Antibiotic Resistance Related to Biofilm Formation in Klebsiella pneumoniae
Abstract
:1. Introduction
2. Antibiotic Resistance
3. Biofilm
3.1. Virulence and Biofilm Formation
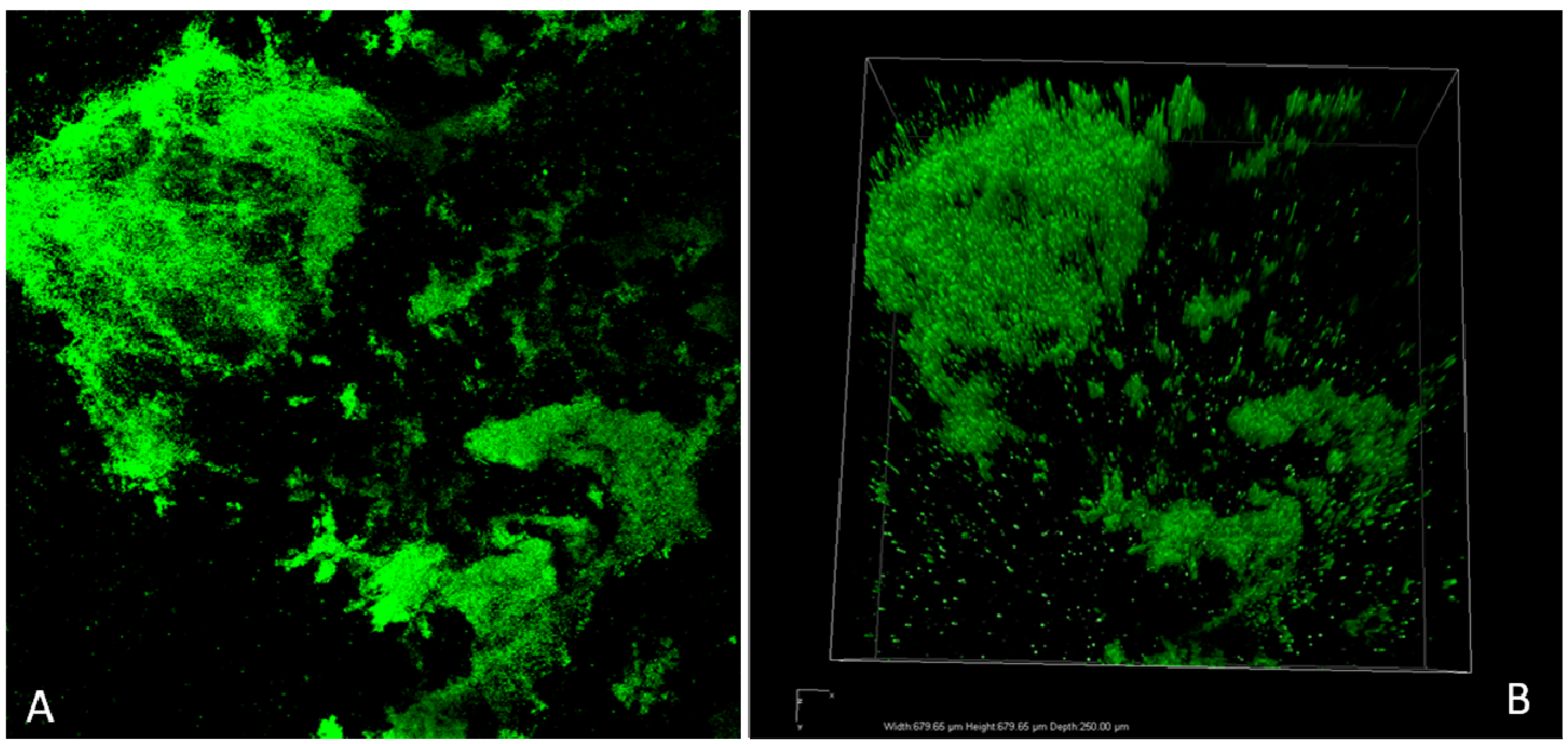
3.2. Involvement in Mixed Biofilms
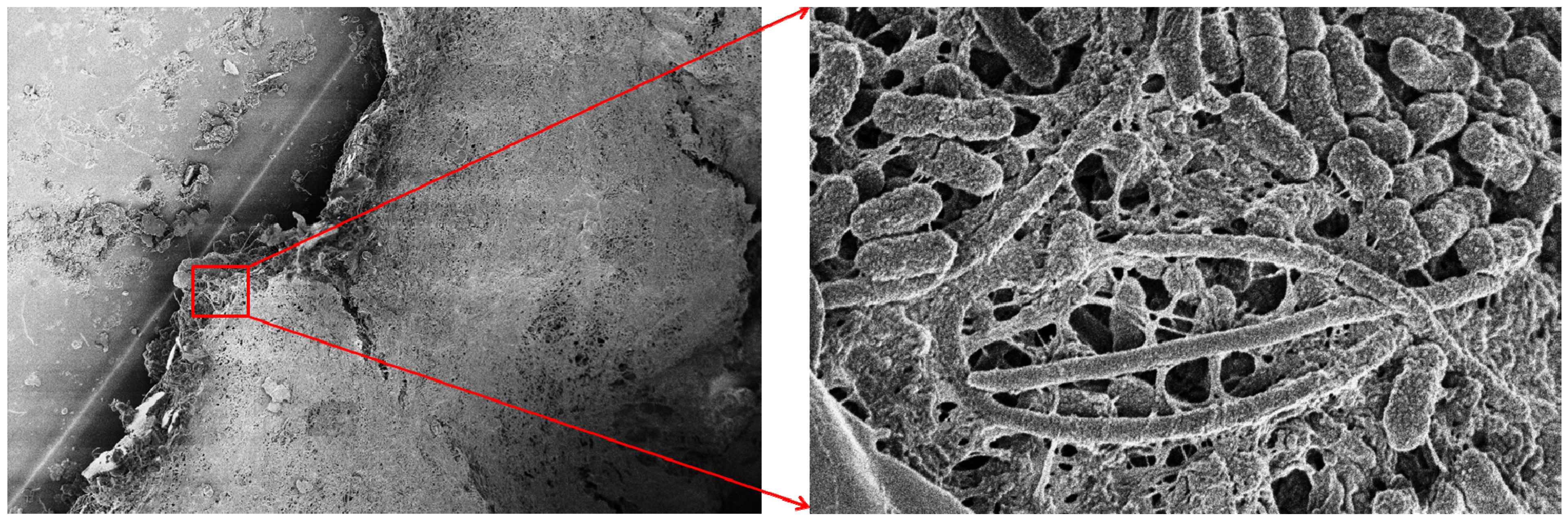
4. Antibiotic Resistance of Biofilm-Growing Strains
5. Correlation between Biofilm and Antibiotic Resistance
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Varaldo, P.E.; Biavasco, F.; Mannelli, S.; Pompei, R.; Proietti, A. Distribution and antibiotic susceptibility of extraintestinal clinical isolates of Klebsiella, Enterobacter and Serratia species. Eur. J. Clin. Microbiol. Infect. Dis. 1988, 4, 495–500. [Google Scholar] [CrossRef]
- Bagley, S.T. Habitat association of Klebsiella species. Infect. Control 1985, 6, 52–58. [Google Scholar] [PubMed]
- Podschun, R.; Ullmann, U. Klebsiella spp. as nosocomial pathogens: Epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 1998, 11, 589–603. [Google Scholar]
- Arakawa, Y.; Wacharotayankun, R.; Nagatsuka, T.; Ito, H.; Kato, N.; Ohta, M. Genomic organization of the Klebsiella pneumoniae cps region responsible for serotype K2 capsular polysaccharide synthesis in virulent strain Chedid. J. Bacteriol. 1995, 177, 1788–1796. [Google Scholar] [PubMed]
- Gerlach, G.F.; Clegg, S.; Allen, B.L. Identification and characterization of the genes encoding the type 3 and type 1 fimbrial adhesins of Klebsiella pneumoniae. J. Clin. Microbiol. 1989, 171, 1262–1270. [Google Scholar]
- Martino, P.D.; Bertin, Y.; Girardeau, J.P.; Livrelli, V.; Joly, B.; Darfeuille Michaud, A. Molecular characterization and adhesive properties of CF29K, and adhesion of Klebsiella pneumoniae strains involved in nosocomial infections. Infect. Immun. 1995, 63, 4336–4344. [Google Scholar] [PubMed]
- Ørskov, I.; Ørskov, F. Serotyping of Klebsiella. Methods Microbiol. 1984, 14, 143–164. [Google Scholar]
- Ehrenwort, L.; Baer, H. The pathogenicity of Klebsiella pneumoniae for mice: The relationship to the quantity and rate of production of type-specific capsular polysaccharide. J. Bacteriol. 1956, 72, 713–717. [Google Scholar] [PubMed]
- Amako, K.; Meno, Y.; Takade, A. Fine structures of the capsules of Klebsiella pneumoniae and Escherichia coli K1. J. Bacteriol. 1988, 170, 4960–4962. [Google Scholar] [PubMed]
- Podschun, R.; Ullmann, U. Klebsiella capsular type K7 in relation to toxicity, susceptibility to phagocytosis and resistance to serum. J. Med. Microbiol. 1992, 36, 250–254. [Google Scholar] [CrossRef] [PubMed]
- Williams, P.; Lambert, P.A.; Brown, M.R.W.; Jones, R.J. The role of the O and K antigens in determining the resistance of Klebsiella aerogenes to serum killing and phagocytosis. J. Gen. Microbiol. 1983, 129, 2181–2191. [Google Scholar] [PubMed]
- Livrelli, V.; de Champs, C.; di Martino, P.; Darfeuille-Michaud, A.; Forestier, C.; Joly, B. Adhesive properties and antibiotic resistance of Klebsiella, Enterobacter, and Serratia clinical isolates involved in nosocomial infections. J. Clin. Microbiol. 1996, 34, 1963–1969. [Google Scholar] [PubMed]
- Jones, G.W.; Isaacson, R.E. Proteinaceous bacterial adhesins and their receptors. Crit. Rev. Microbiol. 1983, 10, 229–260. [Google Scholar] [CrossRef] [PubMed]
- Ottow, J.C.G. Ecology, physiology, and genetics of fimbriae and pili. Annu. Rev. Microbiol. 1975, 29, 79–108. [Google Scholar] [CrossRef] [PubMed]
- Kline, K.A.; Dodson, K.W.; Caparon, M.G.; Hultgren, S.J. A tale of two pili: Assembly and function of pili in bacteria. Trends Microbiol. 2010, 18, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Rosen, D.A.; Pinkner, J.S.; Jones, J.M.; Walker, J.N.; Clegg, S.; Hultgren, S.J. Utilization of an intracellular bacterial community pathway in Klebsiella pneumoniae urinary tract infection and the effects of FimK on type 1 pilus expression. Infect. Immun. 2008, 76, 3337–3345. [Google Scholar] [CrossRef] [PubMed]
- Struve, C.; Bojer, M.; Krogfelt, K.A. Characterization of Klebsiella pneumoniae type 1 fimbriae by detection of phase variation during colonization and infection and impact on virulence. Infect. Immun. 2008, 76, 4055–4065. [Google Scholar] [CrossRef] [PubMed]
- Alcántar-Curiel, M.D.; Blackburn, D.; Saldaña, Z.; Gayosso-Vázquez, C.; Iovine, N.M.; de la Cruz, M.A.; Girón, J.A. Multi-functional analysis of Klebsiella pneumoniae fimbrial types in adherence and biofilm formation. Virulence 2013, 4, 129–138. [Google Scholar]
- Burmolle, M.; Bahl, M.I.; Jensen, L.B.; Sorensen, S.J.; Hansen, L.H. Type 3 fimbriae, encoded by the conjugative plasmid pOLA52, enhance biofilm formation and transfer frequencies in Enterobacteriaceae strains. Microbiology 2008, 154, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Ong, C.L.; Ulett, G.C.; Mabbett, A.N.; Beatson, S.A.; Webb, R.I.; Monaghan, W.; Nimmo, G.R.; Looke, D.F.; McEwan, A.G.; Schembri, M.A. Identification of type 3 fimbriae in uropathogenic Escherichia coli reveals a role in biofilm formation. J. Bacteriol. 2008, 190, 1054–1063. [Google Scholar] [CrossRef] [PubMed]
- Ong, C.L.; Beatson, S.A.; Totsika, M.; Forestier, C.; McEwan, A.G.; Schembri, M.A. Molecular analysis of type 3 fimbrial genes from Escherichia coli, Klebsiella and Citrobacter species. BMC Microbiol. 2010, 10, 183. [Google Scholar] [CrossRef] [PubMed]
- Lederman, E.R.; Crum, N.F. Pyogenic liver abscess with a focus on Klebsiella pneumoniae as a primary pathogen: An emerging disease with unique clinical characteristics. Am. J. Gastroenterol. 2005, 100, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Fung, C.P.; Chang, F.Y.; Lee, S.C.; Hu, B.S.; Kuo, B.I.; Liu, C.Y.; Ho, M.; Siu, L.K. A global emerging disease of Klebsiella pneumoniae liver abscess: Is serotype K1 an important factor for complicated endophthalmitis? Gut 2002, 50, 420–424. [Google Scholar] [CrossRef] [PubMed]
- Laupland, K.B.; Ross, T.; Pitout, J.D.; Church, D.L.; Gregson, D.B. Community-onset urinary tract infections: A population-based assessment. Infection 2007, 35, 150–153. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.H.; Kao, C.Y.; Yang, D.C.; Tseng, C.C.; Wu, A.B.; Teng, C.H.; Wang, M.C.; Wu, J.J. Clinical and microbiological characteristics of Klebsiella pneumoniae from community-acquired recurrent urinary tract infections. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 1533–1539. [Google Scholar] [CrossRef] [PubMed]
- Tu, Y.C.; Lu, M.C.; Chiang, M.K.; Huang, S.P.; Peng, H.L.; Chang, H.Y.; Jan, M.S.; Lai, Y.C. Genetic requirements for Klebsiella pneumoniae-induced liver abscess in an oral infection model. Infect. Immun. 2009, 77, 2657–2671. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.K.; Chung, D.R.; Wie, S.H.; Yoo, J.H.; Park, S.W. Risk factor analysis of invasive liver abscess caused by the K1 serotype Klebsiella pneumoniae. Eur. J. Clin. Microbiol. Infect. Dis. 2009, 28, 109–111. [Google Scholar] [CrossRef] [PubMed]
- Oelschlaeger, T.A.; Tall, B.D. Invasion of cultured human epithelial cells by Klebsiella pneumoniae isolated from the urinary tract. Infect. Immun. 1997, 6, 2950–2958. [Google Scholar]
- Montgomerie, J.Z. Epidemiology of Klebsiella and hospital-associated infections. Rev. Infect. Dis. 1979, 1, 736–753. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A. Hospital-acquired infections in the neonatal intensive care unit-Klebsiella pneumoniae. Semin. Perinatol. 2002, 26, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Ko, W.C.; Paterson, D.L.; Sagnimeni, A.J.; Hansen, D.S.; von Gottberg, A.; Mohapatra, S.; Casellas, J.M.; Goossens, H.; Mulazimoglu, L.; Trenholme, G.; et al. Community acquired Klebsiella pneumoniae bacteremia: Global differences in clinical patterns. Emerg. Infect. Dis. 2002, 8, 160–166. [Google Scholar] [CrossRef] [PubMed]
- Yinnon, A.M.; Butnaru, A.; Raveh, D.; Jerassy, Z.; Rudensky, B. Klebsiella bacteremia: community versus nosocomial infection. Mon. J. Assoc. Physicians 1996, 89, 933–941. [Google Scholar]
- Niveditha, S.; Pramodhini, S.; Umadevi, S.; Kumar, S.; Stephen, S. The isolation and the biofilm formation of uropathogens in the patients with catheter associated urinary tract infections (UTIs). J. Clin. Diagn. Res. 2012, 6, 1478–1482. [Google Scholar] [PubMed]
- Ronald, A. The etiology of urinary tract infection: Traditional and emerging pathogens. Am. J. Med. 2002, 113, 14S–19S. [Google Scholar] [CrossRef] [PubMed]
- Frank, D.N.; Wilson, S.S.; St Amand, A.L.; Pace, N.R. Culture independent microbiological analysis of Foley urinary catheter biofilms. PLoS One 2009, 4, e7811. [Google Scholar] [CrossRef] [PubMed]
- Lye, W.C.; Chan, R.K.T.; Lee, E.J.C.; Kumarasinghe, G. Urinary tract infections in patients with diabetes mellitus. J. Infect. 1992, 24, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Bennett, C.J.; Young, M.N.; Darrington, H. Differences in urinary tract infection in male and female spinal cord injury patients on intermittent catheterization. Paraplegia 1995, 33, 69–72. [Google Scholar] [CrossRef] [PubMed]
- Siegman-Igra, Y.; Golan, H.; Schwartz, D.; Cahaner, Y.; de-Mayo, G.; Orni-Wasserlauf, R. Epidemiology of vascular catheter-related bloodstream infections in a large university hospital in Israel. Scand. J. Infect. Dis. 2000, 32, 411–415. [Google Scholar]
- Singhai, M.; Malik, A.; Shahid, M.; Malik, M.A.; Goyal, R. A study on device-related infections with special reference to biofilm production and antibiotic resistance. J. Glob. Infect. Dis. 2012, 4, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.Y.; Lyles-Banks, R.D.; Lolans, K.; Hines, D.W.; Spear, J.B.; Petrak, R.; Trick, W.E.; Weinstein, R.A.; Hayden, M.K.; Centers for Disease Control and Prevention Epicenters Program. The importance of long-term acute care hospitals in the regional epidemiology of Klebsiella pneumoniae carbapenemase-producing Enterobacteriaceae. Clin. Infect. Dis. 2013, 57, 1246–1252. [Google Scholar]
- Orsi, G.B.; Scorzolini, L.; Franchi, C.; Mondillo, V.; Rosa, G.; Venditti, M. Hospital-acquired infection surveillance in a neurosurgical intensive care unit. J. Hosp. Infect. 2006, 64, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Giani, T.; Pini, B.; Arena, F.; Conte, V.; Bracco, S.; Migliavacca, R.; AMCLI-CRE Survey Participants; Pantosti, A.; Pagani, L.; Luzzaro, F.; et al. Epidemic diffusion of KPC carbapenemase-producing Klebsiella pneumoniae in Italy: Results of the first countrywide survey, 15 May to 30 June 2011. Euro Surveill. 2013, 30, 18–22. [Google Scholar]
- Azzopardi, E.A.; Azzopardi, E.; Camilleri, L.; Villapalos, J.; Boyce, D.E.; Dziewulski, P.; Dickson, W.A.; Whitaker, I.S. Gram negative wound infection in hospitalised adult burn patients-systematic review and metanalysis. PLoS One 2014, 9, e95042. [Google Scholar] [CrossRef] [PubMed]
- Varaldo, P.E.; Nicoletti, G.; Schito, G.C.; Maida, A.; Facinelli, B.; Stefani, S.; Gianrossi, G.; Muresu, E. Circulation in Italy of beta-lactamase-producing strains within the major groups of bacterial pathogens. Eur. J. Epidemiol. 1990, 6, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Guérillot, F.; Carret, G.; Flandrois, J.P. A statistical evaluation of the bactericidal effects of ceftibuten in combination with aminoglycosides and ciprofloxacin. J. Antimicrob. Chemother. 1993, 32, 685–694. [Google Scholar]
- Elkhaïli, H.; Kamili, N.; Linger, L.; Levêque, D.; Pompei, D.; Monteil, H.; Jehl, F. In vitro time-kill curves of cefepime and cefpirome combined with amikacin, gentamicin or ciprofloxacin against Klebsiella pneumoniae producing extended-spectrum beta-lactamase. Chemotherapy 1997, 43, 245–253. [Google Scholar]
- Traub, W.H.; Schwarze, I.; Bauer, D. Nosocomial outbreak of cross-infection due to multiple-antibiotic-resistant Klebsiella pneumoniae: Characterization of the strain and antibiotic susceptibility studies. Chemotherapy 2000, 46, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Patel, G.; Bonomo, R.A. Status report on carbapenemases: Challenges and prospects. Expert Rev. Anti Infect. Ther. 2011, 9, 555–570. [Google Scholar] [CrossRef] [PubMed]
- Voulgari, E.; Poulou, A.; Koumaki, V.; Tsakris, A. Carbapenemase-producing Enterobacteriaceae: Now that the storm is finally here, how will timely detection help us fight back? Future Microbiol. 2013, 8, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Yigit, H.; Queenan, A.M.; Anderson, G.J.; Domenech-Sanchez, A.; Biddle, J.W.; Steward, C.D.; Alberti, S.; Bush, K.; Tenover, F.C. Novel carbapenem-hydrolyzing beta-lactamase, KPC-1, from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2001, 45, 1151–1161. [Google Scholar] [CrossRef] [PubMed]
- Papp-Wallace, K.M.; Bethel, C.R.; Distler, A.M.; Kasuboski, C.; Taracila, M.; Bonomo, R.A. Inhibitor resistance in the KPC-2 beta-lactamase, a preeminent property of this class A beta-lactamase. Antimicrob. Agents Chemother. 2010, 54, 890–897. [Google Scholar]
- Molton, J.S.; Tambyah, P.A.; Ang, B.S.; Ling, M.L.; Fisher, D.A. The global spread of healthcare-associated multidrug-resistant bacteria: A perspective from Asia. Clin. Infect. Dis. 2013, 56, 1310–1318. [Google Scholar] [CrossRef] [PubMed]
- Grundmann, H.; Livermore, D.M.; Giske, C.G.; Canton, R.; Rossolini, G.M.; Campos, J.; Vatopoulos, A.; Gniadkowski, M.; Toth, A.; Pfeifer, Y.; et al. CNSE Working Group. Carbapenem-non-susceptible Enterobacteriaceae in Europe: Conclusions from a meeting of national experts. Euro Surveill. 2010, 15, 19711. [Google Scholar]
- Miriagou, V.; Cornaglia, G.; Edelstein, M.; Galani, I.; Giske, C.G.; Gniadkowski, M.; Malamou-Lada, E.; Martinez-Martinez, L.; Navarro, F.; Nordmann, P.; et al. Acquired carbapenemases in Gram-negative bacterial pathogens: Detection and surveillance issues. Clin. Microbiol. Infect. 2010, 16, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Tzouvelekis, L.S.; Markogiannakis, A.; Psichogiou, M.; Tassios, P.T.; Daikos, G.L. Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: An evolving crisis of global dimensions. Clin. Microbiol. Rev. 2012, 25, 682–707. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, B.; Tsang, A.K.; Yi, Y.; Woo, P.C.; Liu, C.H. Genotypic analysis of Klebsiella pneumoniae isolates in a Beijing Hospital reveals high genetic diversity and clonal population structure of drug-resistant isolates. PLoS One 2013, 8, e57091. [Google Scholar] [CrossRef] [PubMed]
- Le Chevallier, M.W.; Cawthon, C.D.; Lee, R.G. Factors promoting survival of bacteria in chlorinated water supplies. Appl. Environ. Microbiol. 1988, 54, 649–654. [Google Scholar]
- Reid, G.; Charbonneau-Smith, R.; Lam, D.; Kang, Y.S.; Lacerte, M.; Hayes, K.C. Bacterial biofilm formation in the urinary bladder of spinal cord injured patients. Paraplegia 1992, 30, 711–717. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Zhang, Z. Biofilm-forming Klebsiella pneumoniae strains have greater likelihood of producing extended-spectrum beta-lactamases. J. Hosp. Infect. 2008, 68, 369–371. [Google Scholar] [CrossRef] [PubMed]
- Nicolau Korres, A.M.; Aquije, G.M.; Buss, D.S.; Ventura, J.A.; Fernandes, P.M.; Fernandes, A.A. Comparison of biofilm and attachment mechanisms of a phytopathological and clinical isolate of Klebsiella pneumoniae subsp. pneumoniae. Sci. World J. 2013, 10, 925375. [Google Scholar]
- Donelli, G.; Vuotto, C. Biofilm-based infections in long-term care facilities. Future Microbiol. 2014, 9, 175–188. [Google Scholar] [CrossRef] [PubMed]
- Langstraat, J.; Bohse, M.; Clegg, S. Type 3 fimbrial shaft (MrkA) of Klebsiella pneumoniae, but not the fimbrial adhesin (MrkD), facilitates biofilm formation. Infect. Immun. 2001, 69, 5805–5812. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, P.; Cafferini, N.; Joly, B.; Darfeuille-Michaud, A. Klebsiella pneumoniae type 3 pili facilitate adherence and biofilm formation on abiotic surfaces. Res. Microbiol. 2003, 154, 9–16. [Google Scholar]
- Jagnow, J.; Clegg, S. Klebsiella pneumoniae MrkD-mediated biofilm formation on extracellular matrix- and collagen-coated surfaces. Microbiology 2003, 149, 2397–2405. [Google Scholar] [CrossRef] [PubMed]
- Lavender, H.F.; Jagnow, J.R.; Clegg, S. Biofilm formation in vitro and virulence in vivo of mutants of Klebsiella pneumoniae. Infect. Immun. 2004, 72, 4888–4890. [Google Scholar] [CrossRef] [PubMed]
- Boddicker, J.D.; Anderson, R.A.; Jagnow, J.; Clegg, S. Signature-tagged mutagenesis of Klebsiella pneumoniae to identify genes that influence biofilm formation on extracellular matrix material. Infect. Immun. 2006, 74, 4590–4597. [Google Scholar] [CrossRef] [PubMed]
- Schroll, C.; Barken, K.B.; Krogfelt, K.A.; Struve, C. Role of type 1 and type 3 fimbriae in Klebsiella pneumoniae biofilm formation. BMC Microbiol. 2010, 10, 179. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.G.; Clegg, S. Role of MrkJ, a phosphodiesterase, in type 3 fimbrial expression and biofilm formation in Klebsiella pneumoniae. J. Bacteriol. 2010, 192, 3944–3950. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.N.; Mortensen, M.S.; Krogfelt, K.A.; Clegg, S. Role of Klebsiella pneumoniae type 1 and type 3 fimbriae in colonizing silicone tubes implanted into the bladders of mice as a model of catheter-associated urinary tract infections. Infect. Immun. 2013, 81, 3009–3017. [Google Scholar] [CrossRef] [PubMed]
- Balestrino, D.; Ghigo, J.M.; Charbonnel, N.; Haagensen, J.A.; Forestier, C. The characterization of functions involved in the establishment and maturation of Klebsiella pneumoniae in vitro biofilm reveals dual roles for surface exopolysaccharides. Environ. Microbiol. 2008, 10, 685–701. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.C.; Lin, T.L.; Hsieh, P.F.; Yang, H.C.; Wang, J.T. Isolation of genes involved in biofilm formation of a Klebsiella pneumoniae strain causing pyogenic liver abscess. PLoS One 2011, 6, e23500. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.W.; Lam, I.; Chang, H.Y.; Tsai, S.F.; Palsson, B.O.; Charusanti, P. Capsule deletion via a λ-Red knockout system perturbs biofilm formation and fimbriae expression in Klebsiella pneumoniae MGH 78578. BMC Res. Notes 2014, 8, 7–13. [Google Scholar]
- Siebel, M.A.; Characklis, W.G. Observations of binary population biofilms. Biotechnol. Bioeng. 1991, 37, 778–789. [Google Scholar] [CrossRef] [PubMed]
- Skillman, L.C.; Sutherland, I.W.; Jones, M.V. The role of exopolysaccharides in dual species biofilm development. J. Appl. Microbiol. 1998, 85, 13S–18S. [Google Scholar] [CrossRef] [PubMed]
- Stickler, D.; Morris, N.; Moreno, M.C.; Sabbuba, N. Studies on the formation of crystalline bacterial biofilms on urethral catheters. Eur. J. Clin. Microbiol. Infect. Dis. 1998, 17, 649–652. [Google Scholar] [CrossRef] [PubMed]
- Macleod, S.M.; Stickler, D.J. Species interactions in mixed-community crystalline biofilms on urinary catheters. J. Med. Microbiol. 2007, 56, 1549–1557. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.W.; Periasamy, S.; Mukherjee, M.; Xie, C.; Kjelleberg, S.; Rice, S.A. Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J. 2014, 8, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.M.; Yu, J.L.; Liu, G.X.; Hu, L.Y.; Li, L.Q.; Li, F.; Yang, H. Electron microscopic analysis of biofilm on tracheal tubes removed from intubated neonates and the relationship between bilofilm and lower respiratory infection. Zhonghua Er Ke Za Zhi. 2007, 45, 655–660. [Google Scholar] [PubMed]
- Anderl, J.N.; Franklin, M.J.; Stewart, P.S. Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 2000, 44, 1818–1824. [Google Scholar] [CrossRef] [PubMed]
- Zahller, J.; Stewart, P.S. A transmission electron microscopic study of antibiotic action on Klebsiella pneumoniae biofilm. Antimicrob. Agents Chemother. 2002, 46, 2679–2683. [Google Scholar] [CrossRef] [PubMed]
- Anderl, J.N.; Zahller, J.; Roe, F.; Stewart, P.S. Role of nutrient limitation and stationary-phase existence in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 2003, 47, 1251–1256. [Google Scholar] [CrossRef] [PubMed]
- Cernohorská, L.; Votava, M. Determination of minimal regrowth concentration (MRC) in clinical isolates of various biofilm-forming bacteria. Folia Microbiol. (Praha) 2004, 49, 75–78. [Google Scholar]
- Liaqat, I.; Sumbal, F.; Sabri, A.N. Tetracycline and chloramphenicol efficiency against selected biofilm forming bacteria. Curr. Microbiol. 2009, 59, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, A.P.; Extremina, C.; Fonseca, A.F.; Sousa, J.C. Effect of subinhibitory concentration of piperacillin/tazobactam on Pseudomonas aeruginosa. J. Med. Microbiol. 2004, 53, 903–910. [Google Scholar] [CrossRef] [PubMed]
- Rachid, S.; Ohlsen, K.; Witte, W.; Hacker, J.; Ziebuhr, W. Effect of subinhibitory antibiotic concentrations on polysaccharide intercellular adhesion expression in biofilm forming Staphylococcus epidermidis. Antimicrob. Agents Chemother. 2000, 44, 3357–3363. [Google Scholar] [CrossRef] [PubMed]
- Singla, S.; Harjai, K.; Chhibber, S. Susceptibility of different phases of biofilm of Klebsiella pneumoniae to three different antibiotics. J. Antibiot. (Tokyo) 2013, 66, 61–66. [Google Scholar] [CrossRef]
- Bellifa, S.; Hassaine, H.; Balestrino, D.; Charbonnel, N.; M’hamedi, I.; Terki, I.K.; Lachachi, M.; Didi, W.; Forestier, C. Evaluation of biofilm formation of K. pneumoniae isolated from medical devices at the University Hospital of Tlemcen, Algeria. Afr. J. Microbiol. Res. 2013, 7, 5558–5564. [Google Scholar]
- Naparstek, L.; Carmeli, Y.; Navon-Venezia, S.; Banin, E. Biofilm formation and susceptibility to gentamicin and colistin of extremely drug-resistant KPC-producing Klebsiella pneumoniae. J. Antimicrob. Chemother. 2014, 69, 1027–1034. [Google Scholar] [CrossRef] [PubMed]
- Singla, S.; Harjai, K.; Chhibber, S. Artificial Klebsiella pneumoniae biofilm model mimicking in vivo system: Altered morphological characteristics and antibiotic resistance. J. Antibiot. (Tokyo) 2014, 67, 305–309. [Google Scholar] [CrossRef]
- Chen, P.; Seth, A.K.; Abercrombie, J.J.; Mustoe, T.A.; Leung, K.P. Activity of imipenem against Klebsiella pneumoniae biofilms in vitro and in vivo. Antimicrob. Agents Chemother. 2014, 58, 1208–1213. [Google Scholar] [CrossRef] [PubMed]
- Fuursted, K.; Schøler, L.; Hansen, F.; Dam, K.; Bojer, M.S.; Hammerum, A.M.; Dagnæs-Hansen, F.; Olsen, A.; Jasemian, Y.; Struve, C. Virulence of a Klebsiella pneumoniae strain carrying the New Delhi metallo-beta-lactamase-1 (NDM-1). Microbes Infect. 2012, 14, 155–158. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, P.; Shanmugam, N.; Sivaraman, U.; Kumar, S.; Selvaraj, S. Antiobiotic resistance pattern of biofilm-forming uropathogens isolated from catheterized patients in Pondicherry, India. Australas. Med. J. 2012, 5, 344–348. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, C.J., Jr.; Mende, K.; Beckius, M.L.; Akers, K.S.; Romano, D.R.; Wenke, J.C.; Murray, C.K. Biofilm formation by clinical isolates and the implications in chronic infections. BMC Infect. Dis. 2013, 29, 13–47. [Google Scholar]
- Hennequin, C.; Aumeran, C.; Robin, F.; Traore, O.; Forestier, C. Antibiotic resistance and plasmid transfer capacity in biofilm formed with a CTX-M-15-producing Klebsiella pneumoniae isolate. J. Antimicrob. Chemother. 2012, 67, 2123–2130. [Google Scholar] [CrossRef] [PubMed]
- Hennequin, C.; Robin, F.; Cabrolier, N.; Bonnet, R.; Forestier, C. Characterization of a DHA-1-producing Klebsiella pneumoniae strain involved in an outbreak and role of the AmpR regulator in virulence. Antimicrob. Agents Chemother. 2012, 56, 288–294. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Vuotto, C.; Longo, F.; Balice, M.P.; Donelli, G.; Varaldo, P.E. Antibiotic Resistance Related to Biofilm Formation in Klebsiella pneumoniae. Pathogens 2014, 3, 743-758. https://doi.org/10.3390/pathogens3030743
Vuotto C, Longo F, Balice MP, Donelli G, Varaldo PE. Antibiotic Resistance Related to Biofilm Formation in Klebsiella pneumoniae. Pathogens. 2014; 3(3):743-758. https://doi.org/10.3390/pathogens3030743
Chicago/Turabian StyleVuotto, Claudia, Francesca Longo, Maria Pia Balice, Gianfranco Donelli, and Pietro E. Varaldo. 2014. "Antibiotic Resistance Related to Biofilm Formation in Klebsiella pneumoniae" Pathogens 3, no. 3: 743-758. https://doi.org/10.3390/pathogens3030743
APA StyleVuotto, C., Longo, F., Balice, M. P., Donelli, G., & Varaldo, P. E. (2014). Antibiotic Resistance Related to Biofilm Formation in Klebsiella pneumoniae. Pathogens, 3(3), 743-758. https://doi.org/10.3390/pathogens3030743






