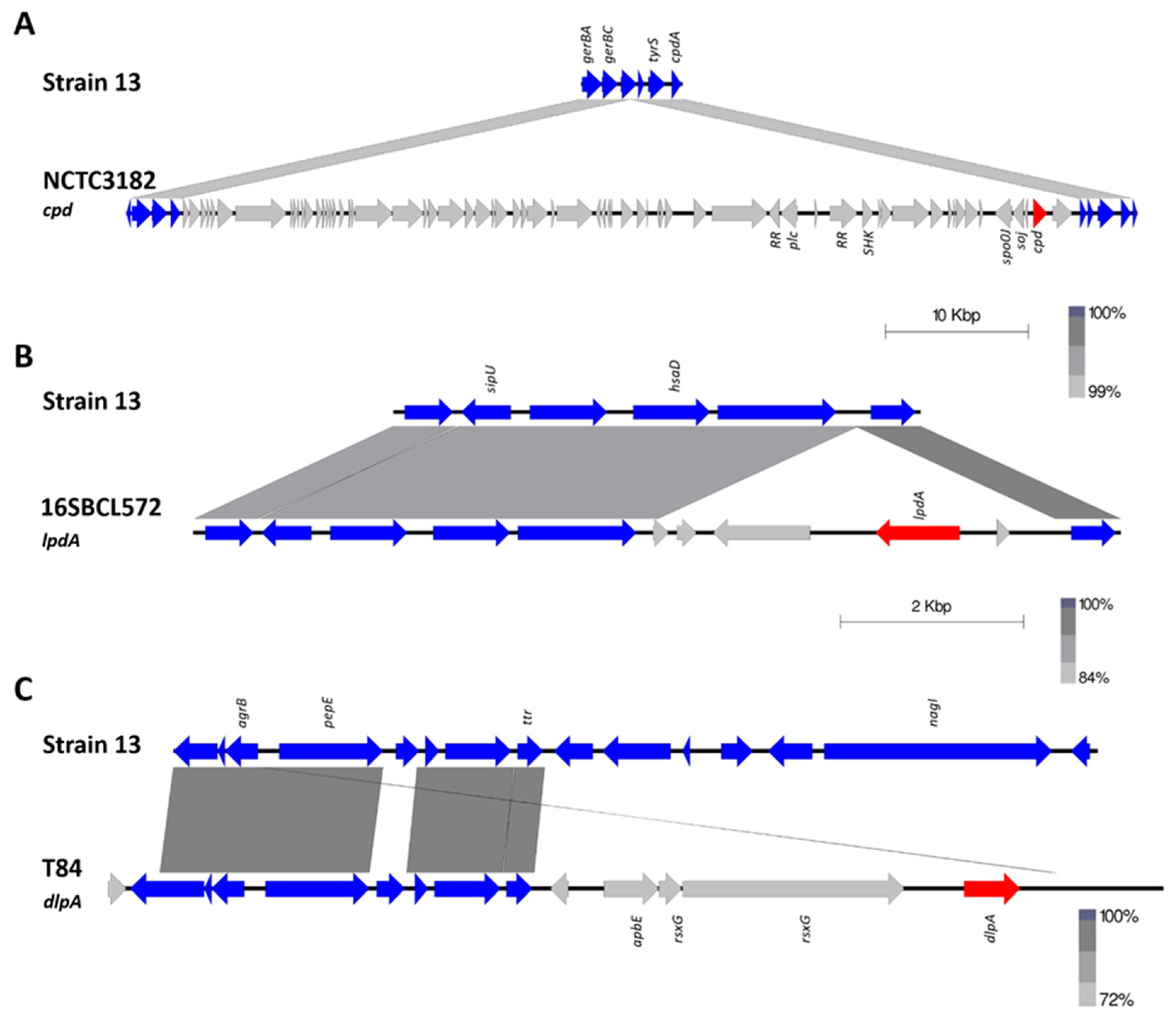
2.2. Novel Protein Sequences with Leukotoxin/Hemolysin Domain
2.3. Novel Protein Sequences with Epsilon Toxin-Like Aerolysin Domain
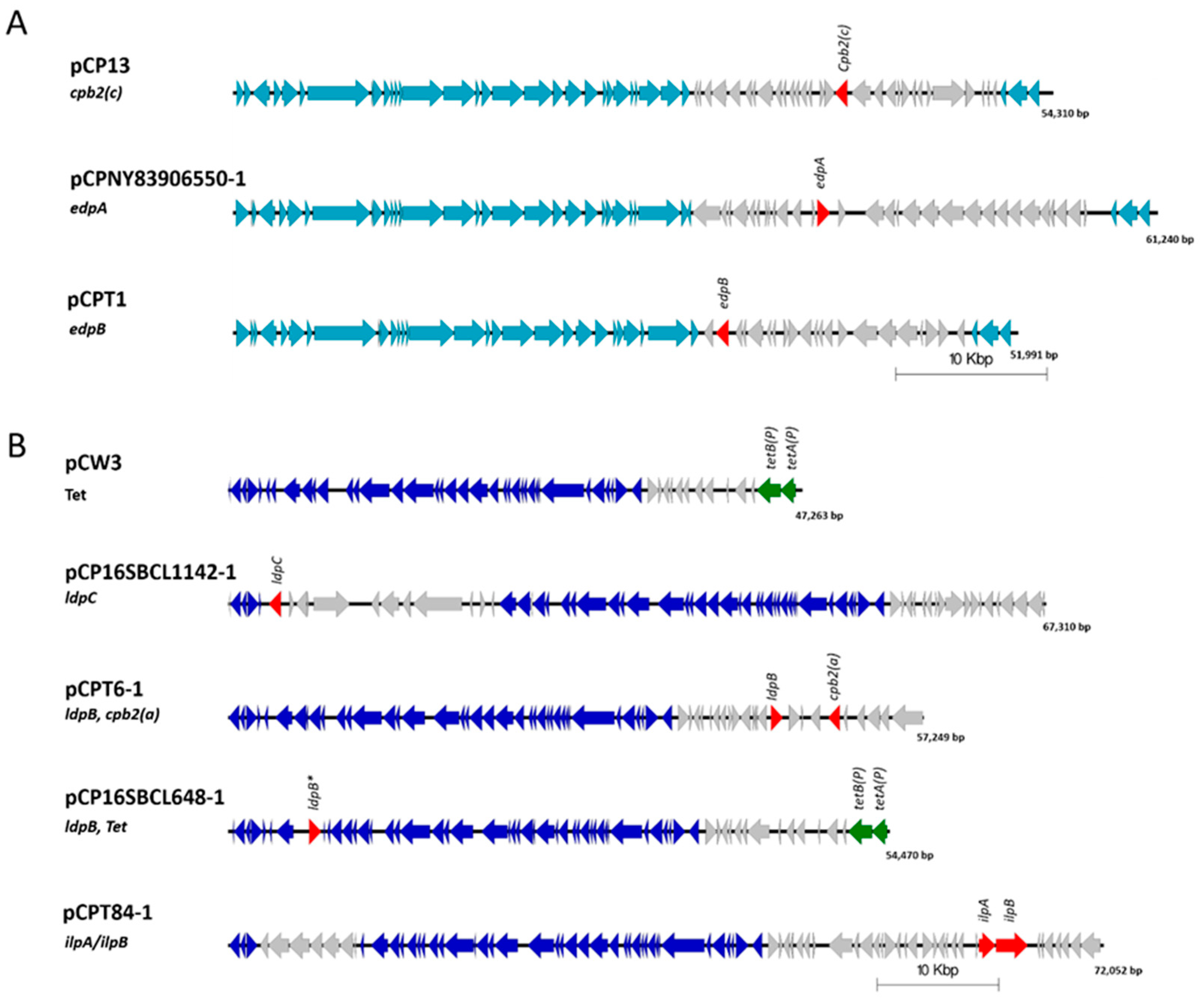
2.4. Novel Protein Sequences with Similarity to Clostridial Binary Toxins
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rood, J.I.; Adams, V.; Lacey, J.; Lyras, D.; McClane, B.A.; Melville, S.B.; Moore, R.J.; Popoff, M.R.; Sarker, M.R.; Songer, J.G.; et al. Expansion of the Clostridium perfringens toxin-based typing scheme. Anaerobe 2018, 53, 5–10. [Google Scholar] [CrossRef]
- Lacey, J.A.; Allnutt, T.R.; Vezina, B.; Van, T.T.H.; Stent, T.; Han, X.; Rood, J.I.; Wade, B.; Keyburn, A.L.; Seemann, T.; et al. Whole genome analysis reveals the diversity and evolutionary relationships between necrotic enteritis-causing strains of Clostridium perfringens. BMC Genom. 2018, 19, 379. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Adams, V.; Bannam, T.L.; Miyamoto, K.; Garcia, J.P.; Uzal, F.A.; Rood, J.I.; McClane, B.A. Toxin plasmids of Clostridium perfringens. Microbiol. Mol. Biol. Rev. MMBR 2013, 77, 208–233. [Google Scholar] [CrossRef] [PubMed]
- Lacey, J.A.; Keyburn, A.L.; Ford, M.E.; Portela, R.W.; Johanesen, P.A.; Lyras, D.; Moore, R.J. Conjugation-mediated horizontal gene transfer of Clostridium perfringens plasmids in the chicken gastrointestinal tract results in the formation of new virulent strains. Appl. Environ. Microbiol. 2017, 83, e01814-17. [Google Scholar] [CrossRef]
- Hughes, M.L.; Poon, R.; Adams, V.; Sayeed, S.; Saputo, J.; Uzal, F.A.; McClane, B.A.; Rood, J.I. Epsilon-toxin plasmids of Clostridium perfringens type D are conjugative. J. Bacteriol. 2007, 189, 7531–7538. [Google Scholar] [CrossRef]
- Brynestad, S.; Sarker, M.R.; McClane, B.A.; Granum, P.E.; Rood, J.I. Enterotoxin plasmid from Clostridium perfringens is conjugative. Infect. Immun. 2001, 69, 3483–3487. [Google Scholar] [CrossRef] [PubMed]
- Gohari, I.M.; Kropinski, A.M.; Weese, S.J.; Parreira, V.R.; Whitehead, A.E.; Boerlin, P.; Prescott, J.F. Plasmid characterization and chromosome analysis of two netF+ Clostridium perfringens isolates associated with foal and canine necrotizing enteritis. PLoS ONE 2016, 11, e0148344. [Google Scholar] [CrossRef]
- Keyburn, A.L.; Boyce, J.D.; Vaz, P.; Bannam, T.L.; Ford, M.E.; Parker, D.; Di Rubbo, A.; Rood, J.I.; Moore, R.J. NetB, a new toxin that is associated with avian necrotic enteritis caused by Clostridium perfringens. PLoS Pathog. 2008, 4, e26. [Google Scholar] [CrossRef]
- Yonogi, S.; Matsuda, S.; Kawai, T.; Yoda, T.; Harada, T.; Kumeda, Y.; Gotoh, K.; Hiyoshi, H.; Nakamura, S.; Kodama, T.; et al. BEC, a novel enterotoxin of Clostridium perfringens found in human clinical isolates from acute gastroenteritis outbreaks. Infect. Immun. 2014, 82, 2390–2399. [Google Scholar] [CrossRef]
- Mehdizadeh Gohari, I.; Parreira, V.R.; Nowell, V.J.; Nicholson, V.M.; Oliphant, K.; Prescott, J.F. A novel pore-forming toxin in type A Clostridium perfringens is associated with both fatal canine hemorrhagic gastroenteritis and fatal foal necrotizing enterocolitis. PLoS ONE 2015, 10, e0122684. [Google Scholar] [CrossRef]
- Nagahama, M.; Ochi, S.; Oda, M.; Miyamoto, K.; Takehara, M.; Kobayashi, K. Recent insights into Clostridium perfringens Beta-toxin. Toxins 2015, 7, 396–406. [Google Scholar] [CrossRef] [PubMed]
- Manich, M.; Knapp, O.; Gibert, M.; Maier, E.; Jolivet-Reynaud, C.; Geny, B.; Benz, R.; Popoff, M.R. Clostridium perfringens Delta toxin is sequence related to Beta toxin, NetB, and Staphylococcus pore-forming toxins, but shows functional differences. PLoS ONE 2008, 3, e3764. [Google Scholar] [CrossRef] [PubMed]
- Keyburn, A.L.; Bannam, T.L.; Moore, R.J.; Rood, J.I. NetB, a pore-forming toxin from necrotic enteritis strains of Clostridium perfringens. Toxins 2010, 2, 1913–1927. [Google Scholar] [CrossRef] [PubMed]
- Titball, R.W.; Naylor, C.E.; Basak, A.K. The Clostridium perfringens alpha-toxin. Anaerobe 1999, 5, 51–64. [Google Scholar] [CrossRef]
- Kitadokoro, K.; Nishimura, K.; Kamitani, S.; Fukui-Miyazaki, A.; Toshima, H.; Abe, H.; Kamata, Y.; Sugita-Konishi, Y.; Yamamoto, S.; Karatani, H.; et al. Crystal structure of Clostridium perfringens enterotoxin displays features of beta-pore-forming toxins. J. Biol. Chem. 2011, 286, 19549–19555. [Google Scholar] [CrossRef] [PubMed]
- Knapp, O.; Maier, E.; Benz, R.; Geny, B.; Popoff, M.R. Identification of the channel-forming domain of Clostridium perfringens Epsilon-toxin (ETX). Biochim. Biophys. Acta 2009, 1788, 2584–2593. [Google Scholar] [CrossRef] [PubMed]
- Titball, R.W.; Leslie, D.L.; Harvey, S.; Kelly, D. Hemolytic and sphingomyelinase activities of Clostridium perfringens alpha-toxin are dependent on a domain homologous to that of an enzyme from the human arachidonic acid pathway. Infect. Immun. 1991, 59, 1872–1874. [Google Scholar]
- Verherstraeten, S.; Goossens, E.; Valgaeren, B.; Pardon, B.; Timbermont, L.; Haesebrouck, F.; Ducatelle, R.; Deprez, P.; Wade, K.R.; Tweten, R.; et al. Perfringolysin O: The underrated Clostridium perfringens toxin? Toxins 2015, 7, 1702–1721. [Google Scholar] [CrossRef] [PubMed]
- Freedman, J.C.; Shrestha, A.; McClane, B.A. Clostridium perfringens enterotoxin: Action, genetics, and translational applications. Toxins 2016, 8, 73. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, J.; Nagahama, M.; Oda, M.; Tsuge, H.; Kobayashi, K. Clostridium perfringens Iota-Toxin: Structure and function. Toxins 2009, 1, 208–228. [Google Scholar] [CrossRef]
- Irikura, D.; Monma, C.; Suzuki, Y.; Nakama, A.; Kai, A.; Fukui-Miyazaki, A.; Horiguchi, Y.; Yoshinari, T.; Sugita-Konishi, Y.; Kamata, Y. Identification and characterization of a new Enterotoxin produced by Clostridium perfringens isolated from food poisoning outbreaks. PLoS ONE 2015, 10, e0138183. [Google Scholar] [CrossRef]
- Gerding, D.N.; Johnson, S.; Rupnik, M.; Aktories, K. Clostridium difficile binary toxin CDT. Gut Microbes 2014, 5, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Perelle, S.; Gibert, M.; Boquet, P.; Popoff, M.R. Characterization of Clostridium perfringens iota-toxin genes and expression in Escherichia coli. Infect. Immun. 1993, 61, 5147–5156. [Google Scholar] [PubMed]
- Neumeyer, T.; Schiffler, B.; Maier, E.; Lang, A.E.; Aktories, K.; Benz, R. Clostridium botulinum C2 toxin Identification of the binding site for cloroquine and related compounds and influence of the binding site on propertise of the C2II channel. J. Biol. Chem. 2008, 283, 3904–3914. [Google Scholar] [CrossRef]
- Kiu, R.; Hall, L.J. An update on the human and animal enteric pathogen Clostridium perfringens. Emerg. Microbes Infect. 2018, 7, 141. [Google Scholar] [CrossRef] [PubMed]
- Lepp, D.; Roxas, B.; Parreira, V.R.; Marri, P.R.; Rosey, E.L.; Gong, J.; Songer, J.G.; Vedantam, G.; Prescott, J.F. Identification of novel pathogenicity loci in Clostridium perfringens strains that cause avian necrotic enteritis. PLoS ONE 2010, 5, e10795. [Google Scholar] [CrossRef]
- Vidor, C.J.; Watts, T.D.; Adams, V.; Bulach, D.; Couchman, E.; Rood, J.I.; Fairweather, N.F.; Awad, M.; Lyras, D. Clostridium sordellii pathogenicity locus plasmid pCS1-1 encodes a novel clostridial conjugation locus. mBio 2018, 9, e01761-17. [Google Scholar] [CrossRef]
- Traore, D.A.K.; Wisniewski, J.A.; Flanigan, S.F.; Conroy, P.J.; Panjikar, S.; Mok, Y.-F.; Lao, C.; Griffin, M.D.W.; Adams, V.; Rood, J.I.; et al. Crystal structure of TcpK in complex with oriT DNA of the antibiotic resistance plasmid pCW3. Nat. Commun. 2018, 9, 3732. [Google Scholar] [CrossRef]
- Bannam, T.L.; Teng, W.L.; Bulach, D.; Lyras, D.; Rood, J.I. Functional identification of conjugation and replication regions of the tetracycline resistance plasmid pCW3 from Clostridium perfringens. J. Bacteriol. 2006, 188, 4942–4951. [Google Scholar] [CrossRef]
- Seike, S.; Miyamoto, K.; Kobayashi, K.; Takehara, M.; Nagahama, M. Clostridium perfringens Delta-toxin induces rapid cell necrosis. PLoS ONE 2016, 11, e0147957. [Google Scholar] [CrossRef]
- Mehdizadeh Gohari, I.; Kropinski, A.M.; Weese, S.J.; Whitehead, A.E.; Parreira, V.R.; Boerlin, P.; Prescott, J.F. NetF-producing Clostridium perfringens: Clonality and plasmid pathogenicity loci analysis. Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2017, 49, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Yonogi, S.; Kanki, M.; Ohnishi, T.; Shiono, M.; Iida, T.; Kumeda, Y. Development and application of a multiplex PCR assay for detection of the Clostridium perfringens enterotoxin-encoding genes cpe and becAB. J. Microbiol. Methods 2016, 127, 172–175. [Google Scholar] [CrossRef] [PubMed]
- Bailey, M.A.; Macklin, K.S.; Krehling, J.T. Use of a Multiplex PCR for the Detection of Toxin-Encoding Genes netB and tpeL in Strains of Clostridium perfringens. ISRN Vet. Sci. 2013, 2013, 865702. [Google Scholar] [CrossRef] [PubMed]
- Badagliacca, P.; Di Provvido, A.; Scattolini, S.; Pompei, G.; Di Giannatale, E. Toxin genotyping of Clostridium perfringens strains using a polymerase chain reaction protocol. Vet. Ital. 2010, 46, 113–118. [Google Scholar] [PubMed]
- Van Asten, A.J.A.M.; van der Wiel, C.W.; Nikolaou, G.; Houwers, D.J.; Gröne, A. A multiplex PCR for toxin typing of Clostridium perfringens isolates. Vet. Microbiol. 2009, 136, 411–412. [Google Scholar] [CrossRef]
- Erol, I.; Goncuoglu, M.; Ayaz, N.D.; Bilir Ormanci, F.S.; Hildebrandt, G. Molecular typing of Clostridium perfringens isolated from turkey meat by multiplex PCR. Lett. Appl. Microbiol. 2008, 47, 31–34. [Google Scholar] [CrossRef]
- Heikinheimo, A.; Korkeala, H. Multiplex PCR assay for toxinotyping Clostridium perfringens isolates obtained from Finnish broiler chickens. Lett. Appl. Microbiol. 2005, 40, 407–411. [Google Scholar] [CrossRef] [PubMed]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinform. Oxf. Engl. 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef]
- Nielsen, H. Predicting secretory proteins with SignalP. Methods Mol. Biol. Clifton NJ 2017, 1611, 59–73. [Google Scholar] [CrossRef]
- Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 2011, 7, e1002195. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. Publ. Protein Soc. 2018, 27, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-C.; Minh, B.Q.; Susko, E.; Roger, A.J. Modeling site heterogeneity with posterior mean site frequency profiles accelerates accurate phylogenomic estimation. Syst. Biol. 2018, 67, 216–235. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinf. 2009, 10, 421. [Google Scholar] [CrossRef]
- Morgulis, A.; Coulouris, G.; Raytselis, Y.; Madden, T.L.; Agarwala, R.; Schäffer, A.A. Database indexing for production MegaBLAST searches. Bioinform. Oxf. Engl. 2008, 24, 1757–1764. [Google Scholar] [CrossRef]
- Zhang, Z.; Schwartz, S.; Wagner, L.; Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. J. Comput. Mol. Cell Biol. 2000, 7, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinform. Oxf. Engl. 2011, 27, 1009–1010. [Google Scholar] [CrossRef] [PubMed]

| Toxin Type | Alpha (plc or cpa) | Beta (cpb) | Epsilon (etx) | Iota (iap and ibp) | Enterotoxin (cpe) | NetB (netB) |
|---|---|---|---|---|---|---|
| A | + | - | - | - | - | - |
| B | + | + | + | - | - | - |
| C | + | + | - | - | +/- | - |
| D | + | - | + | - | +/- | - |
| E | + | - | - | + | +/- | - |
| F * | + | - | - | - | + | - |
| G * | + | - | - | - | - | + |
| Strain | Toxin Type | Host * | Year | Country | Accession | Toxins | Toxin Homologs |
|---|---|---|---|---|---|---|---|
| T43 | A | Turkey, Healthy | 2009 | Finland | SAMN05933484 | plc | dlpA, ilpA/B |
| T46 | A | Turkey, NE | 2010 | Finland | SAMN05933485 | plc | dlpA, ilpA/B |
| T84 | A | Turkey, NE | 2011 | Finland | SAMN05929587 | plc | dlpA, ilpA/B |
| 16SBCL571 | A | Contaminated food | 2015 | France | SAMN09721446 | plc | lpdA |
| 16SBCL572 | A | Contaminated food | 2015 | France | SAMN09721448 | plc | lpdA |
| WER-NE36 | G | Chicken, NE | 2010 | Australia | SAMN07326176 | plc, netB, cpb2 | ldpB |
| EHE-NE7 | G | Chicken, NE | 2002 | Australia | SAMN07326146 | plc, netB, cpb2 | ldpB |
| T6 | A | Turkey, NE | 2005 | Finland | SAMN05929277 | plc, cpb2 | ldpB, lpdC |
| 16SBCL648 | A | - | 2016 | France | SAMN09721463 | plc | ldpB |
| T34 | A | Turkey, NE | 2009 | Finland | SAMN05933483 | plc | ldpC |
| T53 | G | Turkey, Healthy | 2010 | Finland | SAMN05929586 | plc, netb, | ldpC |
| 16SBCL1142 | A | - | 2015 | France | SAMN09721467 | plc | ldpC |
| T22 | A | Turkey, Healthy | 2009 | Finland | SAMN05929282 | plc | ilpA/B |
| NY83906550 | A | Human, Blood | 2012 | USA | SAMN08466960 | plc, cpb2 | edpA |
| NY83905249 | A | Human, Blood | 2010 | USA | SAMN08466959 | plc, cpb2 | edpA |
| 16SBCL600 | A | Contaminated food | 2015 | France | SAMN09721470 | plc | edpA |
| 16SBCL609 | A | Contaminated food | 2015 | France | SAMN09721433 | plc | edpA |
| 16SBCL1126 | F | Contaminated food | 2015 | France | SAMN09721434 | plc, cpe | edpA |
| T1 | A | Turkey, NE | 1998 | Finland | SAMN05928332 | plc, cpb2 | edpB |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lacey, J.A.; Johanesen, P.A.; Lyras, D.; Moore, R.J. In silico Identification of Novel Toxin Homologs and Associated Mobile Genetic Elements in Clostridium perfringens. Pathogens 2019, 8, 16. https://doi.org/10.3390/pathogens8010016
Lacey JA, Johanesen PA, Lyras D, Moore RJ. In silico Identification of Novel Toxin Homologs and Associated Mobile Genetic Elements in Clostridium perfringens. Pathogens. 2019; 8(1):16. https://doi.org/10.3390/pathogens8010016
Chicago/Turabian StyleLacey, Jake A., Priscilla A. Johanesen, Dena Lyras, and Robert J. Moore. 2019. "In silico Identification of Novel Toxin Homologs and Associated Mobile Genetic Elements in Clostridium perfringens" Pathogens 8, no. 1: 16. https://doi.org/10.3390/pathogens8010016
APA StyleLacey, J. A., Johanesen, P. A., Lyras, D., & Moore, R. J. (2019). In silico Identification of Novel Toxin Homologs and Associated Mobile Genetic Elements in Clostridium perfringens. Pathogens, 8(1), 16. https://doi.org/10.3390/pathogens8010016





