Phylogenetic Lineages of PRRSV-2 from Canada Reveal Patterns of Transboundary Spread and Two Novel Sub-Lineages in North America
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Source
2.2. Sequence Classification, Estimation of Genetic Distances, and Sequences Filtering
2.3. Dataset Down-Sampling
2.3.1. Down-Sampling for Recombination Analysis
2.3.2. Down-Sampling for Phylogeographic Analysis
2.3.3. Down-Sampling for Estimation of Divergence Times
2.4. Phylogenetic Analysis
2.5. Recombination Analysis
2.6. Phylogeographic Analysis
2.7. Estimation of Divergence Times
3. Results
3.1. Assignment of Sub-Lineages and Variants
3.2. Phylogenetic Analyses and Sequence Comparisons Warrant Two New PRRSV-2 Sub-Lineages
3.3. Establishment of New PRRSV-2 Genetic Variants
3.4. Evidence of Intra-Sub-Lineage Recombination Was Detected in a Small Number of 1K and 1L Sequences
3.5. Differential Spatial Distribution of Sub-Lineages
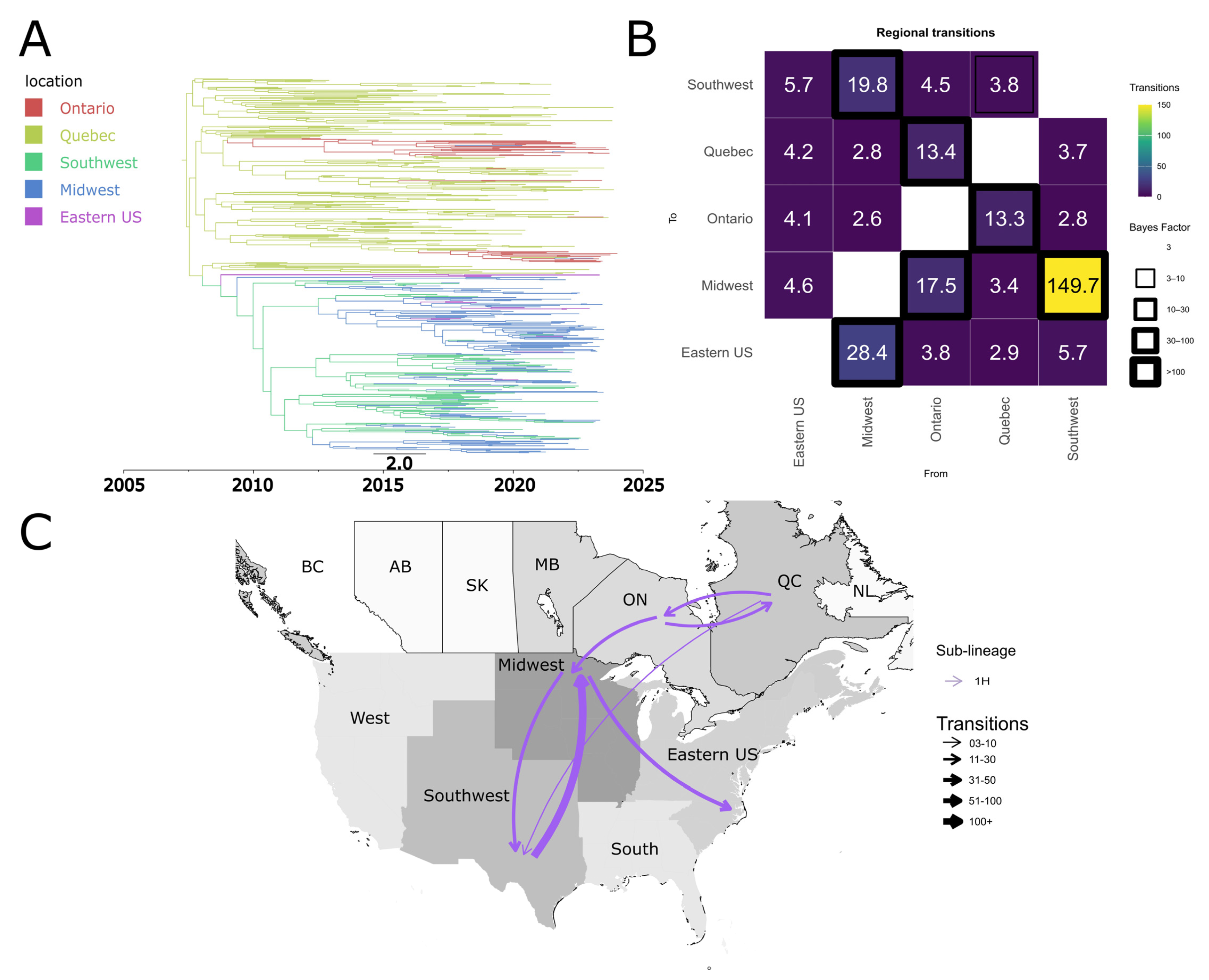
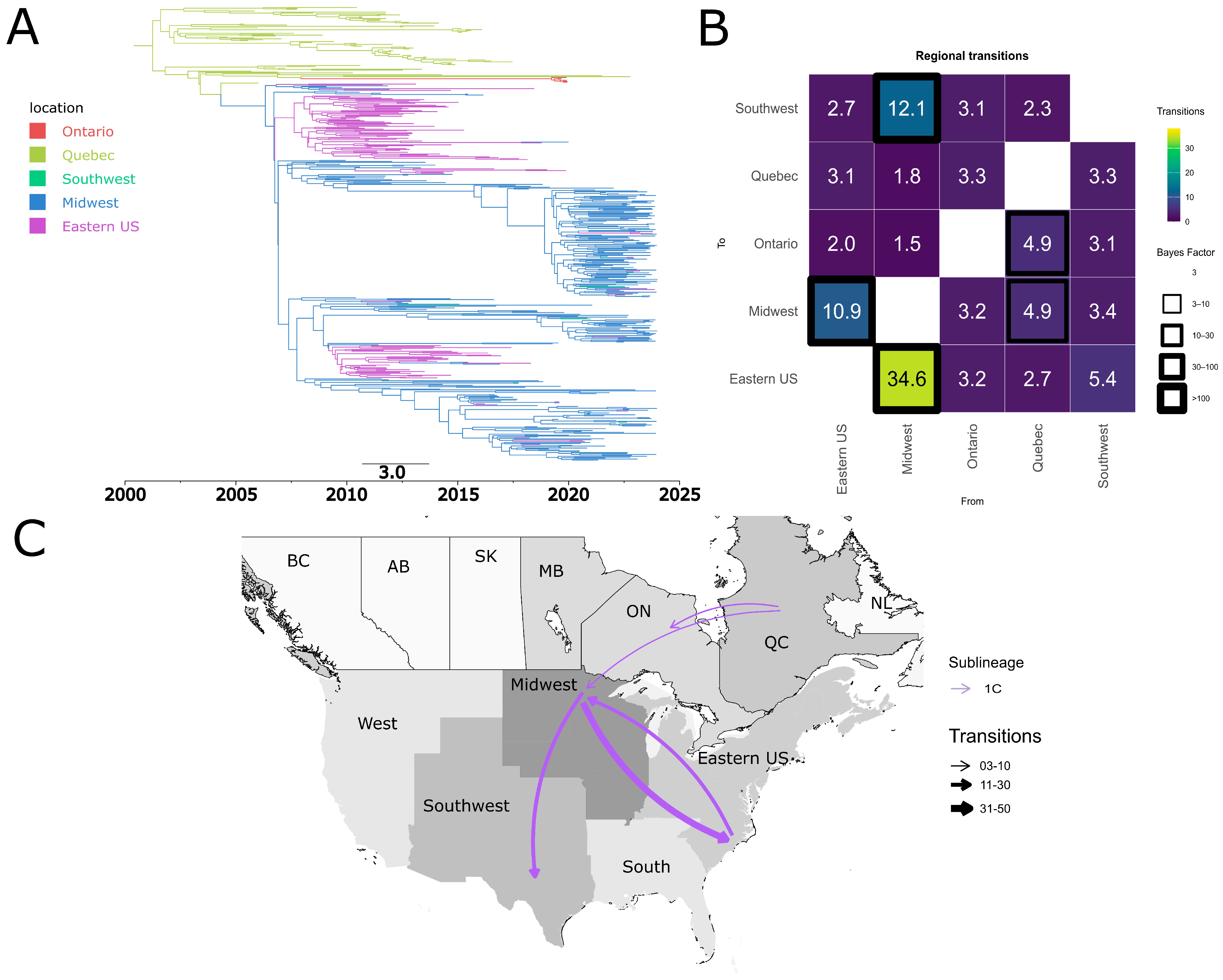
3.6. Quebec and the USA Midwest Are Potential Hubs for the Dispersal of PRRSV-2 Sub-Lineages Between Canada and the United States
3.7. Age of Sub-Lineages 1K, 1L, and Undetermined Clades
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Albina, E. Epidemiology of porcine reproductive and respiratory syndrome (PRRS): An overview. Vet. Microbiol. 1997, 55, 309–316. [Google Scholar] [CrossRef]
- Mussell, A.; Oginskyy, A.; Grier, K.; Morin, M.; Lachance, M.; Whittington, L.; Friendship, R.; Canadian Swine Health. A Risk, Benefit, Strength, Weakness, Opportunity and Threat Analysis for the Control and Possible Eradication of Porcine Reproductive and Respiratory Syndrome (PRRS) Virus Within the Canadian Swine Herd; Canadian Swine Health Board, Ed.; Canadian Swine Health Board: Ottawa, ON, Canada, 2011. [Google Scholar]
- Government-of-Canada. Agriculture and Agri-Food Canada—Hogs and Pork; Government of Canada: Ottawa, ON, Canada, 2025.
- FAO United Nations. FAOSTAT: Trade—Crops and Livestock Products; FAO: Rome, Italy, 2023; Available online: https://www.fao.org/faostat/en/#data/TM (accessed on 23 September 2025).
- USDA. Hogs & Pork—Sector at a Glance; Economic Research Service; U.S. Department of Agriculture: Washington, DC, USA, 2025. Available online: https://www.ers.usda.gov/topics/animal-products/hogs-pork/sector-at-a-glance#Loc6 (accessed on 23 September 2025).
- Snijder, E.J.; Meulenberg, J.J. The molecular biology of arteriviruses. J. Gen. Virol. 1998, 79, 961–980. [Google Scholar] [CrossRef] [PubMed]
- Brinton, M.A.; Gulyaeva, A.A.; Balasuriya, U.B.; Dunowska, M.; Faaberg, K.S.; Goldberg, T.; Leung, F.C.; Nauwynck, H.J.; Snijder, E.J.; Stadejek, T. ICTV virus taxonomy profile: Arteriviridae 2021. J. Gen. Virol. 2021, 102, 001632. [Google Scholar] [CrossRef] [PubMed]
- Van Breedam, W.; Delputte, P.L.; Van Gorp, H.; Misinzo, G.; Vanderheijden, N.; Duan, X.; Nauwynck, H.J. Porcine reproductive and respiratory syndrome virus entry into the porcine macrophage. J. Gen. Virol. 2010, 91, 1659–1667. [Google Scholar] [CrossRef] [PubMed]
- World Organization for Animal Health. PRRS: The Disease, Its Diagnosis, Prevention and Control. Available online: https://www.oie.int/fileadmin/Home/eng/Our_scientific_expertise/docs/pdf/PRRS_guide_web_bulletin.pdf (accessed on 23 September 2025).
- Hanada, K.; Suzuki, Y.; Nakane, T.; Hirose, O.; Gojobori, T. The origin and evolution of porcine reproductive and respiratory syndrome viruses. Mol. Biol. Evol. 2005, 22, 1024–1031. [Google Scholar] [CrossRef]
- Pamornchainavakul, N.; Kikuti, M.; Paploski, I.A.; Makau, D.N.; Rovira, A.; Corzo, C.A.; VanderWaal, K. Measuring how recombination re-shapes the evolutionary history of PRRSV-2: A genome-based phylodynamic analysis of the emergence of a novel PRRSV-2 variant. Front. Vet. Sci. 2022, 9, 846904. [Google Scholar] [CrossRef]
- Cui, X.; Xia, D.; Huang, X.; Sun, Y.; Shi, M.; Zhang, J.; Li, G.; Yang, Y.; Wang, H.; Cai, X. Analysis of recombinant characteristics based on 949 PRRSV-2 genomic sequences obtained from 1991 to 2021 shows that viral multiplication ability contributes to dominant recombination. Microbiol. Spectr. 2022, 10, e02934-22. [Google Scholar] [CrossRef]
- Duffy, S.; Shackelton, L.A.; Holmes, E.C. Rates of evolutionary change in viruses: Patterns and determinants. Nat. Rev. Genet. 2008, 9, 267–276. [Google Scholar] [CrossRef]
- Shi, M.; Lam, T.T.-Y.; Hon, C.-C.; Murtaugh, M.P.; Davies, P.R.; Hui, R.K.-H.; Li, J.; Wong, L.T.-W.; Yip, C.-W.; Jiang, J.-W. Phylogeny-based evolutionary, demographical, and geographical dissection of North American type 2 porcine reproductive and respiratory syndrome viruses. J. Virol. 2010, 84, 8700–8711. [Google Scholar] [CrossRef]
- Paploski, I.A.; Pamornchainavakul, N.; Makau, D.N.; Rovira, A.; Corzo, C.A.; Schroeder, D.C.; Cheeran, M.C.; Doeschl-Wilson, A.; Kao, R.R.; Lycett, S. Phylogenetic structure and sequential dominance of sub-lineages of PRRSV type-2 lineage 1 in the United States. Vaccines 2021, 9, 608. [Google Scholar] [CrossRef]
- Paploski, I.A.D.; Corzo, C.; Rovira, A.; Murtaugh, M.P.; Sanhueza, J.M.; Vilalta, C.; Schroeder, D.C.; VanderWaal, K. Temporal dynamics of co-circulating lineages of porcine reproductive and respiratory syndrome virus. Front. Microbiol. 2019, 10, 2486. [Google Scholar] [CrossRef]
- VanderWaal, K.; Pamornchainavakul, N.; Kikuti, M.; Zhang, J.; Zeller, M.; Trevisan, G.; Rossow, S.; Schwartz, M.; Linhares, D.C.; Holtkamp, D.J. PRRSV-2 variant classification: A dynamic nomenclature for enhanced monitoring and surveillance. mSphere 2025, 10, e00709-24. [Google Scholar] [CrossRef]
- VanderWaal, K.; Pamornchainavakul, N.; Kikuti, M.; Linhares, D.C.; Trevisan, G.; Zhang, J.; Anderson, T.K.; Zeller, M.; Rossow, S.; Holtkamp, D.J. Phylogenetic-based methods for fine-scale classification of PRRSV-2 ORF5 sequences: A comparison of their robustness and reproducibility. Front. Virol. 2024, 4, 1433931. [Google Scholar] [CrossRef]
- Yim-Im, W.; Anderson, T.K.; Paploski, I.A.; VanderWaal, K.; Gauger, P.; Krueger, K.; Shi, M.; Main, R.; Zhang, J. Refining PRRSV-2 genetic classification based on global ORF5 sequences and investigation of their geographic distributions and temporal changes. Microbiol. Spectr. 2023, 11, e02916-23. [Google Scholar] [CrossRef] [PubMed]
- Pamornchainavakul, N.; Paploski, I.A.; Makau, D.N.; Kikuti, M.; Rovira, A.; Lycett, S.; Corzo, C.A.; VanderWaal, K. Mapping the dynamics of contemporary PRRSV-2 evolution and its emergence and spreading hotspots in the US using phylogeography. Pathogens 2023, 12, 740. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Lemey, P.; Brar, M.S.; Suchard, M.A.; Murtaugh, M.P.; Carman, S.; D’Allaire, S.; Delisle, B.; Lambert, M.-È.; Gagnon, C.A. The spread of type 2 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) in North America: A phylogeographic approach. Virology 2013, 447, 146–154. [Google Scholar] [CrossRef]
- Lambert, M.-È.; Delisle, B.; Arsenault, J.; Poljak, Z.; D’Allaire, S. Positioning Quebec ORF5 sequences of porcine reproductive and respiratory syndrome virus (PRRSV) within Canada and worldwide diversity. Infect. Genet. Evol. 2019, 74, 103999. [Google Scholar] [CrossRef]
- Lambert, M.-È.; Arsenault, J.; Côté, J.-C.; D’Allaire, S. A descriptive study on spatial and temporal distributions of genetic clusters of porcine reproductive and respiratory syndrome virus infecting pig sites in Quebec, Canada, between 2010 and 2019. Porc. Health Manag. 2024, 10, 7. [Google Scholar] [CrossRef]
- Shi, M.; Lam, T.T.-Y.; Hon, C.-C.; Hui, R.K.-H.; Faaberg, K.S.; Wennblom, T.; Murtaugh, M.P.; Stadejek, T.; Leung, F.C.-C. Molecular epidemiology of PRRSV: A phylogenetic perspective. Virus Res. 2010, 154, 7–17. [Google Scholar] [CrossRef]
- Perez, A.M.; Linhares, D.C.; Arruda, A.G.; VanderWaal, K.; Machado, G.; Vilalta, C.; Sanhueza, J.M.; Torrison, J.; Torremorell, M.; Corzo, C.A. Individual or common good? Voluntary data sharing to inform disease surveillance systems in food animals. Front. Vet. Sci. 2019, 6, 194. [Google Scholar] [CrossRef]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S.; et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2022, 50, D20–D26. [Google Scholar] [CrossRef] [PubMed]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, S.; Eveleigh, R.J.; Archibald, J.M. Treetrimmer: A method for phylogenetic dataset size reduction. BMC Res. Notes 2013, 6, 145. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T. ModelTest-NG: A new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 2020, 37, 291–294. [Google Scholar] [CrossRef]
- Argimón, S.; Abudahab, K.; Goater, R.J.; Fedosejev, A.; Bhai, J.; Glasner, C.; Feil, E.J.; Holden, M.T.; Yeats, C.A.; Grundmann, H. Microreact: Visualizing and sharing data for genomic epidemiology and phylogeography. Microb. Genom. 2016, 2, e000093. [Google Scholar] [CrossRef]
- Martin, D.; Rybicki, E.P. RDP: Detection of recombination amongst aligned sequences. Bioinformatics 2000, 16, 562–563. [Google Scholar] [CrossRef]
- Padidam, M.; Sawyer, S.; Fauquet, C.M. Possible emergence of new geminiviruses by frequent recombination. Virology 1999, 265, 218–224. [Google Scholar] [CrossRef]
- Martin, D.P.; Posada, D.; Crandall, K.A.; Williamson, C. A modified bootscan algorithm for automated identification of recombinant sequences and recombination breakpoints. AIDS Res. Hum. Retroviruses 2005, 21, 98–102. [Google Scholar] [CrossRef]
- Smith, J.M. Analyzing the mosaic structure of genes. J. Mol. Evol. 1992, 34, 126–129. [Google Scholar] [CrossRef]
- Posada, D.; Crandall, K.A. Evaluation of methods for detecting recombination from DNA sequences: Computer simulations. Proc. Natl. Acad. Sci. USA 2001, 98, 13757–13762. [Google Scholar] [CrossRef]
- Gibbs, M.J.; Armstrong, J.S.; Gibbs, A.J. Sister-scanning: A Monte Carlo procedure for assessing signals in recombinant sequences. Bioinformatics 2000, 16, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Lam, H.M.; Ratmann, O.; Boni, M.F. Improved algorithmic complexity for the 3SEQ recombination detection algorithm. Mol. Biol. Evol. 2018, 35, 247–251. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.P.; Varsani, A.; Roumagnac, P.; Botha, G.; Maslamoney, S.; Schwab, T.; Kelz, Z.; Kumar, V.; Murrell, B. RDP5: A computer program for analyzing recombination in, and removing signals of recombination from, nucleotide sequence datasets. Virus Evol. 2021, 7, veaa087. [Google Scholar] [CrossRef] [PubMed]
- Center SHI. Emerging Diseases, Rapid Response Program. 2025. Available online: https://www.swinehealth.org/rapid-response-to-emerging-disease-program/ (accessed on 23 September 2025).
- Rambaut, A.; Lam, T.T.; Max Carvalho, L.; Pybus, O.G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2016, 2, vew007. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Kass, R.E.; Raftery, A.E. Bayes Factors. J. Am. Stat. Assoc. 1995, 90, 773–795. [Google Scholar] [CrossRef]
- Baele, G.; Lemey, P.; Suchard, M.A. Genealogical working distributions for Bayesian model testing with phylogenetic uncertainty. Syst. Biol. 2016, 65, 250–264. [Google Scholar] [CrossRef]
- Lemey, P.; Rambaut, A.; Drummond, A.J.; Suchard, M.A. Bayesian phylogeography finds its roots. PLoS Comput. Biol. 2009, 5, e1000520. [Google Scholar] [CrossRef]
- Gill, M.S.; Lemey, P.; Faria, N.R.; Rambaut, A.; Shapiro, B.; Suchard, M.A. Improving Bayesian population dynamics inference: A coalescent-based model for multiple loci. Mol. Biol. Evol. 2013, 30, 713–724. [Google Scholar] [CrossRef]
- Drummond, A.J.; Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 2007, 7, 214. [Google Scholar] [CrossRef]
- Murtaugh, M.P.; Stadejek, T.; Abrahante, J.E.; Lam, T.T.; Leung, F.C.-C. The ever-expanding diversity of porcine reproductive and respiratory syndrome virus. Virus Res. 2010, 154, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.H.; Kim, H.; Park, B.; Kim, H. Tracing the genetic history of porcine reproductive and respiratory syndrome viruses derived from the complete ORF 5-7 sequences: A Bayesian coalescent approach. Arch. Virol. 2012, 157, 2143–2151. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Chen, Q.; Wang, L.; Madson, D.; Harmon, K.; Gauger, P.; Zhang, J.; Li, G. Recombination between vaccine and field strains of porcine reproductive and respiratory syndrome virus. Emerg. Infect. Dis. 2019, 25, 2335. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.-Y.; Xia, D.-S.; Luo, L.-Z.; An, T.-Q. Recombination of porcine reproductive and respiratory syndrome virus: Features, possible mechanisms, and future directions. Viruses 2024, 16, 929. [Google Scholar] [CrossRef]
- Kim, S.-C.; Kang, S.C.; Kim, H.-J.; Park, J.; Kim, H.-R.; Park, C.-K.; Kim, W.-I. Nationwide emergence and spread of highly virulent PRRSV-2 mutants in Korea. Porc. Health Manag. 2025, 11, 58. [Google Scholar]
- Zhou, L.; Han, J.; Yang, H. The evolution and diversity of porcine reproductive and respiratory syndrome virus in China. Vet. Microbiol. 2024, 298, 110252. [Google Scholar] [CrossRef]
- Shen, Y.-F.; Arruda, A.G.; Koscielny, M.P.; Cheng, T.-Y. Contrasting PRRSV temporal lineage patterns at the individual farm, production system, and regional levels in Ohio and neighboring states from 2017 to 2021. Prev. Vet. Med. 2024, 226, 106186. [Google Scholar]
- Dee, S.; Deen, J. Evaluation of an industry-based sanitation protocol for transport vehicles contaminated with porcine reproductive and respiratory syndrome virus. J. Swine Health Prod. 2006, 14, 126–132. [Google Scholar] [CrossRef]
- Pileri, E.; Mateu, E. Review on the transmission porcine reproductive and respiratory syndrome virus between pigs and farms and impact on vaccination. Vet. Res. 2016, 47, 108. [Google Scholar] [CrossRef]
- Magee, D.; Suchard, M.A.; Scotch, M. Bayesian phylogeography of influenza A/H3N2 for the 2014–2015 season in the United States using three frameworks of ancestral state reconstruction. PLoS Comput. Biol. 2017, 13, e1005389. [Google Scholar]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Herrera da Silva, J.P.; Paploski, I.A.D.; Charette, R.; Dufresne, L.; Messier, S.; Bolduc, J.; Kikuti, M.; Pamornchainavakul, N.; Corzo, C.A.; VanderWaal, K. Phylogenetic Lineages of PRRSV-2 from Canada Reveal Patterns of Transboundary Spread and Two Novel Sub-Lineages in North America. Pathogens 2026, 15, 346. https://doi.org/10.3390/pathogens15040346
Herrera da Silva JP, Paploski IAD, Charette R, Dufresne L, Messier S, Bolduc J, Kikuti M, Pamornchainavakul N, Corzo CA, VanderWaal K. Phylogenetic Lineages of PRRSV-2 from Canada Reveal Patterns of Transboundary Spread and Two Novel Sub-Lineages in North America. Pathogens. 2026; 15(4):346. https://doi.org/10.3390/pathogens15040346
Chicago/Turabian StyleHerrera da Silva, João P., Igor A. D. Paploski, Robert Charette, Luc Dufresne, Sylvain Messier, Julie Bolduc, Mariana Kikuti, Nakarin Pamornchainavakul, Cesar A. Corzo, and Kimberly VanderWaal. 2026. "Phylogenetic Lineages of PRRSV-2 from Canada Reveal Patterns of Transboundary Spread and Two Novel Sub-Lineages in North America" Pathogens 15, no. 4: 346. https://doi.org/10.3390/pathogens15040346
APA StyleHerrera da Silva, J. P., Paploski, I. A. D., Charette, R., Dufresne, L., Messier, S., Bolduc, J., Kikuti, M., Pamornchainavakul, N., Corzo, C. A., & VanderWaal, K. (2026). Phylogenetic Lineages of PRRSV-2 from Canada Reveal Patterns of Transboundary Spread and Two Novel Sub-Lineages in North America. Pathogens, 15(4), 346. https://doi.org/10.3390/pathogens15040346








