Hsp90 Interacts with the Bacterial Effector NleH1
Abstract
:1. Introduction
2. Results
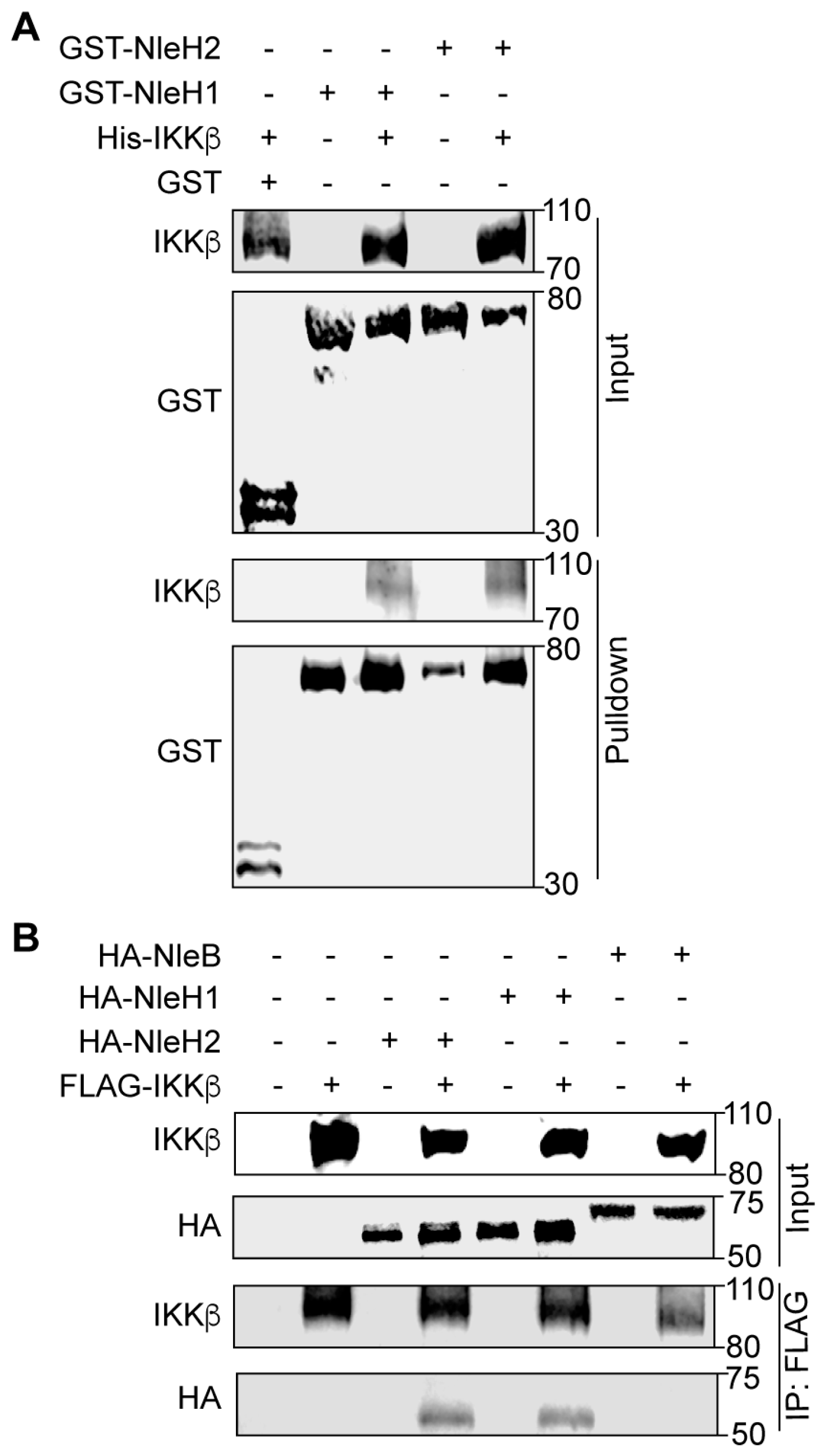
2.1. NleH1 and NleH2 Bind to IKKβ
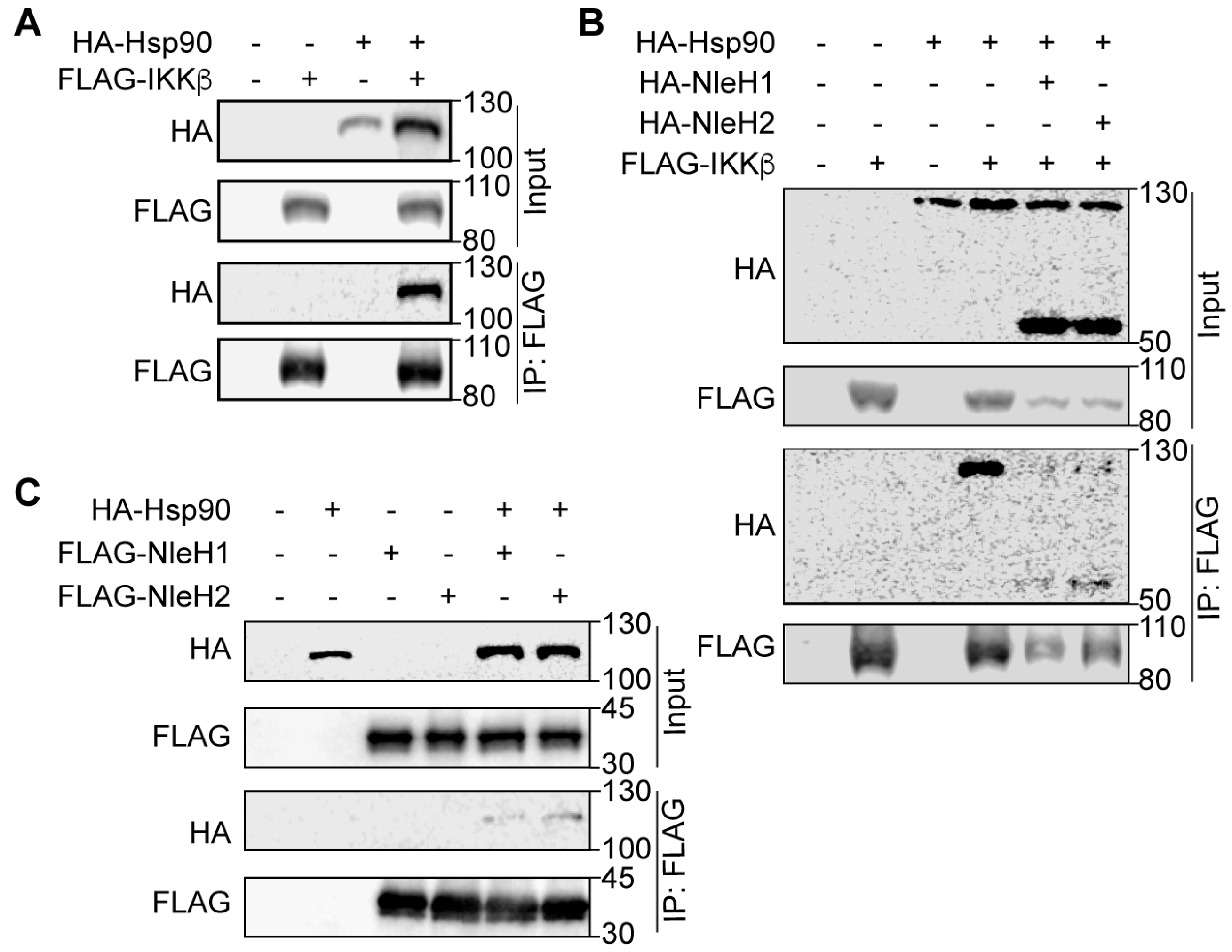
2.2. Hsp90 Interacts with NleH1, NleH2 and IKKβ
2.3. Hsp90 Is a Cofactor of Both NleH1 and IKKβ
3. Discussion
4. Materials and Methods
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Le Negrate, G. Subversion of innate immune responses by bacterial hindrance of nf-kappab pathway. Cell. Microbiol. 2012, 14, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.Y.; Anderson, D.E.; Barnitz, R.A.; Snow, A.; Bidere, N.; Zheng, L.X.; Hegde, V.; Lam, L.T.; Staudt, L.M.; Levens, D.; et al. Ribosomal protein s3: A kh domain subunit in nf-kappa b complexes that mediates selective gene regulation. Cell 2007, 131, 927–939. [Google Scholar] [CrossRef] [PubMed]
- Alkalay, I.; Yaron, A.; Hatzubai, A.; Orian, A.; Ciechanover, A.; Ben-Neriah, Y. Stimulation-dependent i kappa b alpha phosphorylation marks the nf-kappa b inhibitor for degradation via the ubiquitin-proteasome pathway. Proc. Natl. Acad. Sci. USA 1995, 92, 10599–10603. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.; Weaver, A.; Gao, X.; Bern, M.; Hardwidge, P.R.; Lenardo, M.J. Ikkbeta phosphorylation regulates rps3 nuclear translocation and nf-kappab function during infection with Escherichia coli strain o157:H7. Nat. Immunol. 2011, 12, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Wan, F.; Mateo, K.; Callegari, E.; Wang, D.; Deng, W.; Puente, J.; Li, F.; Chaussee, M.S.; Finlay, B.B.; et al. Bacterial effector binding to ribosomal protein s3 subverts nf-kappab function. PLoS Pathog. 2009, 5, e1000708. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.H.; Gao, X.; Singh, G.; Hardwidge, P.R. Escherichia coli virulence protein nleh1 interaction with the v-crk sarcoma virus ct10 oncogene-like protein (crkl) governs nleh1 inhibition of the ribosomal protein s3 (rps3)/nuclear factor kappab (nf-kappab) pathway. J. Biol. Chem. 2013, 288, 34567–34574. [Google Scholar] [CrossRef] [PubMed]
- Csermely, P.; Schnaider, T.; Soti, C.; Prohaszka, Z.; Nardai, G. The 90-kda molecular chaperone family: Structure, function, and clinical applications. A comprehensive review. Pharmacol. Ther. 1998, 79, 129–168. [Google Scholar] [CrossRef]
- Calderwood, S.K. Cdc37 as a co-chaperone to hsp90. Subcell. Biochem. 2015, 78, 103–112. [Google Scholar] [PubMed]
- Taipale, M.; Krykbaeva, I.; Koeva, M.; Kayatekin, C.; Westover, K.D.; Karras, G.I.; Lindquist, S. Quantitative analysis of hsp90-client interactions reveals principles of substrate recognition. Cell 2012, 150, 987–1001. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Cao, P.; Goeddel, D.V. Tnf-induced recruitment and activation of the ikk complex require cdc37 and hsp90. Mol. Cell 2002, 9, 401–410. [Google Scholar] [CrossRef]
- Hinz, M.; Broemer, M.; Arslan, S.C.; Otto, A.; Mueller, E.C.; Dettmer, R.; Scheidereit, C. Signal responsiveness of ikappab kinases is determined by cdc37-assisted transient interaction with hsp90. J. Biol. Chem. 2007, 282, 32311–32319. [Google Scholar] [CrossRef] [PubMed]
- Mazaira, G.I.; Zgajnar, N.R.; Lotufo, C.M.; Daneri-Becerra, C.; Sivils, J.C.; Soto, O.B.; Cox, M.B.; Galigniana, M.D. The nuclear receptor field: A historical overview and future challenges. Nucl. Recept. Res. 2018, 5, 101320. [Google Scholar] [CrossRef] [PubMed]
- Pearl, L.H. Review: The hsp90 molecular chaperone-an enigmatic atpase. Biopolymers 2016, 105, 594–607. [Google Scholar] [CrossRef] [PubMed]
- Goetz, M.P.; Toft, D.O.; Ames, M.M.; Erlichman, C. The hsp90 chaperone complex as a novel target for cancer therapy. Ann. Oncol. 2003, 14, 1169–1176. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.; Devin, A.; Miller, A.; Lin, Y.; Rodriguez, Y.; Neckers, L.; Liu, Z.G. Disruption of hsp90 function results in degradation of the death domain kinase, receptor-interacting protein (rip), and blockage of tumor necrosis factor-induced nuclear factor-kappab activation. J. Biol. Chem. 2000, 275, 10519–10526. [Google Scholar] [CrossRef] [PubMed]
- Broemer, M.; Krappmann, D.; Scheidereit, C. Requirement of hsp90 activity for ikappab kinase (ikk) biosynthesis and for constitutive and inducible ikk and nf-kappab activation. Oncogene 2004, 23, 5378–5386. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.H.; Gao, X.F.; Tsai, K.; Olsen, R.; Wan, F.Y.; Hardwidge, P.R. Functional differences and interactions between the Escherichia coli type III secretion system effectors nleh1 and nleh2. Infect. Immun. 2012, 80, 2133–2140. [Google Scholar] [CrossRef] [PubMed]
- Royan, S.V.; Jones, R.M.; Koutsouris, A.; Roxas, J.L.; Falzari, K.; Weflen, A.W.; Kim, A.; Bellmeyer, A.; Turner, J.R.; Neish, A.S.; et al. Enteropathogenic E. coli non-lee encoded effectors nleh1 and nleh2 attenuate nf-kappab activation. Mol. Microbiol. 2010, 78, 1232–1245. [Google Scholar] [CrossRef] [PubMed]
- Greten, F.R.; Eckmann, L.; Greten, T.F.; Park, J.M.; Li, Z.W.; Egan, L.J.; Kagnoff, M.F.; Karin, M. Ikk beta links inflammation and tumorigenesis in a mouse model of colitis-associated cancer. Cell 2004, 118, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Evaristo, C.; Spranger, S.; Barnes, S.E.; Miller, M.L.; Molinero, L.L.; Locke, F.L.; Gajewski, T.F.; Alegre, M.L. Cutting edge: Engineering active ikk beta in t cells drives tumor rejection. J. Immunol. 2016, 196, 2933–2938. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.S.; Jang, C.Y.; Kim, H.D.; Lee, J.Y.; Ahn, B.Y.; Kim, J. Interaction of hsp90 with ribosomal proteins protects from ubiquitination and proteasome-dependent degradation. Mol. Biol. Cell 2006, 17, 824–833. [Google Scholar] [CrossRef] [PubMed]
- El Qaidi, S.; Chen, K.; Halim, A.; Siukstaite, L.; Rueter, C.; Hurtado-Guerrero, R.; Clausen, H.; Hardwidge, P.R. Nleb/ssek effectors from Citrobacter rodentium, Escherichia coli, and Salmonella enterica display distinct differences in host substrate specificity. J. Biol. Chem. 2017, 292, 11423–11430. [Google Scholar] [CrossRef] [PubMed]

| Strains | ||
| E. coli BL21(DE3) | E. coli F− ompT hsdSB (rB-mB-) gal dcm (DE3) | |
| BL21(DE3)/NleH1-pET42a | GST-EHEC NleH1 | [5] |
| BL21(DE3)/NleH2-pET42a | GST-EHEC NleH2 | [5] |
| BL21(DE3)/IKKβ pET28a | His-IKKβ | This study |
| EHEC EDL933 | Wild-type E. coli O157:H7 isolate | CDC |
| EDL933 ΔnleH1 | EHEC nleH1 deletion | [5] |
| Plasmids | ||
| HA | HA fusion expression | Clontech |
| NleH1-HA | HA fused to E. coli EDL933 NleH1 | [5] |
| NleH2-HA | HA fused to E. coli EDL933 NleH2 | [5] |
| Hsp90-HA | HA fused to Hsp90 | This study |
| 3× FLAG | FLAG expression | Sigma |
| 3× FLAG-IKKβ | FLAG-IKKβ | [4] |
| 3× FLAG-NleH1 | FLAG-EHEC NleH1 | This study |
| 3× FLAG-NleH2 | FLAG-EHEC NleH2 | This study |
| pET42a | Bacterial GST fusion expression | Novagen |
| NleH1-pET42a | GST-EHEC NleH1 | [5] |
| NleH2-pET42a | GST-EHEC NleH2 | [5] |
| pET28a | Bacterial His6 fusion expression | Novagen |
| IKKβ pET28a | His-IKKβ | This study |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, M.; Hardwidge, P.R. Hsp90 Interacts with the Bacterial Effector NleH1. Pathogens 2018, 7, 87. https://doi.org/10.3390/pathogens7040087
Wu M, Hardwidge PR. Hsp90 Interacts with the Bacterial Effector NleH1. Pathogens. 2018; 7(4):87. https://doi.org/10.3390/pathogens7040087
Chicago/Turabian StyleWu, Miaomiao, and Philip R. Hardwidge. 2018. "Hsp90 Interacts with the Bacterial Effector NleH1" Pathogens 7, no. 4: 87. https://doi.org/10.3390/pathogens7040087
APA StyleWu, M., & Hardwidge, P. R. (2018). Hsp90 Interacts with the Bacterial Effector NleH1. Pathogens, 7(4), 87. https://doi.org/10.3390/pathogens7040087





