Antibacterial Efficacy of Pseudomonas aeruginosa Bacteriophages on a Drosophila Infection Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Origin of Bacteria, Virus Isolation
2.2. Phage Bioinformatics
2.3. Phage Biological Characteristics
2.4. Drosophila Infectivity Test of Distinct P. aeruginosa Strains and Phage Protection
3. Results
3.1. Molecular Characteristics of the Viruses
| Genome (bp) | ORF | G+C (%) | Mostly Related Virus | Genus | Classification | AC No: | |
|---|---|---|---|---|---|---|---|
| Klucen469 | 43225 | 52 | 60.8 | YMC11 NC_030923 | Kantovirus-like | Subfamily: Corkvirinae; Order: Autographivirales | PX843241 |
| Radvan531 | 40737 | 46 | 59.8 | VSW-3 NC_041885 | Napahaivirus-like | Family: Autonotaviridae; Order: Autographivirales | PX843244 |
| Kovar531 | 45196 | 70 | 52.4 | oldone MT119371 | Bruynoghevirus | Class: Caudoviricetes | PX843246 |
| Onen484 | 42855 | 57 | 53.7 | Phi73 NC_007806 | Septimatrevirus | Subfamily: Jondennisvirinae; Class: Caudoviricetes | PX843242 |
| Onen526 | 42816 | 61 | 53.2 | SCUT S4 MK165658 | Septimatrevirus | Subfamily: Jondennisvirinae; Class: Caudoviricetes | PX843243 |
| Lasov521 | 63274 | 87 | 60.0 | KPP25 NC_024123 | Kochitakasuvirus | Class: Caudoviricetes | PX843245 |
3.2. Molecular Characteristics of Pseudomonas Phage Klucen469
3.3. Molecular Characteristics of Pseudomonas Radvan531 Phage
3.4. Molecular Characteristics of Pseudomonas Viruses Onen484 and Onen526
3.5. Molecular Characteristics of the Kovar531
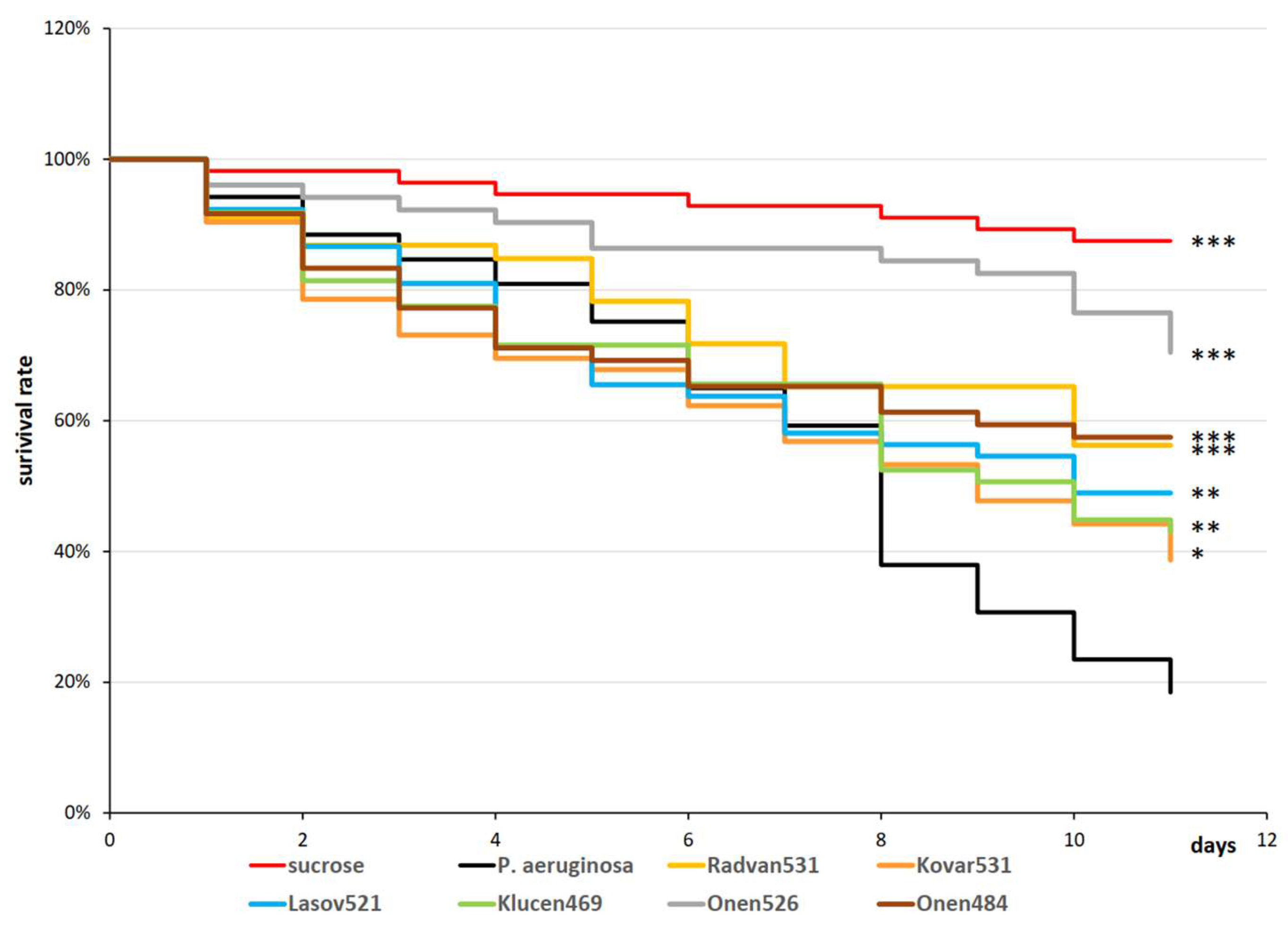
3.6. Efficacy of Drosophila Flies Infection/Curation with Distinct Bacteriophage
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ambreetha, S.; Balachandar, D. Pathogenesis of plant-associated Pseudomonas aeruginosa in Caenorhabditis elegans model. BMC Microbiol. 2022, 22, 269. [Google Scholar] [CrossRef]
- De Oliveira, D.M.P.; Forde, B.M.; Kidd, T.J.; Harris, P.N.A.; Schembri, M.A.; Beatson, S.A.; Paterson, D.L.; Walker, M.J. Antimicrobial resistance in ESKAPE pathogens. Clin. Microbiol. Rev. 2020, 13, e00181-19. [Google Scholar] [CrossRef]
- Subedi, D.; Vijay, A.K.; Kohli, G.S.; Rice, S.A.; Willcox, M. Comparative genomics of clinical strains of Pseudomonas aeruginosa strains isolated from different geographic sites. Sci. Rep. 2018, 8, 15668. [Google Scholar] [CrossRef]
- Bassetti, M.; Vena, A.; Croxatto, A.; Righi, E.; Guery, B. How to manage Pseudomonas aeruginosa infections. Drugs Context 2018, 7, 212527. [Google Scholar] [CrossRef]
- Hatfull, G.F.; Dedrick, R.M.; Schooley, R.T. Phage therapy for antibiotic-resistant bacterial infections. Annu. Rev. Med. 2022, 73, 197–211. [Google Scholar] [CrossRef]
- Pottier, M.; Gravey, F.; Castagnet, S.; Auzou, M.; Langlois, B.; Guérin, F.; Giard, J.-C.; Léon, A.; Le Hello, S. A 10-year microbiological study of Pseudomonas aeruginosa strains revealed the circulation of populations resistant to both carbapenems and quaternary ammonium compounds. Sci. Rep. 2013, 13, 2639. [Google Scholar] [CrossRef]
- Soothill, J.S. Treatment of experimental infections of mice with bacteriophages. J. Med. Microbiol. 1992, 37, 258–261. [Google Scholar] [CrossRef] [PubMed]
- Soothill, J.S. Bacteriophage prevents destruction of skin grafts by Pseudomonas aeruginosa. Burns 1994, 20, 209–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, B.; Xu, M.; Yan, Q.; Liu, S.; Zhu, X.; Sun, Z.; Reed, E.; Ding, L.; Gong, J.; et al. Use of bacteriophage in the treatment of experimental animal bacteremia from imipenem-resistant Pseudomonas aeruginosa. Int. J. Mol. Med. 2006, 17, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Heo, Y.-J.; Lee, Y.-R.; Jung, H.-H.; Lee, J.; Ko, G.; Cho, Y.-H. Antibacterial efficacy of phages against Pseudomonas aeruginosa infections in mice and Drosophila melanogaster. Antimicrob. Agents Chemother. 2009, 53, 2469–2474. [Google Scholar] [CrossRef]
- Hawkins, C.; Harper, D.; Burch, D.; Änggård, E.; Soothill, J. Topical treatment of Pseudomonas aeruginosa otitis of dogs with a bacteriophage mixture: A before/after clinical trial. Vet. Microbiol. 2010, 146, 309–313. [Google Scholar] [CrossRef]
- Gu, J.; Li, X.; Yang, M.; Du, C.; Cui, Z.; Gong, P.; Xia, F.; Song, J.; Zhang, L.; Li, J.; et al. Therapeutic effect of Pseudomonas aeruginosa phage YH30 on mink hemorrhagic pneumonia. Vet. Microbiol. 2016, 190, 5–11. [Google Scholar] [CrossRef] [PubMed]
- Debarbieux, L.; Luduc, D.; Maura, D.; Morello, E.; Criscuolo, A.; Grossi, O.; Balloy, V.; Touqui, L. Bacteriophages can treat and prevent Pseudomonas aeruginosa lung infection. J. Inf. Dis. 2010, 201, 1096–1104. [Google Scholar] [CrossRef]
- Chegini, Z.; Khoshbayan, A.; Moghadam, M.T.; Farahani, I.; Jazireian, P.; Shariati, A. Bacteriophage therapy against Pseudomonas aeruginosa biofilms: A review. Ann. Clin. Microbiol. Antimicrob. 2020, 19, 45. [Google Scholar] [CrossRef]
- Paradis-Bleau, C.; Cloutier, I.; Lemieux, L.; Sanschagrin, F.; Laroche, J.; Auger, M.; Levesque, R.C. Peptidoglycan lytic activity of the Pseudomonas aeruginosa phage fKZ gp144 lytic transglycosylase. FEMS Microbiol. Lett. 2007, 266, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Briers, Y.; Volckaert, G.; Cornelissen, A.; Lagaert, S.; Michiels, C.W.; Hertveldt, K.; Lavigne, R. Muralytic activity and modular structure of the endolysins of Pseudomonas aeruginosa bacteriophages fKZ and EL. Mol. Microbiol. 2007, 65, 1334–1344. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Feng, C.; Ren, J.; Zhuang, X.; Zhang, Y.; Zhu, Y.; Dong, K.; He, P.; Guo, X.; Qin, J. A novel antimicrobial endolysin, LysPA26, against Pseudomonas aeruginosa. Front. Microbiol. 2017, 8, 293. [Google Scholar] [CrossRef]
- Lindberg, H.M.; McKean, K.A.; Wang, I.-N. Phage fitness may help predict phage therapy efficacy. Bacteriophage 2014, 4, e964081. [Google Scholar] [CrossRef]
- Brix, A.; Cafora, M.; Aureli, M.; Pistocchi, A. Animal models to translate phage therapy to human medicine. Int. J. Mol. Sci. 2020, 21, 3715. [Google Scholar] [CrossRef]
- Apidianakis, Y.; Rahme, L.G. Drosophila melanogaster as a model host for studying Pseudomonas aeruginosa infection. Nat. Protoc. 2009, 4, 1285–1294. [Google Scholar] [CrossRef]
- D’ Argenio, D.A.; Gallagher, L.A.; Berg, C.A.; Manoil, C. Drosophila as a model host for Pseudomonas aeruginosa infection. J. Bacter 2001, 183, 1466–1471. [Google Scholar] [CrossRef]
- Limmer, S.; Quintin, J.; Hetru, C.; Ferrandon, D. Virulence on the fly: Drosophila melanogaster as a model genetic organism to decipher host-pathogen interactions. Curr. Drug Targets 2011, 12, 978–999. [Google Scholar] [CrossRef]
- Limmer, S.; Haller, S.; Drenkard, E.; Lee, J.; Yu, S.; Kocks, C.; Ausubel, F.M.; Ferrandon, D. Pseudomonas aeruginosa RhlR is required to neutralize the cellular immune response in a Drosophila melanogaster oral infection model. Proc. Natl. Acad. Sci. USA 2011, 108, 17378–17383. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, H.; Sibley, C.D.; Surette, M.G.; Lewenza, S. Drosophila melanogaster as an animal model for the study of Pseudomonas aeruginosa biofilm infections in vivo. PLoS Pathog. 2011, 7, e1002299. [Google Scholar] [CrossRef] [PubMed]
- Lutter, E.I.; Faria, M.M.P.; Rabin, H.R.; Storey, D.G. Pseudomonas aeruginosa cystic fibrosis isolates from individual patients demonstrate a range of levels of lethality in two Drosophila melanogaster infection models. Infect. Immun. 2008, 76, 1877–1888. [Google Scholar] [CrossRef]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef] [PubMed]
- Petrzik, K.; Brázdová, S. Jojan: A novel virus that lyses Stenotrophomonas maltophilia from dog. Virus Genes 2023, 59, 775–780. [Google Scholar] [CrossRef]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef]
- Delcher, A.L.; Harmon, D.; Kasif, S.; White, O.; Salzberg, S.L. Improved microbial gene identification with GLIMMER. Nucleic Acids Res. 1999, 27, 4636–4641. [Google Scholar] [CrossRef]
- Urbanowicz, M.; Guziński, A.; Szulc, Ż. PhageAI: A new approach to predicting the lifestyle of bacteriophages using proteinBERT and convolutional neural networks. BioRxiv 2025. [Google Scholar] [CrossRef]
- Merrill, B.D.; Ward, A.T.; Grose, J.H.; Hope, S. Software-based analysis of bacteriophage genomes, physical ends, and packaging strategies. BMC Genom. 2016, 17, 679. [Google Scholar] [CrossRef]
- Kropinski, A.M. Practical advice on the one-step growth curve. In Bacteriophages: Methods and Protocols, Volume 3; Methods in Molecular Biology; Clokie, M.R.J., Kropinski, A.M., Lavigne, R., Eds.; Springer Science+Business Media LLC: New York, NY, USA, 2018; Volume 1681, pp. 41–47. [Google Scholar] [CrossRef]
- Nishimura, Y.; Yoshida, T.; Kuronishi, M.; Uehara, H.; Ogata, H.; Goto, S. ViPTree: The viral proteomic tree server. Bioinformatics 2017, 33, 2379–2380. [Google Scholar] [CrossRef] [PubMed]
- Clark, T.G.; Bradburn, M.J.; Love, S.B.; Altman, D.G. Survival analysis part I: Basic concept and first analyses. Brit. J. Cancer 2003, 89, 232–238. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid Annotations using Subsystems Technology. BMC Genom. 2008, 8, 75. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, L.; Stephens, A.; Nam, S.Z.; Rau, D.; Kübler, J.; Lozajic, M.; Gabler, F.; Söding, J.; Lupas, A.N.; Alva, V. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 2018, 430, 2237–2243. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Skaar, K.; Claesson, M.; Odegrip, R.; Högbom, M.; Haggård-Ljungquist, E.; Stenmark, P. Crystal structure of the bacteriophage P2 integrase catalytic domain. FEBS Lett. 2015, 589, 3556–3563. [Google Scholar] [CrossRef]
- Yukgehnaish, K.; Rajandas, H.; Parimannan, S.; Manickam, R.; Marimuthu, K.; Petersen, B.; Clokie, M.R.J.; Millard, A.; Sicheritz-Pontén, T. PhageLeads: Rapid assessment of phage therapeutic suitability using an ensemble machine learning approach. Viruses 2022, 8, 342. [Google Scholar] [CrossRef]
- Krumsiek, J.; Arnold, R.; Rattei, T. Gepard: A rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics 2007, 23, 1026–1028. [Google Scholar] [CrossRef]
- Vidal, M.; Arch, M.; Fuentes, E.; Cardona, P.-J. Drosophila melanogaster experimental model to test new antimicrobials: A methodological approach. Front. Microbiol. 2024, 15, 1478263. [Google Scholar] [CrossRef] [PubMed]
- Chugani, S.A.; Whiteley, M.; Kimberly, M.L.; Greenberg, E.P. QscR, a modulator of quorum-sensing signal synthesis and virulence in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2001, 98, 2752–2757. [Google Scholar] [CrossRef]
- Lutter, E.I.; Purighalla, S.; Duong, J.; Storey, D.G. Lethality and cooperation of Pseudomonas aeruginosa quorum-sensing mutants in Drosophila melanogaster infection models. Microbiology 2012, 158, 2125–2132. [Google Scholar] [CrossRef]
- Essoh, C.; Latino, L.; Midoux, C.; Blouin, Y.; Loukou, G.; Nguetta, S.-P.A.; Lathro, S.; Cablanmian, A.; Kouassi, A.K.; Vergnaud, G.; et al. Investigation of a large collection of Pseudomonas aeruginosa bacteriophages collected from a single environmental source in Abidjan, Côte d’Ivoire. PLoS ONE 2015, 10, e0130548. [Google Scholar] [CrossRef]
- Karumidze, N.; Thomas, J.A.; Kvatadze, N.; Goderdzishvili, M.; Hakala, K.W.; Weintraub, S.T.; Alavidze, Z.; Hardies, S.C. Characterization of lytic Pseudomonas aeruginosa bacteriophages via biological properties and genomic sequences. Appl. Microbiol. Biotechnol. 2012, 94, 1609–1617. [Google Scholar] [CrossRef]
- Peters, D.L.; Lynch, K.H.; Stothard, P.; Dennis, J.J. The isolation and characterization of two Stenotrophomonas maltophilia bacteriophages capable of cross-taxonomic order infectivity. BMC Genom. 2015, 16, 664. [Google Scholar] [CrossRef]
- Namonyo, S.; Carvalho, G.; Guo, J.; Weynberg, K.D. Novel bacteriophages show activity against selected Australian clinical strains of Pseudomonas aeruginosa. Microorganisms 2022, 10, 210. [Google Scholar] [CrossRef]
- Knezevic, P.; Petrovic Fabijan, A.; Gavric, D.; Pejic, J.; Doffkay, Z.; Rakhely, G. Phages from genus Bruynoghevirus and phagetherapy: Pseudomonas phage Delta case. Viruses 2021, 13, 1965. [Google Scholar] [CrossRef] [PubMed]
- Farlow, J.; Freyberger, H.R.; He, Y.; Ward, A.M.; Rutvisuttinunt, W.; Li, T.; Campbell, R.; Jacobs, A.C.; Nikolich, M.P.; Filippov, A.A. Complete genome sequences of 10 phages lytic against multidrug-resistant Pseudomonas aeruginosa. Microbiol. Resour. Announc. 2020, 9, e00503-20. [Google Scholar] [CrossRef] [PubMed]
- Nordstrom, H.R.; Evans, D.R.; Finney, A.G.; Westbrook, K.J.; Zamora, P.F.; Hofstaedter, C.E.; Yassin, M.H.; Pradhan, A.; Iovleva, A.; Ernst, R.K.; et al. Genomic characterization of lytic bacteriophages targeting genetically diverse Pseudomonas aeruginosa clinical isolates. iScience 2022, 25, 104372. [Google Scholar] [CrossRef]
- Bolognini, S.; Ferretti, C.; Campobasso, C.; Trovato, E.; Marchetti, M.; Rindi, L.; Tavanti, A.; Di Luca, M. Therapeutic optimization of Pseudomonas aeruginosa phages: From isolation to directed evolution. Viruses 2025, 17, 938. [Google Scholar] [CrossRef] [PubMed]
- Tkhilaishvili, T.; Wang, L.; Tavanti, A.; Trampuz, A.; Di Luca, M. Antibacterial efficacy of two commercially available bacteriophage formulations, Staphylococcal bacteriophage and PYO bacteriophage, against methicillin-resistant Staphylococcus aureus: Prevention and eradication of biofilm formation and control of a systemic infection of Galleria mellonella larvae. Front. Microbiol. 2020, 11, 110. [Google Scholar] [CrossRef] [PubMed]




| Bacteria/Strain | Collection | Source |
|---|---|---|
| P. aeruginosa POCH2 | - | canine ear swab |
| P. aeruginosa 1959 | CCM | urine |
| P. aeruginosa 1961 | CCM | outer-ear infection |
| P. aeruginosa 1968 | CCM | unknown source |
| P. aeruginosa 3630 | CCM | otitis externa |
| P. aeruginosa 3989 | CCM | waterworth, clinical |
| P. aeruginosa 7930 | CCM | animal room water |
| P. aeruginosa 22644 | DSMZ | infected wound, type strain |
| Bacteria Strain | Klucen469 | Kovar531 | Radvan531 | Onen484 | Onen526 | Lasov521 |
|---|---|---|---|---|---|---|
| P. aeruginosa POCH2 | + | + | + | + | + | + |
| P. aeruginosa 1959 | + | + | + | + | + | + |
| P. aeruginosa 1961 | + | + | - | - | - | + |
| P. aeruginosa 1968 | - | + | - | - | - | - |
| P. aeruginosa 3630 | + | - | + | + | + | + |
| P. aeruginosa 3989 | + | + | + | + | + | + |
| P. aeruginosa 7930 | + | + | + | + | + | + |
| P. aeruginosa 22644 | + | + | + | + | + | + |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Petrzik, K.; Brázdová, S. Antibacterial Efficacy of Pseudomonas aeruginosa Bacteriophages on a Drosophila Infection Model. Pathogens 2026, 15, 411. https://doi.org/10.3390/pathogens15040411
Petrzik K, Brázdová S. Antibacterial Efficacy of Pseudomonas aeruginosa Bacteriophages on a Drosophila Infection Model. Pathogens. 2026; 15(4):411. https://doi.org/10.3390/pathogens15040411
Chicago/Turabian StylePetrzik, Karel, and Sára Brázdová. 2026. "Antibacterial Efficacy of Pseudomonas aeruginosa Bacteriophages on a Drosophila Infection Model" Pathogens 15, no. 4: 411. https://doi.org/10.3390/pathogens15040411
APA StylePetrzik, K., & Brázdová, S. (2026). Antibacterial Efficacy of Pseudomonas aeruginosa Bacteriophages on a Drosophila Infection Model. Pathogens, 15(4), 411. https://doi.org/10.3390/pathogens15040411






