Antimicrobial Resistance in Veterinary Bacterial Pathogens: Resistance Patterns, Zoonotic Risks and One Health Implications
Abstract
1. Introduction
2. Materials and Methods
3. Antimicrobial Use and Emergence of Resistance in Veterinary Medicine
4. Antimicrobial Resistance in Major Veterinary Pathogens
4.1. Staphylococcus spp.
4.2. Pseudomonas aeruginosa
4.3. Enterobacterales
5. Multidrug Resistance in Veterinary Bacterial Isolates
6. Epidemiology and Zoonotic Transmission of AMR
6.1. Epidemiological Trends in Companion Animals and Livestock
6.2. Zoonotic Transmission of Antimicrobial-Resistant Bacteria
6.2.1. Transmission Pathways Between Animals, Humans, and the Environment
6.2.2. Companion Animals and Livestock as Reservoirs of Resistant Bacteria
7. One Health and Environmental Dimensions of AMR
7.1. One Health Framework for AMR Control
7.2. Environmental Reservoirs and Dissemination Pathways
8. Molecular Mechanisms of Antimicrobial Resistance in Veterinary Pathogens
9. Antimicrobial Stewardship and One Health-Based Control Strategies
10. Clinical Management and Alternative Strategies for MDR Infections
10.1. Clinical Challenges in Treating MDR Infections
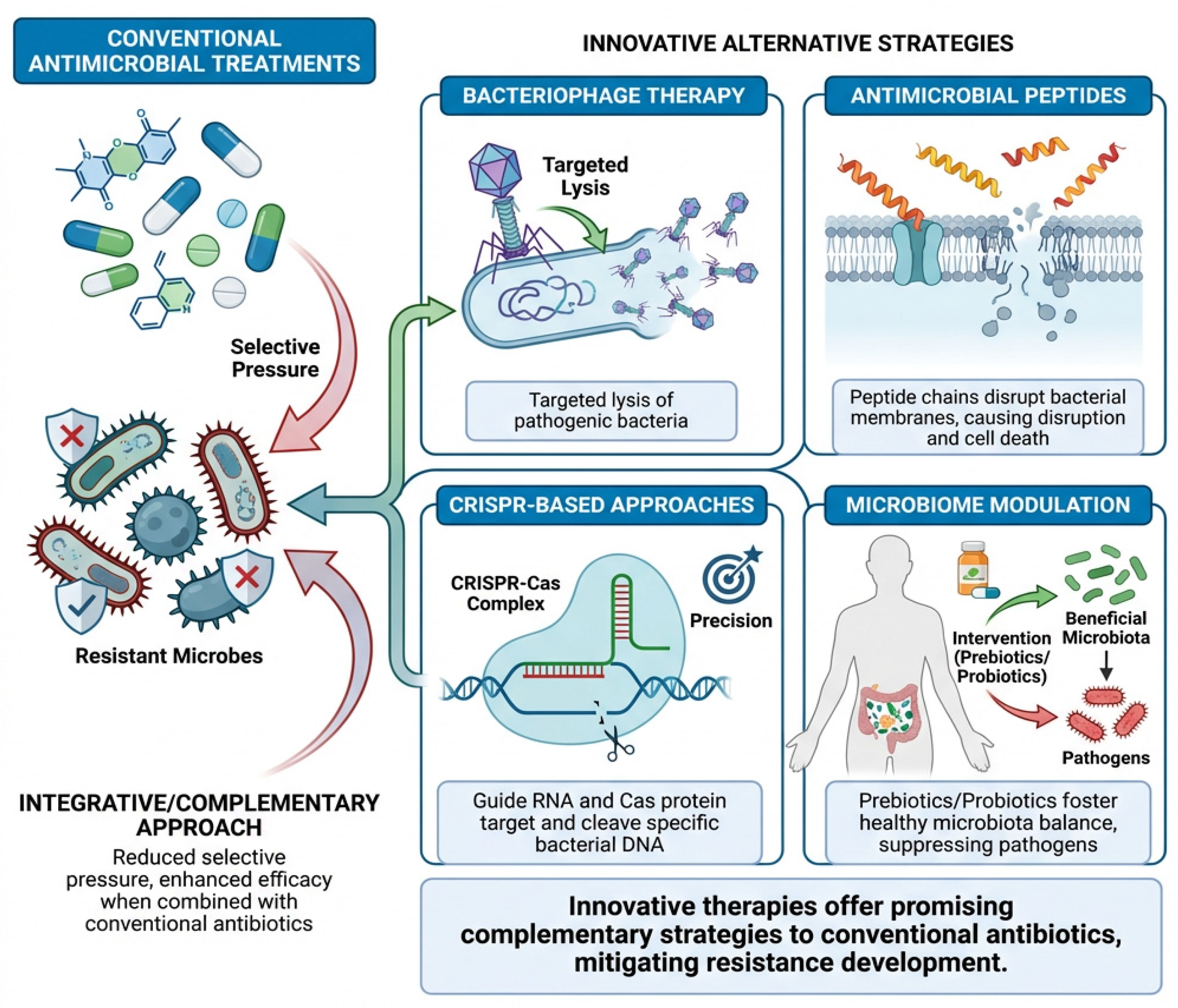
10.2. Alternative Strategies to Combat AMR
10.3. Diagnostic Innovations in Veterinary AMR
11. Global Policy Initiatives and Future Directions in Veterinary AMR
12. Economic Implications of Antimicrobial Resistance in Veterinary Medicine
13. Conclusions and Recommendations
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMR | Antimicrobial resistance |
| MDR | Multidrug-resistant |
| AST | Antimicrobial susceptibility testing |
| MRSA | Methicillin-resistant Staphylococcus aureus |
| ASP | Antimicrobial stewardship programs |
| AMU | Antimicrobial use |
| WHO | World Health Organization |
| WOAH | World Organisation for Animal Health |
| EFSA | European Food Safety Authority |
References
- Abushaheen, M.A.; Fatani, A.J.; Alosaimi, M.; Mansy, W.; George, M.; Acharya, S.; Rathod, S.; Divakar, D.D.; Jhugroo, C.; Vellappally, S.; et al. Antimicrobial resistance, mechanisms and its clinical significance. Disease-a-Month 2020, 66, 100971. [Google Scholar] [CrossRef] [PubMed]
- Best, C.M.; Bard, A.M.; Rees, G.M.; Reyher, K.K. Validation, visibility, vagueness and variation: A qualitative assessment of existing veterinary guidelines for antimicrobial use in cattle and sheep in the UK. PLoS ONE 2023, 18, e0294733. [Google Scholar] [CrossRef]
- Huemer, M.; Shambat, S.; Brugger, S.; Zinkernagel, A. Antibiotic resistance and persistence—Implications for human health and treatment perspectives. EMBO Rep. 2020, 21, e51034. [Google Scholar] [CrossRef]
- Salam, M.; Al-Amin, M.; Salam, M.; Pawar, J.; Akhter, N.; Rabaan, A.; Alqumber, M. Antimicrobial Resistance: A Growing Serious Threat for Global Public Health. Healthcare 2023, 11, 1946. [Google Scholar] [CrossRef]
- Galgano, M.; Pellegrini, F.; Catalano, E.; Capozzi, L.; Del Sambro, L.; Sposato, A.; Lucente, M.S.; Vasinioti, V.I.; Catella, C.; Odigie, A.E.; et al. Acquired Bacterial Resistance to Antibiotics and Resistance Genes: From Past to Future. Antibiotics 2025, 14, 222. [Google Scholar] [CrossRef]
- Uddin, T.; Chakraborty, A.; Khusro, A.; Zidan, B.; Mitra, S.; Emran, T.; Dhama, K.; Ripon, M.; Gajdács, M.; Sahibzada, M.; et al. Antibiotic resistance in microbes: History, mechanisms, therapeutic strategies and future prospects. J. Infect. Public Health 2021, 14, 1750–1766. [Google Scholar] [CrossRef]
- Oliveira, M.; Antunes, W.; Mota, S.; Madureira-Carvalho, Á.; Dinis-Oliveira, R.J.; Dias da Silva, D. An Overview of the Recent Advances in Antimicrobial Resistance. Microorganisms 2024, 12, 1920. [Google Scholar] [CrossRef] [PubMed]
- Popa, I.; Imre, K.; Morar, A.; Iancu, I.; Iorgoni, V.; Bochiș, T.; Pop, C.; Gligor, A.; Florea, T.; Popa, S.A.; et al. Questionnaire-Based Survey Regarding the Rational Usage of Antimicrobial Agents in Food-Producing Animals in Romania. Vet. Sci. 2025, 12, 408. [Google Scholar] [CrossRef]
- Mancuso, G.; Midiri, A.; Gerace, E.; Biondo, C. Bacterial Antibiotic Resistance: The Most Critical Pathogens. Pathogens 2021, 10, 1310. [Google Scholar] [CrossRef]
- Pham, Y.; Wozniak, T.M. Systems thinking to understand the complexity of antimicrobial resistance across One Health: A systematic review of current approaches. One Health 2025, 20, 101081. [Google Scholar] [CrossRef] [PubMed]
- Hoque, R.; Ahmed, S.; Naher, N.; Islam, M.; Rousham, E.; Islam, B.; Hassan, S. Tackling antimicrobial resistance in Bangladesh: A scoping review of policy and practice in human, animal and environment sectors. PLoS ONE 2020, 15, e0227947. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson-Palme, J.; Kristiansson, E.; Larsson, D.G.J. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol. Rev. 2018, 42, fux053. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Wang, B.; Zhang, X.; Zhang, J.; Zhang, H.; Liu, X.; Gao, Z.; Yu, Z. The spread of antibiotic resistance to humans and potential protection strategies. Ecotoxicol. Environ. Saf. 2023, 254, 114734. [Google Scholar] [CrossRef] [PubMed]
- Bassetti, S.; Tschudin-Sutter, S.; Egli, A.; Osthoff, M. Optimizing antibiotic therapies to reduce the risk of bacterial resistance. Eur. J. Intern. Med. 2022, 99, 7–12. [Google Scholar] [CrossRef]
- Lorusso, A.B.; Carrara, J.A.; Barroso, C.D.N.; Tuon, F.F.; Faoro, H. Role of Efflux Pumps on Antimicrobial Resistance in Pseudomonas aeruginosa. Int. J. Mol. Sci. 2022, 23, 15779. [Google Scholar] [CrossRef]
- Jin, M.; Osman, M.; Green, B.A.; Yang, Y.; Ahuja, A.; Lu, Z.; Cazer, C.L. Evidence for the transmission of antimicrobial resistant bacteria between humans and companion animals: A scoping review. One Health 2023, 17, 100593. [Google Scholar] [CrossRef]
- Horodyska, I.; Kasperska, P.; Michalski, K.; Bubak, J.; Herman, I.; Miszczak, M. Natural Microbiota of Dogs and Cats as a Source and Vector of Resistance Genes-Clinical Significance. Int. J. Mol. Sci. 2025, 26, 7717. [Google Scholar] [CrossRef]
- Endale, H.; Mathewos, M.; Abdeta, D. Potential Causes of Spread of Antimicrobial Resistance and Preventive Measures in One Health Perspective-A Review. Infect. Drug Resist. 2023, 16, 7515–7545. [Google Scholar] [CrossRef]
- Hernando-Amado, S.; Coque, T.M.; Baquero, F.; Martínez, J.L. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat. Microbiol. 2019, 4, 1432–1442. [Google Scholar] [CrossRef]
- Kaspar, U.; von Lützau, A.; Schlattmann, A.; Roesler, U.; Köck, R.; Becker, K. Zoonotic multidrug-resistant microorganisms among small companion animals in Germany. PLoS ONE 2018, 13, e0208364. [Google Scholar] [CrossRef]
- Palma, E.; Tilocca, B.; Roncada, P. Antimicrobial Resistance in Veterinary Medicine: An Overview. Int. J. Mol. Sci. 2020, 21, 1914. [Google Scholar] [CrossRef]
- Lenart-Boroń, A.; Stankiewicz, K.; Czernecka, N.; Ratajewicz, A.; Bulanda, K.; Heliasz, M.; Sosińska, D.; Dworak, K.; Ciesielska, D.; Siemińska, I.; et al. Wounds of Companion Animals as a Habitat of Antibiotic-Resistant Bacteria That Are Potentially Harmful to Humans-Phenotypic, Proteomic and Molecular Detection. Int. J. Mol. Sci. 2024, 25, 3121. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, H.I.G.; Silva, V.; de Sousa, T.; Calouro, R.; Saraiva, S.; Igrejas, G.; Poeta, P. Antimicrobial Resistance in European Companion Animals Practice: A One Health Approach. Animals 2025, 15, 1708. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Gascón, A.; Solinís, M.Á.; Isla, A. The Role of PK/PD Analysis in the Development and Evaluation of Antimicrobials. Pharmaceutics 2021, 13, 833. [Google Scholar] [CrossRef]
- Haulisah, N.A.; Hassan, L.; Jajere, S.M.; Ahmad, N.I.; Bejo, S.K. High prevalence of antimicrobial resistance and multidrug resistance among bacterial isolates from diseased pets: Retrospective laboratory data (2015–2017). PLoS ONE 2022, 17, e0277664. [Google Scholar] [CrossRef] [PubMed]
- Larsson, D.G.J.; Flach, C.F. Antibiotic resistance in the environment. Nat. Rev. Microbiol. 2022, 20, 257–269. [Google Scholar] [CrossRef]
- Moran, D. A framework for improved one health governance and policy making for antimicrobial use. BMJ Glob. Health 2019, 4, e001807. [Google Scholar] [CrossRef]
- Berendonk, T.U.; Manaia, C.M.; Merlin, C.; Fatta-Kassinos, D.; Cytryn, E.; Walsh, F.; Bürgmann, H.; Sørum, H.; Norström, M.; Pons, M.N.; et al. Tackling antibiotic resistance: The environmental framework. Nat. Rev. Microbiol. 2015, 13, 310–317. [Google Scholar] [CrossRef]
- Ardakani, Z.; Canali, M.; Aragrande, M.; Tomassone, L.; Simoes, M.; Balzani, A.; Beber, C.L. Evaluating the contribution of antimicrobial use in farmed animals to global antimicrobial resistance in humans. One Health 2023, 17, 100647. [Google Scholar] [CrossRef]
- Iancu, I.; Popa, S.A.; Degi, J.; Gligor, A.; Popa, I.; Iorgoni, V.; Nistor, P.; Imre, K.; Nichita, I.; Herman, V. Aerobic Uterine Pathogens in Dairy Cattle: Surveillance and Antimicrobial Resistance Profiles in Postpartum Endometritis. Antibiotics 2025, 14, 650. [Google Scholar] [CrossRef]
- Capita, R.; Alonso-Calleja, C. Antibiotic-resistant bacteria: A challenge for the food industry. Crit. Rev. Food Sci. Nutr. 2013, 53, 11–48. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.K.C.; Ng, R.W.Y.; Leung, S.S.Y.; Hui, M.; Ip, M. Overcoming the rising incidence and evolving mechanisms of antibiotic resistance by novel drug delivery approaches—An overview. Adv. Drug Deliv. Rev. 2022, 181, 114078. [Google Scholar] [CrossRef]
- Wistrand-Yuen, E.; Knopp, M.; Hjort, K.; Koskiniemi, S.; Berg, O.G.; Andersson, D.I. Evolution of high-level resistance during low-level antibiotic exposure. Nat. Commun. 2018, 9, 1599. [Google Scholar] [CrossRef] [PubMed]
- Tamma, P.D.; Avdic, E.; Li, D.X.; Dzintars, K.; Cosgrove, S.E. Association of Adverse Events With Antibiotic Use in Hospitalized Patients. JAMA Intern. Med. 2017, 177, 1308–1315. [Google Scholar] [CrossRef]
- Kollef, M.H. Broad-spectrum antimicrobials and the treatment of serious bacterial infections: Getting it right up front. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2008, 47, S3–S13. [Google Scholar] [CrossRef] [PubMed]
- Lund, D.; Parras-Moltó, M.; Inda-Díaz, J.S.; Ebmeyer, S.; Larsson, D.G.J.; Johnning, A.; Kristiansson, E. Genetic compatibility and ecological connectivity drive the dissemination of antibiotic resistance genes. Nat. Commun. 2025, 16, 2595. [Google Scholar] [CrossRef]
- Spencer, D.C.; Paton, T.F.; Mulroney, K.T.; Inglis, T.J.J.; Sutton, J.M.; Morgan, H. A fast impedance-based antimicrobial susceptibility test. Nat. Commun. 2020, 11, 5328. [Google Scholar] [CrossRef]
- Ahmed, M.J.; Hossain, M.I.; Hossain Bhuiyan, M.I.; Rahman, M.A.; Kaderi, I.J.; Nirob, A.I.; Rahi, M.A.I.; Mamun, M.; Alam, K.E.; Chowdhury, M.J.; et al. Antimicrobial resistance in meat and meat products from Asia: An urgent one health challenge—A systematic review and meta-analysis. Poult. Sci. 2025, 104, 105811. [Google Scholar] [CrossRef]
- Cooke, E.; Rendi-Wagner, P. Rising antimicrobial resistance in Europe: EMA and ECDC outline urgent actions to achieve EU targets. Lancet Reg. Health—Eur. 2025, 59, 101539. [Google Scholar] [CrossRef]
- Pomba, C.; Rantala, M.; Greko, C.; Baptiste, K.E.; Catry, B.; van Duijkeren, E.; Mateus, A.; Moreno, M.A.; Pyörälä, S.; Ružauskas, M.; et al. Public health risk of antimicrobial resistance transfer from companion animals. J. Antimicrob. Chemother. 2017, 72, 957–968. [Google Scholar] [CrossRef]
- Scarpellini, R.; Assirelli, G.; Giunti, M.; Esposito, E.; Mondo, E.; Piva, S. Monitoring the Prevalence of Antimicrobial Resistance in Companion Animals: Results from Clinical Isolates in an Italian University Veterinary Hospital. Transbound. Emerg. Dis. 2023, 2023, 6695493. [Google Scholar] [CrossRef]
- Elbehiry, A.; Marzouk, E. From Farm to Fork: Antimicrobial-Resistant Bacterial Pathogens in Livestock Production and the Food Chain. Vet. Sci. 2025, 12, 862. [Google Scholar] [CrossRef]
- Karaman, R.; Jubeh, B.; Breijyeh, Z. Resistance of Gram-Positive Bacteria to Current Antibacterial Agents and Overcoming Approaches. Molecules 2020, 25, 2888. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.W.K.; Millar, B.C.; Moore, J.E. Antimicrobial Resistance (AMR). Br. J. Biomed. Sci. 2023, 80, 11387. [Google Scholar] [CrossRef]
- Wang, N.; Guo, X.; Yan, Z.; Wang, W.; Chen, B.; Ge, F.; Ye, B. A Comprehensive Analysis on Spread and Distribution Characteristic of Antibiotic Resistance Genes in Livestock Farms of Southeastern China. PLoS ONE 2016, 11, e0156889. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.P.; May, H.E.; AbuOun, M.; Stubberfield, E.; Gilson, D.; Chau, K.K.; Crook, D.W.; Shaw, L.P.; Read, D.S.; Stoesser, N.; et al. A longitudinal study reveals persistence of antimicrobial resistance on livestock farms is not due to antimicrobial usage alone. Front. Microbiol. 2023, 14, 1070340. [Google Scholar] [CrossRef] [PubMed]
- Pandey, S.; Doo, H.; Keum, G.B.; Kim, E.S.; Kwak, J.; Ryu, S.; Choi, Y.; Kang, J.; Kim, S.; Lee, N.R.; et al. Antibiotic resistance in livestock, environment and humans: One Health perspective. J. Anim. Sci. Technol. 2024, 66, 266–278. [Google Scholar] [CrossRef]
- Marco-Fuertes, A.; Marin, C.; Gimeno-Cardona, C.; Artal-Muñoz, V.; Vega, S.; Montoro-Dasi, L. Multidrug-Resistant Commensal and Infection-Causing Staphylococcus spp. Isolated from Companion Animals in the Valencia Region. Vet. Sci. 2024, 11, 54. [Google Scholar] [CrossRef]
- Alauzet, C.; Lozniewski, A.; Marchandin, H. Metronidazole resistance and nim genes in anaerobes: A review. Anaerobe 2019, 55, 40–53. [Google Scholar] [CrossRef]
- van Duijkeren, E.; Greko, C.; Pringle, M.; Baptiste, K.E.; Catry, B.; Jukes, H.; Moreno, M.A.; Pomba, M.C.; Pyörälä, S.; Rantala, M.; et al. Pleuromutilins: Use in food-producing animals in the European Union, development of resistance and impact on human and animal health. J. Antimicrob. Chemother. 2014, 69, 2022–2031. [Google Scholar] [CrossRef]
- Miklasińska-Majdanik, M. Mechanisms of Resistance to Macrolide Antibiotics among Staphylococcus aureus. Antibiotics 2021, 10, 1406. [Google Scholar] [CrossRef] [PubMed]
- Butaye, P.; Devriese, L.A.; Haesebrouck, F. Antimicrobial growth promoters used in animal feed: Effects of less well known antibiotics on gram-positive bacteria. Clin. Microbiol. Rev. 2003, 16, 175–188. [Google Scholar] [CrossRef]
- Safdar, N.; Saleem, S.; Salman, M.; Tareq, A.H.; Ishaq, S.; Ambreen, S.; Hameed, A.; Habib, M.B.; Ali, T.M. Economic burden of antimicrobial resistance on patients in Pakistan. Front. Public Health 2025, 13, 1481212. [Google Scholar] [CrossRef] [PubMed]
- Robb, A.R.; Ure, R.; Chaput, D.L.; Foster, G. Emergence of novel methicillin resistant Staphylococcus pseudintermedius lineages revealed by whole genome sequencing of isolates from companion animals and humans in Scotland. PLoS ONE 2024, 19, e0305211. [Google Scholar] [CrossRef] [PubMed]
- Srednik, M.E.; Perea, C.A.; Giacoboni, G.I.; Hicks, J.A.; Foxx, C.L.; Harris, B.; Schlater, L.K. Genomic Features of Antimicrobial Resistance in Staphylococcus pseudintermedius Isolated from Dogs with Pyoderma in Argentina and the United States: A Comparative Study. Int. J. Mol. Sci. 2023, 24, 11361. [Google Scholar] [CrossRef]
- Iorgoni, V.; Iancu, I.; Popa, I.; Gligor, A.; Orghici, G.; Sicoe, B.; Badea, C.; Dreghiciu, C.; Luca, I.; Nistor, P.; et al. The First Report of a Pulmonary Abscess Due to Streptococcus intermedius in Rabbits in Romania. Microorganisms 2025, 13, 769. [Google Scholar] [CrossRef]
- Maree, M.; Thi Nguyen, L.T.; Ohniwa, R.L.; Higashide, M.; Msadek, T.; Morikawa, K. Natural transformation allows transfer of SCCmec-mediated methicillin resistance in Staphylococcus aureus biofilms. Nat. Commun. 2022, 13, 2477. [Google Scholar] [CrossRef]
- Popa, I.; Iancu, I.; Iorgoni, V.; Gligor, A.; Imre, K.; Tîrziu, E.; Bochiș, T.; Pop, C.; Degi, J.; Ivan, A.; et al. Antimicrobial Susceptibility and Fluoroquinolone Resistance Patterns of Pseudomonas aeruginosa Isolated from Canine Otitis Externa in Romania. Antibiotics 2026, 15, 144. [Google Scholar] [CrossRef]
- Popa, I.; Iancu, I.; Gligor, A.; Imre, K.; Tîrziu, E.; Bochiș, T.; Pop, C.; Degi, J.; Ivan, A.; Dahma, M.; et al. Prevalence and Antimicrobial Resistance Profiles of E. coli, P. mirabilis, and E. cloacae Complex Isolated from Dogs with Otitis Externa. Antibiotics 2026, 15, 343. [Google Scholar] [CrossRef]
- Guimarães, L.; Teixeira, I.M.; da Silva, I.T.; Antunes, M.; Pesset, C.; Fonseca, C.; Santos, A.L.; Côrtes, M.F.; Penna, B. Epidemiologic case investigation on the zoonotic transmission of Methicillin-resistant Staphylococcus pseudintermedius among dogs and their owners. J. Infect. Public Health 2023, 16, 183–189. [Google Scholar] [CrossRef]
- Wood, S.J.; Kuzel, T.M.; Shafikhani, S.H. Pseudomonas aeruginosa: Infections, Animal Modeling, and Therapeutics. Cells 2023, 12, 199. [Google Scholar] [CrossRef]
- Jousserand, N.; Auvray, F.; Chagneau, C.; Cavalié, L.; Maurey, C.; Drut, A.; Lavoué, R.; Oswald, E. Zoonotic potential of uropathogenic Escherichia coli lineages from companion animals. Vet. Res. 2025, 56, 69. [Google Scholar] [CrossRef] [PubMed]
- Bonnevie, A.; Myrenås, M.; Nilsson, O. ESBL- and pAmpC-producing Enterobacterales from Swedish dogs and cats 2017–2021: A retrospective study. Acta Vet. Scand. 2025, 67, 2. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Dong, Z.; Ai, S.; Chen, S.; Dong, M.; Li, Q.; Zhou, Z.; Liu, H.; Zhong, Z.; Ma, X.; et al. Virulence-related factors and antimicrobial resistance in Proteus mirabilis isolated from domestic and stray dogs. Front. Microbiol. 2023, 14, 1141418. [Google Scholar] [CrossRef]
- Harada, K.; Shimizu, T.; Mukai, Y.; Kuwajima, K.; Sato, T.; Kajino, A.; Usui, M.; Tamura, Y.; Kimura, Y.; Miyamoto, T.; et al. Phenotypic and molecular characterization of antimicrobial resistance in Enterobacter spp. isolates from companion animals in Japan. PLoS ONE 2017, 12, e0174178. [Google Scholar] [CrossRef] [PubMed]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Subramanya, S.H.; Bairy, I.; Metok, Y.; Baral, B.P.; Gautam, D.; Nayak, N. Detection and characterization of ESBL-producing Enterobacteriaceae from the gut of subsistence farmers, their livestock, and the surrounding environment in rural Nepal. Sci. Rep. 2021, 11, 2091. [Google Scholar] [CrossRef]
- Tirziu, E.; Lazăr, R.; Sala, C.; Nichita, I.; Morar, A.; Şereş, M.; Imre, K. Salmonella in raw chicken meat from the Romanian seaside: Frequency of isolation and antibiotic resistance. J. Food Prot. 2015, 78, 1003–1006. [Google Scholar] [CrossRef]
- Swarthout, J.M.; Chan, E.M.G.; Garcia, D.; Nadimpalli, M.L.; Pickering, A.J. Human Colonization with Antibiotic-Resistant Bacteria from Nonoccupational Exposure to Domesticated Animals in Low- and Middle-Income Countries: A Critical Review. Environ. Sci. Technol. 2022, 56, 14875–14890. [Google Scholar] [CrossRef]
- Checcucci, A.; Buscaroli, E.; Modesto, M.; Luise, D.; Blasioli, S.; Scarafile, D.; Di Vito, M.; Bugli, F.; Trevisi, P.; Braschi, I.; et al. The swine waste resistome: Spreading and transfer of antibiotic resistance genes in Escherichia coli strains and the associated microbial communities. Ecotoxicol. Environ. Saf. 2024, 283, 116774. [Google Scholar] [CrossRef]
- Kaspar, U.; von Lützau, K.; Schlattmann, A.; Rösler, U.; Köck, R.; Becker, K. Zoonotic multidrug-resistant microorganisms among non-hospitalized horses from Germany. One Health 2019, 7, 100091. [Google Scholar] [CrossRef] [PubMed]
- Walther, B.; Tedin, K.; Lübke-Becker, A. Multidrug-resistant opportunistic pathogens challenging veterinary infection control. Vet. Microbiol. 2017, 200, 71–78. [Google Scholar] [CrossRef]
- Popa, S.A.; Morar, A.; Ban-Cucerzan, A.; Imre, K. Last decade mini-review of the scientific progresses in the monitoring of the occurrence and antimicrobial susceptibility profile of poultry origin Campylobacter spp. within the European Union countries. Rev. Rom. Med. Vet. 2022, 32, 75–82. [Google Scholar]
- Popa, S.A.; Morar, A.; Ban-Cucerzan, A.; Tîrziu, E.; Herman, V.; Imre, M.; Florea, T.; Morar, D.; Pătrânjan, R.T.; Imre, K. First study on the frequency of isolation and phenotypic antimicrobial resistance profiles of pig- and cattle-origin Campylobacter strains in Romania. Vet. Res. Commun. 2024, 48, 2621–2627. [Google Scholar] [CrossRef] [PubMed]
- Popa, S.A.; Herman, V.; Tîrziu, E.; Morar, A.; Ban-Cucerzan, A.; Imre, M.; Pătrînjan, R.-T.; Imre, K. Public Health Risk of Campylobacter spp. Isolated from Slaughterhouse and Retail Poultry Meat: Prevalence and Antimicrobial Resistance Profiles. Pathogens 2025, 14, 316. [Google Scholar] [CrossRef]
- Zhang, T.; Nickerson, R.; Zhang, W.; Peng, X.; Shang, Y.; Zhou, Y.; Luo, Q.; Wen, G.; Cheng, Z. The impacts of animal agriculture on One Health-Bacterial zoonosis, antimicrobial resistance, and beyond. One Health 2024, 18, 100748. [Google Scholar] [CrossRef]
- De Giusti, M.; Barbato, D.; Lia, L.; Colamesta, V.; Lombardi, A.; Cacchio, D.; Villari, P.; La Torre, G. Collaboration between human and veterinary medicine as a tool to solve public health problems. Lancet Planet. Health 2019, 3, e64–e65. [Google Scholar] [CrossRef]
- Joosten, P.; Ceccarelli, D.; Odent, E.; Sarrazin, S.; Graveland, H.; Van Gompel, L.; Battisti, A.; Caprioli, A.; Franco, A.; Wagenaar, J.A.; et al. Antimicrobial Usage and Resistance in Companion Animals: A Cross-Sectional Study in Three European Countries. Antibiotics 2020, 9, 87. [Google Scholar] [CrossRef]
- Grönthal, T.; Ollilainen, M.; Eklund, M.; Piiparinen, H.; Gindonis, V.; Junnila, J.; Saijonmaa-Koulumies, L.; Liimatainen, R.; Rantala, M. Epidemiology of methicillin resistant Staphylococcus pseudintermedius in guide dogs in Finland. Acta Vet. Scand. 2015, 57, 37. [Google Scholar] [CrossRef]
- van Duijkeren, E.; Kamphuis, M.; van der Mije, I.C.; Laarhoven, L.M.; Duim, B.; Wagenaar, J.A.; Houwers, D.J. Transmission of methicillin-resistant Staphylococcus pseudintermedius between infected dogs and cats and contact pets, humans and the environment in households and veterinary clinics. Vet. Microbiol. 2011, 150, 338–343. [Google Scholar] [CrossRef]
- Salgado-Caxito, M.; Benavides, J.A.; Adell, A.D.; Paes, A.C.; Moreno-Switt, A.I. Global prevalence and molecular characterization of extended-spectrum β-lactamase producing-Escherichia coli in dogs and cats—A scoping review and meta-analysis. One 2021, 12, 100236. [Google Scholar] [CrossRef]
- Dos SRibeiro, C.; van de Burgwal, L.H.M.; Regeer, B.J. Overcoming challenges for designing and implementing the One Health approach: A systematic review of the literature. One Health 2019, 7, 100085. [Google Scholar] [CrossRef]
- Pepin, K.; Carlisle, K.; Anderson, D.; Baker, M.; Chipman, R.; Benschop, J.; French, N.; Greenhalgh, S.; McDougall, S.; Muellner, P.; et al. Steps towards operationalizing One Health approaches. One Health 2024, 18, 100740. [Google Scholar] [CrossRef]
- One Health High-Level Expert Panel (OHHLEP); Adisasmito, W.B.; Almuhairi, S.; Behravesh, C.B.; Bilivogui, P.; Bukachi, S.A.; Casas, N.; Cediel Becerra, N.; Charron, D.F.; Chaudhary, A.; et al. One Health: A new definition for a sustainable and healthy future. PLoS Pathog. 2022, 18, e1010537. [Google Scholar] [CrossRef]
- Milazzo, A.; Liu, J.; Multani, P.; Steele, S.; Hoon, E.; Chaber, A. One Health implementation: A systematic scoping review using the Quadripartite One Health Joint Plan of Action. One Health 2025, 20, 101008. [Google Scholar] [CrossRef] [PubMed]
- Vikesland, P.; Garner, E.; Gupta, S.; Kang, S.; Maile-Moskowitz, A.; Zhu, N. Differential Drivers of Antimicrobial Resistance across the World. Acc. Chem. Res. 2019, 52, 916–924. [Google Scholar] [CrossRef] [PubMed]
- Gillieatt, B.F.; Coleman, N.V. Unravelling the mechanisms of antibiotic and heavy metal resistance co-selection in environmental bacteria. FEMS Microbiol. Rev. 2024, 48, fuae017. [Google Scholar] [CrossRef]
- Wyres, K.; Holt, K. Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr. Opin. Microbiol. 2018, 45, 131–139. [Google Scholar] [CrossRef]
- Marathe, N.P.; Regina, V.R.; Walujkar, S.A.; Charan, S.S.; Moore, E.R.; Larsson, D.G.; Shouche, Y.S. A treatment plant receiving waste water from multiple bulk drug manufacturers is a reservoir for highly multi-drug resistant integron-bearing bacteria. PLoS ONE 2013, 8, e77310. [Google Scholar] [CrossRef]
- Rolo, J.; Worning, P.; Boye Nielsen, J.; Sobral, R.; Bowden, R.; Bouchami, O.; Damborg, P.; Guardabassi, L.; Perreten, V.; Westh, H.; et al. Evidence for the evolutionary steps leading to mecA-mediated β-lactam resistance in staphylococci. PLoS Genet. 2017, 13, e1006674. [Google Scholar] [CrossRef]
- Aedo, S.; Tomasz, A. Role of the Stringent Stress Response in the Antibiotic Resistance Phenotype of Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2016, 60, 2311–2317. [Google Scholar] [CrossRef] [PubMed]
- Bilyk, B.L.; Panchal, V.V.; Tinajero-Trejo, M.; Hobbs, J.K.; Foster, S.J. An Interplay of Multiple Positive and Negative Factors Governs Methicillin Resistance in Staphylococcus aureus. Microbiol. Mol. Biol. Rev. 2022, 86, e0015921. [Google Scholar] [CrossRef]
- Moore, N.M.; Flaws, M.L. Antimicrobial resistance mechanisms in Pseudomonas aeruginosa. Clin. Lab. Sci. 2011, 24, 47–51. [Google Scholar] [CrossRef] [PubMed]
- Lambert, P. Mechanisms of antibiotic resistance in Pseudomonas aeruginosa. J. R. Soc. Med. 2002, 95, 22. [Google Scholar] [PubMed]
- Dyar, O.J.; Huttner, B.; Schouten, J.; Pulcini, C.; ESGAP (ESCMID Study Group for Antimicrobial stewardshiP). What is antimicrobial stewardship? Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2017, 23, 793–798. [Google Scholar] [CrossRef]
- Mendelson, M.; Morris, A.M.; Thursky, K.; Pulcini, C. How to start an antimicrobial stewardship programme in a hospital. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2020, 26, 447–453. [Google Scholar] [CrossRef]
- Gilham, E.L.; Pearce-Smith, N.; Carter, V.; Ashiru-Oredope, D. Assessment of global antimicrobial resistance campaigns conducted to improve public awareness and antimicrobial use behaviours: A rapid systematic review. BMC Public Health 2024, 24, 396. [Google Scholar] [CrossRef]
- Castro-Sánchez, E.; Drumright, L.N.; Gharbi, M.; Farrell, S.; Holmes, A.H. Mapping Antimicrobial Stewardship in Undergraduate Medical, Dental, Pharmacy, Nursing and Veterinary Education in the United Kingdom. PLoS ONE 2016, 11, e0150056. [Google Scholar] [CrossRef]
- de la Fuente-Nunez, C.; Cesaro, A.; Hancock, R.E.W. Antibiotic failure: Beyond antimicrobial resistance. Drug Resist. Updates Rev. Comment. Antimicrob. Anticancer Chemother. 2023, 71, 101012. [Google Scholar] [CrossRef]
- Ban-Cucerzan, A.; Imre, K.; Morar, A.; Marcu, A.; Hotea, I.; Popa, S.-A.; Pătrînjan, R.-T.; Bucur, I.-M.; Gașpar, C.; Plotuna, A.-M.; et al. Persistent Threats: A Comprehensive Review of Biofilm Formation, Control, and Economic Implications in Food Processing Environments. Microorganisms 2025, 13, 1805. [Google Scholar] [CrossRef]
- Mafe, A.; Büsselberg, D. Phage Therapy in Managing Multidrug-Resistant (MDR) Infections in Cancer Therapy: Innovations, Complications, and Future Directions. Pharmaceutics 2025, 17, 820. [Google Scholar] [CrossRef]
- Pallavali, R.R.; Degati, V.L.; Lomada, D.; Reddy, M.C.; Durbaka, V.R.P. Isolation and in vitro evaluation of bacteriophages against MDR-bacterial isolates from septic wound infections. PLoS ONE 2017, 12, e0179245. [Google Scholar] [CrossRef]
- van der Meer, J.W.M.; Vandenbroucke-Grauls, C.M.J.E. Bacteriofagen in de strijd tegen multiresistente bacteriën [Bacteriophages in the battle against multidrug resistant bacteria]. Ned. Tijdschr. Voor Geneeskd. 2018, 162, D2433. [Google Scholar]
- Herman, V.; Roșiu, D.; Cătana, N.; Degi, J.; Iancu, I.; Mițiți, I.; Ciobanu, G.; Grema, C.F.; Pascu, C. Evaluation of propolis for antibacterial activity in vitro. Rev. Romana Med. Vet. 2018, 28, 13–17. [Google Scholar]
- Theuretzbacher, U.; Piddock, L.J.V. Non-traditional Antibacterial Therapeutic Options and Challenges. Cell Host Microbe 2019, 26, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, C.; Sarkar, P.; Issa, R.; Haldar, J. Alternatives to Conventional Antibiotics in the Era of Antimicrobial Resistance. Trends Microbiol. 2019, 27, 323–338. [Google Scholar] [CrossRef] [PubMed]
- MacNair, C.; Rutherford, S.; Tan, M. Alternative therapeutic strategies to treat antibiotic-resistant pathogens. Nat. Rev. Microbiol. 2023, 22, 262–275. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.; Draz, M.; Gu, M.; Yu, W.; Ruan, Z.; Luo, Q. Novel Strategy to Combat Antibiotic Resistance: A Sight into the Combination of CRISPR/Cas9 and Nanoparticles. Pharmaceutics 2021, 13, 352. [Google Scholar] [CrossRef]
- Yoon, B.; Kim, J. CRISPR–Cas-Mediated Reprogramming Strategies to Overcome Antimicrobial Resistance. Pharmaceutics 2026, 18, 95. [Google Scholar] [CrossRef]
- Alidriss, O.M.; AlSudais, H.; Alhumaidan, O.S.; Altwaijry, H.D.; Bakhsh, A.; Almuhanna, Y.; Alkudmani, Z.S.; Alqarni, I.A.; Alenazi, D.; Aljasham, A.T.; et al. Targeted Drug Delivery Strategies in Overcoming Antimicrobial Resistance: Advances and Future Directions. Pharmaceutics 2025, 17, 1426. [Google Scholar] [CrossRef]
- Citorik, R.; Mimee, M.; Lu, T. Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases. Nat. Biotechnol. 2014, 32, 1141–1145. [Google Scholar] [CrossRef] [PubMed]
- Makabenta, J.; Nabawy, A.; Li, C.; Schmidt-Malan, S.; Patel, R.; Rotello, V. Nanomaterial-based therapeutics for antibiotic-resistant bacterial infections. Nat. Rev. Microbiol. 2020, 19, 23–36. [Google Scholar] [CrossRef] [PubMed]
- Khaledi, A.; Weimann, A.; Schniederjans, M.; Asgari, E.; Kuo, T.; Oliver, A.; Cabot, G.; Kola, A.; Gastmeier, P.; Hogardt, M.; et al. Predicting antimicrobial resistance in Pseudomonas aeruginosa with machine learning-enabled molecular diagnostics. EMBO Mol. Med. 2020, 12, e10264. [Google Scholar] [CrossRef]
- Sherry, N.; Horan, K.; Ballard, S.; Da Silva, A.; Gorrie, C.; Schultz, M.; Stevens, K.; Valcanis, M.; Sait, M.; Stinear, T.; et al. An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance. Nat. Commun. 2023, 14, 60. [Google Scholar] [CrossRef]
- Chevalier, J.; Hansen, M.; Blumel, B.; Chuchu, V.; Gleeson, B.; Ferreyra, C.; Khan, S.; Nichols, B.; Grantz, K. Modeling the impact and cost of a culture-dependent molecular test for antimicrobial resistance in resource-limited settings. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2025, 82, 25–32. [Google Scholar] [CrossRef] [PubMed]
- James, R.; Hardefeldt, L.; Ierano, C.; Charani, E.; Dowson, L.; Elkins, S.; Thursky, K. Antimicrobial stewardship from a One Health perspective. Nat. Rev. Microbiol. 2025, 24, 146–162. [Google Scholar] [CrossRef]
- Bertagnolio, S.; Dobreva, Z.; Centner, C.M.; Olaru, I.D.; Donà, D.; Burzo, S.; Huttner, B.D.; Chaillon, A.; Gebreselassie, N.; Wi, T.; et al. WHO global research priorities for antimicrobial resistance in human health. Lancet Microbe 2024, 5, 100902. [Google Scholar] [CrossRef]
- Lesho, E.P.; Laguio-Vila, M. The Slow-Motion Catastrophe of Antimicrobial Resistance and Practical Interventions for All Prescribers. Mayo Clin. Proc. 2019, 94, 1040–1047. [Google Scholar] [CrossRef]
- Founou, R.C.; Founou, L.L.; Essack, S.Y. Clinical and economic impact of antibiotic resistance in developing countries: A systematic review and meta-analysis. PLoS ONE 2017, 12, e0189621. [Google Scholar] [CrossRef]
- Samtiya, M.; Matthews, K.R.; Dhewa, T.; Puniya, A.K. Antimicrobial Resistance in the Food Chain: Trends, Mechanisms, Pathways, and Possible Regulation Strategies. Foods 2022, 11, 2966. [Google Scholar] [CrossRef]
- Innes, G.K.; Randad, P.R.; Korinek, A.; Davis, M.F.; Price, L.B.; So, A.D.; Heaney, C.D. External Societal Costs of Antimicrobial Resistance in Humans Attributable to Antimicrobial Use in Livestock. Annu. Rev. Public Health 2020, 41, 141–157. [Google Scholar] [CrossRef]
- Acosta, A.; Roland-Holst, D.; Nicolli, F.; Tirkaso, W.; Rocha, J.S.; Song, J. A One Health framework to assess the economic returns on investment in livestock antimicrobial stewardship. One Health 2025, 21, 101188. [Google Scholar] [CrossRef]





| Antimicrobial Class | Examples Commonly Used in Veterinary Medicine | Major Bacterial Targets | Frequently Reported Resistance Trends |
|---|---|---|---|
| β-lactams | Amoxicillin, ampicillin, cephalexin, ceftiofur | Staphylococcus spp., E. coli, Proteus spp. | Increasing resistance reported in multiple regions including Europe and Asia, particularly in livestock-associated isolates [38,39] |
| Fluoroquinolones | Enrofloxacin, marbofloxacin, ciprofloxacin | Pseudomonas aeruginosa, E. coli, Enterobacter spp. | Rising resistance in Gram-negative bacteria across companion and food animals in regions with high antimicrobial usage [40] |
| Aminoglycosides | Gentamicin, amikacin, neomycin | Pseudomonas aeruginosa, E. coli, Staphylococcus spp. | Moderate but increasing resistance in hospital and veterinary clinical isolates [41] |
| Tetracyclines | Tetracycline, doxycycline, oxytetracycline | E. coli, Staphylococcus spp. | High resistance prevalence globally, especially in intensive livestock production systems [42] |
| Macrolides | Erythromycin, tylosin, tilmicosin | Staphylococcus spp., respiratory pathogens | Increasing resistance in Gram-positive pathogens, particularly in Europe and North America [43] |
| sulfonamide-diaminopyrimidine combinations | Sulfamethoxazole-trimethoprim | E. coli, Proteus mirabilis | Widespread resistance reported globally with regional variability depending on antimicrobial usage practices [44] |
| Phenicols | Florfenicol, chloramphenicol | Respiratory and systemic pathogens | Variable resistance patterns, higher in regions with intensive livestock farming [45,46] |
| Polymyxins | Colistin | Gram-negative bacteria, especially E. coli | Emergence of resistance reported in livestock, raising significant One Health concerns [47] |
| Lincosamides | Lincomycin, clindamycin | Staphylococcus spp., anaerobes | Variable resistance reported, more common in companion animals [48] |
| Nitroimidazoles | Metronidazole | Anaerobic bacteria | Generally low resistance, but regional variability exists [49] |
| Pleuromutilins | Tiamulin, valnemulin | Gram-positive respiratory pathogens | Low to moderate resistance, mainly in food-producing animals [50] |
| Streptogramins | Virginiamycin | Gram-positive bacteria (Staphylococcus spp., Enterococcus spp.) | Resistance reported, often associated with cross-resistance to macrolides and lincosamides [51]. |
| Cyclic polypeptides | Bacitracin | Gram-positive bacteria | Resistance reported in veterinary isolates [52] |
| Bacterial Pathogen | Common Infections in Animals | Frequently Reported Resistance | Clinical Implications |
|---|---|---|---|
| Staphylococcus spp. | Skin, otitis, wounds, postoperative infections | Resistance to β-lactams (including methicillin), macrolides, tetracyclines, and fluoroquinolones widely reported in companion animals [60] | MDR complicates treatment in companion animals |
| Pseudomonas aeruginosa | Otitis externa, wound infections, urinary tract infections | Intrinsic resistance to multiple classes; acquired resistance to β-lactams, fluoroquinolones, and aminoglycosides reported in clinical isolates [61] | Intrinsic + acquired resistance limits therapy |
| E. coli | Urinary tract infections, septicemia, gastrointestinal infections | High prevalence of resistance to β-lactams, fluoroquinolones, tetracyclines, and sulfonamides; ESBL-producing strains reported globally [62,63] | Increasing prevalence of MDR isolates in animals |
| Proteus mirabilis | Urinary tract infections, wound infections | Resistance to β-lactams, fluoroquinolones, and aminoglycosides reported, with regional variability [64] | Emerging resistance patterns reported in veterinary isolates |
| Enterobacter spp. | Opportunistic infections, urinary tract infections | Resistance to β-lactams (including AmpC), cephalosporins, and fluoroquinolones reported in veterinary isolates [65] | Intrinsic + acquired resistance contributes to treatment difficulties |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Popa, I.; Iancu, I.; Popa, S.A.; Gligor, A.; Imre, K.; Tîrziu, E.; Bochiș, T.; Pop, C.; Degi, J.; Ivan, A.A.; et al. Antimicrobial Resistance in Veterinary Bacterial Pathogens: Resistance Patterns, Zoonotic Risks and One Health Implications. Pathogens 2026, 15, 525. https://doi.org/10.3390/pathogens15050525
Popa I, Iancu I, Popa SA, Gligor A, Imre K, Tîrziu E, Bochiș T, Pop C, Degi J, Ivan AA, et al. Antimicrobial Resistance in Veterinary Bacterial Pathogens: Resistance Patterns, Zoonotic Risks and One Health Implications. Pathogens. 2026; 15(5):525. https://doi.org/10.3390/pathogens15050525
Chicago/Turabian StylePopa, Ionela, Ionica Iancu, Sebastian Alexandru Popa, Alexandru Gligor, Kalman Imre, Emil Tîrziu, Timeea Bochiș, Călin Pop, Janos Degi, Andrei Alexandru Ivan, and et al. 2026. "Antimicrobial Resistance in Veterinary Bacterial Pathogens: Resistance Patterns, Zoonotic Risks and One Health Implications" Pathogens 15, no. 5: 525. https://doi.org/10.3390/pathogens15050525
APA StylePopa, I., Iancu, I., Popa, S. A., Gligor, A., Imre, K., Tîrziu, E., Bochiș, T., Pop, C., Degi, J., Ivan, A. A., Dahma, M., Plotuna, A.-M., Pentea, M., Herman, V., & Nichita, I. (2026). Antimicrobial Resistance in Veterinary Bacterial Pathogens: Resistance Patterns, Zoonotic Risks and One Health Implications. Pathogens, 15(5), 525. https://doi.org/10.3390/pathogens15050525










