Cross-Protection in PRRSV: Mechanisms, Limitations, and Implications for Vaccine Design
Abstract
1. Introduction
2. Cross-Species Protection Between PRRSV-1 and PRRSV-2: An Underappreciated Phenomenon
3. European PRRSV(PRRSV-1): Apparent Vaccine Efficacy and Hidden Gaps
| PRRSV-1 Lineage/Subtype | Representative Strain(s) | Geographic Distribution | Vaccine Anchor(s) | Homologous Protection | Heterologous Protection | Key Evidence Gaps | Key References |
|---|---|---|---|---|---|---|---|
| Subtype 1 (PRRSV-1-1) | Lelystad | Europe (widespread) | Lelystad-based MLVs | High (clinical & virological) | Partial (respiratory endpoints only) | Estimates confounded by low-virulence challenge strains; reproductive protection against heterologous strains largely untested | [46,48,49,50,52] |
| Subtype 1 (recent variants) | AUT15-33, PR40 | Europe (regional) | Subtype 1-based MLVs | Limited data | Partial (clinical only) | No systematic reproductive challenge data; strain-specific variability poorly defined | [54,55,58,62,63] |
| Subtype 2 (PRRSV-1-2) | WestSib13 | Eastern Europe | None | Not evaluated | Not evaluated | Absence of homologous and heterologous protection studies; very limited sequence availability | [41] |
| Subtype 3 (PRRSV-1-3) | Lena, SU1-Bel | Eastern Europe (experimental model) | None (commercial) | High after reinfection | Partial with subtype 1 MLVs (clinical only) | No Lena-based MLV; inactivated vaccines ineffective despite NAb induction | [40,57,58,60,61,62,63,64] |
| Autogenous PRRSV-1 vaccines | Farm-specific isolates | Europe | Inactivated autogenous vaccines | Variable (viremia reduction) | Limited | Protection is strain- and farm-specific; minimal cross-strain applicability | [65,66,67,68] |
4. North American PRRSV (PRRSV-2): High Diversity and Constrained Cross-Protection
| PRRSV-2 Lineage | Representative Strain(s) | Geographic Distribution | Vaccine Anchor(s) | Homologous Protection | Heterologous Protection | Key Evidence Gaps | Key References |
|---|---|---|---|---|---|---|---|
| Lineage 5 (L5) | VR-2332, NADC-8 | North America; global dissemination | VR-2332-based MLVs (e.g., Ingelvac PRRS MLV) | High (clinical & virological) | Variable across lineages | Little data against non-L5 historical isolates; limited relevance to current dominant strains | [47,93] |
| Lineage 1 (overall) | NADC-20, NADC-30, NA174 | North America, Asia | L1D-based MLV (Prevacent PRRS) | Not evaluated | Partial, strain-dependent | Dominant modern lineage lacks homologous challenge studies | [23,72,75] |
| Lineage 1C.5 | 1-4-4 L1C.5 | North America (dominant since 2019) | None | Not evaluated | Not evaluated | No homologous or controlled heterologous vaccine data despite high prevalence and virulence | [23,73] |
| Lineage 8 (HP-PRRSV) | JXA1, TJ, HuN4 | China, Southeast Asia | HP-PRRSV-based MLVs | High (homologous) | Partial against some L1 and L2 strains | Limited reproductive protection data; inconsistent outcomes across studies | [77,78,80,81,82,83,84,94] |
| Lineage 3 | QYYZ, SD53 | China, Taiwan | None (limited L7 vaccine data) | Not evaluated | Limited; poor neutralization by L5/L8 vaccines | High pathogenicity but scarce vaccine evaluation | [85,86,87,88,89,90,91] |
| Minor lineages (L2, L4, L6–L7, L9) | NADC-34, MN184A | Regional | L7-based MLV (limited use) | Limited data | Limited data | Sparse experimental evaluation; unclear epidemiological relevance | [91,92] |
5. Determinants of Protective Immunity
5.1. T-Cell-Mediated Immunity in Broad and Heterologous Protection Against PRRSV
5.2. Neutralizing Antibodies in PRRSV: Delayed Kinetics but Critical for Protection
6. Key Factors Modulating PRRSV Homologous and Heterologous Protection
6.1. Impact of Multiple Exposures and Booster Vaccination
6.2. Strain-Specific Effect: Pathogenicity, and Immune Induction
6.3. Structural Determinants, Complement, and Rational Immunogen Design
6.4. Vaccination Context and Host Factors
6.5. Preexisting Immunity to PRRSV: Potential Drawbacks
7. Summary and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Shi, M.; Lam, T.T.-Y.; Hon, C.-C.; Hui, R.K.-H.; Faaberg, K.S.; Wennblom, T.; Murtaugh, M.P.; Stadejek, T.; Leung, F.C.-C. Molecular epidemiology of PRRSV: A phylogenetic perspective. Virus Res. 2010, 154, 7–17. [Google Scholar] [CrossRef]
- Zhou, L.; Ge, X.; Yang, H. Porcine Reproductive and Respiratory Syndrome Modified Live Virus Vaccine: A “Leaky” Vaccine with Debatable Efficacy and Safety. Vaccines 2021, 9, 362. [Google Scholar] [CrossRef]
- Yim-Im, W.; Anderson, T.K.; Paploski, I.A.D.; VanderWaal, K.; Gauger, P.; Krueger, K.; Shi, M.; Main, R.; Zhang, J. Refining PRRSV-2 genetic classification based on global ORF5 sequences and investigation of their geographic distributions and temporal changes. Microbiol. Spectr. 2023, 11, e0291623. [Google Scholar] [CrossRef]
- Yim-Im, W.; Anderson, T.K.; Böhmer, J.; Baliellas, J.; Stadejek, T.; Gauger, P.C.; Krueger, K.M.; Vermeulen, C.J.; Buter, R.; Kazlouski, A.; et al. Refining genetic classification of global porcine reproductive and respiratory syndrome virus type 1 (PRRSV-1) and investigating their geographic and temporal distributions. Vet. Microbiol. 2025, 302, 110413. [Google Scholar] [CrossRef]
- Kick, A.R.; Grete, A.F.; Crisci, E.; Almond, G.W.; Käser, T. Testable Candidate Immune Correlates of Protection for Porcine Reproductive and Respiratory Syndrome Virus Vaccination. Vaccines 2023, 11, 594. [Google Scholar] [CrossRef]
- Sirisereewan, C.; Byrne, J.J.; Sulaiman, L.; Williams, A.; Hause, B.M.; Ferreira, J.B.; Almond, G.W.; Gabriel, B.; De Groot, A.S.; Käser, T.; et al. Establishment of Immune Biobank for Vaccine Immunogenicity Prediction Using In Vitro and In Silico Methods Against Porcine Reproductive and Respiratory Syndrome Virus. Vaccines 2025, 13, 1052. [Google Scholar] [CrossRef]
- Davis, S.K.; Jia, F.; Wright, Q.G.; Islam, M.T.; Bean, A.; Layton, D.; Williams, D.T.; Lynch, S.E. Defining correlates of protection for mammalian livestock vaccines against high-priority viral diseases. Front. Immunol. 2024, 15, 1397780. [Google Scholar] [CrossRef]
- Yoon, K.-J.; Zimmerman, J.J.; Swenson, S.L.; McGinley, M.J.; Eernisse, K.A.; Brevik, A.; Rhinehart, L.L.; Frey, M.L.; Hill, H.T.; Platt, K.B. Characterization of the humoral immune response to porcine reproductive and respiratory syndrome (PRRS) virus infection. J. Vet. Diagn. Investig. 1995, 7, 305–312. [Google Scholar] [CrossRef]
- Crisci, E.; Kick, A.R.; Cortes, L.M.; Byrne, J.J.; Amaral, A.F.; Love, K.; Tong, H.; Zhang, J.; Gauger, P.C.; Pittman, J.S.; et al. Challenges and lessons learned from a field trial on the understanding of the porcine respiratory disease complex. Vaccines 2025, 13, 740. [Google Scholar] [CrossRef]
- Kick, A.R.; Amaral, A.F.; Cortes, L.M.; Fogle, J.E.; Crisci, E.; Almond, G.W.; Käser, T. The T-Cell Response to Type 2 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV). Viruses 2019, 11, 796. [Google Scholar] [CrossRef]
- Mateu, E.; Diaz, I. The challenge of PRRS immunology. Vet. J. 2008, 177, 345–351. [Google Scholar] [CrossRef] [PubMed]
- van Woensel, P.A.M.; Liefkens, K.; Demaret, S. Effect on viraemia of an American and a European serotype PRRSV vaccine after challenge with European wild-type strains of the virus. Vet. Rec. 1998, 142, 510–512. [Google Scholar] [CrossRef]
- Labarque, G.; Van Gucht, S.; Van Reeth, K.; Nauwynck, H.; Pensaert, M. Respiratory tract protection upon challenge of pigs vaccinated with attenuated porcine reproductive and respiratory syndrome virus vaccines. Vet. Microbiol. 2003, 95, 187–197. [Google Scholar] [CrossRef]
- Bøtner, A.; Strandbygaard, B.; Sørensen, K.J.; Have, P.; Madsen, K.G.; Madsen, E.S.; Alexandersen, S. Appearance of acute PRRS-like symptoms in sow herds after vaccination with a modified live PRRS vaccine. Vet. Rec. 1997, 141, 497–499. [Google Scholar] [CrossRef] [PubMed]
- Balka, G.; Hornyák, Á.; Bálint, Á.; Kiss, I.; Kecskeméti, S.; Bakonyi, T.; Rusvai, M. Genetic diversity of porcine reproductive and respiratory syndrome virus strains circulating in Hungarian swine herds. Vet. Microbiol. 2008, 127, 128–135. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Stadejek, T.; Stankevicius, A.; Murtaugh, M.P.; Oleksiewicz, M.B. Molecular evolution of PRRSV in Europe: Current state of play. Vet. Microbiol. 2013, 165, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Jakab, S.; Kaszab, E.; Marton, S.; Bányai, K.; Bálint, Á.; Nemes, I.; Szabó, I. Genetic diversity of imported PRRSV-2 strains, 2005–2020, Hungary. Front. Vet. Sci. 2022, 9, 986850. [Google Scholar] [CrossRef]
- Raev, S.; Yuzhakov, A.; Bulgakov, A.; Kostina, L.; Gerasianinov, A.; Verkhovsky, O.; Zaberezhny, A.; Aliper, T. An Outbreak of a Respiratory Disorder at a Russian Swine Farm Associated with the Co-Circulation of PRRSV1 and PRRSV2. Viruses 2020, 12, 1169. [Google Scholar] [CrossRef]
- Kukushkin, A.S.; Baybikov, T.Z.; Scherbakov, A.V.; Timina, A.M.; Baborenko, E.P.; Puzankova, O.S.; Pronin, I.A.; Fomin, A.E. First Outbreak of Atypical Porcine Reproductive and Respiratory Syndrome in Russia Caused by Highly Pathogenic Chinese-Like Prrs Virus. In Proceedings of the NA PRRSV Symposium 2008, Chicago, IL, USA, 5–6 December 2008; pp. 23–26. [Google Scholar]
- Dewey, C.; Charbonneau, G.; Carman, S.; Hamel, A.; Nayar, G.; Friendship, R.; Eernisse, K.; Swenson, S. Lelystad-like strain of porcine reproductive and respiratory syndrome virus (PRRSV) identified in Canadian swine. Can. Vet. J. Rev. Vet. Can. 2000, 41, 493–494. [Google Scholar]
- Ropp, S.L.; Mahlum Wees, C.E.; Fang, Y.; Nelson, E.A.; Rossow, K.D.; Bien, M.; Arndt, B.; Preszler, S.; Steen, P.; Christopher-Hennings, J.; et al. Characterization of Emerging European-Like Porcine Reproductive and Respiratory Syndrome Virus Isolates in the United States. J. Virol. 2004, 78, 3684–3703. [Google Scholar] [CrossRef]
- Sun, Q.; Xu, H.; An, T.; Cai, X.; Tian, Z.; Zhang, H. Recent Progress in Studies of Porcine Reproductive and Respiratory Syndrome Virus 1 in China. Viruses 2023, 15, 1528. [Google Scholar] [CrossRef]
- Chandra, S.; Cezar, G.; Rupasinghe, K.; Magalhães, E.; Silva, G.S.; Almeida, M.; Crim, B.; Burrough, E.; Gauger, P.; Madson, D.; et al. Harnessing sequencing data for porcine reproductive and respiratory syndrome virus (PRRSV): Tracking genetic evolution dynamics and emerging sequences in US swine industry. Front. Vet. Sci. 2025, 12, 1571020. [Google Scholar] [CrossRef]
- Robinson, S.R.; Li, J.; Nelson, E.A.; Murtaugh, M.P. Broadly neutralizing antibodies against the rapidly evolving porcine reproductive and respiratory syndrome virus. Virus Res. 2015, 203, 56–65. [Google Scholar] [CrossRef]
- Han, K.; Seo, H.W.; Park, C.; Chae, C. Vaccination of sows against type 2 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) before artificial insemination protects against type 2 PRRSV challenge but does not protect against type 1 PRRSV challenge in late gestation. Vet. Res. 2014, 45, 12. [Google Scholar] [CrossRef]
- Han, K.; Seo, H.W.; Shin, J.H.; Oh, Y.; Kang, I.; Park, C.; Chae, C. Effect of the modified live porcine reproductive and respiratory syndrome virus (PRRSV) vaccine on European and North American PRRSV shedding in semen from infected boars. Clin. Vaccine Immunol. 2011, 18, 1600–1607. [Google Scholar] [CrossRef]
- Park, C.; Choi, K.; Jeong, J.; Chae, C. Cross-protection of a new type 2 porcine reproductive and respiratory syndrome virus (PRRSV) modified live vaccine (Fostera PRRS) against heterologous type 1 PRRSV challenge in growing pigs. Vet. Microbiol. 2015, 177, 87–94. [Google Scholar] [CrossRef]
- Oh, T.; Kim, H.; Park, K.H.; Yang, S.; Jeong, J.; Kim, S.; Kang, I.; Park, S.-J.; Chae, C. Comparison of 4 commercial modified-live porcine reproductive and respiratory syndrome virus (PRRSV) vaccines against heterologous Korean PRRSV-1 and PRRSV-2 challenge. Can. J. Vet. Res. 2019, 83, 57–67. [Google Scholar]
- Jeong, J.; Park, C.; Oh, T.; Park, K.H.; Yang, S.; Kang, I.; Park, S.-J.; Chae, C. Cross-protection of a modified-live porcine reproductive and respiratory syndrome virus (PRRSV)-2 vaccine against a heterologous PRRSV-1 challenge in late-term pregnancy gilts. Vet. Microbiol. 2018, 223, 119–125. [Google Scholar] [CrossRef]
- Lakshmanappa, Y.S.; Shang, P.; Renu, S.; Dhakal, S.; Hogshead, B.; Xiao, Y.; Wang, T.; Fang, Y.; Renukaradhya, G.J. Concurrent but consecutive vaccination of modified live PRRSV-1 and PRRSV-2 provides better protection in nursery pigs. Vet. Microbiol. 2025, 302, 110391. [Google Scholar] [CrossRef]
- Franzo, G.; Tucciarone, C.M.; Legnardi, M.; Melegari, I.; Berjaoui, S.; Gatta, G.; Poletto, F.; Valente, R.; Marcone, D.; Lorusso, A.; et al. Porcine reproductive and respiratory syndrome virus 2 in Europe: Neither wild nor tamed. Porc. Health Manag. 2025, 11, 62. [Google Scholar] [CrossRef]
- Martínez-Lobo, F.J.; Díez-Fuertes, F.; Segalés, J.; García-Artiga, C.; Simarro, I.; Castro, J.M.; Prieto, C. Comparative pathogenicity of type 1 and type 2 isolates of porcine reproductive and respiratory syndrome virus (PRRSV) in a young pig infection model. Vet. Microbiol. 2011, 154, 58–68. [Google Scholar] [CrossRef]
- Pan, J.; Zeng, M.; Zhao, M.; Huang, L. Research Progress on the detection methods of porcine reproductive and respiratory syndrome virus. Front. Microbiol. 2023, 14, 1097905. [Google Scholar] [CrossRef]
- Pamornchainavakul, N.; Paploski, I.A.D.; Makau, D.N.; Kikuti, M.; Rovira, A.; Lycett, S.; Corzo, C.A.; VanderWaal, K. Mapping the Dynamics of Contemporary PRRSV-2 Evolution and Its Emergence and Spreading Hotspots in the U.S. Using Phylogeography. Pathogens 2023, 12, 740. [Google Scholar] [CrossRef]
- Kim, T.; Park, C.; Choi, K.; Jeong, J.; Kang, I.; Park, S.-J.; Chae, C. Comparison of Two Commercial Type 1 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) Modified Live Vaccines against Heterologous Type 1 and Type 2 PRRSV Challenge in Growing Pigs. Clin. Vaccine Immunol. 2015, 22, 631–640. [Google Scholar] [CrossRef]
- Park, C.; Kim, T.; Choi, K.; Jeong, J.; Kang, I.; Park, S.-J.; Chae, C. Two Commercial Type 1 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV)-Modified Live Vaccines Reduce Seminal Shedding of Type 1 PRRSV but not Type 2 PRRSV in Infected Boars. Transbound. Emerg. Dis. 2017, 64, 194–203. [Google Scholar] [CrossRef]
- Choi, K.; Park, C.; Jeong, J.; Kang, I.; Park, S.-J.; Chae, C. Comparison of commercial type 1 and type 2 PRRSV vaccines against heterologous dual challenge. Vet. Rec. 2016, 178, 291. [Google Scholar] [CrossRef] [PubMed]
- Roca, M.; Gimeno, M.; Bruguera, S.; Segalés, J.; Díaz, I.; Galindo-Cardiel, I.J.; Martínez, E.; Darwich, L.; Fang, Y.; Maldonado, J.; et al. Effects of challenge with a virulent genotype II strain of porcine reproductive and respiratory syndrome virus on piglets vaccinated with an attenuated genotype I strain vaccine. Vet. J. 2012, 193, 92–96. [Google Scholar] [CrossRef] [PubMed]
- Weesendorp, E.; Rebel, J.M.J.; Popma-De Graaf, D.J.; Fijten, H.P.D.; Stockhofe-Zurwieden, N. Lung pathogenicity of European genotype 3 strain porcine reproductive and respiratory syndrome virus (PRRSV) differs from that of subtype 1 strains. Vet. Microbiol. 2014, 174, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Yuzhakov, A.G.; Raev, S.A.; Skrylev, A.N.; Mishin, A.M.; Grebennikova, T.V.; Verkhovsky, O.A.; Zaberezhny, A.D.; Trus, I.; Nauwynck, H.J.; Aliper, T.I. Genetic and pathogenic characterization of a Russian subtype 2 PRRSV-1 isolate. Vet. Microbiol. 2017, 211, 22–28. [Google Scholar] [CrossRef]
- Karniychuk, U.U.; Geldhof, M.; Vanhee, M.; Van Doorsselaere, J.; Saveleva, T.A.; Nauwynck, H.J. Pathogenesis and antigenic characterization of a new East European subtype 3 porcine reproductive and respiratory syndrome virus isolate. BMC Vet. Res. 2010, 6, 30. [Google Scholar] [CrossRef]
- Eck, M.; Durán, M.G.; Ricklin, M.E.; Locher, S.; Sarraseca, J.; Rodríguez, M.J.; McCullough, K.C.; Summerfield, A.; Zimmer, G.; Ruggli, N. Virus replicon particles expressing porcine reproductive and respiratory syndrome virus proteins elicit immune priming but do not confer protection from viremia in pigs. Vet. Res. 2016, 47, 33. [Google Scholar] [CrossRef] [PubMed]
- Balasch, M.; Fort, M.; Taylor, L.P.; Díaz, I.; Mateu, E.; Calvert, J.G. Immune response development after vaccination of 1-day-old naïve pigs with a Porcine Reproductive and Respiratory Syndrome 1-based modified live virus vaccine. Porc. Health Manag. 2019, 5, 2. [Google Scholar] [CrossRef]
- Yuzhakov, A.G.; Raev, S.A.; Shchetinin, A.M.; Gushchin, V.A.; Alekseev, K.P.; Stafford, V.V.; Komina, A.K.; Zaberezhny, A.D.; Gulyukin, A.M.; Aliper, T.I. Full-genome analysis and pathogenicity of a genetically distinct Russian PRRSV-1 Tyu16 strain. Vet. Microbiol. 2020, 247, 108784. [Google Scholar] [CrossRef] [PubMed]
- Genís, S.; Kvisgaard, L.K.; Larsen, L.E.; Taylor, L.P.; Calvert, J.G.; Balasch, M. Assessment of the Impact of the Recombinant Porcine Reproductive and Respiratory Syndrome Virus Horsens Strain on the Reproductive Performance in Pregnant Sows. Pathogens 2020, 9, 772. [Google Scholar] [CrossRef]
- Scortti, M.; Prieto, C.; Simarro, I.; Castro, J.M. Reproductive performance of gilts following vaccination and subsequent heterologous challenge with European strains of porcine reproductive and respiratory syndrome virus. Theriogenology 2006, 66, 1884–1893. [Google Scholar] [CrossRef] [PubMed]
- Lager, K.M.; Mengeling, W.L.; Brockmeier, S.L. Evaluation of protective immunity in gilts inoculated with the NADC-8 isolate of porcine reproductive and respiratory syndrome virus (PRRSV) and challenge-exposed with an antigenically distinct PRRSV isolate. Am. J. Vet. Res. 1999, 60, 1022–1027. [Google Scholar] [CrossRef]
- Labarque, G.; Reeth, K.V.; Nauwynck, H.; Drexler, C.; Gucht, S.V.; Pensaert, M. Impact of genetic diversity of European-type porcine reproductive and respiratory syndrome virus strains on vaccine efficacy. Vaccine 2004, 22, 4183–4190. [Google Scholar] [CrossRef]
- Prieto, C.; Álvarez, E.; Martínez-Lobo, F.J.; Simarro, I.; Castro, J.M. Similarity of European porcine reproductive and respiratory syndrome virus strains to vaccine strain is not necessarily predictive of the degree of protective immunity conferred. Vet. J. 2008, 175, 356–363. [Google Scholar] [CrossRef]
- Kroll, J.; Piontkowski, M.; Rathkjen, P.H.; Orveillon, F.-X.; Kraft, C.; Duran, O.G. Long duration of immunity against a type 1 heterologous PRRS virus challenge in pigs immunised with a novel PRRS MLV vaccine: A randomised controlled study. Porc. Health Manag. 2018, 4, 11. [Google Scholar] [CrossRef]
- Geldhof, M.F.; Vanhee, M.; Van Breedam, W.; Van Doorsselaere, J.; Karniychuk, U.U.; Nauwynck, H.J. Comparison of the efficacy of autogenous inactivated Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) vaccines with that of commercial vaccines against homologous and heterologous challenges. BMC Vet. Res. 2012, 8, 182. [Google Scholar] [CrossRef]
- Kreutzmann, H.; Stadler, J.; Knecht, C.; Sassu, E.L.; Ruczizka, U.; Zablotski, Y.; Vatzia, E.; Balka, G.; Zaruba, M.; Chen, H.-W.; et al. Phenotypic Characterization of a Virulent PRRSV-1 Isolate in a Reproductive Model With and Without Prior Heterologous Modified Live PRRSV-1 Vaccination. Front. Vet. Sci. 2022, 9, 820233. [Google Scholar] [CrossRef] [PubMed]
- Scortti, M.; Prieto, C.; Alvarez, E.; Simarro, I.; Castro, J.M. Failure of an inactivated vaccine against porcine reproductive and respiratory syndrome to protect gilts against a heterologous challenge with PRRSV. Vet. Rec. 2007, 161, 809–813. [Google Scholar] [PubMed]
- Canelli, E.; Catella, A.; Borghetti, P.; Ferrari, L.; Ogno, G.; De Angelis, E.; Bonilauri, P.; Guazzetti, S.; Nardini, R.; Martelli, P. Efficacy of a modified-live virus vaccine in pigs experimentally infected with a highly pathogenic porcine reproductive and respiratory syndrome virus type 1 (HP-PRRSV-1). Vet. Microbiol. 2018, 226, 89–96. [Google Scholar] [CrossRef]
- Kreutzmann, H.; Dürlinger, S.; Knecht, C.; Koch, M.; Cabana, M.; Torrent, G.; Balasch, M.; Taylor, L.P.; Balka, G.; Gerner, W.; et al. Efficacy of a Modified Live Virus Vaccine against Porcine Reproductive and Respiratory Syndrome Virus 1 (PRRSV-1) Administered to 1-Day-Old Piglets in Front of Heterologous PRRSV-1 Challenge. Pathogens 2021, 10, 1342. [Google Scholar] [CrossRef] [PubMed]
- Mateu, E.; Cortey, M.; Serena, M.S.; Domingo-Carreño, I.; Alberch, M.; Aguirre, L.; Diaz, I.; Martín, M.; Sanchez-Carvajal, J.M.; Rodriguez-Gomez, I.M.; et al. Efficacy of an intranasally administered live attenuated PRRSV-2 vaccine against challenge with a highly virulent PRRSV-1 strain. Front. Vet. Sci. 2025, 12, 1619052. [Google Scholar] [CrossRef]
- Morgan, S.B.; Graham, S.P.; Salguero, F.J.; Sánchez Cordón, P.J.; Mokhtar, H.; Rebel, J.M.J.; Weesendorp, E.; Bodman-Smith, K.B.; Steinbach, F.; Frossard, J.P. Increased pathogenicity of European porcine reproductive and respiratory syndrome virus is associated with enhanced adaptive responses and viral clearance. Vet. Microbiol. 2013, 163, 13–22. [Google Scholar] [CrossRef]
- Bonckaert, C.; van der Meulen, K.; Rodríguez-Ballarà, I.; Pedrazuela Sanz, R.; Martinez, M.F.; Nauwynck, H.J. Modified-live PRRSV subtype 1 vaccine UNISTRAIN® PRRS provides a partial clinical and virological protection upon challenge with East European subtype 3 PRRSV strain Lena. Porc. Health Manag. 2016, 2, 12. [Google Scholar] [CrossRef]
- Renson, P.; Fablet, C.; Le Dimna, M.; Mahé, S.; Touzain, F.; Blanchard, Y.; Paboeuf, F.; Rose, N.; Bourry, O. Preparation for emergence of an Eastern European porcine reproductive and respiratory syndrome virus (PRRSV) strain in Western Europe: Immunization with modified live virus vaccines or a field strain confers partial protection. Vet. Microbiol. 2017, 204, 133–140. [Google Scholar] [CrossRef]
- Weesendorp, E.; Stockhofe-Zurwieden, N.; Nauwynck, H.J.; Popma-De Graaf, D.J.; Rebel, J.M.J. Characterization of immune responses following homologous reinfection of pigs with European subtype 1 and 3 porcine reproductive and respiratory syndrome virus strains that differ in virulence. Vet. Microbiol. 2016, 182, 64–74. [Google Scholar] [CrossRef]
- Sautter, C.A.; Trus, I.; Nauwynck, H.; Summerfield, A. No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome. Viruses 2019, 11, 829. [Google Scholar] [CrossRef]
- Trus, I.; Bonckaert, C.; van der Meulen, K.; Nauwynck, H.J. Efficacy of an attenuated European subtype 1 porcine reproductive and respiratory syndrome virus (PRRSV) vaccine in pigs upon challenge with the East European subtype 3 PRRSV strain Lena. Vaccine 2014, 32, 2995–3003. [Google Scholar] [CrossRef] [PubMed]
- Trus, I.; Frydas, I.S.; Reddy, V.R.A.P.; Bonckaert, C.; Li, Y.; Kvisgaard, L.K.; Larsen, L.E.; Nauwynck, H.J. Immunity raised by recent European subtype 1 PRRSV strains allows better replication of East European subtype 3 PRRSV strain Lena than that raised by an older strain. Vet. Res. 2016, 47, 15. [Google Scholar] [CrossRef]
- Stadejek, T.; Larsen, L.E.; Podgórska, K.; Bøtner, A.; Botti, S.; Dolka, I.; Fabisiak, M.; Heegaard, P.M.H.; Hjulsager, C.K.; Huć, T.; et al. Pathogenicity of three genetically diverse strains of PRRSV Type 1 in specific pathogen free pigs. Vet. Microbiol. 2017, 209, 13–19. [Google Scholar] [CrossRef]
- Kick, A.R.; Wolfe, Z.C.; Amaral, A.F.; Cortes, L.M.; Almond, G.W.; Crisci, E.; Gauger, P.C.; Pittman, J.; Käser, T. Maternal Autogenous Inactivated Virus Vaccination Boosts Immunity to PRRSV in Piglets. Vaccines 2021, 9, 106. [Google Scholar] [CrossRef] [PubMed]
- Geldhof, M.F.; Van Breedam, W.; De Jong, E.; Lopez Rodriguez, A.; Karniychuk, U.U.; Vanhee, M.; Van Doorsselaere, J.; Maes, D.; Nauwynck, H.J. Antibody response and maternal immunity upon boosting PRRSV-immune sows with experimental farm-specific and commercial PRRSV vaccines. Vet. Microbiol. 2013, 167, 260–271. [Google Scholar] [CrossRef]
- Vanhee, M.; Delputte, P.L.; Delrue, I.; Geldhof, M.F.; Nauwynck, H.J. Development of an experimental inactivated PRRSV vaccine that induces virus-neutralizing antibodies. Vet. Res. 2009, 40, 63. [Google Scholar] [CrossRef]
- Pertich, A.; Barna, Z.; Makai, O.; Farkas, J.; Molnár, T.; Bálint, Á.; Szabó, I.; Albert, M. Elimination of porcine reproductive and respiratory syndrome virus infection using an inactivated vaccine in combination with a roll-over method in a Hungarian large-scale pig herd. Acta Vet. Scand. 2022, 64, 12. [Google Scholar] [CrossRef]
- Zhu, D.; Liu, G.; Li, H.; Li, F.; Xu, X.; Fu, Y.; Chen, P.; Zhang, G.; Sun, Y. Molecular Epidemiology, Lineage Evolutionary Dynamics, and Antigenic Variation Analysis of Type II PRRSV in China During 2024–2025. Transbound. Emerg. Dis. 2025, 2025, 2054759. [Google Scholar] [CrossRef]
- Shi, M.; Lam, T.T.-Y.; Hon, C.-C.; Murtaugh, M.P.; Davies, P.R.; Hui, R.K.-H.; Li, J.; Wong, L.T.-W.; Yip, C.-W.; Jiang, J.-W.; et al. Phylogeny-Based Evolutionary, Demographical, and Geographical Dissection of North American Type 2 Porcine Reproductive and Respiratory Syndrome Viruses. J. Virol. 2010, 84, 8700–8711. [Google Scholar] [CrossRef]
- Li, X.; Galliher-Beckley, A.; Pappan, L.; Trible, B.; Kerrigan, M.; Beck, A.; Hesse, R.; Blecha, F.; Nietfeld, J.C.; Rowland, R.R.; et al. Comparison of host immune responses to homologous and heterologous type II porcine reproductive and respiratory syndrome virus (PRRSV) challenge in vaccinated and unvaccinated pigs. BioMed Res. Int. 2014, 2014, 416727. [Google Scholar] [CrossRef] [PubMed]
- Paploski, I.A.D.; Pamornchainavakul, N.; Makau, D.N.; Rovira, A.; Corzo, C.A.; Schroeder, D.C.; Cheeran, M.C.-J.; Doeschl-Wilson, A.; Kao, R.R.; Lycett, S.; et al. Phylogenetic Structure and Sequential Dominance of Sub-Lineages of PRRSV Type-2 Lineage 1 in the United States. Vaccines 2021, 9, 608. [Google Scholar] [CrossRef] [PubMed]
- Wiarda, J.E.; Anderson, S.J.; Kim, H.; Chang, T.; Hanson, L.T.; Zanella, E.L.; Arruda, B.; Brand, M.W.; Hau, S.J.; Zhang, J.; et al. United States PRRSV 1-4-4 L1C.5 isolate demonstrates similar pathogenicity to a historic Chinese highly pathogenic PRRSV. Virology 2026, 615, 110748. [Google Scholar] [CrossRef]
- Trevisan, G.; Sparks, J.; Zeller, M.; Tong, H.; Li, G.; Zhang, J.; Gauger, P.C.; Rademacher, C.; Main, R.; Silva, A.P.P.; et al. Emergence of a PRRSV Strain Recombined From Two Modified-Live Virus Vaccines and Its Elimination From a Breeding Herd. Transbound. Emerg. Dis. 2025, 2025, 5770608. [Google Scholar] [CrossRef]
- Proctor, J.; Wolf, I.; Brodsky, D.; Cortes, L.M.; Frias-De-Diego, A.; Almond, G.W.; Crisci, E.; Negrão Watanabe, T.T.; Hammer, J.M.; Käser, T. Heterologous vaccine immunogenicity, efficacy, and immune correlates of protection of a modified-live virus porcine reproductive and respiratory syndrome virus vaccine. Front. Microbiol. 2022, 13, 977796. [Google Scholar] [CrossRef]
- Ruedas-Torres, I.; Rodríguez-Gómez, I.M.; Sánchez-Carvajal, J.M.; Larenas-Muñoz, F.; Pallarés, F.J.; Carrasco, L.; Gómez-Laguna, J. The jigsaw of PRRSV virulence. Vet. Microbiol. 2021, 260, 109168. [Google Scholar] [CrossRef]
- Yu, X.; Zhou, Z.; Cao, Z.; Wu, J.; Zhang, Z.; Xu, B.; Wang, C.; Hu, D.; Deng, X.; Han, W.; et al. Assessment of the Safety and Efficacy of an Attenuated Live Vaccine Based on Highly Pathogenic Porcine Reproductive and Respiratory Syndrome Virus. Clin. Vaccine Immunol. 2015, 22, 493–502. [Google Scholar] [CrossRef]
- Leng, X.; Li, Z.; Xia, M.; He, Y.; Wu, H. Evaluation of the Efficacy of an Attenuated Live Vaccine against Highly Pathogenic Porcine Reproductive and Respiratory Syndrome Virus in Young Pigs. Clin. Vaccine Immunol. 2012, 19, 1199–1206. [Google Scholar] [CrossRef]
- Madapong, A.; Saeng-chuto, K.; Boonsoongnern, A.; Tantituvanont, A.; Nilubol, D. Cell-mediated immune response and protective efficacy of porcine reproductive and respiratory syndrome virus modified-live vaccines against co-challenge with PRRSV-1 and PRRSV-2. Sci. Rep. 2020, 10, 1649. [Google Scholar] [CrossRef]
- Galliher-Beckley, A.; Li, X.; Bates, J.T.; Madera, R.; Waters, A.; Nietfeld, J.; Henningson, J.; He, D.; Feng, W.; Chen, R.; et al. Pigs immunized with Chinese highly pathogenic PRRS virus modified live vaccine are protected from challenge with North American PRRSV strain NADC-20. Vaccine 2015, 33, 3518–3525. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, X.; Guo, T.; Qiao, S.; Guo, Z.; Li, R.; Jin, Q.; Hu, X.; Xing, G.; Deng, R.; et al. Efficacy of a live attenuated highly pathogenic PRRSV vaccine against a NADC30-like strain challenge: Implications for ADE of PRRSV. BMC Vet. Res. 2021, 17, 260. [Google Scholar] [CrossRef]
- Zhou, L.; Yang, B.; Xu, L.; Jin, H.; Ge, X.; Guo, X.; Han, J.; Yang, H. Efficacy evaluation of three modified-live virus vaccines against a strain of porcine reproductive and respiratory syndrome virus NADC30-like. Vet. Microbiol. 2017, 207, 108–116. [Google Scholar] [CrossRef]
- Zhang, H.; Xia, M.; Wang, W.; Ju, D.; Cao, L.; Wu, B.; Wang, X.; Wu, Y.; Song, N.; Hu, J.; et al. An Attenuated Highly Pathogenic Chinese PRRS Viral Vaccine Confers Cross Protection to Pigs against Challenge with the Emerging PRRSV NADC30-Like Strain. Virol. Sin. 2018, 33, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Li, Z.; Li, J.; Kong, Y.; Yang, L.; Mah, C.K.; Liu, G.; Yu, B.; Wang, K. Efficacy evaluation of three modified-live PRRS vaccines against a local strain of highly pathogenic porcine reproductive and respiratory syndrome virus. Vet. Microbiol. 2019, 229, 117–123. [Google Scholar] [CrossRef] [PubMed]
- Kuo, K.-L.; Lin, W.-H.; Chiou, M.-T.; Zhang, J.; Lin, C.-N. Co-circulation of lineage 1 and lineage 3 porcine reproductive and respiratory syndrome virus type 2 (PRRSV-2) in Taiwan during 2018–2024. Vet. Microbiol. 2025, 306, 110567. [Google Scholar] [CrossRef]
- Tian, L.; Wang, X.; Rong, Z.; Zhang, M.; Sun, Y.; Zhou, R.; Ma, J.; Zhang, C.; Liu, S.; Cao, N.; et al. Pathogenic and transcriptomic differences among porcine reproductive and respiratory syndrome viruses from distinct lineages in piglets. Vet. Res. 2025, 56, 224. [Google Scholar] [CrossRef]
- Zhou, L.; Kang, R.; Zhang, Y.; Yu, J.; Xie, B.; Chen, C.; Li, X.; Chen, B.; Liang, L.; Zhu, J.; et al. Emergence of two novel recombinant porcine reproductive and respiratory syndrome viruses 2 (lineage 3) in Southwestern China. Vet. Microbiol. 2019, 232, 30–41. [Google Scholar] [CrossRef]
- Han, G.; Xu, H.; Wang, Y.; Liu, Z.; He, F. Efficacy Evaluation of Two Commercial Vaccines Against a Recombinant PRRSV2 Strain ZJnb16-2 From Lineage 8 and 3 in China. Pathogens 2020, 9, 59. [Google Scholar] [CrossRef]
- Han, G.; Xu, H.; Wang, K.; He, F. Emergence of Two different recombinant PRRSV strains with low neutralizing antibody susceptibility in China. Sci. Rep. 2019, 9, 2490. [Google Scholar] [CrossRef]
- Hou, F.-H.; Lee, W.-C.; Liao, J.-W.; Chien, M.-S.; Kuo, C.-J.; Chung, H.-P.; Chia, M.-Y. Evaluation of a type 2 modified live porcine reproductive and respiratory syndrome vaccine against heterologous challenge of a lineage 3 highly virulent isolate in pigs. PeerJ 2020, 8, e8840. [Google Scholar] [CrossRef]
- Chiu, H.-J.; Chang, S.-W.; Lin, H.; Chuang, Y.-C.; Kuo, K.-L.; Lin, C.-H.; Chiou, M.-T.; Lin, C.-N. Lineage 7 Porcine Reproductive and Respiratory Syndrome Vaccine Demonstrates Cross-Protection Against Lineage 1 and Lineage 3 Strains. Vaccines 2025, 13, 102. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; Wei, Z.; Khan, M.; Zhou, Z.; Zhang, J.; Huang, X.; Yang, Y.; Wang, S.; Wang, H.; Cai, X.; et al. Refining Lineage Classification and Updated RFLP Patterns of PRRSV-2 Revealed Viral Spatiotemporal Distribution Characteristics in China in 1991–2023. Transbound. Emerg. Dis. 2025, 2025, 9977088. [Google Scholar] [CrossRef]
- Zuckermann, F.A.; Garcia, E.A.; Luque, I.D.; Christopher-Hennings, J.; Doster, A.; Brito, M.; Osorio, F. Assessment of the efficacy of commercial porcine reproductive and respiratory syndrome virus (PRRSV) vaccines based on measurement of serologic response, frequency of gamma-IFN-producing cells and virological parameters of protection upon challenge. Vet. Microbiol. 2007, 123, 69–85. [Google Scholar] [CrossRef]
- Tian, Z.-J.; An, T.-Q.; Zhou, Y.-J.; Peng, J.-M.; Hu, S.-P.; Wei, T.-C.; Jiang, Y.-F.; Xiao, Y.; Tong, G.-Z. An attenuated live vaccine based on highly pathogenic porcine reproductive and respiratory syndrome virus (HP-PRRSV) protects piglets against HP-PRRS. Vet. Microbiol. 2009, 138, 34–40. [Google Scholar] [CrossRef]
- Chung, C.J.; Cha, S.-H.; Grimm, A.L.; Chung, G.; Gibson, K.A.; Yoon, K.-J.; Parish, S.M.; Ho, C.-S.; Lee, S.S. Recognition of Highly Diverse Type-1 and -2 Porcine Reproductive and Respiratory Syndrome Viruses (PRRSVs) by T-Lymphocytes Induced in Pigs after Experimental Infection with a Type-2 PRRSV Strain. PLoS ONE 2016, 11, e0165450. [Google Scholar] [CrossRef]
- Correas, I.; Osorio, F.A.; Steffen, D.; Pattnaik, A.K.; Vu, H.L.X. Cross reactivity of immune responses to porcine reproductive and respiratory syndrome virus infection. Vaccine 2017, 35, 782–788. [Google Scholar] [CrossRef]
- Díaz, I.; Gimeno, M.; Darwich, L.; Navarro, N.; Kuzemtseva, L.; López, S.; Galindo, I.; Segalés, J.; Martín, M.; Pujols, J.; et al. Characterization of homologous and heterologous adaptive immune responses in porcine reproductive and respiratory syndrome virus infection. Vet. Res. 2012, 43, 30. [Google Scholar] [CrossRef]
- Li, Y.; Díaz, I.; Martín-Valls, G.; Beyersdorf, N.; Mateu, E. Systemic CD4 cytotoxic T cells improve protection against PRRSV-1 transplacental infection. Front. Immunol. 2023, 13, 1020227. [Google Scholar] [CrossRef] [PubMed]
- Chung, C.J.; Cha, S.-H.; Grimm, A.L.; Ajithdoss, D.; Rzepka, J.; Chung, G.; Yu, J.; Davis, W.C.; Ho, C.-S. Pigs that recover from porcine reproduction and respiratory syndrome virus infection develop cytotoxic CD4+CD8+ and CD4+CD8- T-cells that kill virus infected cells. PLoS ONE 2018, 13, e0203482. [Google Scholar] [CrossRef]
- Lager, K.M.; Mengeling, W.L.; Brockmeier, S.L. Duration of homologous porcine reproductive and respiratory syndrome virus immunity in pregnant swine. Vet. Microbiol. 1997, 58, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Meier, W.A.; Galeota, J.; Osorio, F.A.; Husmann, R.J.; Schnitzlein, W.M.; Zuckermann, F.A. Gradual development of the interferon-γ response of swine to porcine reproductive and respiratory syndrome virus infection or vaccination. Virology 2003, 309, 18–31. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, L.; Martelli, P.; Saleri, R.; De Angelis, E.; Cavalli, V.; Bresaola, M.; Benetti, M.; Borghetti, P. Lymphocyte activation as cytokine gene expression and secretion is related to the porcine reproductive and respiratory syndrome virus (PRRSV) isolate after in vitro homologous and heterologous recall of peripheral blood mononuclear cells (PBMC) from pigs vaccinated and exposed to natural infection. Vet. Immunol. Immunopathol. 2013, 151, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Hammer, J.M.; Gutierrez, A.H.; Huntimer, L.; Gabriel, B.; Martin, W.D.; Hammer, S.E.; Käser, T.; De Groot, A.S. T cell epitope content comparison using EpiCC correlates with vaccine efficacy against heterologous porcine reproductive and respiratory syndrome virus type 2 strains. Front. Microbiol. 2025, 16, 1625309. [Google Scholar] [CrossRef] [PubMed]
- Bordet, E.; Blanc, F.; Tiret, M.; Crisci, E.; Bouguyon, E.; Renson, P.; Maisonnasse, P.; Bourge, M.; Leplat, J.-J.; Giuffra, E.; et al. Porcine Reproductive and Respiratory Syndrome Virus Type 1.3 Lena Triggers Conventional Dendritic Cells 1 Activation and T Helper 1 Immune Response Without Infecting Dendritic Cells. Front. Immunol. 2018, 9, 2299. [Google Scholar] [CrossRef]
- Li, X.; Galliher-Beckley, A.; Wang, L.; Nietfeld, J.; Feng, W.; Shi, J. Comparison of Immune Responses in Pigs Infected with Chinese Highly Pathogenic PRRS Virus Strain HV and North American Strain NADC-20. Open Virol. J. 2017, 11, 73–82. [Google Scholar] [CrossRef]
- Zhang, W.; Wang, X.; Zhang, H.; Pan, Y.; Ma, W.; Xu, Y.; Tian, Z.; Xia, C.; Fu, L.; Wang, Y. Comparison of pathogenicity and host responses of emerging porcine reproductive and respiratory syndrome virus variants in piglets. J. Virol. 2024, 98, e01542-23. [Google Scholar] [CrossRef]
- Charerntantanakul, W.; Pongjaroenkit, S. Co-administration of saponin quil A and PRRSV-1 modified-live virus vaccine up-regulates gene expression of type I interferon-regulated gene, type I and II interferon, and inflammatory cytokines and reduces viremia in response to PRRSV-2 challenge. Vet. Immunol. Immunopathol. 2018, 205, 24–34. [Google Scholar] [CrossRef]
- Plagemann, P.G.W. Neutralizing Antibody Formation in Swine Infected with Seven Strains of Porcine Reproductive and Respiratory Syndrome Virus as Measured by Indirect ELISA with Peptides Containing the GP5 Neutralization Epitope. Viral Immunol. 2006, 19, 285–293. [Google Scholar] [CrossRef]
- Li, X.; Pei, Z.; Bai, Y.; Wang, L.; Shi, J.; Tian, K. Phenotypic characterization of porcine IFNγ-producing lymphocytes in porcine reproductive and respiratory syndrome virus vaccinated and challenged pigs. Virol. Sin. 2018, 33, 524–530. [Google Scholar] [CrossRef]
- Chen, X.; Zhao, J.; Ji, P.; Li, X.; Niu, H.; Jiao, D.; Zhang, L.; Zhu, Q.; Liu, X.; Hiscox, J.A.; et al. A porcine monoclonal antibody targeting a conserved GP4 epitope protects against In vivo infection via the induction of broad-spectrum PRRSV neutralization. Adv. Sci. 2025, 12, e08875. [Google Scholar] [CrossRef]
- Weiland, E.; Wieczorek-Krohmer, M.; Kohl, D.; Conzelmann, K.K.; Weiland, F. Monoclonal antibodies to the GP5 of porcine reproductive and respiratory syndrome virus are more effective in virus neutralization than monoclonal antibodies to the GP4. Vet. Microbiol. 1999, 66, 171–186. [Google Scholar] [CrossRef] [PubMed]
- Burkard, C.; Opriessnig, T.; Mileham, A.J.; Stadejek, T.; Ait-Ali, T.; Lillico, S.G.; Whitelaw, C.B.A.; Archibald, A.L. Pigs lacking the scavenger receptor cysteine-rich domain 5 of CD163 are resistant to porcine reproductive and respiratory syndrome virus 1 infection. J. Virol. 2018, 92, e00415-18. [Google Scholar] [CrossRef]
- Xu, H.; Liu, Z.; Zheng, S.; Han, G.; He, F. CD163 antibodies inhibit PRRSV infection via receptor blocking and transcription suppression. Vaccines 2020, 8, 592. [Google Scholar] [CrossRef]
- Cancel-Tirado, S.M.; Evans, R.B.; Yoon, K.-J. Monoclonal antibody analysis of porcine reproductive and respiratory syndrome virus epitopes associated with antibody-dependent enhancement and neutralization of virus infection. Vet. Immunol. Immunopathol. 2004, 102, 249–262. [Google Scholar] [CrossRef]
- Delputte, P.L.; Meerts, P.; Costers, S.; Nauwynck, H.J. Effect of virus-specific antibodies on attachment, internalization and infection of porcine reproductive and respiratory syndrome virus in primary macrophages. Vet. Immunol. Immunopathol. 2004, 102, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.R.; Rahe, M.C.; Gray, D.K.; Martins, K.V.; Murtaugh, M.P. Porcine reproductive and respiratory syndrome virus neutralizing antibodies provide in vivo cross-protection to PRRSV1 and PRRSV2 viral challenge. Virus Res. 2018, 248, 13–23. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhai, T.; Li, M.; Zhang, K.; Li, J.; Zheng, X.; Tian, C.; Chen, R.; Dong, J.; Zhou, E.-M.; et al. A broadly neutralizing monoclonal antibody induces broad protection against heterogeneous PRRSV strains in piglets. Vet. Res. 2021, 52, 45. [Google Scholar] [CrossRef] [PubMed]
- Lopez, O.J.; Oliveira, M.F.; Garcia, E.A.; Kwon, B.J.; Doster, A.; Osorio, F.A. Protection against porcine reproductive and respiratory syndrome virus (PRRSV) infection through passive transfer of PRRSV-neutralizing antibodies is dose dependent. Clin. Vaccine Immunol. 2007, 14, 269–275. [Google Scholar] [CrossRef]
- Martin-Valls, G.E.; Li, Y.; Clilverd, H.; Soto, J.; Cortey, M.; Mateu, E. Levels of neutralizing antibodies against resident farm strain or vaccine strain are not indicators of protection against PRRSV-1 vertical transmission under farm conditions. BMC Vet. Res. 2023, 19, 217. [Google Scholar] [CrossRef]
- Popescu, L.N.; Trible, B.R.; Chen, N.; Rowland, R.R.R. GP5 of porcine reproductive and respiratory syndrome virus (PRRSV) as a target for homologous and broadly neutralizing antibodies. Vet. Microbiol. 2017, 209, 90–96. [Google Scholar] [CrossRef]
- Martínez-Lobo, F.J.; Díez-Fuertes, F.; Simarro, I.; Castro, J.M.; Prieto, C. The ability of porcine reproductive and respiratory syndrome virus isolates to induce broadly reactive neutralizing antibodies correlates with In vivo protection. Front. Immunol. 2021, 12, 691145. [Google Scholar] [CrossRef]
- Martínez-Lobo, F.J.; Díez-Fuertes, F.; Simarro, I.; Castro, J.M.; Prieto, C. Porcine Reproductive and Respiratory Syndrome Virus isolates differ in their susceptibility to neutralization. Vaccine 2011, 29, 6928–6940. [Google Scholar] [CrossRef]
- Vu, H.L.X.; Pattnaik, A.K.; Osorio, F.A. Strategies to broaden the cross-protective efficacy of vaccines against porcine reproductive and respiratory syndrome virus. Vet. Microbiol. 2017, 206, 29–34. [Google Scholar] [CrossRef]
- Young, J.E.; Dvorak, C.M.T.; Graham, S.P.; Murtaugh, M.P. Isolation of porcine reproductive and respiratory syndrome virus GP5-specific, neutralizing monoclonal antibodies from hyperimmune sows. Front. Immunol. 2021, 12, 638493. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Li, S.; Tian, Y.; Li, X.; Li, S.; Li, J.; Qiu, M.; Sun, Z.; Xiao, Y.; Yan, X.; et al. Chimeric HP-PRRSV2 containing an ORF2-6 consensus sequence induces antibodies with broadly neutralizing activity and confers cross protection against virulent NADC30-like isolate. Vet. Res. 2021, 52, 74. [Google Scholar] [CrossRef] [PubMed]
- Trible, B.R.; Popescu, L.N.; Monday, N.; Calvert, J.G.; Rowland, R.R.R. A single amino acid deletion in the matrix protein of porcine reproductive and respiratory syndrome virus confers resistance to a polyclonal swine antibody with broadly neutralizing activity. J. Virol. 2015, 89, 6515–6520. [Google Scholar] [CrossRef]
- Baker, B.; Thacker, E.; Thacker, B.; Vincent, A. A Preliminary Investigation into Possible PRRSV Anergy Induction from Repeated Immunization with a Modified Live Vaccine; University Digital Conservancy: Minneapolis, MN, USA, 1999. [Google Scholar]
- Hsueh, F.-C.; Wang, S.-Y.; Lin, W.-H.; Lin, C.-F.; Tsai, C.-Y.; Huang, C.-W.; Sun, N.; Chiou, M.-T.; Lin, C.-N. Correlation of neutralizing antibodies (NAbs) between sows and piglets and evaluation of protectability associated with maternally derived NAbs in pigs against circulating porcine reproductive and respiratory syndrome virus (PRRSV) under field conditions. Vaccines 2021, 9, 414. [Google Scholar] [CrossRef] [PubMed]
- Renson, P.; Fablet, C.; Andraud, M.; Normand, V.; Lebret, A.; Paboeuf, F.; Rose, N.; Bourry, O. Maternally-derived neutralizing antibodies reduce vaccine efficacy against porcine reproductive and respiratory syndrome virus infection. Vaccine 2019, 37, 4318–4324. [Google Scholar] [CrossRef] [PubMed]
- Balasch, M.; Fort, M.; Taylor, L.P.; Calvert, J.G. Vaccination of 1-day-old pigs with a porcine reproductive and respiratory syndrome virus (PRRSV) modified live attenuated virus vaccine is able to overcome maternal immunity. Porc. Health Manag. 2018, 4, 25. [Google Scholar] [CrossRef]
- Toman, M.; Celer, V.; Kavanová, L.; Levá, L.; Frolichova, J.; Ondráčková, P.; Kudláčková, H.; Nechvátalová, K.; Salat, J.; Faldyna, M. Dynamics and Differences in Systemic and Local Immune Responses After Vaccination With Inactivated and Live Commercial Vaccines and Subsequent Subclinical Infection With PRRS Virus. Front. Immunol. 2019, 10, 1689. [Google Scholar] [CrossRef]
- Ruggeri, J.; Ferlazzo, G.; Boniotti, M.B.; Capucci, L.; Guarneri, F.; Barbieri, I.; Alborali, G.L.; Amadori, M. Characterization of the IgA response to PRRS virus in pig oral fluids. PLoS ONE 2020, 15, e0229065. [Google Scholar] [CrossRef]
- Wu, F.; Ourmanov, I.; Kirmaier, A.; Leviyang, S.; LaBranche, C.; Huang, J.; Whitted, S.; Matsuda, K.; Montefiori, D.; Hirsch, V.M. SIV infection duration largely determines broadening of neutralizing antibody response in macaques. J. Clin. Investig. 2020, 130, 5413–5424. [Google Scholar] [CrossRef]
- Fiers, J.; Tignon, M.; Cay, A.B.; Simons, X.; Maes, D. Porcine Reproductive and Respiratory Syndrome virus (PRRSv): A Cross-Sectional Study on ELISA Seronegative, Multivaccinated Sows. Viruses 2022, 14, 1944. [Google Scholar] [CrossRef]
- Díaz, I.; Genís-Jorquera, B.; Martín-Valls, G.E.; Mateu, E. Using commercial ELISAs to assess humoral response in sows repeatedly vaccinated with modified live porcine reproductive and respiratory syndrome virus. Vet. Rec. 2020, 186, 123. [Google Scholar] [CrossRef]
- Albina, E.; Carrat, C.; Charley, B. Interferon-alpha response to swine arterivirus (PoAV), the porcine reproductive and respiratory syndrome virus. J. Interferon Cytokine Res. 1998, 18, 485–490. [Google Scholar] [CrossRef]
- Van Reeth, K.; Labarque, G.; Nauwynck, H.; Pensaert, M. Differential production of proinflammatory cytokines in the pig lung during different respiratory virus infections: Correlations with pathogenicity. Res. Vet. Sci. 1999, 67, 47–52. [Google Scholar] [CrossRef]
- Kim, W.-I.; Lee, D.-S.; Johnson, W.; Roof, M.; Cha, S.-H.; Yoon, K.-J. Effect of genotypic and biotypic differences among PRRS viruses on the serologic assessment of pigs for virus infection. Vet. Microbiol. 2007, 123, 1–14. [Google Scholar] [CrossRef]
- Lee, S.-M.; Schommer, S.K.; Kleiboeker, S.B. Porcine reproductive and respiratory syndrome virus field isolates differ in in vitro interferon phenotypes. Vet. Immunol. Immunopathol. 2004, 102, 217–231. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.; Xie, J.; Vanderheijden, N.; Nauwynck, H.J. Isolation and characterization of a new population of nasal surface macrophages and their susceptibility to PRRSV-1 subtype 1 (LV) and subtype 3 (Lena). Vet. Res. 2020, 51, 21. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, J.; Tian, Y.; Liu, J.; Zhu, J.; Chen, N.; Shang, S. A potent CD8 T-cell response may be associated with partial cross-protection conferred by an attenuated Chinese HP-PRRSV vaccine against NADC30-like PRRSV challenge. J. Gen. Virol. 2023, 104, 001850. [Google Scholar] [CrossRef] [PubMed]
- Ostrowski, M.; Galeota, J.A.; Jar, A.M.; Platt, K.B.; Osorio, F.A.; Lopez, O.J. Identification of neutralizing and nonneutralizing epitopes in the porcine reproductive and respiratory syndrome virus GP5 ectodomain. J. Virol. 2002, 76, 4241–4250. [Google Scholar] [CrossRef]
- Vu, H.L.X.; Kwon, B.; Yoon, K.-J.; Laegreid, W.W.; Pattnaik, A.K.; Osorio, F.A. Immune evasion of porcine reproductive and respiratory syndrome virus through glycan shielding involves both glycoprotein 5 as well as glycoprotein 3. J. Virol. 2011, 85, 5555–5564. [Google Scholar] [CrossRef]
- Wei, Z.; Lin, T.; Sun, L.; Li, Y.; Wang, X.; Gao, F.; Liu, R.; Chen, C.; Tong, G.; Yuan, S. N-linked glycosylation of GP5 of porcine reproductive and respiratory syndrome virus is critically important for virus replication In vivo. J. Virol. 2012, 86, 9941–9951. [Google Scholar] [CrossRef]
- Choi, J.-C.; Kim, M.-S.; Choi, H.-Y.; Kang, Y.-L.; Choi, I.-Y.; Jung, S.-W.; Jeong, J.-Y.; Kim, M.-C.; Cho, A.Y.; Lee, J.-H.; et al. Porcine Reproductive and Respiratory Syndrome Virus Engineered by Serine Substitution on the 44th Amino Acid of GP5 Resulted in a Potential Vaccine Candidate with the Ability to Produce High Levels of Neutralizing Antibody. Vet. Sci. 2023, 10, 191. [Google Scholar] [CrossRef]
- Thaa, B.; Sinhadri, B.C.; Tielesch, C.; Krause, E.; Veit, M. Signal peptide cleavage from GP5 of PRRSV: A minor fraction of molecules retains the decoy epitope, a presumed molecular cause for viral persistence. PLoS ONE 2013, 8, e65548. [Google Scholar] [CrossRef]
- Das, P.B.; Dinh, P.X.; Ansari, I.H.; de Lima, M.; Osorio, F.A.; Pattnaik, A.K. The minor envelope glycoproteins GP2a and GP4 of porcine reproductive and respiratory syndrome virus interact with the receptor CD163. J. Virol. 2010, 84, 1731–1740. [Google Scholar] [CrossRef]
- Wang, J.; Yan, J.; Wang, S.; Chen, R.; Xing, Y.; Liu, Q.; Gao, S.; Zhu, Y.; Li, J.; Zhou, Y.; et al. An Expeditious Neutralization Assay for Porcine Reproductive and Respiratory Syndrome Virus Based on a Recombinant Virus Expressing Green Fluorescent Protein. Curr. Issues Mol. Biol. 2024, 46, 1047–1063. [Google Scholar] [CrossRef]
- Jusa, E.R.; Inaba, Y.; Kouno, M.; Hirose, O.; Shibata, I.; Kubota, M.; Yasuhara, H. Slow-reacting and complement-requiring neutralizing antibody in swine infected with porcine reproductive and respiratory syndrome (PRRS) virus. J. Vet. Med. Sci. 1996, 58, 749–753. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cui, J.; O’Connell, C.M.; Hagen, C.; Sawicki, K.; Smyth, J.A.; Verardi, P.H.; Van Kruiningen, H.J.; Garmendia, A.E. Broad protection of pigs against heterologous PRRSV strains by a GP5-mosaic DNA vaccine prime/GP5-mosaic rVaccinia (VACV) vaccine boost. Vaccines 2020, 8, 106. [Google Scholar] [CrossRef] [PubMed]
- Jeong, C.-G.; Khatun, A.; Nazki, S.; Kim, S.-C.; Noh, Y.-H.; Kang, S.-C.; Lee, D.-U.; Yang, M.-S.; Shabir, N.; Yoon, I.-J.; et al. Evaluation of the cross-protective efficacy of a chimeric PRRSV vaccine against two genetically diverse PRRSV2 field strains in a reproductive model. Vaccines 2021, 9, 1258. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Ni, Y.-Y.; Piñeyro, P.; Cossaboom, C.M.; Subramaniam, S.; Sanford, B.J.; Dryman, B.A.; Huang, Y.-W.; Meng, X.-J. Broadening the heterologous cross-neutralizing antibody inducing ability of porcine reproductive and respiratory syndrome virus by breeding the GP4 or M genes. PLoS ONE 2013, 8, e66645. [Google Scholar] [CrossRef]
- Zhang, L.; Chen, J.; Shen, C.; Wang, G.; Lu, Z.; Zeng, D.; Gao, Y.; Chen, H.; Xia, N.; Chen, Y. Vaccination with Deglycosylated Modified Hemagglutinin Broadly Protects against Influenza Virus Infection in Mice and Ferrets. Vaccines 2022, 10, 1304. [Google Scholar] [CrossRef]
- Cheng, C.-W.; Wu, C.-Y.; Wang, S.-W.; Chen, J.-Y.; Kung, C.-C.; Liao, K.-S.; Wong, C.-H. Low-sugar universal mRNA vaccine against coronavirus variants with deletion of glycosites in the S2 or stem of SARS-CoV-2 spike messenger RNA (mRNA). Proc. Natl. Acad. Sci. USA 2023, 120, e2314392120. [Google Scholar] [CrossRef]
- Pamornchainavakul, N.; Paploski, I.A.D.; Makau, D.N.; Baker, J.P.; Huang, J.; Ferreira, C.P.; Corzo, C.A.; Rovira, A.; Cheeran, M.C.-J.; Lycett, S.; et al. Experimental evidence of vaccine-driven evolution of porcine reproductive and respiratory syndrome virus type 2. Virus Evol. 2025, 11, veaf056. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Soriano, Á.; Martínez-Lobo, F.J.; Garza-Moreno, L.; Castillo-Pérez, J.; Caballero, E.; Castro, J.M.; Simarro, I.; Prieto, C. Determination of the frequency of individuals with broadly cross-reactive neutralizing antibodies against PRRSV in the sow population under field conditions. Porc. Health Manag. 2024, 10, 26. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, H.; Li, W.; Feng, X.; Han, F.; Zhang, Y.; Chen, J.; Liu, D.; Xia, P. Activating Fc Gamma Receptors and Viral Receptors Are Required for Antibody-Dependent Enhancement of Porcine Reproductive and Respiratory Syndrome Virus Infection. Vet. Sci. 2022, 9, 470. [Google Scholar] [CrossRef]
- Gu, W.; Guo, L.; Yu, H.; Niu, J.; Huang, M.; Luo, X.; Li, R.; Tian, Z.; Feng, L.; Wang, Y. Involvement of CD16 in antibody-dependent enhancement of porcine reproductive and respiratory syndrome virus infection. J. Gen. Virol. 2015, 96, 1712–1722. [Google Scholar] [CrossRef]
- Zhang, L.; Li, W.; Sun, Y.; Kong, L.; Xu, P.; Xia, P.; Zhang, G. Antibody-Mediated Porcine Reproductive and Respiratory Syndrome Virus Infection Downregulates the Production of Interferon-α and Tumor Necrosis Factor-α in Porcine Alveolar Macrophages via Fc Gamma Receptor I and III. Viruses 2020, 12, 187. [Google Scholar] [CrossRef]
- Li, Y.; Xu, L.; Jiao, D.; Zheng, Z.; Chen, Z.; Jing, Y.; Li, Z.; Ma, Z.; Feng, Y.; Guo, X.; et al. Genomic similarity and antibody-dependent enhancement of immune serum potentially affect the protective efficacy of commercial MLV vaccines against NADC30-like PRRSV. Virol. Sin. 2023, 38, 813–826. [Google Scholar] [CrossRef] [PubMed]
- Yoon, K.-J.; Wu, L.-L.; Zimmerman, J.J.; Hill, H.T.; Platt, K.B. Antibody-Dependent Enhancement (ADE) of Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) Infection in Pigs. Viral Immunol. 1996, 9, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Nan, Y.; Yu, Y.; Yang, Z.; Zhang, Y.-J. Variable interference with interferon signal transduction by different strains of porcine reproductive and respiratory syndrome virus. Vet. Microbiol. 2013, 166, 493–503. [Google Scholar] [CrossRef] [PubMed]
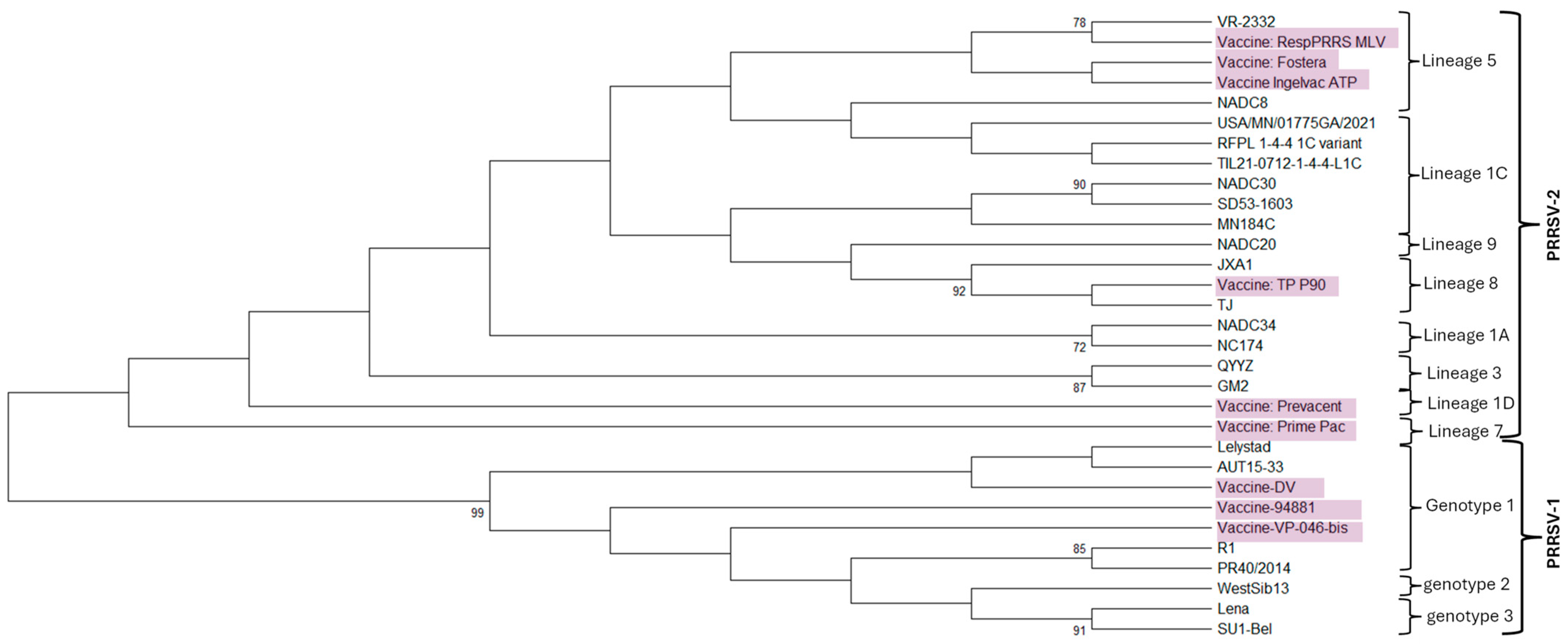
| Subtype/Lineage | Specific Vaccine Available | Sublineage | Representative Strain | GenBank Accession |
|---|---|---|---|---|
| PRRSV-1 | ||||
| Subtype 1-1 | yes | na | Lelystad (LV) | M96262 |
| Spanish R1 | OM893828 | |||
| AUT15-33 | MT000052 | |||
| PR40/2014 | MF346695 | |||
| MLV-DV | KJ127878.1 | |||
| Unistrain-VP-046-bis | MK134483.1 | |||
| 94881 | KT988004.1 | |||
| Subtype 1-2 | no | WestSib13 | KX668221.1 | |
| Subtype 1-3 | no | Lena | JF802085 | |
| SU1-Bel | KP889243 | |||
| PRRSV-2 | ||||
| Lineage 1 | yes | 1A | NC174 | PP171544 |
| 1C | NADC30 | JN654459.1 | ||
| 1C.5 | USA/MN/01775GA/2021 | OR634972.1 | ||
| RFLP 1-4-4 variant | MW887655 | |||
| UIL21-0712 | PQ810800 | |||
| L1D | Prevacent® PRRS | KU131568 | ||
| L1A | NADC34 | MF326985 | ||
| L1H | USA/81793-6/2019 | OR634975 | ||
| Lineage 3 | no | 3.5 | QYYZ | JQ308798 |
| 3.5 | GM2 | JN662424.1 | ||
| Lineage 5 | yes | 5A | VR-2332 | EF536003.1 |
| RespPRRS MLV | AF066183 | |||
| 5B | NADC-8 | AF396833 | ||
| Lineage 7 | yes | na | Prime Pac® PRRS RR | DQ779791 |
| Lineage 8 | yes | 8.7 | JXA1 | EF112445 |
| 8.7 | TJ | EU860248 | ||
| 8.7 | TP P90 | GU232737 | ||
| 8C | Fostera® PRRS | KP300938 | ||
| 8A | Ingelvac PRRS® ATP | DQ988080 | ||
| Lineage 9 | no | na | NADC-20 | JX069953 |
| Primary Immunization/Exposure | Challenge Virus | Experimental Model | Protection Endpoints Assessed | Primary Protection Domain | Main Outcome | Key References |
|---|---|---|---|---|---|---|
| PRRSV-2 MLV | PRRSV-1 | Growing pigs | Clinical signs, viremia, lung lesions | Clinical > Virological | Partial protection: reduced clinical severity and viremia | [27,28] |
| PRRSV-2 MLV | PRRSV-1 | Pregnant sows/gilts | Reproductive performance, viremia | Reproductive (partial) | Partial protection: improved reproductive outcomes but incomplete virological protection | [29] |
| PRRSV-2 MLV | PRRSV-1 | Boars | Virus shedding in semen | None | Limited protection: reduced PRRSV-2 shedding only | [26] |
| PRRSV-2 MLV | PRRSV-1 | Nursery pigs | Clinical disease, viremia | Variable/endpoint-dependent | Inconsistent: protection varied by strain and endpoint | [28] |
| PRRSV-1 MLV | PRRSV-2 | Growing pigs | Clinical disease, viremia, shedding | None | No or minimal protection | [35,36] |
| PRRSV-1 MLV | PRRSV-2 | Boars | Semen shedding | None | No protection against PRRSV-2 shedding | [36] |
| PRRSV-1 MLV | PRRSV-2 | Nursery pigs | Clinical signs | Clinical only | Partial clinical protection (no virological protection) | [38] |
| PRRSV-1 MLV followed by PRRSV-2 MLV (consecutive) | PRRSV-1 + PRRSV-2 | Nursery pigs | Clinical signs, viremia | Bidirectional (partial) | Improved bidirectional protection compared with single-species vaccination | [30] |
| Natural PRRSV-2 infection | PRRSV-1 | In vitro (MARC-145 cells) | Viral replication | Virological (in vitro) | Inhibition of PRRSV-1 replication after PRRSV-2 pre-infection | |
| Natural PRRSV-1 infection | PRRSV-2 | In vitro (MARC-145 cells) | Viral replication | None (in vitro) | No inhibition of PRRSV-2 replication | |
| PRRSV-2 MLV | PRRSV-1 | Late-gestation sows | Reproductive outcomes | None | No protection | [25] |
| PRRSV-1 or PRRSV-2 MLV | PRRSV-1 + PRRSV-2 dual challenge | Growing pigs | Clinical signs, viremia | Asymmetric | Limited and asymmetric protection, favoring PRRSV-2 priming | [37] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Raev, S.A.; Cai, L.; Muro, N.; Madera, R.; Wang, L.; Shi, J. Cross-Protection in PRRSV: Mechanisms, Limitations, and Implications for Vaccine Design. Pathogens 2026, 15, 345. https://doi.org/10.3390/pathogens15040345
Raev SA, Cai L, Muro N, Madera R, Wang L, Shi J. Cross-Protection in PRRSV: Mechanisms, Limitations, and Implications for Vaccine Design. Pathogens. 2026; 15(4):345. https://doi.org/10.3390/pathogens15040345
Chicago/Turabian StyleRaev, Sergei A., Limeng Cai, Nina Muro, Rachel Madera, Lihua Wang, and Jishu Shi. 2026. "Cross-Protection in PRRSV: Mechanisms, Limitations, and Implications for Vaccine Design" Pathogens 15, no. 4: 345. https://doi.org/10.3390/pathogens15040345
APA StyleRaev, S. A., Cai, L., Muro, N., Madera, R., Wang, L., & Shi, J. (2026). Cross-Protection in PRRSV: Mechanisms, Limitations, and Implications for Vaccine Design. Pathogens, 15(4), 345. https://doi.org/10.3390/pathogens15040345








