Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal
Abstract
1. Introduction
2. Results
2.1. Detection of Fecal Indicator Bacteria, Human-Fecal Markers, and Index Viruses
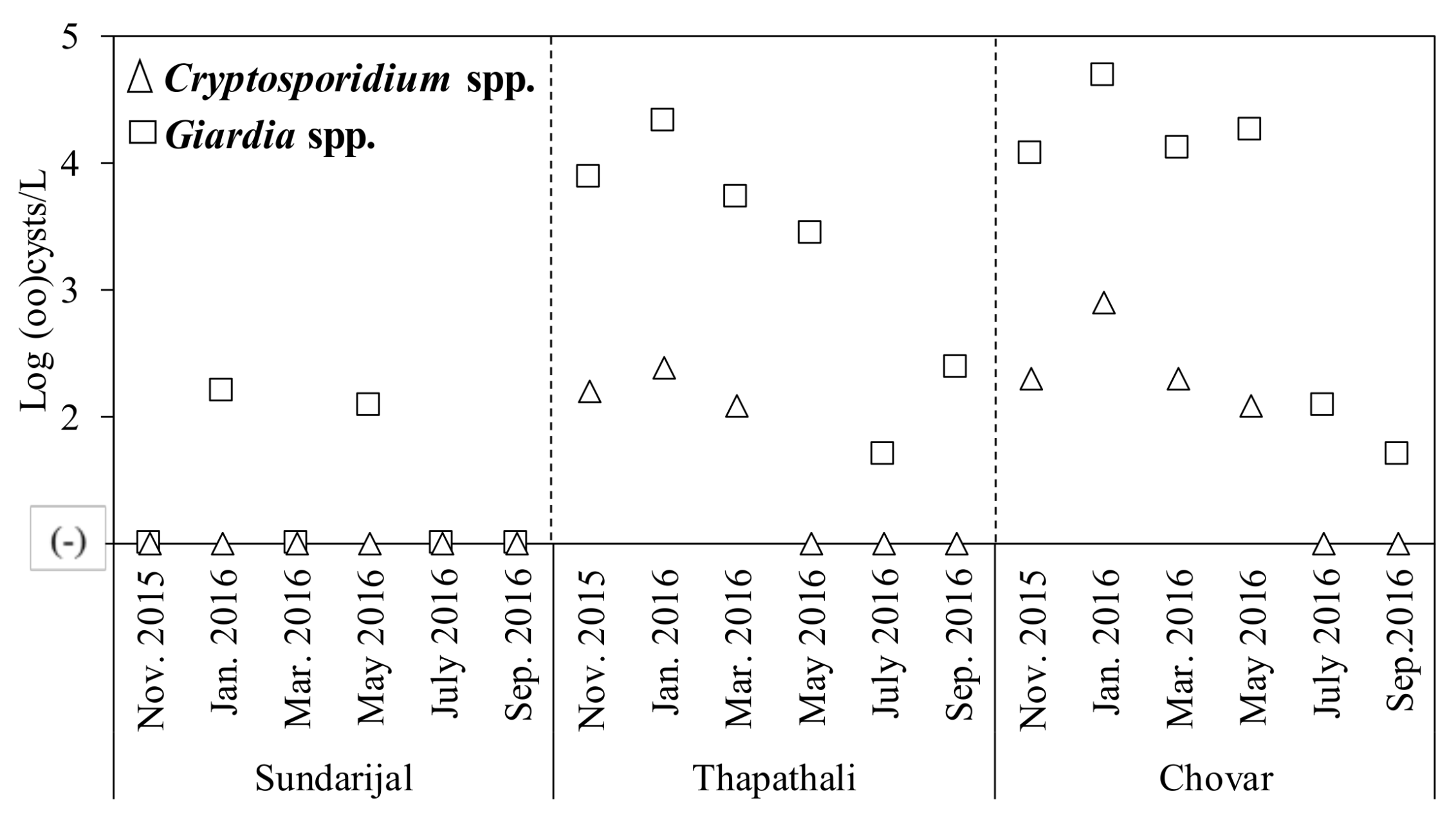
2.2. Detection of Protozoa
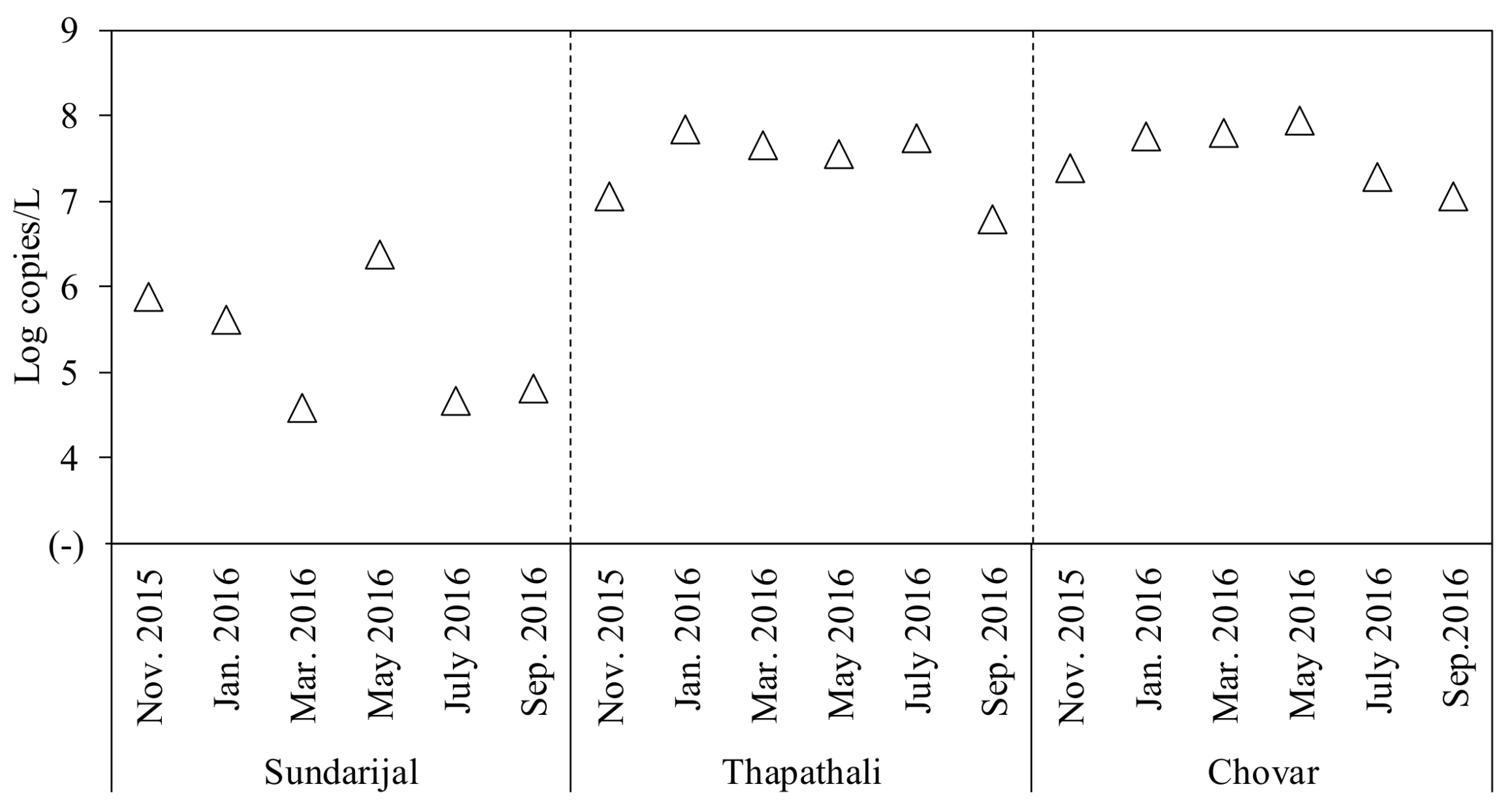
2.3. Detection of Human Enteric Viruses
2.4. Relationships Between Pathogens and Indicators
3. Discussion
4. Materials and Methods
4.1. Collection of River Water Samples
4.2. Detection of Total Coliforms and E. coli
4.3. Detection of Bacteria Using qPCR
4.4. Virus and Protozoan Concentration Methods
4.5. Detection of Cryptosporidium and Giardia (Oo)Cysts Using Fluorescence Microscopy
4.6. Detection of Viral DNA Using qPCR
4.7. Detection of Viral RNA Using RT-qPCR
4.8. Statistical Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gautam, R.; Shrestha, J.K.; Shrestha, G.K.C. Assessment of river water intrusion at the periphery of Bagmati River in Kathmandu Valley. Nepal J. Sci. Technol. 2013, 14, 137–146. [Google Scholar] [CrossRef]
- Paudel, A. Environmental management of the Bagmati river basin. In UNEP EIA Training Resource Manual; UNEP: Nairobi, Kenya, 2002; pp. 269–279. Available online: https://www.iaia.org/pdf/case-studies/BasmatiRiverBasin.pdf (accessed on 2 April 2018).
- Kannel, P.R.; Lee, S.; Lee, Y.S.; Kanel, S.R.; Pelletier, G.J. Application of automated QUAL2Kw for water quality modeling and management in the Bagmati River. Nepal Ecol. Modell. 2007, 202, 503–517. [Google Scholar] [CrossRef]
- Udmale, P.; Ishidaira, H.; Thapa, B.R.; Shakya, N.M. The status of domestic water demand: Supply deficit in the Kathmandu Valley, Nepal. Water 2016, 8, 196. [Google Scholar] [CrossRef]
- Mehta, K.R.; Kushwaha, U.K.S. An Assessment of Aquatic Biodiversity of River Bagmati Nepal. Ecol. Evol. Biol. 2016, 1, 35–40. [Google Scholar] [CrossRef]
- WHO. Guidelines for Safe Recreational Water. Coastal and Fresh Waters; World Health Organization: Geneva, Switzerland, 2003; Volume 1, p. 253. [Google Scholar]
- Shrestha, S.; Haramoto, E.; Sherchand, J.B.; Hada, S.; Rajbhandari, S.; Shindo, J. Prevalence of protozoa and indicator bacteria in wastewater irrigation sources in Kathmandu Valley, Nepal: Cases from Kirtipur, Bhaktapur, and Madhyapur Thimi municipalities. J. Water Environ. Technol. 2016, 14, 149–157. [Google Scholar] [CrossRef]
- Thakur, J.K.; Neupane, M.; Mohanan, A.A. Water poverty in upper Bagmati River basin in Nepal. Water Sci. 2017, 31, 93–108. [Google Scholar] [CrossRef]
- Shrestha, N.; Lamsal, A.; Regmi, R.K.; Mishra, B.K. Current Status of Water Environment in Kathmandu Valley, Nepal; United Nations University: Tokyo, Japan, 2015; pp. 1–5. [Google Scholar]
- Kitajima, M.; Rachmadi, A.T.; Iker, B.C.; Haramoto, E.; Gerba, C.P. Temporal variations in genotype distribution of human sapoviruses and Aichi virus 1 in wastewater in southern Arizona, United States. J. Appl. Microbiol. 2018, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Regmi, R.K.; Mishra, B.K. Water and Urban Initiative: Current Water Quality Status of Rivers in the Kathmandu Valley; United Nations University: Tokyo, Japan, 2016; pp. 1–5. [Google Scholar]
- Haramoto, E.; Yamada, K.; Nishida, K. Prevalence of protozoa, viruses, coliphages and indicator bacteria in groundwater and river water in the Kathmandu Valley, Nepal. Trans. R. Soc. Trop. Med. Hyg. 2011, 105, 711–716. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M. Quantification and genotyping of Aichi virus 1 in water samples in the Kathmandu Valley, Nepal. Food Environ. Virol. 2017, 9, 350–353. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, S.; Shrestha, S.; Shindo, J.; Sherchand, J.B.; Haramoto, E. Virological quality of irrigation water sources and pepper mild mottle virus and tobacco mosaic virus as index of pathogenic virus contamination level. Food Environ. Virol. 2018, 10, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, S.; Haramoto, E.; Sherchand, J.B.; Rajbhandari, S.; Prajapati, M.; Shindo, J. Seasonal variation of microbial quality of irrigation water in different sources in the Kathmandu Valley, Nepal. Naresuan Univ. Eng. J. 2016, 11, 57–62. [Google Scholar] [CrossRef]
- Ghaju Shrestha, R.; Tanaka, Y.; Malla, B.; Bhandari, D.; Tandukar, S.; Inoue, D.; Sei, K.; Sherchand, J.B.; Haramoto, E. Next-generation sequencing identification of pathogenic bacterial genes and their relationship with fecal indicator bacteria in different water sources in the Kathmandu Valley, Nepal. Sci. Total. Environ. 2017, 601–602, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Castillo, F.; Loera-Muro, A.; Jacques, M.; Garneau, P.; Avelar-Gonzalez, F.J.; Harel, J.; Guerrero-Barrera, A.L. Waterborne pathogens: Detection methods and challenges. Pathogens 2015, 4, 307–334. [Google Scholar] [CrossRef] [PubMed]
- Deshmukh, R.A.; Joshi, K.; Bhand, S.; Roy, U. Recent developments in detection and enumeration of waterborne bacteria: A retrospective minireview. MicrobiologyOpen 2016, 5, 901–922. [Google Scholar] [CrossRef] [PubMed]
- Girones, A.; Ferrus, M.A.; Alonso, J.L.; Rodriguez-manzano, J.; Calgua, B.; Correa Ade, A.; Hundesa, A.; Carratala, A.; Bofill-Mas, S. Molecular detection of pathogens in water—The pros and cons of molecular techniques. Water Res. 2010, 44, 4325–4339. [Google Scholar] [CrossRef] [PubMed]
- Harwood, V.J.; Butler, J.; Parrish, D.; Wagner, V. Isolation of fecal coliform bacteria from the diamondback terrapin (Malaclemys terrapin centrata). Appl. Environ. Microbiol. 1999, 65, 865–867. [Google Scholar] [PubMed]
- Harwood, V.J.; Staley, C.; Badgley, B.D.; Borges, K.; Korajkic, A. Microbial source tracking markers for detection of fecal contamination in environmental waters: Relationships between pathogens and human health outcomes. FEMS Microbiol. Rev. 2014, 38, 1–40. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Goonetilleke, A.; Powell, D.; Chauhan, K.; Gardner, T. Comparison of molecular markers to detect fresh sewage in environmental waters. Water Res. 2009, 43, 4908–4917. [Google Scholar] [CrossRef] [PubMed]
- McQuaig, S.; Griffith, J.; Harwood, V.J. Association of fecal indicator bacteria with human viruses and microbial source tracking markers at coastal beaches impacted by nonpoint source pollution. Appl. Environ. Microbiol. 2012, 78, 6423–6432. [Google Scholar] [CrossRef] [PubMed]
- Fong, T.T.; Lipp, E.K. Enteric viruses of humans and animals in aquatic environments: Health risks, detection, and potential water quality assessment tools. Microbiol. Mol. Biol. Rev. 2005, 69, 357–371. [Google Scholar] [CrossRef] [PubMed]
- Albinana-Gimenez, N.; Clemente-Casares, P.; Bofill-Mas, S.; Hundesa, A.; Ribas, F.; Girones, R. Distribution of human polyomaviruses, adenoviruses, and hepatitis E virus in the environment and in a drinking-water treatment plant. Environ. Sci. Technol. 2006, 40, 7416–7422. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Quantitative Microbial Risk Assessment: Application for Water Safety Management; World Health Organization: Geneva, Switzerland, 2016; p. 187. [Google Scholar] [CrossRef]
- Sherchand, J.B.; Nakagomi, O.; Dove, W.; Nakagomi, T.; Yokoo, M.; Pandey, B.D.; Cuevas, L.E.; Hart, C.A.; Cunliffe, N.A. Molecular epidemiology of rotavirus diarrhea among children aged <5 Years in Nepal: Predominance of emergent G12 strains during 2 years. J. Infect. Dis. 2009, 200, S182–S187. [Google Scholar] [CrossRef] [PubMed]
- Sherchand, J.B.; Schluter, W.W.; Sherchand, J.B.; Tandukar, S.; Dhakwa, J.R.; Choudhary, G.R.; Mahaseth, C.M. Prevalence of group A genotype human rotavirus among children with diarrhoea in Nepal, 2009–2011. WHO Southeast Asian J Public Health 2012, 1, 432–440. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Ao, Y.; Liu, N.; Li, L.; Duan, Z. Salivirus in children and its association with childhood acute gastroenteritis: A paired case-control study. PLoS ONE 2015, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Cardemil, C.V.; Sherchand, J.B.; Shrestha, L.; Sharma, A.; Gary, H.E.; Estivariz, C.F.; Diez-Valcarce, M.; Ward, M.L.; Bowen, M.D.; Vinjé, J.; et al. Pathogen-specific burden of outpatient diarrhea in infants in Nepal: A multisite prospective case-control study. J. Pediatr. Infect. Dis. Soc. 2017, 6, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, A.C.; Gyhrs, M.L.; Nielsen, L.P.; Pedersen, C.; Böttiger, B. Gastroenteritis and the novel picornaviruses, Aichi virus, cosavirus, saffold virus, and salivirus in young children. J. Clin. Virol. 2013, 57, 239–242. [Google Scholar] [CrossRef] [PubMed]
- Hoa-Tran, T.N.; Nakagomi, T.; Sano, D.; Sherchand, J.B.; Pandey, B.D.; Cunliffe, N.A.; Nakagomi, O. Molecular epidemiology of noroviruses detected in Nepalese children with acute diarrhea between 2005 and 2011: Increase and predominance of minor genotype GII.13. Infect. Genet. Evol. 2015, 30, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Bosch, A.; Abad, F.X.; Pintó, R.M. Human pathogenic viruses in the marine environment. In Oceans and Health: Pathogens in the Marine Environment; Springer: Boston, MA, USA, 2005; pp. 109–131. [Google Scholar]
- Tandukar, S.; Sherchand, J.B.; Xue, J.; Uprety, S.; Sherchan, S.P.; Bhandari, D.; Malla, B.; Ghaju Shrestha, R.; Parajuli, L.; Poudel, S.; et al. Prevalence and associated risk factors of Giardia duodenalis infection among school-going children in Nepal. J. Parasitol. Res. 2018, 117, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Hunter, P.R.; Nichols, G. Epidemiology and clinical features of Cryptosporidium infection in immunocompromised patients. Clin. Microbiol. Rev. 2002, 15, 145–154. [Google Scholar] [CrossRef] [PubMed]
- Ono, K.; Rai, S.K.; Chikahira, M.; Fujimoto, T.; Shibata, H.; Wada, Y.; Tsuji, H.; Oda, Y.; Rai, G.; Shrestha, C.D.; et al. Seasonal distribution of enteropathogens detected from diarrheal stool and water samples collected in Kathmandu, Nepal. Southeast Asian J. Trop. Med. Public Health 2001, 32, 520–526. [Google Scholar] [PubMed]
- Sharma, S. Restoration project along the banks of Bagmati River corridor of Kathmandu Valley for enforcing sustainable land use planning and development researcher. In Final Report: Bagmati Conservation/Restoration Project; Alcoa Foundation Conservation and Sustainability Fellowship Programme: Kathmandu, Nepal, 2009; pp. 1–82. [Google Scholar]
- U.S. Environmental Protection Agency. Recreational Water Quality Criteria; U.S. Environmental Protection Agency: Washington, DC, USA, 2012; pp. 1–69.
- Frahm, E.; Obst, U. Application of the fluorogenic probe technique (TaqMan PCR) to the detection of Enterococcus spp. and Escherichia coli in water samples. J. Microbiol. Methods 2003, 52, 123–131. [Google Scholar] [CrossRef]
- Ludwig, W.; Schleifer, K.H. How quantitative is quantitative PCR with respect to cell counts? Syst. Appl. Microbiol. 2000, 23, 556–562. [Google Scholar] [CrossRef]
- Bernhard, A.E.; Field, K.G. Identification of nonpoint sources of fecal pollution in coastal waters by using host-specific 16S ribosomal DNA genetic markers from fecal anaerobes. Appl. Environ. Microbiol. 2000, 66, 1587–1594. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Katayama, H.; Asami, M.; Akiba, M. Development of a novel method for simultaneous concentration of viruses and protozoa from a single water sample. J. Virol. Methods. 2012, 182, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, M.; Haramoto, E.; Iker, B.C.; Gerba, C.P. Occurrence of Cryptosporidium, Giardia, and Cyclospora in influent and effluent water at wastewater treatment plants in Arizona. Sci. Total Environ. 2014, 484, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 2018, 135, 168–186. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, M.; Haramoto, E.; Phanuwan, C.; Katayama, H. Prevalence and genetic diversity of Aichi viruses in wastewater and river water in Japan. Appl. Environ. Microbiol. 2011, 77, 2184–2187. [Google Scholar] [CrossRef] [PubMed]
- Shieh, Y.S.C.; Wait, D.; Tai, L.; Sobsey, M.D. Methods to remove inhibitors in sewage and other fecal wastes for enterovirus detection by the polymerase chain reaction. J. Virol. Methods. 1995, 54, 51–66. [Google Scholar] [CrossRef]
- Katayama, H.; Shimasaki, A.; Ohgaki, S. Development of a virus concentration method and its application to detection of enterovirus and Norwalk virus from coastal seawater. Appl. Environ. Microbiol. 2002, 68, 1033–1039. [Google Scholar] [CrossRef] [PubMed]
- Stöcker, A.; de Carvalho Dominguez Souza, B.F.; Ribeiro, T.C.M.; Netto, E.M.; Araujo, L.O.; Corrêa, J.I.; Almeida, P.S.; de Mattos, A.P.; da Costa Ribeiro, H., Jr.; Pedral-Sampaio, D.B.; et al. Cosavirus infection in persons with and without gastroenteritis, Brazil. Emerg. Infect. Dis. 2012, 18, 656–659. [Google Scholar] [CrossRef]
- Kageyama, T.; Kojima, S.; Shinohara, M.; Shinohara, M.; Uchida, K.; Fukushi, S.; Hoshino, F.B.; Takeda, N.; Katayama, K. Broadly reactive and highly sensitive assay for Norwalk-like viruses based on real-time quantitative reverse transcription-PCR. J. Clin. Microbiol. 2003, 41, 1548–1557. [Google Scholar] [CrossRef] [PubMed]
- Jothikumar, N.; Kang, G.; Hill, V.R. Broadly reactive TaqMan assay for real-time RT-PCR detection of rotavirus in clinical and environmental samples. J. Virol. Methods. 2009, 155, 126–131. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Otagiri, M. Development of a Reverse Transcription-Quantitative PCR Assay for Detection of Salivirus/Klassevirus. Appl. Environ. Microbiol. 2013, 79, 3529–3532. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Kishida, N.; Konno, Y.; Katayama, H.; Asami, M.; Akiba, M. Occurrence of pepper mild mottle virus in drinking water sources in Japan. Appl. Environ. Microbiol. 2013, 79, 7413–7418. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Breitbart, M.; Lee, W.H.; Run, J.-Q.; Wei, C.L.; Soh, S.W.L.; Hibberd, M.L.; Liu, E.T.; Rohwer, F.; Ruan, Y. RNA viral community in human feces: Prevalence of plant pathogenic viruses. PLoS Biol. 2005, 4, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Balique, F.; Colson, P.; Barry, A.O.; Nappez, C.; Ferretti, A.; Al Moussawi, K.; Ngounga, T.; Lepidi, H.; Ghigo, E.; Mege, J.-L.; et al. Tobacco mosaic virus in the lungs of mice following intra-tracheal inoculation. PLoS ONE 2013, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, M.; Oka, T.; Takagi, H.; Tohya, Y.; Katayama, H.; Takeda, N.; Katayama, K. Development and application of a broadly reactive real-time reverse transcription-PCR assay for detection of murine noroviruses. J. Virol. Methods 2010, 169, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Thapa, B.R.; Ishidaira, H.; Pandey, V.P.; Shakya, N.M. A multi-model approach for analyzing water balance dynamics in Kathmandu Valley, Nepal. J. Hydrol. Reg. Stud. 2017, 9, 149–162. [Google Scholar] [CrossRef]
| Types | Microorganisms Tested | No. of Positive Samples (%) | Concentrations Among Positive Samples | ||||
|---|---|---|---|---|---|---|---|
| Sundarijal (n = 6) | Thapathali (n = 6) | Chovar (n = 6) | Total (n = 18) | Range (min–max) | Unit | ||
| Indicator bacteria | Total coliforms | 6 (100) | 6 (100) | 6 (100) | 18 (100) | 4.8–10.3 | Log MPN/100-mL |
| E. coli | 6 (100) | 6 (100) | 6 (100) | 18 (100) | 3.5–10.0 6.2–8.6 | Log MPN/100-mL Log copies/L | |
| Enterococcus spp. | 5 (83) | 6 (100) | 6 (100) | 17 (94) | 5.6–10.1 | Log copies/L | |
| Potential index viruses | TMV | 5 (83) | 6 (100) | 5 (83) | 16 (89) | 5.1–9.0 | Log copies/L |
| PMMoV | 3 (50) | 6 (100) | 5 (83) | 14 (78) | 4.4–8.0 | Log copies/L | |
| Human-fecal markers | BKPyVs | 0 (0) | 5 (83) | 5 (83) | 10 (56) | 6.0–7.1 | Log copies/L |
| JCPyVs | 4 (67) | 5 (83) | 0 (0) | 9 (50) | 6.0–7.0 | Log copies/L | |
| Human Bacteroidales | 4 (67) | 6 (100) | 6 (100) | 16 (89) | 6.5–9.8 | Log copies/L | |
| Types | Microorganisms Tested | No. of Positive Samples (%) | Concentrations Among Positive Samples | ||||
|---|---|---|---|---|---|---|---|
| Sundarijal (n = 6) | Thapathali (n = 6) | Chovar (n = 6) | Total (n = 18) | Range (min–max) | Unit | ||
| Protozoa | Cryptosporidium spp. | 0 (0) | 3 (50) | 4 (67) | 7 (39) | 2.1–2.9 | Log oocysts/L |
| Giardia spp. | 2 (33) | 6 (100) | 6 (100) | 14 (78) | 1.6–4.7 | Log cysts/L | |
| Human enteric viruses | AiV-1 | 1 (17) | 6 (100) | 5 (83) | 12 (67) | 4.6–6.6 | Log copies/L |
| EVs | 2 (33) | 6 (100) | 5 (83) | 13 (72) | 5.4–7.5 | Log copies/L | |
| HCoSVs | 0 (0) | 5 (83) | 4 (67) | 9 (50) | 5.7–6.7 | Log copies/L | |
| HuAdVs | 0 (0) | 6 (100) | 6 (100) | 12 (67) | 6.2–7.7 | Log copies/L | |
| NoVs-GI | 0 (0) | 4 (67) | 1 (17) | 5 (28) | 4.4–5.0 | Log copies/L | |
| NoVs-GII | 2 (33) | 6 (100) | 3(33) | 11 (61) | 4.9–5.7 | Log copies/L | |
| RVAs | 0 (0) | 1 (17) | 2 (33) | 3 (17) | 4.5–5.0 | Log copies/L | |
| SaliVs | 2 (33) | 6 (100) | 5 (83) | 13 (72) | 4.4–8.3 | Log copies/L | |
| Types | Indicators | R Value | ||
|---|---|---|---|---|
| Total Human Enteric Viruses | Cryptosporidium spp. | Giardia spp. | ||
| Fecal indicator bacteria | Total coliforms | 0.56 * | 0.20 | 0.14 |
| E. coli | 0.69 * | 0.24 | 0.17 | |
| Enterococcus spp. | 0.73 * | 0.30 | 0.30 | |
| Human-fecal markers | BKPyVs | 0.81 * | 0.06 | 0.13 |
| JCPyVs | 0.74 * | 0.09 | 0.01 | |
| Human Bacteroidales | 0.71 * | 0.11 | 0.09 | |
| Index viruses | TMV | 0.49 * | 0.01 | 0.13 |
| PMMoV | 0.76 * | 0.12 | 0.21 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tandukar, S.; Sherchand, J.B.; Bhandari, D.; Sherchan, S.P.; Malla, B.; Ghaju Shrestha, R.; Haramoto, E. Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal. Pathogens 2018, 7, 38. https://doi.org/10.3390/pathogens7020038
Tandukar S, Sherchand JB, Bhandari D, Sherchan SP, Malla B, Ghaju Shrestha R, Haramoto E. Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal. Pathogens. 2018; 7(2):38. https://doi.org/10.3390/pathogens7020038
Chicago/Turabian StyleTandukar, Sarmila, Jeevan B. Sherchand, Dinesh Bhandari, Samendra P. Sherchan, Bikash Malla, Rajani Ghaju Shrestha, and Eiji Haramoto. 2018. "Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal" Pathogens 7, no. 2: 38. https://doi.org/10.3390/pathogens7020038
APA StyleTandukar, S., Sherchand, J. B., Bhandari, D., Sherchan, S. P., Malla, B., Ghaju Shrestha, R., & Haramoto, E. (2018). Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal. Pathogens, 7(2), 38. https://doi.org/10.3390/pathogens7020038







