ATG7 Limits Basal Antiviral Gene Expression and Moderately Promotes VSV Replication in Mammalian Non-Immune Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasmid Constructs
2.2. Cell Lines, CRISPR Gene Editing, and Gene Complementation
2.3. Viruses and 50% Tissue Culture Infectious Dose (TCID50) Assay of Virus Infectivity
2.4. VSV Entry Assay
2.5. Firefly Luciferase Reporter Assay of Viral RNA Replication Using VSV-Luc
2.6. RNA Analysis
2.7. Immunoblotting
2.8. Statistical Analysis
3. Results
3.1. ATG7 Expression Is Required for Basal as Well as VSV-Induced LC3B Lipidation in MEF and HeLa Cultures
3.2. ATG7 Moderately Promotes VSV Propagation in MEF and HeLa Cultures
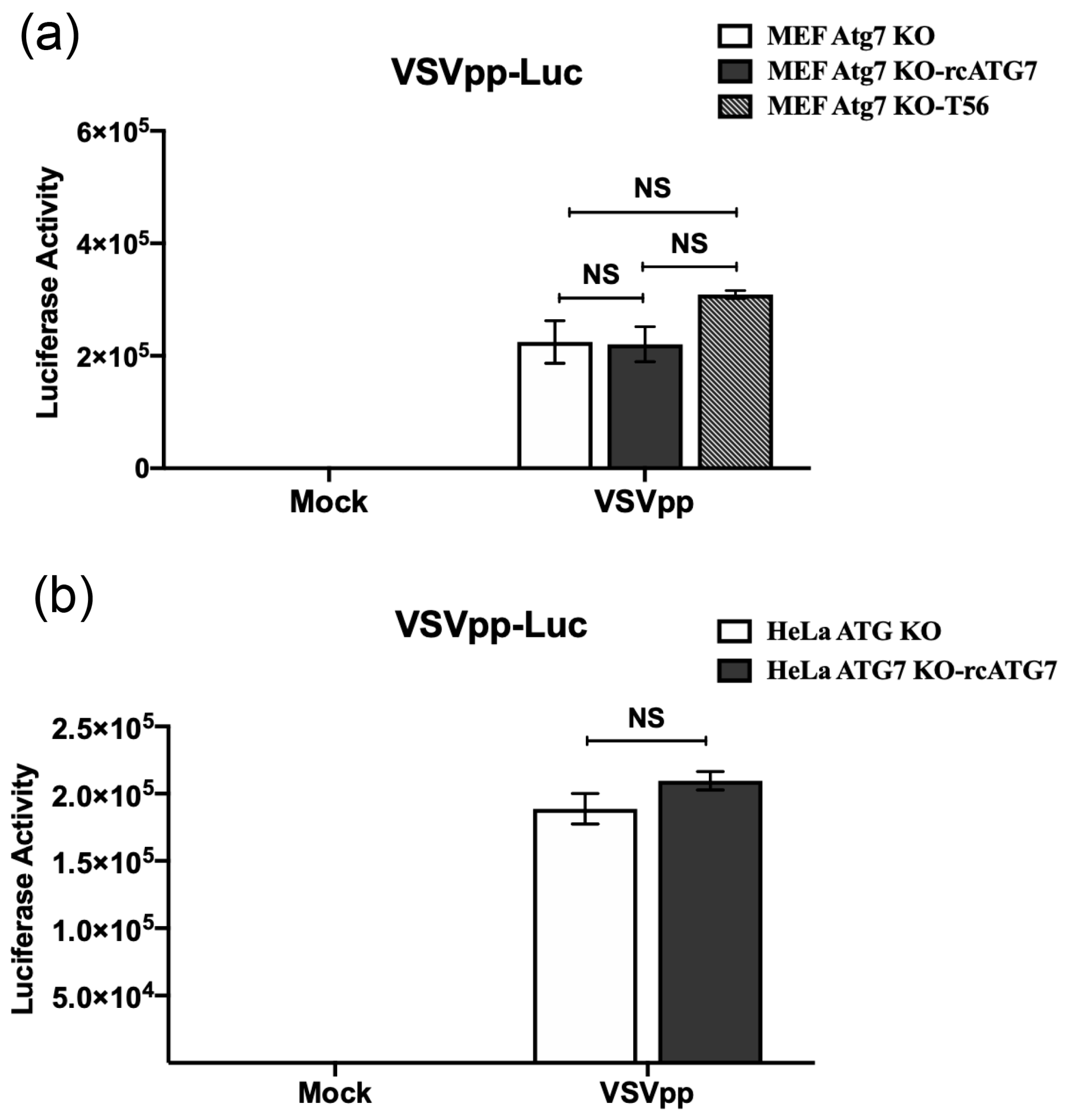
3.3. ATG7 Deficiency Does Not Affect Cellular Entry of VSV
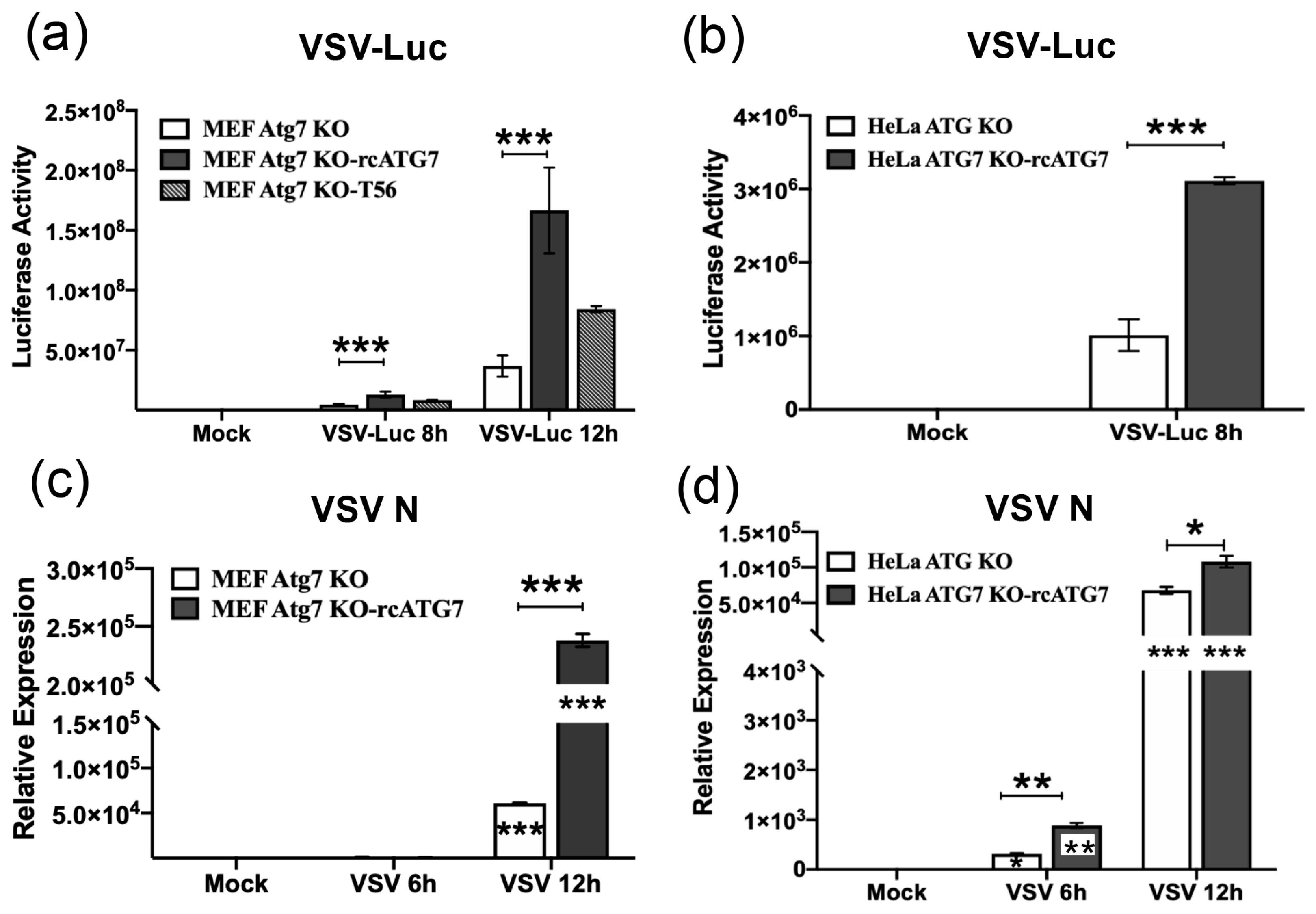
3.4. VSV RNA Replication Takes Place More Efficiently in ATG7-Expressing Cells
3.5. ATG7 Suppresses Baseline as Well as VSV-Induced ISG Expression in MEF- and HeLa-Based Models
3.6. ATG7 Deficiency Is Associated with Moderate Yet Significant Reduction in Cellular Permissiveness for VSV Replication in Huh7.5-TLR3 Cells, Which Are Defective in RIG-I Signaling
3.7. Greater Basal Expression of IFITs Is Correlated with Reduced VSV Replication in ATG7-Deficient Huh7.5-TLR3 Cells, Which Do Not Mount an Antiviral ISG Response to VSV Infection
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hale, A.N.; Ledbetter, D.J.; Gawriluk, T.R.; Rucker, I.; Edmund, B. Autophagy: Regulation and role in development. Autophagy 2013, 9, 951–972. [Google Scholar] [CrossRef]
- Lee, H.K.; Iwasaki, A. Autophagy and antiviral immunity. Curr. Opin. Immunol. 2008, 20, 23–29. [Google Scholar] [CrossRef]
- Chen, T.; Tu, S.; Ding, L.; Jin, M.; Chen, H.; Zhou, H. The role of autophagy in viral infections. J. Biomed. Sci. 2023, 30, 5. [Google Scholar] [CrossRef] [PubMed]
- Collier, J.J.; Suomi, F.; Olahova, M.; McWilliams, T.G.; Taylor, R.W. Emerging roles of ATG7 in human health and disease. EMBO Mol. Med. 2021, 13, e14824. [Google Scholar] [CrossRef] [PubMed]
- Palese, P.; Zheng, H.; Engelhardt, O.G.; Pleschka, S.; García-Sastre, A. Negative-strand RNA viruses: Genetic engineering and applications. Proc. Natl. Acad. Sci. USA 1996, 93, 11354–11358. [Google Scholar] [CrossRef]
- Burns, J.C.; Friedmann, T.; Driever, W.; Burrascano, M.; Yee, J.-K. Vesicular stomatitis virus G glycoprotein pseudotyped retroviral vectors: Concentration to very high titer and efficient gene transfer into mammalian and nonmammalian cells. Proc. Natl. Acad. Sci. USA 1993, 90, 8033–8037. [Google Scholar] [CrossRef]
- Barsoum, J.; Brown, R.; McKee, M.; Boyce, F.M. Efficient transduction of mammalian cells by a recombinant baculovirus having the vesicular stomatitis virus G glycoprotein. Hum. Gene Ther. 1997, 8, 2011–2018. [Google Scholar] [CrossRef] [PubMed]
- Letchworth, G.; Rodriguez, L.; Del Cbarrera, J. Vesicular stomatitis. Vet. J. 1999, 157, 239–260. [Google Scholar] [CrossRef]
- Whitt, M.A. Generation of VSV pseudotypes using recombinant DeltaG-VSV for studies on virus entry, identification of entry inhibitors, and immune responses to vaccines. J. Virol. Methods 2010, 169, 365–374. [Google Scholar] [CrossRef]
- Lester, S.N.; Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 2014, 426, 1246–1264. [Google Scholar] [CrossRef]
- Li, K.; Lee, J.Z. The regulation of innate antiviral immunity by TRIM56. Infect. Dis. Immun. 2025, 5, 120–126. [Google Scholar] [CrossRef]
- Bonjardim, C.A.; Ferreira, P.C.; Kroon, E.G. Interferons: Signaling, antiviral and viral evasion. Immunol. Lett. 2009, 122, 1–11. [Google Scholar] [CrossRef]
- Raftery, N.; Stevenson, N.J. Advances in anti-viral immune defence: Revealing the importance of the IFN JAK/STAT pathway. Cell. Mol. Life Sci. 2017, 74, 2525–2535. [Google Scholar] [CrossRef]
- Devaraj, S.G.; Wang, N.; Chen, Z.; Chen, Z.; Tseng, M.; Barretto, N.; Lin, R.; Peters, C.J.; Tseng, C.T.; Baker, S.C.; et al. Regulation of IRF-3-dependent innate immunity by the papain-like protease domain of the severe acute respiratory syndrome coronavirus. J. Biol. Chem. 2007, 282, 32208–32221. [Google Scholar] [CrossRef]
- Green, J.A.; Stanton, G.J.; Goode, J.; Baron, S. Vesicular stomatitis virus plaque production in monolayer cultures with liquid overlay medium: Description and adaptation to a one-day, human interferon-plaque. J. Clin. Microbiol. 1976, 4, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Jounai, N.; Takeshita, F.; Kobiyama, K.; Sawano, A.; Miyawaki, A.; Xin, K.-Q.; Ishii, K.J.; Kawai, T.; Akira, S.; Suzuki, K. The Atg5–Atg12 conjugate associates with innate antiviral immune responses. Proc. Natl. Acad. Sci. USA 2007, 104, 14050–14055. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.K.; Lund, J.M.; Ramanathan, B.; Mizushima, N.; Iwasaki, A. Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 2007, 315, 1398–1401. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Li, N.L.; Zeng, Y.; Liu, B.; Kumthip, K.; Wang, T.T.; Huo, D.; Ingels, J.F.; Lu, L.; Shang, J.; et al. The Molecular Chaperone GRP78 Contributes to Toll-like Receptor 3-mediated Innate Immune Response to Hepatitis C Virus in Hepatocytes. J. Biol. Chem. 2016, 291, 12294–12309. [Google Scholar] [CrossRef]
- Kumthip, K.; Yang, D.; Li, N.L.; Zhang, Y.; Fan, M.; Sethuraman, A.; Li, K. Pivotal role for the ESCRT-II complex subunit EAP30/SNF8 in IRF3-dependent innate antiviral defense. PLoS Pathog. 2017, 13, e1006713. [Google Scholar] [CrossRef]
- Wang, D.; Wang, R.; Li, K. Impaired Antiviral Responses to Extracellular Double-Stranded RNA and Cytosolic DNA, but Not to Interferon-α Stimulation, in TRIM56-Deficient Cells. Viruses 2022, 14, 89. [Google Scholar] [CrossRef]
- Li, Z.A.; Bajpai, A.K.; Wang, R.; Liu, Y.; Webby, R.J.; Wilk, E.; Gu, W.; Schughart, K.; Li, K.; Lu, L. Systems genetics of influenza A virus-infected mice identifies TRIM21 as a critical regulator of pulmonary innate immune response. Virus Res. 2024, 342, 199335. [Google Scholar] [CrossRef]
- Ahmed, M.; McKenzie, M.O.; Puckett, S.; Hojnacki, M.; Poliquin, L.; Lyles, D.S. Ability of the matrix protein of vesicular stomatitis virus to suppress beta interferon gene expression is genetically correlated with the inhibition of host RNA and protein synthesis. J. Virol. 2003, 77, 4646–4657. [Google Scholar] [CrossRef]
- Shelly, S.; Lukinova, N.; Bambina, S.; Berman, A.; Cherry, S. Autophagy is an essential component of Drosophila immunity against vesicular stomatitis virus. Immunity 2009, 30, 588–598. [Google Scholar] [CrossRef]
- Lamiable, O.; Arnold, J.; de Faria, I.; Olmo, R.P.; Bergami, F.; Meignin, C.; Hoffmann, J.A.; Marques, J.T.; Imler, J.L. Analysis of the Contribution of Hemocytes and Autophagy to Drosophila Antiviral Immunity. J. Virol. 2016, 90, 5415–5426. [Google Scholar] [CrossRef]
- Ogmundsdottir, M.H.; Fock, V.; Sooman, L.; Pogenberg, V.; Dilshat, R.; Bindesboll, C.; Ogmundsdottir, H.M.; Simonsen, A.; Wilmanns, M.; Steingrimsson, E. A short isoform of ATG7 fails to lipidate LC3/GABARAP. Sci. Rep. 2018, 8, 14391. [Google Scholar] [CrossRef]
- Akagi, T.; Sasai, K.; Hanafusa, H. Refractory nature of normal human diploid fibroblasts with respect to oncogene-mediated transformation. Proc. Natl. Acad. Sci. USA 2003, 100, 13567–13572. [Google Scholar] [CrossRef]
- Wang, J.; Liu, B.; Wang, N.; Lee, Y.M.; Liu, C.; Li, K. TRIM56 is a virus- and interferon-inducible E3 ubiquitin ligase that restricts pestivirus infection. J. Virol. 2011, 85, 3733–3745. [Google Scholar] [CrossRef] [PubMed]
- Khabir, M.; Aliche, A.Z.; Sureau, C.; Blanchet, M.; Labonte, P. Hepatitis Delta Virus Alters the Autophagy Process To Promote Its Genome Replication. J. Virol. 2020, 94, e01936-19. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Liang, Y.; Devaraj, S.; Wang, J.; Lemon, S.M.; Li, K. Toll-like receptor 3 mediates establishment of an antiviral state against hepatitis C virus in hepatoma cells. J. Virol. 2009, 83, 9824–9834. [Google Scholar] [CrossRef]
- Selleck, E.M.; Orchard, R.C.; Lassen, K.G.; Beatty, W.L.; Xavier, R.J.; Levine, B.; Virgin, H.W.; Sibley, L.D. A Noncanonical Autophagy Pathway Restricts Toxoplasma gondii Growth in a Strain-Specific Manner in IFN-gamma-Activated Human Cells. mBio 2015, 6, e01157-01115. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Rijnbrand, R.; Jangra, R.K.; Devaraj, S.G.; Qu, L.; Ma, Y.; Lemon, S.M.; Li, K. Ubiquitination and proteasomal degradation of interferon regulatory factor-3 induced by Npro from a cytopathic bovine viral diarrhea virus. Virology 2007, 366, 277–292. [Google Scholar] [CrossRef]
- Cureton, D.K.; Massol, R.H.; Saffarian, S.; Kirchhausen, T.L.; Whelan, S.P. Vesicular stomatitis virus enters cells through vesicles incompletely coated with clathrin that depend upon actin for internalization. PLoS Pathog. 2009, 5, e1000394. [Google Scholar] [CrossRef]
- Jayakar, H.R.; Whitt, M.A. Identification of two additional translation products from the matrix (M) gene that contribute to vesicular stomatitis virus cytopathology. J. Virol. 2002, 76, 8011–8018. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Qiu, C.; Biswas, N.; Jin, J.; Watkins, S.C.; Montelaro, R.C.; Coyne, C.B.; Wang, T. Correlation of the tight junction-like distribution of Claudin-1 to the cellular tropism of hepatitis C virus. J. Biol. Chem. 2008, 283, 8643–8653. [Google Scholar] [CrossRef]
- Wang, N.; Dong, Q.; Li, J.; Jangra, R.K.; Fan, M.; Brasier, A.R.; Lemon, S.M.; Pfeffer, L.M.; Li, K. Viral induction of the zinc finger antiviral protein is IRF3-dependent but NF-κB-independent. J. Biol. Chem. 2010, 285, 6080–6090. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Swaminathan, S. Human IFIT proteins inhibit lytic replication of KSHV: A new feed-forward loop in the innate immune system. PLoS Pathog. 2019, 15, e1007609. [Google Scholar] [CrossRef]
- Kim, D.; Rai, N.K.; Burrows, A.; Kim, S.; Tripathi, A.; Weinberg, S.E.; Dutta, R.; Sen, G.C.; Min, B. IFN-Induced Protein with Tetratricopeptide Repeats 2 Limits Autoimmune Inflammation by Regulating Myeloid Cell Activation and Metabolic Activity. J. Immunol. 2023, 210, 721–731. [Google Scholar] [CrossRef]
- Li, K. Regulation of interferon regulatory factor 3-dependent innate immunity by the HCV NS3/4A protease. Methods Mol. Biol. 2009, 510, 211–226. [Google Scholar] [CrossRef]
- Liu, H.S.; Wang, Y.P.; Lin, P.W.; Chu, M.L.; Lan, S.H.; Wu, S.Y.; Lee, Y.R.; Chang, H.Y. The role of Atg5 gene in tumorigenesis under autophagy deficiency conditions. Kaohsiung J. Med. Sci. 2024, 40, 631–641. [Google Scholar] [CrossRef]
- Neznanov, N.; Dragunsky, E.M.; Chumakov, K.M.; Neznanova, L.; Wek, R.C.; Gudkov, A.V.; Banerjee, A.K. Different effect of proteasome inhibition on vesicular stomatitis virus and poliovirus replication. PLoS ONE 2008, 3, e1887. [Google Scholar] [CrossRef]
- Liu, B.; Li, N.L.; Shen, Y.; Bao, X.; Fabrizio, T.; Elbahesh, H.; Webby, R.J.; Li, K. The C-Terminal Tail of TRIM56 Dictates Antiviral Restriction of Influenza A and B Viruses by Impeding Viral RNA Synthesis. J. Virol. 2016, 90, 4369–4382. [Google Scholar] [CrossRef]
- Shen, Y.; Li, N.L.; Wang, J.; Liu, B.; Lester, S.; Li, K. TRIM56 is an essential component of the TLR3 antiviral signaling pathway. J. Biol. Chem. 2012, 287, 36404–36413. [Google Scholar] [CrossRef]
- Lay Mendoza, M.F.; Acciani, M.D.; Levit, C.N.; Santa Maria, C.; Brindley, M.A. Monitoring Viral Entry in Real-Time Using a Luciferase Recombinant Vesicular Stomatitis Virus Producing SARS-CoV-2, EBOV, LASV, CHIKV, and VSV Glycoproteins. Viruses 2020, 12, 1457. [Google Scholar] [CrossRef]
- Glasner, D.R.; Todd, C.; Cook, B.; D’Urso, A.; Khosla, S.; Estrada, E.; Wagner, J.D.; Bartels, M.D.; Hung, C.T.; Ford, P.; et al. The IFIT2-IFIT3 antiviral complex targets short 5′ untranslated regions on viral mRNAs for translation inhibition. Nat. Microbiol. 2025, 10, 2934–2948. [Google Scholar] [CrossRef]
- Pichlmair, A.; Lassnig, C.; Eberle, C.A.; Gorna, M.W.; Baumann, C.L.; Burkard, T.R.; Burckstummer, T.; Stefanovic, A.; Krieger, S.; Bennett, K.L.; et al. IFIT1 is an antiviral protein that recognizes 5′-triphosphate RNA. Nat. Immunol. 2011, 12, 624–630. [Google Scholar] [CrossRef]
- Fensterl, V.; Wetzel, J.L.; Ramachandran, S.; Ogino, T.; Stohlman, S.A.; Bergmann, C.C.; Diamond, M.S.; Virgin, H.W.; Sen, G.C. Interferon-induced Ifit2/ISG54 protects mice from lethal VSV neuropathogenesis. PLoS Pathog. 2012, 8, e1002712. [Google Scholar] [CrossRef] [PubMed]
- Sumpter, R., Jr.; Loo, Y.M.; Foy, E.; Li, K.; Yoneyama, M.; Fujita, T.; Lemon, S.M.; Gale, M., Jr. Regulating intracellular antiviral defense and permissiveness to hepatitis C virus RNA replication through a cellular RNA helicase, RIG-I. J. Virol. 2005, 79, 2689–2699. [Google Scholar] [CrossRef] [PubMed]
- Nishida, Y.; Arakawa, S.; Fujitani, K.; Yamaguchi, H.; Mizuta, T.; Kanaseki, T.; Komatsu, M.; Otsu, K.; Tsujimoto, Y.; Shimizu, S. Discovery of Atg5/Atg7-independent alternative macroautophagy. Nature 2009, 461, 654–658. [Google Scholar] [CrossRef]
- Ahmad, S.; Mu, X.; Yang, F.; Greenwald, E.; Park, J.W.; Jacob, E.; Zhang, C.Z.; Hur, S. Breaching Self-Tolerance to Alu Duplex RNA Underlies MDA5-Mediated Inflammation. Cell 2018, 172, 797–810.e713. [Google Scholar] [CrossRef] [PubMed]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Tong, X.; Wang, R.; Liu, Y.; Potts, M.B.; Pruett-Miller, S.M.; Whitt, M.A.; Gu, W.; Li, K. ATG7 Limits Basal Antiviral Gene Expression and Moderately Promotes VSV Replication in Mammalian Non-Immune Cells. Pathogens 2026, 15, 404. https://doi.org/10.3390/pathogens15040404
Tong X, Wang R, Liu Y, Potts MB, Pruett-Miller SM, Whitt MA, Gu W, Li K. ATG7 Limits Basal Antiviral Gene Expression and Moderately Promotes VSV Replication in Mammalian Non-Immune Cells. Pathogens. 2026; 15(4):404. https://doi.org/10.3390/pathogens15040404
Chicago/Turabian StyleTong, Xiaohan, Ruixue Wang, Yaxin Liu, Malia B. Potts, Shondra M. Pruett-Miller, Michael A. Whitt, Weikuan Gu, and Kui Li. 2026. "ATG7 Limits Basal Antiviral Gene Expression and Moderately Promotes VSV Replication in Mammalian Non-Immune Cells" Pathogens 15, no. 4: 404. https://doi.org/10.3390/pathogens15040404
APA StyleTong, X., Wang, R., Liu, Y., Potts, M. B., Pruett-Miller, S. M., Whitt, M. A., Gu, W., & Li, K. (2026). ATG7 Limits Basal Antiviral Gene Expression and Moderately Promotes VSV Replication in Mammalian Non-Immune Cells. Pathogens, 15(4), 404. https://doi.org/10.3390/pathogens15040404






