Epigallocatechin Gallate Remodelling of Hfq Amyloid-Like Region Affects Escherichia coli Survival
Abstract
1. Introduction
2. Results
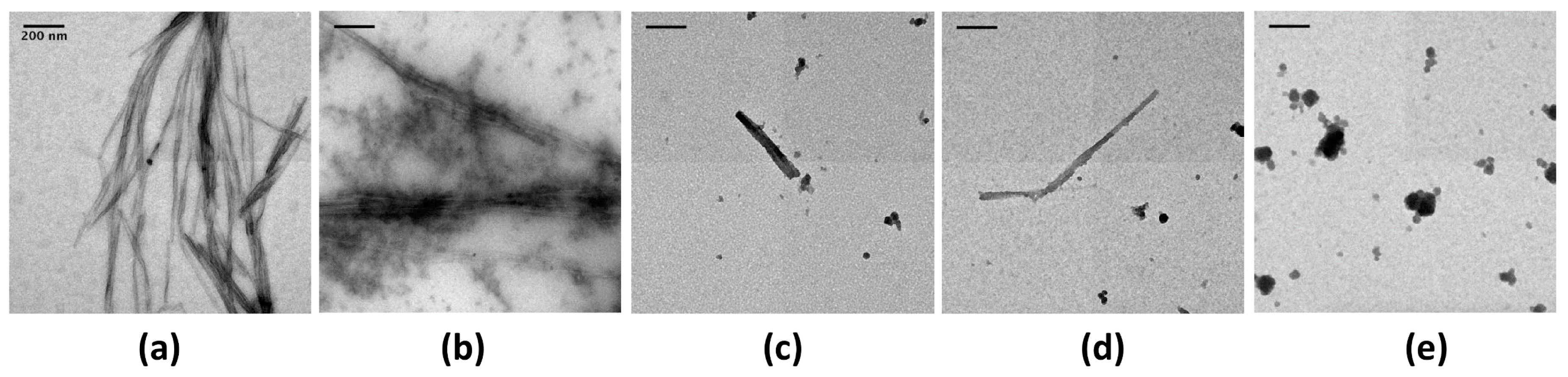
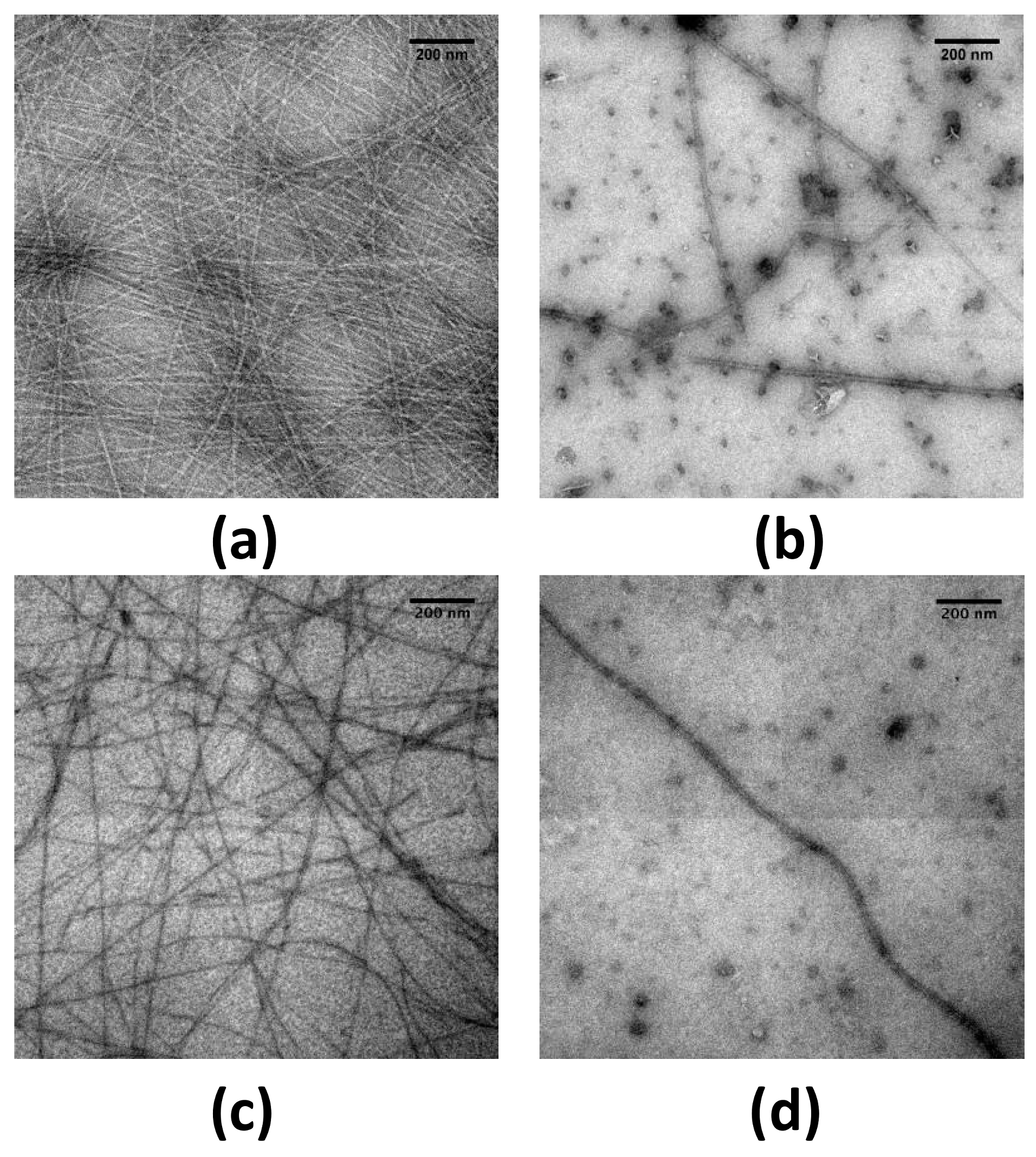
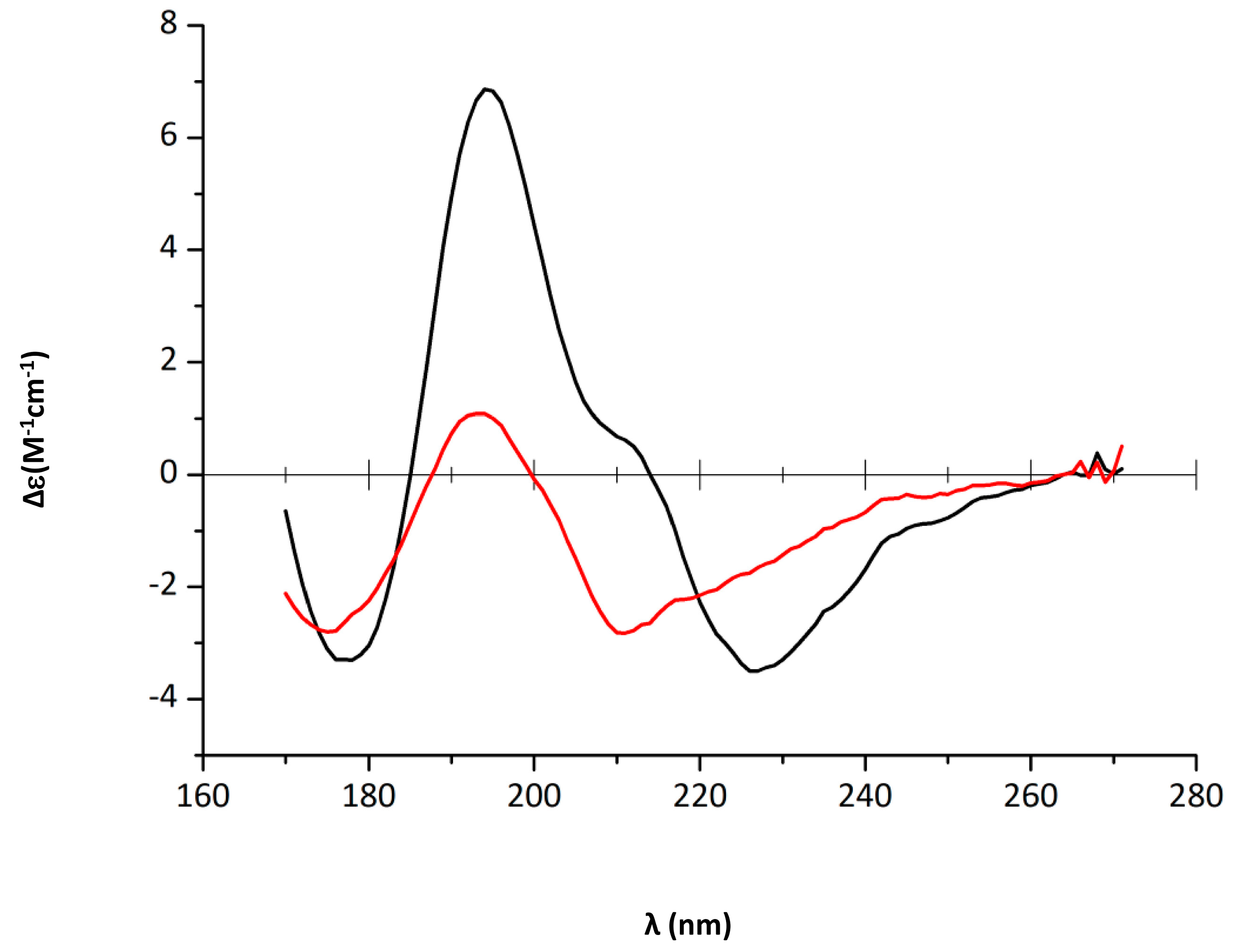
2.1. EGCG Remodels Hfq CTR Amyloid Fibrils
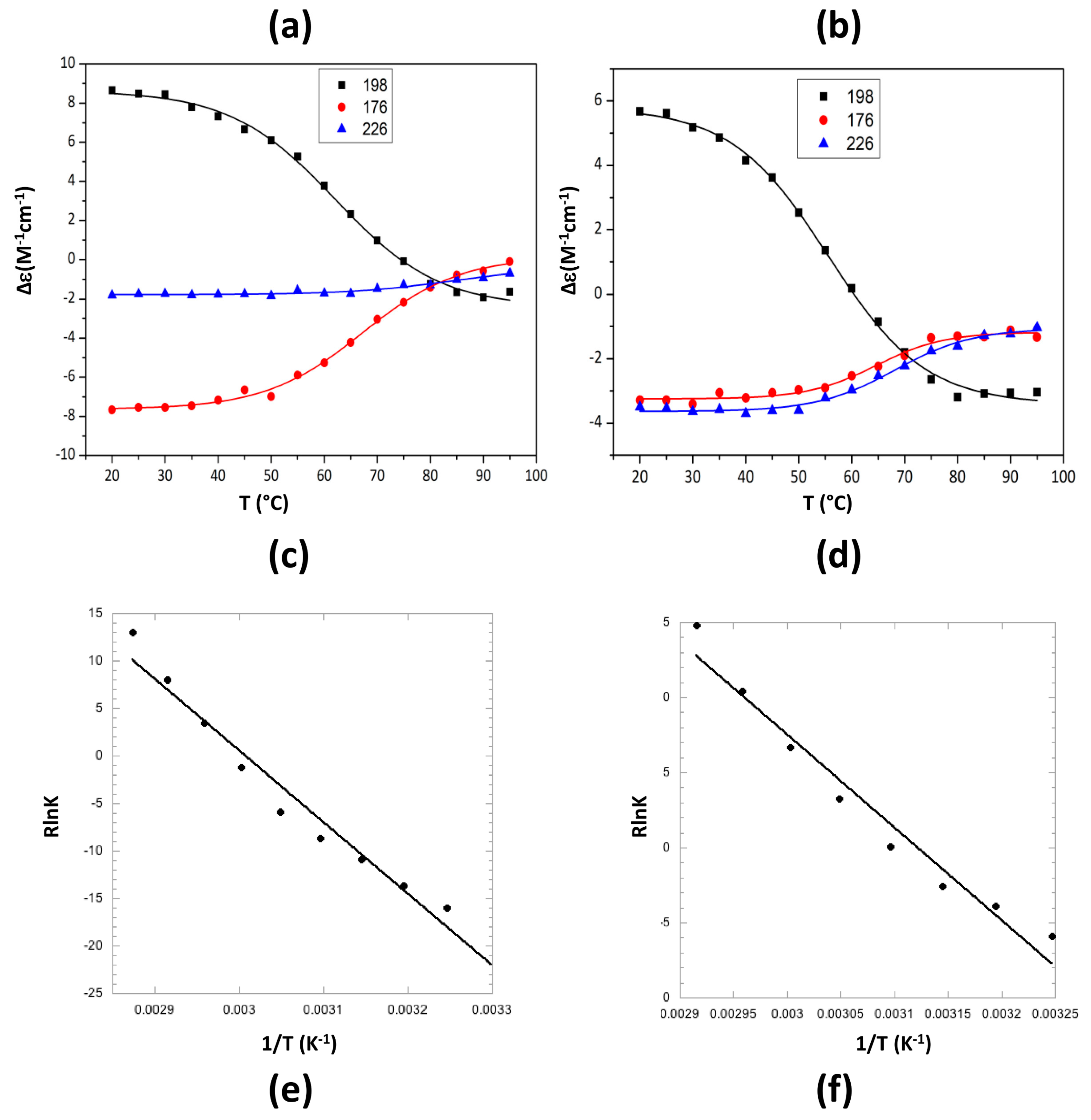
2.2. Characterization of Hfq Amyloid Region Stability in the Presence of EGCG
2.3. EGCG Affects Bacterial Survival Due to Its Interaction with Hfq CTR
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. Hfq CTR Peptides
4.3. TEM Imaging of Protein Fibrils
4.4. Synchrotron Radiation Circular Dichroism (SRCD)
4.5. Construction of E. coli Strains
4.6. Effect of EGCG on E. coli Survival
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Munita, J.M.; Arias, C.A. Mechanisms of Antibiotic Resistance. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Poole, K. Stress responses as determinants of antimicrobial resistance in Gram-negative bacteria. Trends Microbiol. 2012, 20, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Dorman, C.J. Nucleoid-associated proteins and bacterial physiology. Adv. Appl. Microbiol. 2009, 67, 47–64. [Google Scholar] [CrossRef] [PubMed]
- Badaut, C.; Williams, R.; Arluison, V.; Bouffartigues, E.; Robert, B.; Buc, H.; Rimsky, S. The degree of oligomerization of the H-NS nucleoid structuring protein is related to specific binding to DNA. J. Biol. Chem. 2002, 277, 41657–41666. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.A.; Pickard, D.; Higgins, C.F.; Khan, A.; Chatfield, S.N.; Ali, T.; Dorman, C.J.; Hormaeche, C.E.; Dougan, G. Role of hns in the virulence phenotype of pathogenic salmonellae. Mol. Microbiol. 1994, 13, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Schechter, L.M.; Jain, S.; Akbar, S.; Lee, C.A. The small nucleoid-binding proteins H-NS, HU, and Fis affect hilA expression in Salmonella enterica serovar Typhimurium. Infect. Immun. 2003, 71, 5432–5435. [Google Scholar] [CrossRef] [PubMed]
- Porcheron, G.; Dozois, C.M. Interplay between iron homeostasis and virulence: Fur and RyhB as major regulators of bacterial pathogenicity. Vet. Microbiol. 2015, 179, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Kavita, K.; de Mets, F.; Gottesman, S. New aspects of RNA-based regulation by Hfq and its partner sRNAs. Curr. Opin. Microbiol. 2018, 42, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.F.; Han, K.; Chandler, C.E.; Tjaden, B.; Ernst, R.K.; Lory, S. Probing the sRNA regulatory landscape of P. aeruginosa: Post-transcriptional control of determinants of pathogenicity and antibiotic susceptibility. Mol. Microbiol. 2017, 106, 919–937. [Google Scholar] [CrossRef] [PubMed]
- Gottesman, S.; Storz, G. Bacterial small RNA regulators: Versatile roles and rapidly evolving variations. Cold Spring Harb. Perspect. Biol. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Storz, G.; Vogel, J.; Wassarman, K.M. Regulation by small RNAs in bacteria: Expanding frontiers. Mol. Cell. 2011, 43, 880–891. [Google Scholar] [CrossRef] [PubMed]
- Aiba, H. Mechanism of RNA silencing by Hfq-binding small RNAs. Curr. Opin. Microbiol. 2007, 10, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Arluison, V.; Hohng, S.; Roy, R.; Pellegrini, O.; Regnier, P.; Ha, T. Spectroscopic observation of RNA chaperone activities of Hfq in post-transcriptional regulation by a small non-coding RNA. Nucleic Acids Res. 2007, 35, 999–1006. [Google Scholar] [CrossRef] [PubMed]
- Storz, G.; Opdyke, J.A.; Zhang, A. Controlling mRNA stability and translation with small, noncoding RNAs. Curr. Opin. Microbiol. 2004, 7, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Sittka, A.; Pfeiffer, V.; Tedin, K.; Vogel, J. The RNA chaperone Hfq is essential for the virulence of Salmonella typhimurium. Mol. Microbiol. 2007, 63, 193–217. [Google Scholar] [CrossRef] [PubMed]
- Feliciano, J.R.; Grilo, A.M.; Guerreiro, S.I.; Sousa, S.A.; Leitao, J.H. Hfq: A multifaceted RNA chaperone involved in virulence. Future Microbiol. 2016, 11, 137–151. [Google Scholar] [CrossRef] [PubMed]
- Kendall, M.M.; Gruber, C.C.; Rasko, D.A.; Hughes, D.T.; Sperandio, V. Hfq virulence regulation in enterohemorrhagic Escherichia coli O157:H7 strain 86-24. J. Bacteriol. 2011, 193, 6843–6851. [Google Scholar] [CrossRef] [PubMed]
- Dersch, P.; Khan, M.A.; Muhlen, S.; Gorke, B. Roles of Regulatory RNAs for Antibiotic Resistance in Bacteria and Their Potential Value as Novel Drug Targets. Front. Microbiol. 2017, 8, 803. [Google Scholar] [CrossRef] [PubMed]
- El-Mowafi, S.A.; Alumasa, J.N.; Ades, S.E.; Keiler, K.C. Cell-based assay to identify inhibitors of the Hfq-sRNA regulatory pathway. Antimicrob. Agents Chemother. 2014, 58, 5500–5509. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, C.J.; Wilusz, J. Eukaryotic Lsm proteins: Lessons from bacteria. Nat. Struct. Mol. Biol. 2005, 12, 1031–1036. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, C.J.; Wilusz, J. Lsm proteins and Hfq: Life at the 3’ end. RNA Biol. 2013, 10, 592–601. [Google Scholar] [CrossRef] [PubMed]
- Brennan, R.G.; Link, T.M. Hfq structure, function and ligand binding. Curr. Opin. Microbiol. 2007, 10, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Vogel, J.; Luisi, B.F. Hfq and its constellation of RNA. Nat. Rev. Microbiol. 2011, 9, 578–589. [Google Scholar] [CrossRef] [PubMed]
- Link, T.M.; Valentin-Hansen, P.; Brennan, R.G. Structure of Escherichia coli Hfq bound to polyriboadenylate RNA. Proc. Natl. Acad. Sci. USA 2009, 106, 19292–19297. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, M.A.; Pearson, R.F.; Moller, T.; Valentin-Hansen, P.; Brennan, R.G. Structures of the pleiotropic translational regulator Hfq and an Hfq- RNA complex: A bacterial Sm-like protein. EMBO J. 2002, 21, 3546–3556. [Google Scholar] [CrossRef] [PubMed]
- Dimastrogiovanni, D.; Frohlich, K.S.; Bandyra, K.J.; Bruce, H.A.; Hohensee, S.; Vogel, J.; Luisi, B.F. Recognition of the small regulatory RNA RydC by the bacterial Hfq protein. eLife 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Jiang, K.; Zhang, C.; Guttula, D.; Liu, F.; van Kan, J.A.; Lavelle, C.; Kubiak, K.; Malabirade, A.; Lapp, A.; Arluison, V.; et al. Effects of Hfq on the conformation and compaction of DNA. Nucleic Acids Res. 2015, 43, 4332–4341. [Google Scholar] [CrossRef] [PubMed]
- Malabirade, A.; Jiang, K.; Kubiak, K.; Diaz-Mendoza, A.; Liu, F.; van Kan, J.A.; Berret, J.F.; Arluison, V.; van der Maarel, J.R.C. Compaction and condensation of DNA mediated by the C-terminal domain of Hfq. Nucleic Acids Res. 2017, 45, 7299–7308. [Google Scholar] [CrossRef] [PubMed]
- Sauter, C.; Basquin, J.; Suck, D. Sm-like proteins in Eubacteria: The crystal structure of the Hfq protein from Escherichia coli. Nucleic Acids Res. 2003, 31, 4091–4098. [Google Scholar] [CrossRef] [PubMed]
- Nikulin, A.; Stolboushkina, E.; Perederina, A.; Vassilieva, I.; Blaesi, U.; Moll, I.; Kachalova, G.; Yokoyama, S.; Vassylyev, D.; Garber, M.; et al. Structure of Pseudomonas aeruginosa Hfq protein. Acta Crystallogr. D Biol. Crystallogr. 2005, 61, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Sauer, E.; Weichenrieder, O. Structural basis for RNA 3’-end recognition by Hfq. Proc. Natl. Acad. Sci. USA 2011, 108, 13065–13070. [Google Scholar] [CrossRef] [PubMed]
- Stanek, K.A.; Patterson-West, J.; Randolph, P.S.; Mura, C. Crystal structure and RNA-binding properties of an Hfq homolog from the deep-branching Aquificae: Conservation of the lateral RNA-binding mode. Acta Crystallogr. D Struct. Biol. 2017, 73, 294–315. [Google Scholar] [CrossRef] [PubMed]
- Arluison, V.; Mura, C.; Guzman, M.R.; Liquier, J.; Pellegrini, O.; Gingery, M.; Regnier, P.; Marco, S. Three-dimensional Structures of Fibrillar Sm Proteins: Hfq and Other Sm-like Proteins. J. Mol. Biol. 2006, 356, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Fortas, E.; Piccirilli, F.; Malabirade, A.; Militello, V.; Trepout, S.; Marco, S.; Taghbalout, A.; Arluison, V. New insight into the structure and function of Hfq C-terminus. Biosci. Rep. 2015, 35. [Google Scholar] [CrossRef] [PubMed]
- Taghbalout, A.; Yang, Q.; Arluison, V. The Escherichia coli RNA processing and degradation machinery is compartmentalized within an organized cellular network. Biochem. J. 2014, 458, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Diestra, E.; Cayrol, B.; Arluison, V.; Risco, C. Cellular electron microscopy imaging reveals the localization of the Hfq protein close to the bacterial membrane. PLoS ONE 2009, 4, e8301. [Google Scholar] [CrossRef] [PubMed]
- Guillier, M.; Gottesman, S.; Storz, G. Modulating the outer membrane with small RNAs. Genes Dev. 2006, 20, 2338–2348. [Google Scholar] [CrossRef] [PubMed]
- Parker, A.; Gottesman, S. Small RNA Regulation of TolC, the Outer Membrane Component of Bacterial Multidrug Transporters. J. Bacteriol. 2016, 198, 1101–1113. [Google Scholar] [CrossRef] [PubMed]
- Palhano, F.L.; Lee, J.; Grimster, N.P.; Kelly, J.W. Toward the molecular mechanism(s) by which EGCG treatment remodels mature amyloid fibrils. J. Am. Chem. Soc. 2013, 135, 7503–7510. [Google Scholar] [CrossRef] [PubMed]
- Maury, C.P. The emerging concept of functional amyloid. J. Intern. Med. 2009, 265, 329–334. [Google Scholar] [CrossRef] [PubMed]
- Stains, C.I.; Mondal, K.; Ghosh, I. Molecules that target beta-amyloid. ChemMedChem 2007, 2, 1674–1692. [Google Scholar] [CrossRef] [PubMed]
- Malabirade, A.; Morgado-Brajones, J.; Trepout, S.; Wien, F.; Marquez, I.; Seguin, J.; Marco, S.; Velez, M.; Arluison, V. Membrane association of the bacterial riboregulator Hfq and functional perspectives. Sci. Rep. 2017, 7, 10724. [Google Scholar] [CrossRef] [PubMed]
- Malabirade, A.; Partouche, D.; El Hamoui, O.; Turbant, F.; Geinguenaud, F.; Recouvreux, P.; Bizien, T.; Busi, F.; Wien, F.; Arluison, V. Revised role for Hfq bacterial regulator on DNA topology. Sci. Rep. 2018, 8, 16792. [Google Scholar] [CrossRef] [PubMed]
- Gobeaux, F.; Wien, F. Reversible assembly of a drug peptide into amyloid fibrils: A dynamic circular dichroism study. Langmuir 2018. [Google Scholar] [CrossRef] [PubMed]
- Iwata, K.; Matsuura, T.; Sakurai, K.; Nakagawa, A.; Goto, Y. High-resolution crystal structure of beta2-microglobulin formed at pH 7.0. J. Biochem. 2007, 142, 413–419. [Google Scholar] [CrossRef] [PubMed]
- De Meester, P.; Brick, P.; Lloyd, L.F.; Blow, D.M.; Onesti, S. Structure of the Kunitz-type soybean trypsin inhibitor (STI): Implication for the interactions between members of the STI family and tissue-plasminogen activator. Acta Crystallogr. D Biol. Crystallogr. 1998, 54, 589–597. [Google Scholar] [CrossRef] [PubMed]
- Micsonai, A.; Wien, F.; Bulyaki, E.; Kun, J.; Moussong, E.; Lee, Y.H.; Goto, Y.; Refregiers, M.; Kardos, J. BeStSel: A web server for accurate protein secondary structure prediction and fold recognition from the circular dichroism spectra. Nucleic Acids Res. 2018, 46, W315–W322. [Google Scholar] [CrossRef] [PubMed]
- Jensen, K.F. The Escherichia coli K-12 “wild types” W3110 and MG1655 have an rph frameshift mutation that leads to pyrimidine starvation due to low pyrE expression levels. J. Bacteriol. 1993, 175, 3401–3407. [Google Scholar] [CrossRef] [PubMed]
- Snitsarev, V.; Young, M.N.; Miller, R.M.; Rotella, D.P. The spectral properties of (-)-epigallocatechin 3-O-gallate (EGCG) fluorescence in different solvents: Dependence on solvent polarity. PLoS ONE 2013, 8, e79834. [Google Scholar] [CrossRef] [PubMed]
- Davey, H.M. Life, death, and in-between: Meanings and methods in microbiology. Appl. Environ. Microbiol. 2011, 77, 5571–5576. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Bak, G.; Lee, J.; Kim, K.S. Systematic analysis of the role of bacterial Hfq-interacting sRNAs in the response to antibiotics. J. Antimicrob. Chemother. 2015, 70, 1659–1668. [Google Scholar] [CrossRef] [PubMed]
- Reygaert, W.C. The antimicrobial possibilities of green tea. Front. Microbiol. 2014, 5, 434. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Oh, Y.J.; Lim, J.; Youn, M.; Lee, I.; Pak, H.K.; Park, W.; Jo, W.; Park, S. AFM study of the differential inhibitory effects of the green tea polyphenol (-)-epigallocatechin-3-gallate (EGCG) against Gram-positive and Gram-negative bacteria. Food Microbiol. 2012, 29, 80–87. [Google Scholar] [CrossRef] [PubMed]
- Gradisar, H.; Pristovsek, P.; Plaper, A.; Jerala, R. Green tea catechins inhibit bacterial DNA gyrase by interaction with its ATP binding site. J. Med. Chem. 2007, 50, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Martinez, M.D.; Navarro-Peran, E.; Cabezas-Herrera, J.; Ruiz-Gomez, J.; Garcia-Canovas, F.; Rodriguez-Lopez, J.N. Antifolate activity of epigallocatechin gallate against Stenotrophomonas maltophilia. Antimicrob. Agents Chemother. 2005, 49, 2914–2920. [Google Scholar] [CrossRef] [PubMed]
- Serra, D.O.; Mika, F.; Richter, A.M.; Hengge, R. The green tea polyphenol EGCG inhibits E. coli biofilm formation by impairing amyloid curli fibre assembly and downregulating the biofilm regulator CsgD via the sigma(E) -dependent sRNA RybB. Mol. Microbiol. 2016, 101, 136–151. [Google Scholar] [CrossRef] [PubMed]
- Klein, G.; Raina, S. Regulated Control of the Assembly and Diversity of LPS by Noncoding sRNAs. BioMed Res. Int. 2015, 2015, 153561. [Google Scholar] [CrossRef] [PubMed]
- Cech, G.M.; Szalewska-Palasz, A.; Kubiak, K.; Malabirade, A.; Grange, W.; Arluison, V.; Wegrzyn, G. The Escherichia Coli Hfq Protein: An Unattended DNA-Transactions Regulator. Front. Mol. Biosci. 2016, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Liu, F.F.; Dong, X.Y.; Sun, Y. Thermodynamic analysis of the molecular interactions between amyloid beta-peptide 42 and (-)-epigallocatechin-3-gallate. J. Phys. Chem. B 2010, 114, 11576–11583. [Google Scholar] [CrossRef] [PubMed]
- Nakano, S.; Megro, S.I.; Hase, T.; Suzuki, T.; Isemura, M.; Nakamura, Y.; Ito, S. Computational Molecular Docking and X-ray Crystallographic Studies of Catechins in New Drug Design Strategies. Molecules 2018, 23. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Razqan, G.S.; Kwon, D.H. Antibacterial activity of epigallocatechin-3-gallate (EGCG) and its synergism with beta-lactam antibiotics sensitizing carbapenem-associated multidrug resistant clinical isolates of Acinetobacter baumannii. Phytomedicine 2017, 24, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.; Kim, J.H.; Lee, C.K.; Oh, C.H.; Song, H.J. The Antimicrobial Activity of (-)-Epigallocatehin-3-Gallate and Green Tea Extracts against Pseudomonas aeruginosa and Escherichia coli Isolated from Skin Wounds. Ann. Dermatol. 2014, 26, 564–569. [Google Scholar] [CrossRef] [PubMed]
- Refregiers, M.; Wien, F.; Ta, H.P.; Premvardhan, L.; Bac, S.; Jamme, F.; Rouam, V.; Lagarde, B.; Polack, F.; Giorgetta, J.L.; et al. DISCO synchrotron-radiation circular-dichroism endstation at SOLEIL. J. Synchrotron. Radiat. 2012, 19, 831–835. [Google Scholar] [CrossRef] [PubMed]
- Wien, F.; Wallace, B.A. Calcium fluoride micro cells for synchrotron radiation circular dichroism spectroscopy. Appl. Spectrosc. 2005, 59, 1109–1113. [Google Scholar] [CrossRef] [PubMed]
- Micsonai, A.; Wien, F.; Kernya, L.; Lee, Y.H.; Goto, Y.; Refregiers, M.; Kardos, J. Accurate secondary structure prediction and fold recognition for circular dichroism spectroscopy. Proc. Natl. Acad. Sci. USA 2015, 112, E3095–E3103. [Google Scholar] [CrossRef] [PubMed]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Kim, S.H.; Kim, H.; Yeom, J.; Ko, K.; Park, W.; Park, S. AFM probing the mechanism of synergistic effects of the green tea polyphenol (-)-epigallocatechin-3-gallate (EGCG) with cefotaxime against extended-spectrum beta-lactamase (ESBL)-producing Escherichia coli. PLoS ONE 2012, 7, e48880. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.S.; Oh, J.J.; Oh, K.H. Synergistic anti-bacterial and proteomic effects of epigallocatechin gallate on clinical isolates of imipenem-resistant Klebsiella pneumoniae. Phytomedicine 2011, 18, 941–946. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.H.; Asano, N.; Hu, Z.Q.; Shimamura, T. Restoration of antibacterial activity of beta-lactams by epigallocatechin gallate against beta-lactamase-producing species depending on location of beta-lactamase. J. Pharm. Pharmacol. 2003, 55, 735–740. [Google Scholar] [CrossRef] [PubMed]
- Fanaki, N.H.; Kassem, M.A.; Fawzi, M.A.; Dabbous, F.; El-Nakeeb, M.A. Influence of aqueous green tea extract on the antimicrobial activity of some antibiotics against multiresistant clinical isolates. Egypt. J. Med. Microbiol. 2008, 17, 449–460. [Google Scholar]
- Steinmann, J.; Buer, J.; Pietschmann, T.; Steinmann, E. Anti-infective properties of epigallocatechin-3-gallate (EGCG), a component of green tea. Br. J. Pharmacol. 2013, 168, 1059–1073. [Google Scholar] [CrossRef] [PubMed]


| Time (h) | Helix 1 | Helix 2 | Anti 1 | Anti 2 | Anti 3 | Para | Turn | Others | NRMSD |
|---|---|---|---|---|---|---|---|---|---|
| CTR11 with EGCG 5 mM | |||||||||
| 0 | 0.0 | 0.60 | 10.6 | 13.6 | 0.00 | 10.8 | 17.4 | 46.9 | 0.04649 |
| 4 | 5.1 | 5.8 | 0 | 4.8 | 11.4 | 12.1 | 15.4 | 45.3 | 0.06050 |
| Temp. °C | Helix 1 | Helix 2 | Anti 1 | Anti 2 | Anti 3 | Para | Turn | Others | NRMSD |
|---|---|---|---|---|---|---|---|---|---|
| CTR11 without EGCG | |||||||||
| 20 | 0.00 | 0.00 | 15.37 | 35.87 | 0.00 | 0.00 | 5.18 | 43.58 | 0.0403 |
| 25 | 0.00 | 0.00 | 15.31 | 35.95 | 0.00 | 0.00 | 5.38 | 43.36 | 0.0396 |
| 30 | 0.00 | 0.00 | 15.19 | 35.71 | 0.00 | 0.00 | 5.1 | 44.00 | 0.0404 |
| 35 | 0.00 | 0.00 | 14.87 | 33.75 | 0.00 | 0.00 | 5.46 | 45.92 | 0.0416 |
| 40 | 0.00 | 0.00 | 14.47 | 33.17 | 0.00 | 0.00 | 5.90 | 46.46 | 0.0406 |
| 45 | 0.00 | 0.00 | 14.01 | 31.45 | 0.00 | 0.00 | 6.35 | 48.19 | 0.0383 |
| 50 | 0.00 | 0.00 | 13.96 | 29.01 | 0.00 | 0.00 | 5.84 | 51.20 | 0.0434 |
| 55 | 0.00 | 0.00 | 12.15 | 27.17 | 1.15 | 0.00 | 7.57 | 51.96 | 0.0337 |
| 60 | 0.00 | 0.00 | 10.57 | 19.23 | 3.92 | 4.07 | 8.59 | 53.62 | 0.0360 |
| 65 | 0.00 | 0.00 | 8.73 | 13.38 | 7.00 | 9.34 | 10.06 | 51.49 | 0.0348 |
| 70 | 0.00 | 0.89 | 7.23 | 10.79 | 9.95 | 10.91 | 11.82 | 48.41 | 0.0352 |
| 75 | 0.00 | 1.85 | 5.77 | 8.62 | 11.43 | 11.72 | 13.07 | 47.53 | 0.0594 |
| 80 | 0.00 | 3.38 | 3.51 | 5.24 | 12.78 | 11.54 | 15.95 | 47.61 | 0.1453 |
| 85 | 2.33 | 3.29 | 3.75 | 7.39 | 13.70 | 7.38 | 15.53 | 46.62 | 0.0795 |
| 90 | 3.42 | 3.00 | 3.52 | 7.11 | 13.90 | 6.27 | 16.26 | 46.52 | 0.0780 |
| CTR11 with EGCG 5 mM | |||||||||
| 20 | 0.00 | 0.65 | 10.62 | 13.64 | 0.00 | 10.79 | 17.35 | 46.95 | 0.0467 |
| 25 | 0.00 | 0.85 | 10.27 | 13.10 | 0.00 | 11.97 | 17.12 | 46.69 | 0.0465 |
| 30 | 0.00 | 0.80 | 10.03 | 12.41 | 0.00 | 13.67 | 16.53 | 46.57 | 0.0496 |
| 35 | 0.14 | 0.77 | 10.08 | 12.61 | 0.06 | 11.63 | 17.07 | 47.63 | 0.0459 |
| 40 | 0.00 | 0.84 | 9.53 | 9.72 | 0.68 | 15.34 | 16.69 | 47.20 | 0.0595 |
| 45 | 0.95 | 0.70 | 9.52 | 9.69 | 2.01 | 12.69 | 16.66 | 47.79 | 0.0504 |
| 50 | 1.97 | 0.71 | 8.72 | 8.88 | 3.36 | 12.59 | 15.94 | 47.82 | 0.0554 |
| 55 | 3.22 | 0.56 | 7.85 | 8.86 | 5.68 | 10.53 | 15.60 | 47.69 | 0.0600 |
| 60 | 2.55 | 1.19 | 6.93 | 7.65 | 7.59 | 11.92 | 15.59 | 46.58 | 0.0750 |
| 65 | 2.41 | 1.13 | 6.19 | 7.57 | 9.72 | 10.31 | 15.75 | 46.91 | 0.0827 |
| 70 | 3.76 | 1.60 | 5.18 | 7.43 | 10.90 | 9.85 | 15.22 | 46.08 | 0.0921 |
| 75 | 2.74 | 1.91 | 4.30 | 7.84 | 12.47 | 6.95 | 16.54 | 47.26 | 0.0777 |
| 80 | 1.10 | 1.75 | 4.03 | 8.24 | 13.42 | 6.41 | 16.94 | 48.12 | 0.0629 |
| 85 | 0.00 | 1.69 | 3.91 | 9.64 | 14.08 | 4.95 | 16.79 | 48.94 | 0.0620 |
| 90 | 0.00 | 1.82 | 3.94 | 9.80 | 14.67 | 4.74 | 16.80 | 48.23 | 0.0655 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Partouche, D.; Turbant, F.; El Hamoui, O.; Campidelli, C.; Bombled, M.; Trépout, S.; Wien, F.; Arluison, V. Epigallocatechin Gallate Remodelling of Hfq Amyloid-Like Region Affects Escherichia coli Survival. Pathogens 2018, 7, 95. https://doi.org/10.3390/pathogens7040095
Partouche D, Turbant F, El Hamoui O, Campidelli C, Bombled M, Trépout S, Wien F, Arluison V. Epigallocatechin Gallate Remodelling of Hfq Amyloid-Like Region Affects Escherichia coli Survival. Pathogens. 2018; 7(4):95. https://doi.org/10.3390/pathogens7040095
Chicago/Turabian StylePartouche, David, Florian Turbant, Omar El Hamoui, Camille Campidelli, Marianne Bombled, Sylvain Trépout, Frank Wien, and Véronique Arluison. 2018. "Epigallocatechin Gallate Remodelling of Hfq Amyloid-Like Region Affects Escherichia coli Survival" Pathogens 7, no. 4: 95. https://doi.org/10.3390/pathogens7040095
APA StylePartouche, D., Turbant, F., El Hamoui, O., Campidelli, C., Bombled, M., Trépout, S., Wien, F., & Arluison, V. (2018). Epigallocatechin Gallate Remodelling of Hfq Amyloid-Like Region Affects Escherichia coli Survival. Pathogens, 7(4), 95. https://doi.org/10.3390/pathogens7040095





