Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts
Abstract
:1. Introduction
2. Results and Discussion
2.1. Prevalence of Staphylococcus aureus and MRSA in Beef Livers, Beef and Pork
| Prevalence of Staphylococcus aureus | |||
|---|---|---|---|
| Beef | Pork np/n (%) | ||
| Beef Liver *np/n (%) | Beef (Other Cuts) np/n (%) | Total np/n (%) | |
| 40/50 (80%) | 23/46 (50%) | 63/96 (65.6%) | 43/99 (43.3%) |
2.2. Antimicrobial Resistance Screening
| Antimicrobial Resistance | ||||
|---|---|---|---|---|
| Antibiotic | Beef | Pork np/n (%) | ||
| Beef Livers * np/n (%) | Beef (Other Cuts) np/n (%) | Total np/n (%) | ||
| azithromycin ** | 5/143 (3.5) | 31/76 (40.8) | 36/219 (16.4) | 75/115 (65.2) |
| ciprofloxacin ** | 1/143 (0.7) | 15/76 (19.7) | 16/219 (7.3) | 47/115 (40.9) |
| gentamicin ** | 8/143 (5.6) | 22/76 (28.9) | 30/219 (13.7) | 68/115 (59.1) |
| oxacillin ** | 10/143 (6.9) | 6/76 (7.9) | 16/219 (7.3) | 56/115 (48.7) |
| cefoxitin ** | 11/143 (7.7) | 8/76 (10.5) | 19/219 (8.7) | 47/115 (40.9) |
| tetracycline ** | 21/143 (14.7) | 33/76 (43.4) | 54/219 (24.7) | 109/115 (94.8) |
| vancomycin ** | 12/143 (8.4) | 6/76 (7.9) | 18/219 (8.2) | 49/115 (42.6) |
| doxycycline ** | 10/143 (6.9) | 26/76 (34.2) | 36/219 (16.4) | 96/115 (83.5) |
| trimethoprirm/ sulfamethazole ** | 1/143 (0.7) | 2/76 (2.6) | 3/219 (1.4) | 22/115 (19.1) |
| clindamycin ** | 1/143 (0.7) | 9/76 (11.8) | 10/219 (4.5) | 25/115 (21.7) |
| penicillin ** | 31/143 (21.7) | 48/76 (63.2) | 79/219 (36.1) | 102/115 (88.7) |
| ampicillin ns | 142/143 (99.3) | 76/76 (100) | 218/219 (99.5) | 115/115 (100) |
| kanamycin ** | 4/143 (2.8) | 23/76 (30.3) | 27/219 (12.3) | 62/115 (53.9) |
| erythromycin ** | 8/143 (5.6) | 31/76 (40.8) | 39/219 (17.8) | 63/115 (54.8) |
| rifampin * | 21/143 (14.7) | 6/76 (7.9) | 27/219 (12.3) | 25/115 (21.7) |
| chloramphenicol ** | 3/143 (2.1) | 6/76 (7.9) | 9/219 (4.1) | 35/115 (30.4) |
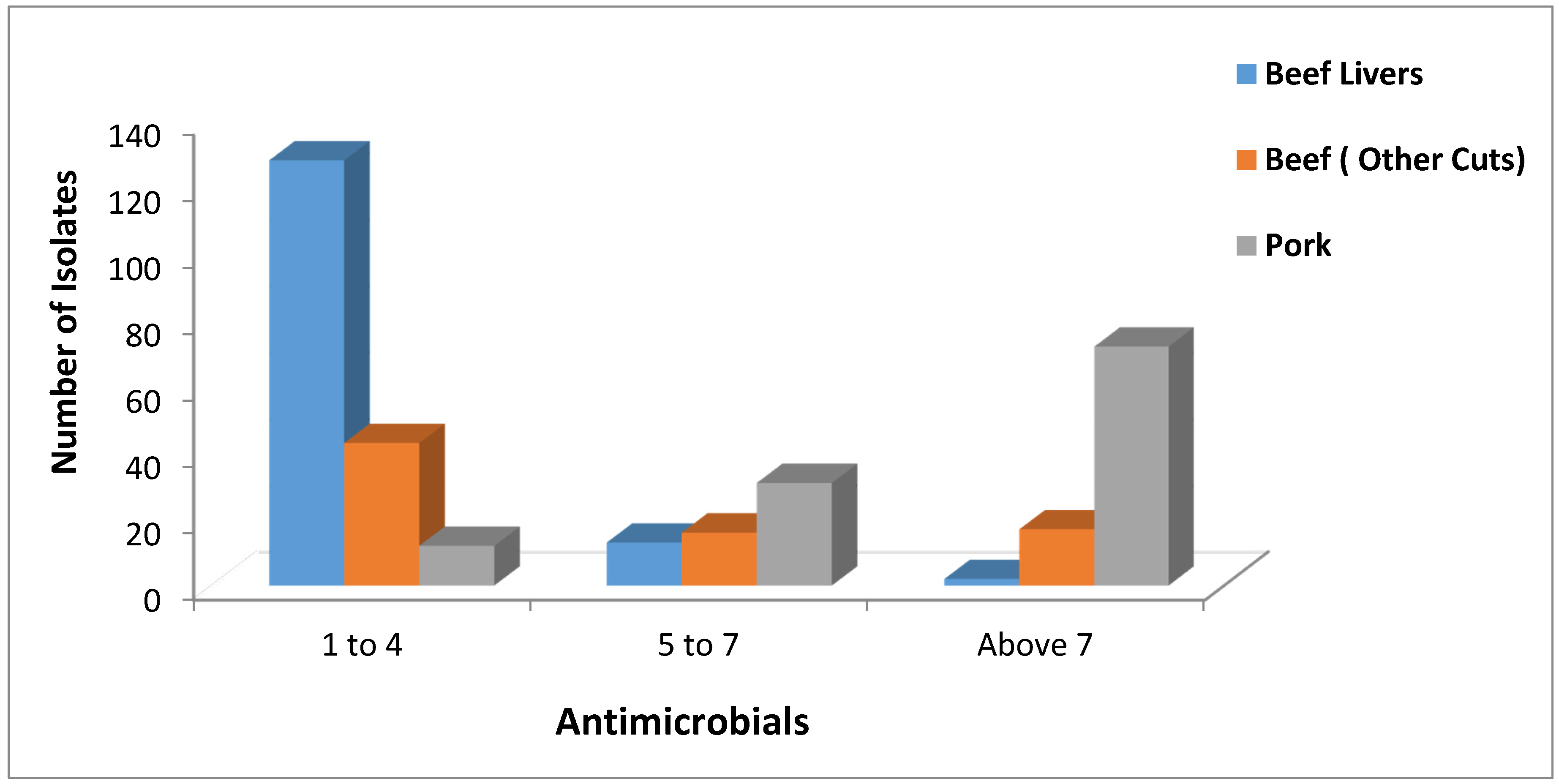
2.3. Toxin Gene Possession Screening
| Prevalence of Toxin Genes | ||||
|---|---|---|---|---|
| Toxin Gene | Beef | Pork np/n (%) | ||
| Beef Livers * np/n (%) | Beef ( Other Cuts) np/n (%) | Total np/n (%) | ||
| sea ns | 0/143 (0.0) | 2/76 (2.6) | 2/219 (0.9) | 0/115 (0) |
| seb-sec ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 0/115 (0) |
| sec ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 1/115 (0.9) |
| sed ns | 0/143 (0.0) | 2/76 (2.6) | 2/219 (0.9) | 0/115 (0) |
| see ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 0/115 (0) |
| seg ** | 4/143 (2.8) | 5/76 (6.6) | 9/219 (4.1) | 18/115 (15.7) |
| seh ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 36/115 (31.3) |
| sei ** | 2/143 (1.4) | 11/76 (14.5) | 13/219 (5.9) | 8/115 (6.9) |
| sej ns | 1/143 (0.7) | 1/76 (1.32) | 2/219 (0.9) | 0/115 (0) |
| tst *** | 4/143 (2.8) | 0/76 (0.0) | 4/219 (1.8) | 15/115 (13.0) |
| eta ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 0/115 (0) |
| etb ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 0/115 (0) |
| lukE-lukD *** | 12/143 (8.4) | 24/76 (31.6) | 36/219 (16.4) | 41/115 (35.7) |
| lukM ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 0/115 (0) |
| hla *** | 141/143 (98.6) | 63/76 (82.9) | 204/219 (93.2) | 100/115 (86.9) |
| hlb *** | 128/143 (89.5) | 38/76 (50.0) | 166/219 (75.8) | 41/115 (35.7) |
| hld *** | 141/143 (98.6) | 63/76 (82.9) | 204/219 (93.2) | 100/115 (86.9) |
| lukS-lukF ns | 0/143 (0.0) | 0/76 (0.0) | 0/219 (0) | 1/115 (0.9) |
2.4. Molecular Typing Using Spa Typing and PFGE
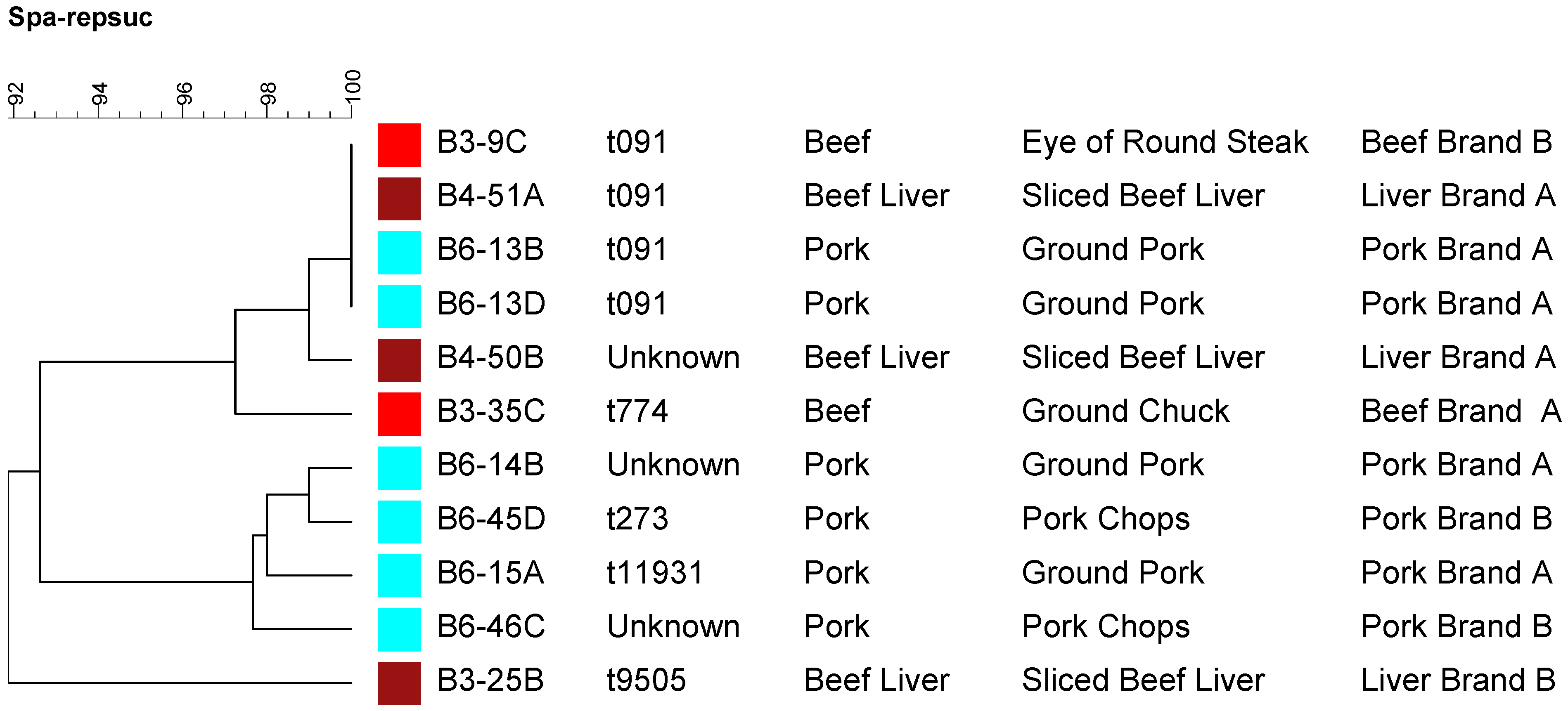
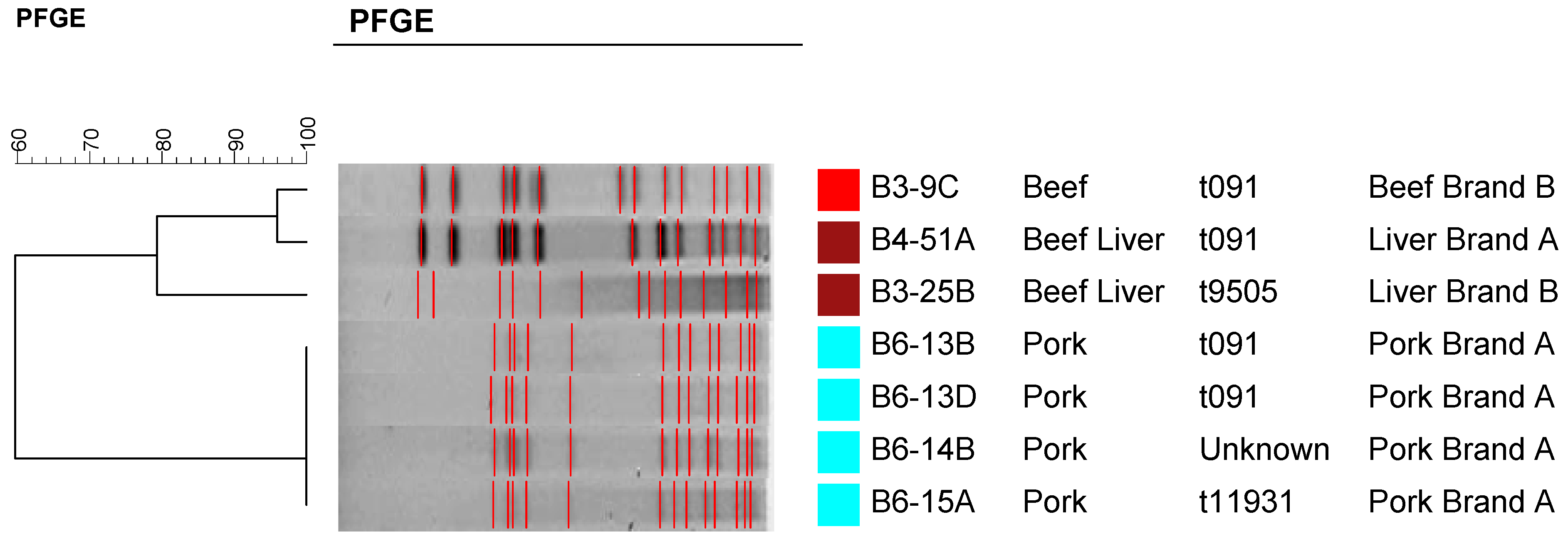
3. Experimental Section
3.1. Isolation of Staphylococcus aureus from Retail Meat Samples
3.2. DNA Extraction
3.3. PCR Identification
3.5. Antimicrobial Resistance Screening
| Antimicrobial Class | Antimicrobials | Conc. 1 (µg/mL) | Conc. 2 (µg/mL) (Break point) | Conc. 3 (µg/mL) | Conc. 4 (µg/mL) |
|---|---|---|---|---|---|
| β-Lactams | penicillin | 0.125 | 0.25 | 0.5 | 1 |
| ampicillin | 0.25 | 0.5 | 1 | 2 | |
| oxacillin + 2% Nacl | 2 | 4 | 8 | 16 | |
| cefoxitin + 2% Nacl | 4 | 8 | 16 | 32 | |
| Tetracyclines | tetracycline | 8 | 16 | 32 | 64 |
| doxycycline | 8 | 16 | 32 | 64 | |
| Macrolides | azithromycin | 4 | 8 | 16 | 32 |
| erythromycin | 4 | 8 | 16 | 32 | |
| Aminoglycosides | kanamycin | 32 | 64 | 128 | 256 |
| gentamicin | 8 | 16 | 32 | 64 | |
| Fluoroquinolones | ciprofloxacin | 2 | 4 | 8 | 16 |
| Lincosamides | clindamycin | 2 | 4 | 8 | 16 |
| Phenicols | chloramphenicol | 16 | 32 | 64 | 128 |
| Glycopeptides | vancomycin | 16 | 32 | 64 | 128 |
| Rifamycines | rifampin | 2 | 4 | 8 | 16 |
| Sulfonamides | trimethoprim/sulfamethoxazole | 2/38 | 4/76 | 8/152 | 16/304 |
3.6. Prevalence of Toxin Genes
| Toxin Gene | Size (bp) | Primer sequences (5'-3') | Multiplex PCR Reaction Set | Ref. |
|---|---|---|---|---|
| sea | 521 | GCA GGG AAC AGC TTT AGG C | A | [48] |
| GTT CTG TAG AAG TAT GAA ACA CG | ||||
| seb-sec | 665 | ATG TAA TTT TGA TAT TCG CAG TG | A | [48] |
| TGC AGG CAT CAT ATC ATA CCA | ||||
| sec | 284 | CTT GTA TGT ATG GAG GAA TAA CAA | A | [48] |
| TGC AGG CAT CAT ATC ATA CCA | ||||
| sed | 385 | GTG GTG AAA TAG ATA GGA CTG C | A | [48] |
| ATA TGA AGG TGC TCT GTG G | ||||
| see | 171 | TAC CAA TTA ACT TGT GGA TAG AC | A | [48] |
| CTC TTT GCA CCT TAC CGC | ||||
| seg | 328 | CGT CTC CAC CTG TTG AAG G | A | [48] |
| CCA AGT GAT TGT CTA TTG TCG | ||||
| seh | 359 | CAA CTG CTG ATT TAG CTC AG | B | [48] |
| GTC GAA TGA GTA ATC TCT AGG | ||||
| sei | 466 | CAA CTC GAA TTT TCA ACA GGT AC | B | [48] |
| CAG GCA GTC CAT CTC CTG | ||||
| sej | 142 | CAT CAG AAC TGT TGT TCC GCT AG | B | [48] |
| CTG AAT TTT ACC ATC AAA GGT AC | ||||
| tst | 560 | GCT TGC GAC AAC TGC TAC AG | B | [48] |
| TGG ATC CGT CAT TCA TTG TTA A | ||||
| eta | 93 | GCA GGT GTT GAT TTA GCA TT | B | [26] |
| AGA TGT CCC TAT TTT TGC TG | ||||
| etb | 226 | ACA AGC AAA AGA ATA CAG CG | B | [26] |
| GTT TTT GGC TGC TTC TCT TG | ||||
| lukS-lukF | 433 | ATC ATT AGG TAA AAT GTC TGG ACA TGA TCC A | C | [27] |
| GCA TCA AST GTA TTG GAT AGC AAA AGC | ||||
| lukE-lukD | 269 | TGA AAA AGG TTC AAA GTT GAT ACG AG | C | [27] |
| TGT ATT CGA TAG CAA AAG CAG TGC A | ||||
| lukM | 780 | TGG ATG TTA CCT ATG CAA CCT AC | C | [27] |
| GTT CGT TTC CAT ATA ATG AAT CAC TAC | ||||
| hla | 209 | CTG ATT ACT ATC CAA GAA ATT CGA TTG | C | [27] |
| CTT TCC AGC CTA CTT TTT TAT CAG T | ||||
| hlb | 309 | GTG CAC TTA CTG ACA ATA GTG C | C | [27] |
| GTT GAT GAG TAG CTA CCT TCA GT | ||||
| hld | 111 | AAG AAT TTT TAT CTT AAT TAA GGA AGG AGT G | C | [27] |
| TTA GTG AAT TTG TTC ACT GTG TCG A |
3.7. Molecular Typing of Staphylococcus aureus Isolates
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Normanno, G.; Firinu, A.; Virgilio, S.; Mula, G.; Dambrosio, A.; Poggiu, A.; Decastelli, L.; Mioni, R.; Scuota, S.; Bolzoni, G.; et al. Coagulase-positive Staphylococci and Staphylococcus aureus in food products marketed in Italy. Int. J. Food Microbiol. 2005, 98, 73–79. [Google Scholar] [PubMed]
- Lee, J.H. Methicillin (oxacillin)-resistant Staphylococcus aureus strains isolated from major food animals and their potential transmission to humans. Appl. Environ. Microbiol. 2003, 69, 6489–6494. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.E.; Iandolo, J.J. Staphylococcus. In Encyclopedia of Food Microbiology; Robinson, R.K., Batt, C.A., Patel, P.D., Eds.; Academic press: New York, NY, USA, 2000; pp. 2062–2065. [Google Scholar]
- Doyle, M.E.; Hartmann, F.A.; Lee Wong, A.C. Methicillin-resistant staphylococci: Implications for our food supply? Anim. Health Res. Rev. 2012, 13, 157–180. [Google Scholar] [CrossRef] [PubMed]
- Verkade, E.; Kluytmans, J. Livestock-associated Staphylococcus aureus CC398: Animal reservoirs and human infections. Infect. Genet. Evol. 2014, 21, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Kadariya, J.; Smith, T.C.; Thapaliya, D. Staphylococcus aureus and staphylococcal food-borne disease: An ongoing challenge in public health. BioMed. Res. Int. 2014, 2014, 827965. [Google Scholar] [CrossRef] [PubMed]
- Genigeorgis, C.A. Present state of knowledge on staphylococcal intoxication. Int. J. Food Microbiol. 1989, 9, 327–360. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Iandolo, J.J.; Stewart, G.C. The enterotoxin D plasmid of Staphylococcus aureus encodes a second enterotoxin determinant (sej). FEMS Microbial. Lett. 1998, 168, 227–233. [Google Scholar] [CrossRef]
- Tirado, C.; Schimdt, K. WHO surveillance programme for control of foodborne infections and intoxications: Preliminary results and trends across greater Europe. J. Infect. 2001, 43, 80–84. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control (CDC). CDC Estimates of Foodborne Illness in the United States. 2011. Available online: http://www.cdc.gov/foodborneburden/2011-foodborne-estimates.html (accessed on 16 May 2013). [Google Scholar]
- Kitai, S.; Shimizu, A.; Kawano, J.; Sato, E.; Nakano, C.; Uji, T.; Kitagawa, H. Characterization of methicillin-resistant Staphylococcus aureus isolated from retail raw chicken meat in Japan. J. Vet. Med. Sci. 2005, 67, 107–110. [Google Scholar] [CrossRef] [PubMed]
- De Neeling, A.J.; van Leeuwen, W.J.; Schouls, L.M.; Schot, C.S.; van Veen-Rutgers, A.; Beunders, A.J.; Buiting, A.G.; Hol, C.; Ligtvoet, E.E.; Petit, P.L.; et al. Resistance of staphylococci in The Netherlands: Surveillance by an electronic network during 1989–1995. J. Antimicrob. Chemoth. 1998, 41, 93–101. [Google Scholar] [CrossRef]
- Khanna, T.; Friendship, R.; Dewey, C.; Weese, J.S. Methicillin resistant Staphylococcus aureus colonization in pigs and pig farmers. Vet. Microbiol. 2008, 128, 298–303. [Google Scholar] [CrossRef] [PubMed]
- Voss, A.; Loeffen, F.; Bakker, J.; Klaassen, C.; Wulf, M. Methicillin resistant Staphylococcus aureus in pig farming. Emerg. Infect. Dis. 2005, 11, 1965–1966. [Google Scholar] [CrossRef] [PubMed]
- Van den Eede, A.; Hermans, K.; Lipinska, U.; Struelens, M.; Deplano, A.; Denis, O.; Gasthuys, F.; Haesebrouck, F.; Martens, A. Nasal carriage of methicillin-resistance Staphylococcus aureus in the equine population: Prevalence, typing and antimicrobial resistance. In Proceedings of the 2nd Symposium on Antimicrobial Resistance in Animals and the Environment, Tours, France, 17–19 December 2007.
- Pu, S.; Wang, F.; Ge, B. Isolation and characterization of methicillin-resistant Staphylococcus aureus strains from Louisiana retail meats. Appl. Environ. Microb. 2009, 75, 265–267. [Google Scholar] [CrossRef]
- Kelman, A.; Soong, Y.A.; Dupuy, N.; Shafer, D.; Richbourg, W.; Johnson, K.; Meng, J. Antimicrobial susceptibility of Staphylococcus aureus from retail ground meats. J. Food Prot. 2011, 74, 1625–1629. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, K.; Wang, X.; Donabedian, S.; Zervos, M.; da Rocha, L.; Zhang, Y. Methicillin-resistant Staphylococcus aureus in retail meat, Detroit, Michigan, USA. Emerg. Infect. Dis. 2011, 17, 1135. [Google Scholar] [CrossRef] [PubMed]
- Hanson, B.M.; Dressler, A.E.; Harper, A.L.; Scheibel, R.P.; Wardyn, S.E.; Roberts, L.K.; Smith, T.C. Prevalence of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus (MRSA) on retail meat in Iowa. J. Infect. Public Health 2011, 4, 169–174. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, A.M.; Hanson, B.M.; Farina, S.A.; Wu, J.Y.; Simmering, J.E.; Wardyn, S.E.; Smith, T.C. MRSA in conventional and alternative retail pork products. PLoS ONE 2012, 7, e30092. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.R.; Davis, J.A.; Barrett, J.B. Prevalence and characterization of methicillin-resistant Staphylococcus aureus isolates from retail meat and humans in Georgia. J. Clin. Microbiol. 2013, 51, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Buyukcangaz, E.; Velasco, V.; Sherwood, J.S.; Stepan, R.M.; Koslofsky, R.J.; Logue, C.M. Molecular typing of Staphylococcus aureus and methicillin-resistant S. aureus (MRSA) isolated from animals and retail meat in North Dakota, United States. Foodborne Pathog. Dis. 2013, 10, 608–617. [Google Scholar] [CrossRef] [PubMed]
- Velasco, V.; Sherwood, J.S.; Rojas-García, P.P.; Logue, C.M. Multiplex real-time PCR for detection of Staphylococcus aureus, mecA and Panton-Valentine Leukocidin (PVL) genes from selective enrichments from animals and retail meat. PloS ONE 2014, 9, e97617. [Google Scholar] [CrossRef] [PubMed]
- David, M.Z.; Daum, R.S. Community-associated methicillin-resistant Staphylococcus aureus: Epidemiology and clinical consequences of an emerging epidemic. Clin. Microbiol. Rev. 2010, 23, 616–687. [Google Scholar] [CrossRef] [PubMed]
- Pu, S.; Wang, F.; Ge, B. Characterization of toxin genes and antimicrobial susceptibility of Staphylococcus aureus isolates from Louisiana retail meats. Foodborne Pathog. Dis. 2011, 8, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Mehrotra, M.; Wang, G.; Johnson, W.M. Multiplex PCR for detection of genes for Staphylococcu aureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. J. Clin. Microbiol. 2000, 38, 1032–1035. [Google Scholar] [PubMed]
- Jarraud, S.; Mougel, C.; Thioulouse, J.; Lina, G.; Meugnier, H.; Forey, F.; Nesme, X.; Etienne, J.; Vandenesch, F. Relationships between Staphylococcus aureus genetic background, virulence factors, agr groups (alleles), and human disease. Infect. Immun. 2002, 70, 631–641. [Google Scholar] [CrossRef] [PubMed]
- Gunaydin, B.; Aslantas, O.; Demir, C. Detection of superantigenic toxin genes in Staphylococcus aureus strains from subclinical bovine mastitis. Trop. Anim. Health Prod. 2011, 43, 1633–1637. [Google Scholar] [PubMed]
- Hwang, S.Y.; Kim, S.H.; Jang, E.J.; Kwon, N.H.; Park, Y.K.; Koo, H.C.; Jung, W.K.; Kim, J.M.; Park, W.H. Novel multiplex PCR for the detection of the Staphylococcus aureus superantigen and its application to raw meat isolates in Korea. Int. J. Food Microbiol. 2007, 117, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Jay, J.M. Further studies on staphylococci in meats, III. Occurrence and characteristics of coagulase-positive strains from a variety of nonfrozen market cuts. Appl. Microbiol. 1962, 10, 247–251. [Google Scholar] [PubMed]
- Waters, A.E.; Contente-Cuomo, T.; Buchhagen, J.; Liu, C.M.; Watson, L.; Pearce, K.; Foster, J.T.; Bowers, J.; Driebe, E.M.; Engelthaler, D.M.; et al. Multidrug-resistant Staphylococcus aureus in US meat and poultry. Clin. Infect. Dis. 2011, 52, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Van Loo, I.H.; Diederen, B.M.; Savelkoul, P.H.; Woudenberg, J.H.; Roosendaal, R.; van Belkum, A.; Lemmens-den Toom, N.; Verhulst, C.; van Keulen, P.H.; Kluytmans, J.A. Methicillin-resistant Staphylococcus aureus in meat products, the Netherlands. Emerg. Infect. Dis. 2007, 13, 1753–1755. [Google Scholar] [CrossRef] [PubMed]
- HSlifierz, M.J.; Friendship, R.M.; Weese, J.S. Methicillin-resistant Staphylococcus aureus in commercial swine herds is associated with disinfectant and zinc usage. Appl. Environ. Microbiol. 2015, 81, 2690–2695. [Google Scholar] [CrossRef] [PubMed]
- Pereira, V.; Lopes, C.; Castro, A.; Silva, J.; Gibbs, P.; Teixeira, P. Characterization for enterotoxin production, virulence factors, and antibiotic susceptibility of Staphylococcus aureus isolates from various foods in Portugal. Food Microbiol. 2009, 26, 278–282. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, L.G.; Holden, M.T.G.; Lindsay, H.; Webb, C.R.; Brown, D.F.J.; Curran, M.D.; Walpole, E.; Brooks, K.; Pickard, D.J.; Teale, C.; et al. Methicillin-resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: A descriptive study. Lancet 2011, 11, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Shore, A.C.; Deasy, E.C.; Slickers, P.; Brennan, G.; Connell, B.O.; Monecke, S.; Ehricht, R.; Coleman, D.C. Detection of staphylococcal cassette chromosome mec Type XI carrying highly divergent mecA, mecI, mecR1, blaZ, and ccr genes in human clinical isolates of clonal complex 130 Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Ch. 2011, 55, 3765–3773. [Google Scholar] [CrossRef]
- Snyder, H.L.; Niebuhr, S.E.; Dickson, J.S. Transfer of Methicillin-Resistant Staphylococcus aureus from Retail Pork Products onto Food Contact Surfaces and the Potential for Consumer Exposure. J. Food Protect. 2013, 76, 2087–2092. [Google Scholar] [CrossRef]
- Shukla, S.K.; Karow, M.E.; Brady, J.M.; Stemper, M.E.; Kislow, J.; Moore, N.; Wroblewski, K.; Chyou, P.-H.; Warshauer, D.M.; Reed, K.D.; et al. Virulence genes and genotypic associations in nasal carriage, community-associated methicillin-susceptible and methicillin-resistant USA400 Staphylococcus aureus isolates. J. Clin. Microbiol. 2010, 48, 3582–3592. [Google Scholar] [CrossRef] [PubMed]
- Hawken, P.; Weese, J.S.; Friendship, R.; Warriner, K. Longitudinal study of clostridium difficile and methicillin-resistant Staphylococcus aureus associated with pigs from weaning through to the end of processing. J. Food Protect. 2013, 76, 624–630. [Google Scholar] [CrossRef]
- Boost, M.; Ho, J.; Guardabassi, L.; O’Donoghue, M. Colonization of butchers with livestock-associated methicillin-resistant Staphylococcus aureus. Zoonoses Public Health 2013, 60, 572–576. [Google Scholar] [CrossRef] [PubMed]
- Ghafir, Y.; China, B.; Dierick, K.; de Zutter, L.; Daube, G. A seven-year survey of Campylobacter contamination in meat at different production stages in Belgium. Int. J. Food Microbiol. 2007, 116, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Noormohamed, A.; Fakhr, M.K. A higher prevalence rate of Campylobacter in retail beef livers compared to other beef and pork meat cuts. Int. J. Environ. Res. Public Health 2013, 10, 2058–2068. [Google Scholar] [CrossRef] [PubMed]
- Noormohamed, A.; Fakhr, M.K. Incidence and antimicrobial resistance profiling of Campylobacter in retail chicken livers and gizzards. Foodborne Pathog. Dis. 2012, 9, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Marmur, J. A procedure for the isolation of deoxyribonucleic acid from microorganisms. J. Mol. Bio. 1961, 3, 208–218. [Google Scholar]
- Martineau, F.; Picard, F.J.; Roy, P.H.; Ouellette, M.; Bergeron, M.G. Species-specific and ubiquitous-DNA-based assays for rapid identification of Staphylococcus aureus. J. Clin. Microbiol. 1998, 36, 618–623. [Google Scholar] [PubMed]
- Murakami, K.; Minamide, W.; Wada, K.; Nakamura, E.; Teraoka, H.; Watanabe, S. Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J. Clin. Microbiol. 1991, 29, 2240–2244. [Google Scholar]
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. 2011; 31, 1–172. [Google Scholar]
- Monday, S.R.; Bohach, G.A. Use of multiplex PCR to detect classical and newly described pyrogenic toxin genes in staphylococcal isolates. J. Clin. Microbiol. 1999, 37, 3411–3414. [Google Scholar]
- Shopsin, B.; Gomez, M.; Montgomery, S.O.; Smith, D.H.; Waddington, M.; Dodge, D.E.; Bost, D.E.; Riehman, M.; Naidich, S.; Kreiswirth, B.N. Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J. Clin. Microbiol. 1999, 37, 3556–3563. [Google Scholar] [PubMed]
- McDougal, L.K.; Steward, C.D.; Killgore, G.E.; Chaitram, J.M.; McAllister, S.K.; Tenover, F.C. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: Establishing a national database. J. Clin. Microbiol. 2003, 41, 5113–5120. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdalrahman, L.S.; Wells, H.; Fakhr, M.K. Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts. Pathogens 2015, 4, 182-198. https://doi.org/10.3390/pathogens4020182
Abdalrahman LS, Wells H, Fakhr MK. Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts. Pathogens. 2015; 4(2):182-198. https://doi.org/10.3390/pathogens4020182
Chicago/Turabian StyleAbdalrahman, Lubna S., Harrington Wells, and Mohamed K. Fakhr. 2015. "Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts" Pathogens 4, no. 2: 182-198. https://doi.org/10.3390/pathogens4020182
APA StyleAbdalrahman, L. S., Wells, H., & Fakhr, M. K. (2015). Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts. Pathogens, 4(2), 182-198. https://doi.org/10.3390/pathogens4020182






