HBV-Induced Pyruvate Increases Lactylation of Pyruvate Kinase M2 (PKM2) at K206 to Promote Liver Fibrosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Reagents
2.2. siRNAs, Plasmids, and Transfection
2.3. RNA Isolation, Reverse Transcription, and RT-qPCR
2.4. Western Blot
2.5. Immunoprecipitation (IP)
2.6. Cell Viability Assay
2.7. PKM2 Crosslinking
2.8. Nuclear and Cytoplasmic Protein Extraction
2.9. Measurement of Lactate and Pyruvate Kinase Activity
2.10. Human Subject Study Ethics Approval and Animal Study Approval
2.11. Statistical Analysis
3. Results
3.1. HBV Infection and Pyruvate Treatment Increased Protein Lactylation
3.2. Pyruvate Induced Profibrotic Gene Expression in LX2 Cells Through Enhanced PKM2 Lactylation
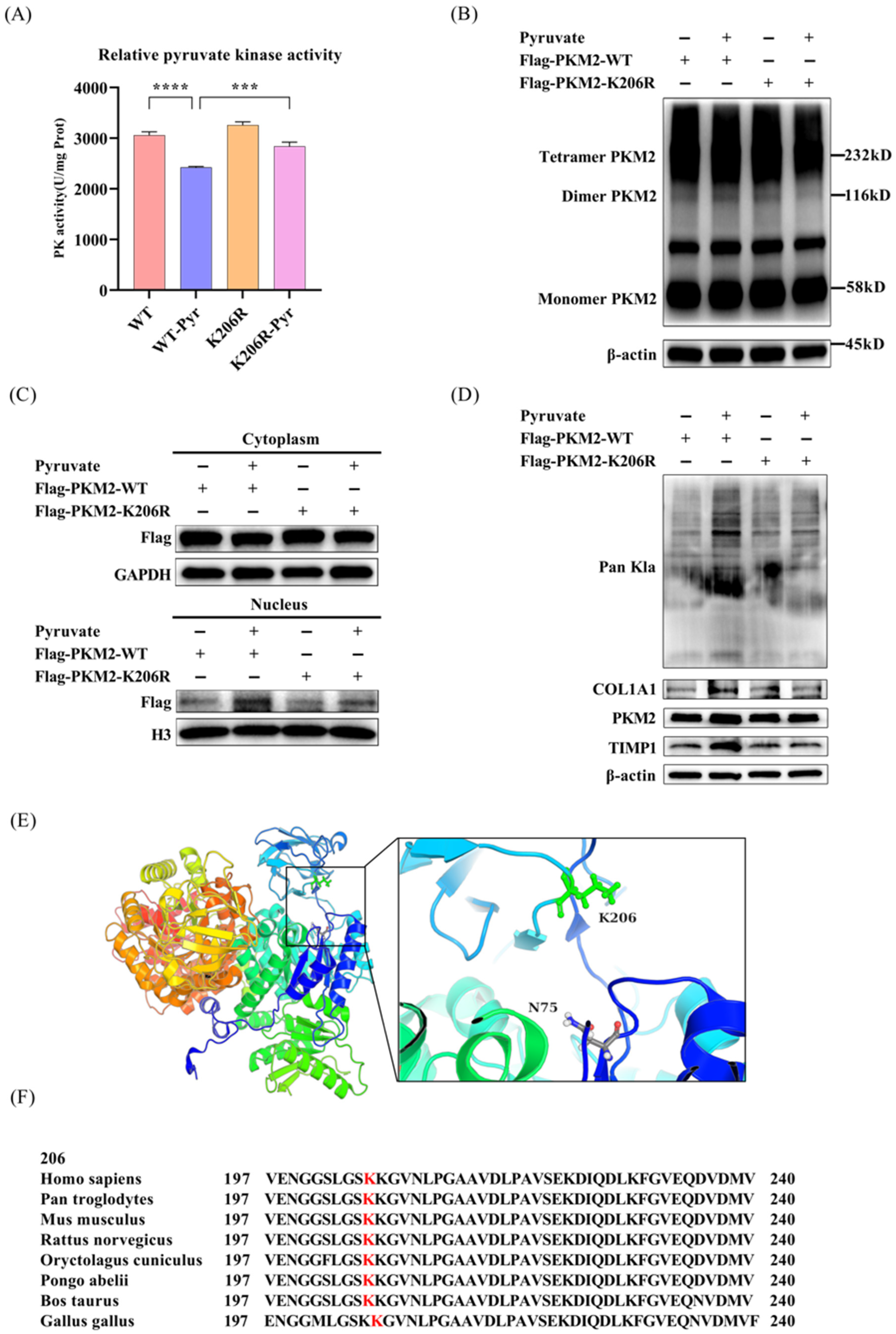
3.3. Pyruvate Promotes PKM2 Lactylation at Lysine 206
3.4. The K206R Mutation Reversed Both the Inhibition of PKM2 Kinase Activity and Blunted the Induction of Pro-Fibrotic Gene Expression Induced by Pyruvate
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jeng, W.J.; Papatheodoridis, G.; Lok, A.S.F. Hepatitis B. Lancet 2023, 401, 1039–1052. [Google Scholar] [CrossRef] [PubMed]
- Iannacone, M.; Guidotti, L.G. Immunobiology and pathogenesis of hepatitis B virus infection. Nat. Rev. Immunol. 2022, 22, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Hammerich, L.; Tacke, F. Hepatic inflammatory responses in liver fibrosis. Nat. Rev. Gastroenterol. Hepatol. 2023, 20, 633–646. [Google Scholar] [CrossRef] [PubMed]
- Horn, P.; Tacke, F. Metabolic reprogramming in liver fibrosis. Cell Metab. 2024, 36, 1439–1455. [Google Scholar] [CrossRef]
- Hou, L.S.; Zhang, Y.W.; Li, H.; Wang, W.; Huan, M.L.; Zhou, S.Y.; Zhang, B.L. The regulatory role and mechanism of autophagy in energy metabolism-related hepatic fibrosis. Pharmacol. Ther. 2022, 234, 108117. [Google Scholar] [CrossRef]
- Ezhilarasan, D. Mitochondria: A critical hub for hepatic stellate cells activation during chronic liver diseases. Hepatobiliary Pancreat. Dis. Int. 2021, 20, 315–322. [Google Scholar] [CrossRef]
- Chang, T.T.; Liaw, Y.F.; Wu, S.S.; Schiff, E.; Han, K.H.; Lai, C.L.; Safadi, R.; Lee, S.S.; Halota, W.; Goodman, Z.; et al. Long-Term Entecavir Therapy Results in the Reversal of Fibrosis/Cirrhosis and Continued Histological Improvement in Patients with Chronic Hepatitis B. Hepatology 2010, 52, 886–893. [Google Scholar] [CrossRef]
- Marcellin, P.; Gane, E.; Buti, M.; Afdhal, N.; Sievert, W.; Jacobson, I.M.; Washington, M.K.; Germanidis, G.; Flaherty, J.F.; Schall, R.A.; et al. Regression of cirrhosis during treatment with tenofovir disoproxil fumarate for chronic hepatitis B: A 5-year open-label follow-up study. Lancet 2013, 381, 468–475. [Google Scholar] [CrossRef]
- Sun, Y.M.; Wu, X.N.; Zhou, J.L.; Meng, T.T.; Wang, B.Q.; Chen, S.Y.; Liu, H.; Wang, T.L.; Zhao, X.Y.; Wu, S.S.; et al. Persistent Low Level of Hepatitis B Virus Promotes Fibrosis Progression During Therapy. Clin. Gastroenterol. Hepatol. 2020, 18, 2582–2591. [Google Scholar] [CrossRef]
- Lamontagne, R.J.; Casciano, J.C.; Bouchard, M.J. A broad investigation of the HBV-mediated changes to primary hepatocyte physiology reveals HBV significantly alters metabolic pathways. Metabolism 2018, 83, 50–59. [Google Scholar] [CrossRef]
- Zhou, L.; He, R.; Fang, P.N.; Li, M.Q.; Yu, H.S.; Wang, Q.M.; Yu, Y.; Wang, F.B.; Zhang, Y.; Chen, A.D.; et al. Hepatitis B virus rigs the cellular metabolome to avoid innate immune recognition. Nat. Commun. 2021, 12, 98. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.Q.; Li, S.L.; Holmes, J.A.; Tu, Z.; Li, Y.J.; Cai, D.C.; Liu, X.; Li, W.T.; Yang, C.H.; Jiao, B.H.; et al. MicroRNA 130a Regulates both Hepatitis C Virus and Hepatitis B Virus Replication through a Central Metabolic Pathway. J. Virol. 2018, 92, e02009-17. [Google Scholar] [CrossRef]
- Duan, X.Q.; Li, S.S.; Li, Q.Y.; Wen, W.X.; Mezzetti, O.; Warner, C.; Xu, M.; Jeyarajan, A.J.; Li, Y.J.; Shi, Y.Q.; et al. HBV induces liver fibrosis through the generation of reactive oxygen species in a pyruvate-dependent manner. Hepatology 2025. [Google Scholar] [CrossRef]
- Rui, L. Energy Metabolism in the Liver. Compr. Physiol. 2014, 4, 177–197. [Google Scholar] [CrossRef]
- Zhang, D.; Tang, Z.Y.; Huang, H.; Zhou, G.L.; Cui, C.; Weng, Y.J.; Liu, W.C.; Kim, S.; Lee, S.; Perez-Neut, M.; et al. Metabolic regulation of gene expression by histone lactylation. Nature 2019, 574, 575–580. [Google Scholar] [CrossRef]
- Li, L.N.; Li, W.W.; Xiao, L.S.; Lai, W.N. Lactylation signature identifies liver fibrosis phenotypes and traces fibrotic progression to hepatocellular carcinoma. Front. Immunol. 2024, 15, 1433393. [Google Scholar] [CrossRef]
- Rho, H.; Terry, A.R.; Chronis, C.; Hay, N. Hexokinase 2-mediated gene expression via histone lactylation is required for hepatic stellate cell activation and liver fibrosis. Cell Metab. 2023, 35, 1406–1423. [Google Scholar] [CrossRef]
- Alquraishi, M.; Puckett, D.L.; Alani, D.S.; Humidat, A.S.; Frankel, V.D.; Donohoe, D.R.; Whelan, J.; Bettaieb, A. Pyruvate kinase M2: A simple molecule with complex functions. Free Radic. Biol. Med. 2019, 143, 176–192. [Google Scholar] [CrossRef]
- Chen, M.; Zhang, J.; Manley, J.L. Turning on a Fuel Switch of Cancer: hnRNP Proteins Regulate Alternative Splicing of Pyruvate Kinase mRNA. Cancer Res. 2010, 70, 8977–8980. [Google Scholar] [CrossRef] [PubMed]
- Christofk, H.R.; Vander Heiden, M.G.; Wu, N.; Asara, J.M.; Cantley, L.C. Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature 2008, 452, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.L.; Wang, H.Z.; Yang, J.J.; Liu, X.W.; Liu, Z.R. Pyruvate Kinase M2 Regulates Gene Transcription by Acting as a Protein Kinase. Mol. Cell 2012, 45, 598–609. [Google Scholar] [CrossRef]
- Hsu, M.C.; Hung, W.C. Pyruvate kinase M2 fuels multiple aspects of cancer cells: From cellular metabolism, transcriptional regulation to extracellular signaling. Mol. Cancer 2018, 17, 35. [Google Scholar] [CrossRef]
- Iqbal, M.A.; Gupta, V.; Gopinath, P.; Mazurek, S.; Bamezai, R.N.K. Pyruvate kinase M2 and cancer: An updated assessment. FEBS Lett. 2014, 588, 2685–2692. [Google Scholar] [CrossRef]
- Tamada, M.; Suematsu, M.; Saya, H. Pyruvate Kinase M2: Multiple Faces for Conferring Benefits on Cancer Cells. Clin. Cancer Res. 2012, 18, 5554–5561. [Google Scholar] [CrossRef]
- Yang, W.W.; Xia, Y.; Hawke, D.; Li, X.J.; Liang, J.; Xing, D.M.; Aldape, K.; Hunter, T.; Yung, W.K.A.; Lu, Z.M. PKM2 Phosphorylates Histone H3 and Promotes Gene Transcription and Tumorigenesis. Cell 2012, 150, 685–696. [Google Scholar] [CrossRef]
- Christofk, H.R.; Vander Heiden, M.G.; Harris, M.H.; Ramanathan, A.; Gerszten, R.E.; Wei, R.; Fleming, M.D.; Schreiber, S.L.; Cantley, L.C. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 2008, 452, 230–233. [Google Scholar] [CrossRef]
- Satyanarayana, G.; Turaga, R.C.; Sharma, M.; Wang, S.M.; Mishra, F.; Peng, G.D.; Deng, X.N.; Yang, J.; Liu, Z.R. Pyruvate kinase M2 regulates fibrosis development and progression by controlling glycine auxotrophy in myofibroblasts. Theranostics 2021, 11, 9331–9341. [Google Scholar] [CrossRef] [PubMed]
- Prakasam, G.; Iqbal, M.A.; Bamezai, R.N.K.; Mazurek, S. Posttranslational Modifications of Pyruvate Kinase M2: Tweaks that Benefit Cancer. Front. Oncol. 2018, 8, 22. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Z.; Yang, P.L.; Yu, T.Y.; Gao, M.; Liu, D.; Zhang, J.; Lu, C.H.; Chen, X.L.; Zhang, X.; Liu, Y. Lactylation of PKM2 Suppresses Inflammatory Metabolic Adaptation in Pro-inflammatory Macrophages. Int. J. Biol. Sci. 2022, 18, 6210–6225. [Google Scholar] [CrossRef]
- Dong, Y.T.; Liu, Y.W.; Liu, X.; Ma, H.M.; Liu, Y.H.; Lv, G.Y.; Niu, J.Q. Multi-omics profiling of primary hepatic stellate cells from advanced liver fibrosis patients reveals distinctive molecular signatures. J. Gastroenterol. Hepatol. 2023, 38, 1416–1425. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.J.; Yan, C.; Ma, J.Q.; Peng, P.P.; Ren, X.L.; Cai, S.L.; Shen, X.; Wu, Y.C.; Zhang, S.; Wang, X.Y.; et al. Lactylome analysis suggests lactylation-dependent mechanisms of metabolic adaptation in hepatocellular carcinoma. Nat. Metab. 2023, 5, 61–79. [Google Scholar] [CrossRef]
- Zheng, D.D.; Jiang, Y.C.; Qu, C.; Yuan, H.; Hu, K.S.; He, L.; Chen, P.; Li, J.Y.; Tu, M.X.; Lin, L.H.; et al. Pyruvate Kinase M2 Tetramerization Protects against Hepatic Stellate Cell Activation and Liver Fibrosis. Am. J. Pathol. 2020, 190, 2267–2281. [Google Scholar] [CrossRef]
- Katrinli, S.; Ozdil, K.; Sahin, A.; Ozturk, O.; Kir, G.; Baykal, A.T.; Akgun, E.; Sarac, O.S.; Sokmen, M.; Doganay, H.L.; et al. Proteomic profiling of HBV infected liver biopsies with different fibrotic stages. Proteome Sci. 2017, 15, 7. [Google Scholar] [CrossRef] [PubMed]
- Steelman, L.S.; Chappell, W.H.; Abrams, S.L.; Kempf, C.R.; Long, J.; Laidler, P.; Mijatovic, S.; Maksimovic-Ivanic, D.; Stivala, F.; Mazzarino, M.C.; et al. Roles of the Raf/MEK/ERK and PI3K/PTEN/Akt/mTOR pathways in controlling growth and sensitivity to therapy-implications for cancer and aging. Aging 2011, 3, 192–222. [Google Scholar] [CrossRef]
- Wang, X.Y.; Wei, Z.Q.; Jiang, Y.F.; Meng, Z.J.; Lu, M.J. mTOR Signaling: The Interface Linking Cellular Metabolism and Hepatitis B Virus Replication. Virol. Sin. 2021, 36, 1303–1314. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.F.; Zhang, X.; Fang, Y.; Xu, Z.Y.; Yu, Y.; Zhang, L.L.; Yang, Y.P.; Li, S.; Wang, Y.P.; Jiang, C.; et al. Identification of a lactylation-related gene signature as the novel biomarkers for early diagnosis of acute myocardial infarction. Int. J. Biol. Macromol. 2024, 282, 137431. [Google Scholar] [CrossRef]
- Liu, J.; Zhao, F.; Qu, Y. Lactylation: A Novel Post-Translational Modification with Clinical Implications in CNS Diseases. Biomolecules 2024, 14, 1175. [Google Scholar] [CrossRef]
- Su, J.; Zheng, Z.; Bian, C.; Chang, S.; Bao, J.; Yu, H.; Xin, Y.; Jiang, X. Functions and mechanisms of lactylation in carcinogenesis and immunosuppression. Front. Immunol. 2023, 14, 1253064. [Google Scholar] [CrossRef]
- Jiang, R.; Ren, W.-J.; Wang, L.-Y.; Zhang, W.; Jiang, Z.-H.; Zhu, G.-Y. Targeting Lactate: An Emerging Strategy for Macrophage Regulation in Chronic Inflammation and Cancer. Biomolecules 2024, 14, 1202. [Google Scholar] [CrossRef]
- Luo, Y.; Yang, Z.; Yu, Y.; Zhang, P. HIF1α lactylation enhances KIAA1199 transcription to promote angiogenesis and vasculogenic mimicry in prostate cancer. Int. J. Biol. Macromol. 2022, 222, 2225–2243. [Google Scholar] [CrossRef] [PubMed]
- Qu, H.; Liu, J.; Zhang, D.; Xie, R.; Wang, L.; Hong, J. Glycolysis in Chronic Liver Diseases: Mechanistic Insights and Therapeutic Opportunities. Cells 2023, 12, 1930. [Google Scholar] [CrossRef] [PubMed]
- Hitosugi, T.; Kang, S.K.; Heiden, M.G.V.; Chung, T.W.; Elf, S.; Lythgoe, K.; Dong, S.Z.; Lonial, S.; Wang, X.; Chen, G.Z.; et al. Tyrosine Phosphorylation Inhibits PKM2 to Promote the Warburg Effect and Tumor Growth. Sci. Signal. 2009, 2, ra73. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Li, F.Z.; Han, H.C.; Chen, Y.; Mao, Z.B.; Luo, J.Y.; Zhao, Y.M.; Zheng, B.; Gu, W.; Zhao, W.H. Parkin Regulates the Activity of Pyruvate Kinase M2. J. Biol. Chem. 2016, 291, 10307–10317. [Google Scholar] [CrossRef] [PubMed]
- Feitelson, M.A.; Reis, H.; Tufan, N.L.; Sun, B.; Pan, J.B.; Lian, Z.R. Putative roles of hepatitis B x antigen in the pathogenesis of chronic liver disease. Cancer Lett. 2009, 286, 69–79. [Google Scholar] [CrossRef]
- Chen, C.F.; Feng, X.; Liao, H.Y.; Jin, W.J.; Zhang, J.; Wang, Y.; Gong, L.L.; Liu, J.J.; Yuan, X.H.; Zhao, B.B.; et al. Regulation of T cell proliferation by JMJD6 and PDGF-BB during chronic hepatitis B infection. Sci. Rep. 2014, 4, 6359. [Google Scholar] [CrossRef]




| Gene Name | Forward (5′-3′) | Reverse (5′-3′) |
|---|---|---|
| β-actin | GATGAGATTGGCATGGCTTT | GTCACCTTCACCGTTCCAGT |
| PKM2 | ACGAGAACATCCTGTGGCTG | GCTCGACCCCAAACTTCAGA |
| COL1A1 | AACATGACCAAAAACCAAAAGTG | CATTGTTTCCTGTGTCTTCTGG |
| TIMP1 | GAGAGACACCAGAGAACCCAC | CCACAGCAACAACAGGATGC |
| Non-HBV (n = 7) | CHB-I (n = 28) | CHB-F (n = 27) | p Value | Effect Size | Power | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P1 | P2 | P3 | P1 | P2 | P3 | P1 | P2 | P3 | ||||
| Age (years) | 46.0 (30.0–50.0) | 36.0 (30.5–42.5) | 48.0 (43.0–51.0) | 0.180 | 0.442 | 0.0001 | 1.09 | 0.13 | 1.15 | 0.71 | 0.06 | 0.99 |
| Male (n, %) | 4 (57.1) | 13 (46.4) | 17 (63.0) | 0.617 | 0.781 | 0.223 | 0.09 | 0.05 | 0.17 | 0.08 | 0.06 | 0.24 |
| ALT (U/L) | 23.0 (17.0–29.0) | 39.0 (27.5–121.3) | 24.0 (16.0–33.0) | 0.006 | 0.798 | 0.0002 | 0.93 | 0.32 | 0.78 | 0.55 | 0.11 | 0.79 |
| AST (U/L) | 21.0 (19.0–24.0) | 29.0 (22.0–62.0) | 23.0 (20.0–26.0) | 0.018 | 0.442 | 0.005 | 0.98 | 0.36 | 0.80 | 0.60 | 0.13 | 0.43 |
| TBIL (μmol/L) | 13.7 (10.4–28.6) | 16.2 (12.5–19.1) | 12.7 (10.7–16.0) | 0.805 | 0.565 | 0.077 | 0.14 | 0.45 | 0.39 | 0.06 | 0.17 | 0.28 |
| Lactate (μmol/mL) | 2.1 (1.6–2.3) | 5.8 (4.2–9.1) | 9.8 (6.9–12.2) | 0.0064 | <0.0001 | 0.0046 | 2.35 | 3.29 | 1.07 | 1.00 | 1.00 | 0.97 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Wen, W.; Du, Q.; Li, S.; Yang, Y.; Wang, X.; Li, S.; Li, Y.; Li, S.; Yang, C.; Xie, H.; et al. HBV-Induced Pyruvate Increases Lactylation of Pyruvate Kinase M2 (PKM2) at K206 to Promote Liver Fibrosis. Pathogens 2026, 15, 431. https://doi.org/10.3390/pathogens15040431
Wen W, Du Q, Li S, Yang Y, Wang X, Li S, Li Y, Li S, Yang C, Xie H, et al. HBV-Induced Pyruvate Increases Lactylation of Pyruvate Kinase M2 (PKM2) at K206 to Promote Liver Fibrosis. Pathogens. 2026; 15(4):431. https://doi.org/10.3390/pathogens15040431
Chicago/Turabian StyleWen, Wenxian, Qin Du, Shuhan Li, Youmin Yang, Xianding Wang, Shasha Li, Yujia Li, Shilin Li, Chunhui Yang, He Xie, and et al. 2026. "HBV-Induced Pyruvate Increases Lactylation of Pyruvate Kinase M2 (PKM2) at K206 to Promote Liver Fibrosis" Pathogens 15, no. 4: 431. https://doi.org/10.3390/pathogens15040431
APA StyleWen, W., Du, Q., Li, S., Yang, Y., Wang, X., Li, S., Li, Y., Li, S., Yang, C., Xie, H., Duan, X., & Chen, L. (2026). HBV-Induced Pyruvate Increases Lactylation of Pyruvate Kinase M2 (PKM2) at K206 to Promote Liver Fibrosis. Pathogens, 15(4), 431. https://doi.org/10.3390/pathogens15040431







