Assessment of the Microbiological Quality of Raw Milk Sold Through Vending Machines at the Farm Level in Switzerland
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Total Viable Counts (TVCs)
2.3. ß-Glucuronidase-Positive Escherichia coli (E. coli)
2.4. Coagulase-Positive Staphylococcus spp. (Presumptive S. aureus)
2.5. Campylobacter spp.
2.6. Yersinia enterocolitica
2.7. Listeria monocytogenes
2.8. Salmonella spp.
2.9. Screening for STEC
2.10. Screening for Methicillin-Resistant Staphylococcus aureus (MRSA)
2.11. Screening for ESBL-Producing Enterobacterales
2.12. DNA Extraction and Whole-Genome-Sequencing (WGS)
2.13. Descriptive Statistics
3. Results
3.1. Sample Collection
3.2. Total Viable Counts (TVCs)
3.3. Escherichia coli (E. coli)
3.4. Coagulase-Positive Staphylococcus spp. (S. aureus)
3.5. Campylobacter spp.
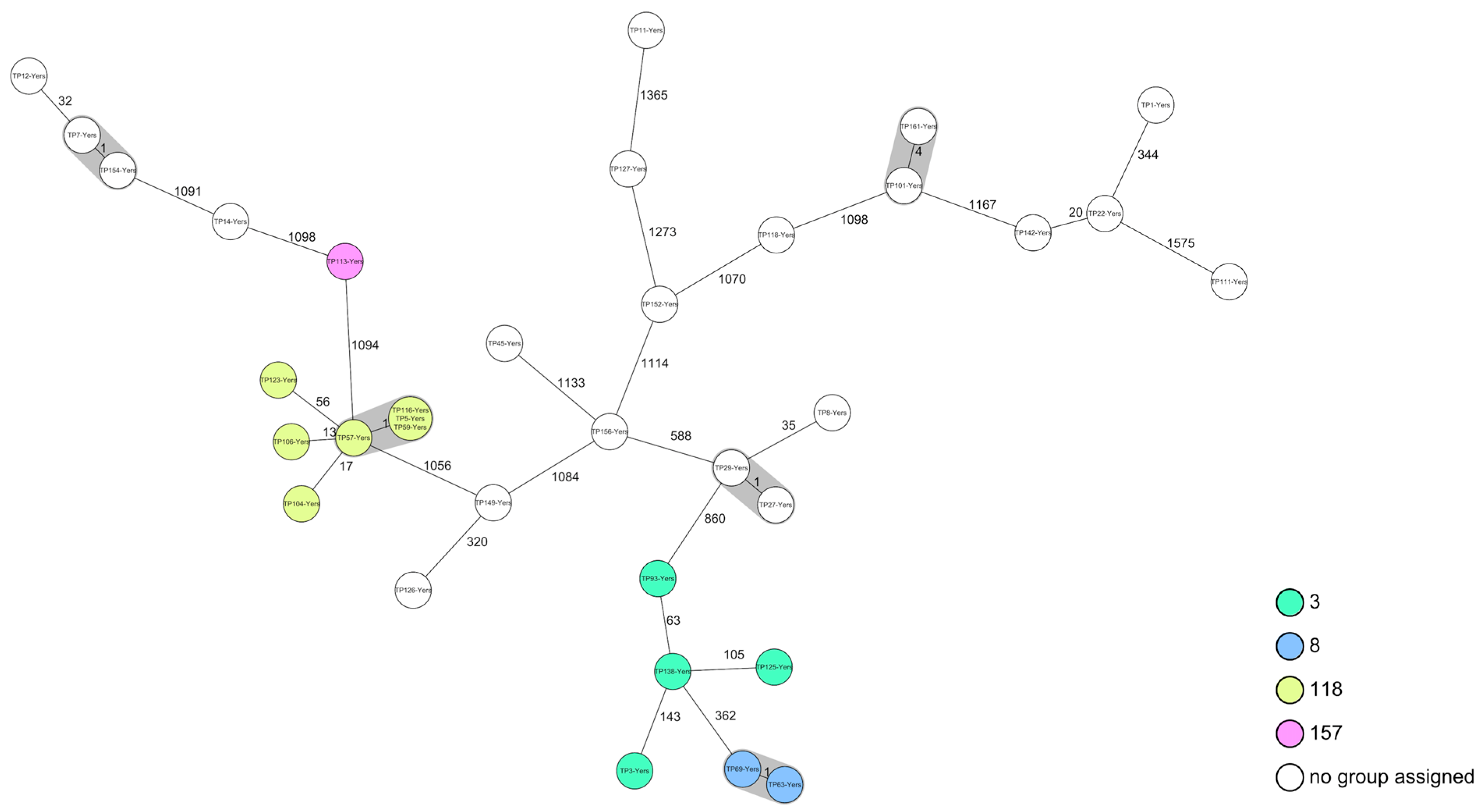
3.6. Yersinia enterocolitica
3.7. Listeria monocytogenes
3.8. Salmonella
3.9. Shiga-Toxin-Producing E. coli (STEC)
3.10. Methicillin-Resistant Staphylococcus aureus (MRSA)
3.11. ESBL-Producing Enterobacterales
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CFUs | Colony-forming units |
| EFSA | European Food Safety Authority |
| ESBL | Extended-spectrum beta-lactamase |
| HPAI | Highly pathogenic avian influenza |
| MALDI-TOF | Matrix-assisted laser desorption/ionisation time-of-flight |
| MLST | Multilocus sequence typing |
| MRSA | Methicillin-resistant Staphylococcus aureus |
| PCR | Polymerase chain reaction |
| STEC | Shiga toxin-producing E. coli |
| TBE | Tick-borne encephalitis |
| TVC | Total viable count |
| WGS | Whole-genome sequencing |
References
- Schweizerische Bundesrat. Milchprüfungsverordnung (MiPV, SR 916.351.0); Schweizerische Bundesrat: Bern, Switzerland, 2010.
- Williams, E.N.; Van Doren, J.M.; Leonard, C.L.; Datta, A.R. Prevalence of Listeria monocytogenes, Salmonella spp., Shiga Toxin-Producing Escherichia coli, and Campylobacter spp. in Raw Milk in the United States between 2000 and 2019: A Systematic Review and Meta-Analysis. J. Food Prot. 2023, 86, 100014. [Google Scholar] [CrossRef]
- Barata, A.R.; Nunes, B.; Oliveira, R.; Guedes, H.; Almeida, C.; Saavedra, M.J.; Da Silva, G.J.; Almeida, G. Occurrence and Seasonality of Campylobacter spp. in Portuguese Dairy Farms. Int. J. Food Microbiol. 2022, 383, 109961. [Google Scholar] [CrossRef]
- Jaakkonen, A.; Castro, H.; Hallanvuo, S.; Ranta, J.; Rossi, M.; Isidro, J.; Lindström, M.; Hakkinen, M. Longitudinal Study of Shiga Toxin-Producing Escherichia coli and Campylobacter jejuni on Finnish Dairy Farms and in Raw Milk. Appl. Environ. Microbiol. 2019, 85, e02910-18. [Google Scholar] [CrossRef]
- Artursson, K.; Schelin, J.; Thisted Lambertz, S.; Hansson, I.; Olsson Engvall, E. Foodborne Pathogens in Unpasteurized Milk in Sweden. Int. J. Food Microbiol. 2018, 284, 120–127. [Google Scholar] [CrossRef]
- Wysok, B.; Wiszniewska-Łaszczych, A.; Uradziński, J.; Szteyn, J. Prevalence and Antimicrobial Resistance of Campylobacter in Raw Milk in the Selected Areas of Poland. Pol. J. Vet. Sci. 2011, 14. [Google Scholar] [CrossRef] [PubMed]
- International Comission on Microbiological Specifications for Foods. Microbial Ecology of Food Commoditis; Springer: Berlin/Heidelberg, Germany, 1980; Volume 2. [Google Scholar]
- Claeys, W.L.; Cardoen, S.; Daube, G.; De Block, J.; Dewettinck, K.; Dierick, K.; De Zutter, L.; Huyghebaert, A.; Imberechts, H.; Thiange, P.; et al. Raw or Heated Cow Milk Consumption: Review of Risks and Benefits. Food Control 2013, 31, 251–262. [Google Scholar] [CrossRef]
- EFSA. Scientific Opinion on the Public Health Risks Related to the Consumption of Raw Drinking Milk. EFSA J. 2015, 13, 3940. [Google Scholar] [CrossRef]
- Bundesamt für Lebensmittelsicherheit und Veterinärwesen (BLV). Bericht zur Überwachung von Zoonosen und Lebensmittelbedingten Krankheitsausbrüchen 2024; Bundesamt für Lebensmittelsicherheit und Veterinärwesen: Bern, Switzerland, 2025. [Google Scholar]
- De Klerk, J.N.; Robinson, P.A. Drivers and Hazards of Consumption of Unpasteurised Bovine Milk and Milk Products in High-Income Countries. PeerJ 2022, 10, e13426. [Google Scholar] [CrossRef] [PubMed]
- Committee on Infectious Diseases; Committee on Nutrition; Brady, M.T.; Byington, C.L.; Davies, H.D.; Edwards, K.M.; Glode, M.P.; Jackson, M.A.; Keyserling, H.L.; Maldonado, Y.A.; et al. Consumption of Raw or Unpasteurized Milk and Milk Products by Pregnant Women and Children. Pediatrics 2014, 133, 175–179. [Google Scholar] [CrossRef]
- Eidgenössische Departement des Innern (EDI). Verordnung Lebensmittel Tierischer Herkunft (VLtH, SR 817.022.108); Eidgenössische Departement des Innern (EDI): Bern, Switzerland, 2016.
- Giacometti, F.; Bonilauri, P.; Serraino, A.; Peli, A.; Amatiste, S.; Arrigoni, N.; Bianchi, M.; Bilei, S.; Cascone, G.; Comin, D.; et al. Four-Year Monitoring of Foodborne Pathogens in Raw Milk Sold by Vending Machines in Italy. J. Food Prot. 2013, 76, 1902–1907. [Google Scholar] [CrossRef] [PubMed]
- Giacometti, F.; Serraino, A.; Finazzi, G.; Daminelli, P.; Losio, M.N.; Tamba, M.; Garigliani, A.; Mattioli, R.; Riu, R.; Zanoni, R.G. Field Handling Conditions of Raw Milk Sold in Vending Machines: Experimental Evaluation of the Behaviour of Listeria monocytogenes, Escherichia coli O157:H7, Salmonella typhimurium and Campylobacter jejuni. Ital. J. Anim. Sci. 2012, 11, e24. [Google Scholar] [CrossRef]
- Costard, S.; Espejo, L.; Groenendaal, H.; Zagmutt, F.J. Outbreak-Related Disease Burden Associated with Consumption of Unpasteurized Cow’s Milk and Cheese, United States, 2009–2014. Emerg. Infect. Dis. 2017, 23, 957–964. [Google Scholar] [CrossRef]
- Bachmann, H.-P.; Bisig, W.; Fröhlich-Wyder, M.-T. Chancen und Risiken für Rohmilchprodukte: Eine Wissenschaftliche Synthese zur Konferenz 2023 vom FACEnetwork; Agroscope: Bern, Switzerland, 2024; Volume 197. [Google Scholar]
- Bachmann, H.-P.; Fröhlich-Wyder, M.T.; Bisig, W. Rohmilch Und Rohmilchprodukte Beeinflussen Die Menschliche Gesundheit – Eine Literaturbesprechung. Agrar. Schweiz 2020, 11, 124–130. [Google Scholar] [CrossRef]
- Brick, T.; Ege, M.; Boeren, S.; Böck, A.; Von Mutius, E.; Vervoort, J.; Hettinga, K. Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins. Nutrients 2017, 9, 963. [Google Scholar] [CrossRef]
- Ahmad Punoo, H. Validation of Milk Product Pasteurization by Alkaline Phosphatase Activity. Concepts Dairy Vet. Sci. 2018, 1, 78–89. [Google Scholar] [CrossRef]
- Loss, G.; Apprich, S.; Waser, M.; Kneifel, W.; Genuneit, J.; Büchele, G.; Weber, J.; Sozanska, B.; Danielewicz, H.; Horak, E.; et al. The Protective Effect of Farm Milk Consumption on Childhood Asthma and Atopy: The GABRIELA Study. J. Allergy Clin. Immunol. 2011, 128, 766–773.e4. [Google Scholar] [CrossRef]
- Loss, G.; Depner, M.; Ulfman, L.H.; Van Neerven, R.J.J.; Hose, A.J.; Genuneit, J.; Karvonen, A.M.; Hyvärinen, A.; Kaulek, V.; Roduit, C.; et al. Consumption of Unprocessed Cow’s Milk Protects Infants from Common Respiratory Infections. J. Allergy Clin. Immunol. 2015, 135, 56–62.e2. [Google Scholar] [CrossRef]
- Sozańska, B. Raw Cow’s Milk and Its Protective Effect on Allergies and Asthma. Nutrients 2019, 11, 469. [Google Scholar] [CrossRef] [PubMed]
- Marchand, S.; De Block, J.; De Jonghe, V.; Coorevits, A.; Heyndrickx, M.; Herman, L. Biofilm Formation in Milk Production and Processing Environments; Influence on Milk Quality and Safety. Compr. Rev. Food Sci. Food Saf. 2012, 11, 133–147. [Google Scholar] [CrossRef]
- Godic Torkar, K.; Kirbiš, A.; Vadnjal, S.; Biasizzo, M.; Galicic, A.; Jevšnik, M. The Microbiological Quality of Slovenian Raw Milk from Vending Machines and Their Hygienic-Technical Conditions. Br. Food J. 2017, 119, 377–389. [Google Scholar] [CrossRef]
- Zulauf, M.; Zweifel, C.; Stephan, R. Microbiological Quality of Raw Milk Sold Directly from Farms to Consumers in Switzerland. J. Food Saf. Food Qual.-Arch. Für Leb. 2018, 69, 140–144. [Google Scholar] [CrossRef]
- Agristat. Milchstatistik Schweiz 2024; Schweizer Bauernverband: Bern, Switzerland, 2025; Available online: https://www.sbv-usp.ch/de/milchstatistik-der-schweiz-2024 (accessed on 26 October 2025).
- Kaiser, F.; Cardenas, S.; Yinda, K.C.; Mukesh, R.K.; Ochwoto, M.; Gallogly, S.; Wickenhagen, A.; Bibby, K.; De Wit, E.; Morris, D.; et al. Highly Pathogenic Avian Influenza A(H5N1) Virus Stability in Irradiated Raw Milk and Wastewater and on Surfaces, United States. Emerg. Infect. Dis. 2025, 31. [Google Scholar] [CrossRef] [PubMed]
- Dhakal, J.; Bhat, S.; James, J.; Otwey, R.Y.; Chapagain, S.; Singh, P. Highly Pathogenic Avian Influenza (HPAI) H5N1 in Raw Pet Foods and Milk: A Growing Threat to Both Companion Animals and Human Health, and Potential Raw Pet Food Industry Liability. J. Food Prot. 2025, 88, 100628. [Google Scholar] [CrossRef]
- Burrough, E.R.; Magstadt, D.R.; Petersen, B.; Timmermans, S.J.; Gauger, P.C.; Zhang, J.; Siepker, C.; Mainenti, M.; Li, G.; Thompson, A.C.; et al. Highly Pathogenic Avian Influenza A(H5N1) Clade 2.3.4.4b Virus Infection in Domestic Dairy Cattle and Cats, United States, 2024. Emerg. Infect. Dis. 2024, 30, 1335. [Google Scholar] [CrossRef]
- Mostafa, A.; Naguib, M.M.; Nogales, A.; Barre, R.S.; Stewart, J.P.; García-Sastre, A.; Martinez-Sobrido, L. Avian Influenza A (H5N1) Virus in Dairy Cattle: Origin, Evolution, and Cross-Species Transmission. mBio 2024, 15, e02542-24. [Google Scholar] [CrossRef]
- Tomassone, L.; Martello, E.; Mannelli, A.; Vicentini, A.; Gossner, C.M.; Leonardi-Bee, J. A Systematic Review on the Prevalence of Tick-Borne Encephalitis Virus in Milk and Milk Products in Europe. Zoonoses Public Health 2025, 72, 248–258. [Google Scholar] [CrossRef] [PubMed]
- Johler, S.; Weder, D.; Bridy, C.; Huguenin, M.-C.; Robert, L.; Hummerjohann, J.; Stephan, R. Outbreak of Staphylococcal Food Poisoning among Children and Staff at a Swiss Boarding School Due to Soft Cheese Made from Raw Milk. J. Dairy Sci. 2015, 98, 2944–2948. [Google Scholar] [CrossRef] [PubMed]
- Ohno, Y.; Sekizuka, T.; Kuroda, M.; Ikeda, T. Outbreaks of Campylobacteriosis Caused by Drinking Raw Milk in Japan: Evidence of Relationship Between Milk and Patients by Using Whole Genome Sequencing. Foodborne Pathog. Dis. 2023, 20, 375–380. [Google Scholar] [CrossRef]
- Kenyon, J.; Inns, T.; Aird, H.; Swift, C.; Astbury, J.; Forester, E.; Decraene, V. Campylobacter Outbreak Associated with Raw Drinking Milk, North West England, 2016. Epidemiol. Infect. 2020, 148, e13. [Google Scholar] [CrossRef]
- Davis, K.R.; Dunn, A.C.; Burnett, C.; McCullough, L.; Dimond, M.; Wagner, J.; Smith, L.; Carter, A.; Willardson, S.; Nakashima, A.K. Campylobacter jejuni Infections Associated with Raw Milk Consumption — Utah, 2014. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 301–305. [Google Scholar] [CrossRef]
- Nichols, M.; Conrad, A.; Whitlock, L.; Stroika, S.; Strain, E.; Weltman, A.; Johnson, L.; DeMent, J.; Reporter, R.; Williams, I. Short Communication: Multistate Outbreak of Listeria Monocytogenes Infections Retrospectively Linked to Unpasteurized Milk Using Whole-Genome Sequencing. J. Dairy Sci. 2020, 103, 176–178. [Google Scholar] [CrossRef]
- Napoleoni, M.; Villa, L.; Barco, L.; Busani, L.; Cibin, V.; Lucarelli, C.; Tiengo, A.; Dionisi, A.; Conti, F.; Da Silva Nunes, F.; et al. A Strong Evidence Outbreak of Salmonella Enteritidis in Central Italy Linked to the Consumption of Contaminated Raw Sheep Milk Cheese. Microorganisms 2021, 9, 2464. [Google Scholar] [CrossRef]
- Weinstein, E.; Lamba, K.; Bond, C.; Peralta, V.; Needham, M.; Beam, S.; Arroyo, F.; Kiang, D.; Chen, Y.; Shah, S.; et al. Outbreak of Salmonella Typhimurium Infections Linked to Commercially Distributed Raw Milk — California and Four Other States, September 2023–March 2024. MMWR Morb. Mortal. Wkly. Rep. 2025, 74, 433–438. [Google Scholar] [CrossRef]
- ISO 4833-1:2013; Microbiology of the Food Chain—Horizontal Method for the Enumeration of Microorganisms—Part 1: Colony Count at 30 °C by the Pour Plate Technique. International Organization for Standardization: Geneva, Switzerland, 2013.
- ISO 16649-2:2001; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of β-Glucuronidase-Positive Escherichia coli—Part 2: Colony-Count Technique at 44 °C Using 5-Bromo-4-Chloro-3-Indolyl β-D-Glucuronide. International Organization for Standardization: Geneva, Switzerland, 2001.
- ISO 6888-2:2021; Microbiology of the Food Chain—Horizontal Method for the Enumeration of Coagulase-Positive Staphylococci (Staphylococcus aureus and Other Species). International Organization for Standardization: Geneva, Switzerland, 2021.
- ISO 10272-1:2017; Microbiology of the Food Chain—Horizontal Method for Detection and Enumeration of Campylobacter spp. International Organization for Standardization: Geneva, Switzerland.
- ISO 10273:2017; Microbiology of the Food Chain—Horizontal Method for the Detection of Pathogenic Yersinia enterocolitica. International Organization for Standardization: Geneva, Switzerland, 2017.
- ISO 11290-1:2017; Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria Monocytogenes and of Listeria spp. International Organization for Standardization: Geneva, Switzerland, 2017.
- ISO 6579-1:2017; Microbiology of the Food Chain—Horizontal Method for the Detection, Enumeration and Serotyping of Salmonella—Part 1: Detection of Salmonella spp. International Organization for Standardization: Geneva, Switzerland, 2017.
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Souvorov, A.; Agarwala, R.; Lipman, D.J. SKESA: Strategic k-Mer Extension for Scrupulous Assemblies. Genome Biol. 2018, 19, 153. [Google Scholar] [CrossRef]
- Seemann, T. Shovill. Available online: https://github.com/tseemann/shovill (accessed on 13 January 2026).
- Konstantinidis, K.T.; Tiedje, J.M. Genomic Insights That Advance the Species Definition for Prokaryotes. Proc. Natl. Acad. Sci. USA 2005, 102, 2567–2572. [Google Scholar] [CrossRef]
- Jain, C.; Rodriguez-R, L.M.; Phillippy, A.M.; Konstantinidis, K.T.; Aluru, S. High Throughput ANI Analysis of 90K Prokaryotic Genomes Reveals Clear Species Boundaries. Nat. Commun. 2018, 9, 5114. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Liu, B.; Zheng, D.; Chen, L.; Yang, J. VFDB 2025: An Integrated Resource for Exploring Anti-Virulence Compounds. Nucleic Acids Res. 2025, 53, D871–D877. [Google Scholar] [CrossRef] [PubMed]
- Feldgarden, M.; Brover, V.; Gonzalez-Escalona, N.; Frye, J.G.; Haendiges, J.; Haft, D.H.; Hoffmann, M.; Pettengill, J.B.; Prasad, A.B.; Tillman, G.E.; et al. AMRFinderPlus and the Reference Gene Catalog Facilitate Examination of the Genomic Links among Antimicrobial Resistance, Stress Response, and Virulence. Sci. Rep. 2021, 11, 12728. [Google Scholar] [CrossRef] [PubMed]
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.-L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: Antibiotic Resistome Surveillance with the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2019, 48, D517–D525. [Google Scholar] [CrossRef]
- Ruppitsch, W.; Pietzka, A.; Prior, K.; Bletz, S.; Fernandez, H.L.; Allerberger, F.; Harmsen, D.; Mellmann, A. Defining and Evaluating a Core Genome Multilocus Sequence Typing Scheme for Whole-Genome Sequence-Based Typing of Listeria Monocytogenes. J. Clin. Microbiol. 2015, 53, 2869–2876. [Google Scholar] [CrossRef]
- Enright, M.C.; Day, N.P.J.; Davies, C.E.; Peacock, S.J.; Spratt, B.G. Multilocus Sequence Typing for Characterization of Methicillin-Resistant and Methicillin-Susceptible Clones of Staphylococcus aureus. J. Clin. Microbiol. 2000, 38, 1008–1015. [Google Scholar] [CrossRef] [PubMed]
- Leopold, S.R.; Goering, R.V.; Witten, A.; Harmsen, D.; Mellmann, A. Bacterial Whole-Genome Sequencing Revisited: Portable, Scalable, and Standardized Analysis for Typing and Detection of Virulence and Antibiotic Resistance Genes. J. Clin. Microbiol. 2014, 52, 2365–2370. [Google Scholar] [CrossRef]
- Nennig, M.; Llarena, A.-K.; Herold, M.; Mossong, J.; Penny, C.; Losch, S.; Tresse, O.; Ragimbeau, C. Investigating Major Recurring Campylobacter Jejuni Lineages in Luxembourg Using Four Core or Whole Genome Sequencing Typing Schemes. Front. Cell. Infect. Microbiol. 2021, 10, 608020. [Google Scholar] [CrossRef]
- Stevens, M.J.A.; Horlbog, J.A.; Diethelm, A.; Stephan, R.; Nüesch-Inderbinen, M. Characteristics and Comparative Genome Analysis of Yersinia Enterocolitica and Related Species Associated with Human Infections in Switzerland 2019–2023. Infect. Genet. Evol. 2024, 123, 105652. [Google Scholar] [CrossRef]
- Alikhan, N.-F.; Zhou, Z.; Sergeant, M.J.; Achtman, M. A Genomic Overview of the Population Structure of Salmonella. PLoS Genet. 2018, 14, e1007261. [Google Scholar] [CrossRef]
- Wirth, T.; Falush, D.; Lan, R.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.R.; Maiden, M.C.J.; Ochman, H.; et al. Sex and Virulence in Escherichia coli: An Evolutionary Perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef] [PubMed]
- Eidgenössisches Departement des Innern (EDI). Verordnung Des EDI Über Die Hygiene Beim Umgang Mit Lebensmitteln (HyV, SR 817.024.1); Eidgenössisches Departement des Innern (EDI): Bern, Switzerland, 2016.
- Bundesamt für Lebensmittelsicherheit und Veterinärwesen (BLV). Die Amtliche Milchprüfung 2024; Bundesamt für Lebensmittelsicherheit und Veterinärwesen: Bern, Switzerland, 2025. [Google Scholar]
- Mikulec, N.; Špoljarić, J.; Plavljanić, D.; Darrer, M.; Oštarić, F.; Gajdoš Kljusurić, J.; Sarim, K.M.; Zdolec, N.; Kazazić, S. Microbiota Composition in Raw Drinking Milk from Vending Machines: A Case Study in Croatia. Fermentation 2025, 11, 55. [Google Scholar] [CrossRef]
- Brooks, J.D.; Flint, S.H. Biofilms in the Food Industry: Problems and Potential Solutions. Int. J. Food Sci. Technol. 2008, 43, 2163–2176. [Google Scholar] [CrossRef]
- Latorre, A.A.; Van Kessel, J.S.; Karns, J.S.; Zurakowski, M.J.; Pradhan, A.K.; Boor, K.J.; Jayarao, B.M.; Houser, B.A.; Daugherty, C.S.; Schukken, Y.H. Biofilm in Milking Equipment on a Dairy Farm as a Potential Source of Bulk Tank Milk Contamination with Listeria Monocytogenes. J. Dairy Sci. 2010, 93, 2792–2802. [Google Scholar] [CrossRef]
- Sharma, M.; Anand, S.K. Characterization of Constitutive Microflora of Biofilms in Dairy Processing Lines. Food Microbiol. 2002, 19, 627–636. [Google Scholar] [CrossRef]
- Cherif-Antar, A.; Moussa–Boudjemâa, B.; Didouh, N.; Medjahdi, K.; Mayo, B.; Flórez, A.B. Diversity and Biofilm-Forming Capability of Bacteria Recovered from Stainless Steel Pipes of a Milk-Processing Dairy Plant. Dairy Sci. Technol. 2016, 96, 27–38. [Google Scholar] [CrossRef]
- Bardbari, A.M.; Arabestani, M.R.; Karami, M.; Keramat, F.; Alikhani, M.Y.; Bagheri, K.P. Correlation between Ability of Biofilm Formation with Their Responsible Genes and MDR Patterns in Clinical and Environmental Acinetobacter Baumannii Isolates. Microb. Pathog. 2017, 108, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Idland, L.; Bø-Granquist, E.G.; Aspholm, M.; Lindbäck, T. The Ability of Shiga Toxin-Producing Escherichia Coli to Grow in Raw Cow’s Milk Stored at Low Temperatures. Foods 2022, 11, 3411. [Google Scholar] [CrossRef]
- Wang, G.; Zhao, T.; Doyle, M.P. Survival and Growth of Escherichia Coli O157:H7 in Unpasteurized and Pasteurized Milk. J. Food Prot. 1997, 60, 610–613. [Google Scholar] [CrossRef]
- Fujikawa, H.; Morozumi, S. Modeling Staphylococcus Aureus Growth and Enterotoxin Production in Milk. Food Microbiol. 2006, 23, 260–267. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, J.; Jin, J.; Li, X.; Zhang, H.; Shi, X.; Zhao, C. Prevalence, Antibiotic Resistance, and Enterotoxin Genes of Staphylococcus Aureus Isolated from Milk and Dairy Products Worldwide: A Systematic Review and Meta-Analysis. Food Res. Int. 2022, 162, 111969. [Google Scholar] [CrossRef]
- Riva, A.; Borghi, E.; Cirasola, D.; Colmegna, S.; Borgo, F.; Amato, E.; Pontello, M.M.; Morace, G. Methicillin-Resistant Staphylococcus Aureus in Raw Milk: Prevalence, SCCmec Typing, Enterotoxin Characterization, and Antimicrobial Resistance Patterns. J. Food Prot. 2015, 78, 1142–1146. [Google Scholar] [CrossRef]
- Kozak, O.; Jan, P.; Gazzarin, C. Global Dairy Sector: Past Development and Outlook: Insights from the International Farm Comparison Network; Agroscope: Posieux, Switzerland, 2024; Volume 193. [Google Scholar]
- Stephan, R.; Annemüller, C.; Hassan, A.A.; Lämmler, C. Characterization of Enterotoxigenic Staphylococcus Aureus Strains Isolated from Bovine Mastitis in North-East Switzerland. Vet. Microbiol. 2001, 78, 373–382. [Google Scholar] [CrossRef]
- Käppeli, N.; Morach, M.; Corti, S.; Eicher, C.; Stephan, R.; Johler, S. Staphylococcus Aureus Related to Bovine Mastitis in Switzerland: Clonal Diversity, Virulence Gene Profiles, and Antimicrobial Resistance of Isolates Collected throughout 2017. J. Dairy Sci. 2019, 102, 3274–3281. [Google Scholar] [CrossRef]
- Hoekstra, J.; Zomer, A.L.; Rutten, V.P.M.G.; Benedictus, L.; Stegeman, A.; Spaninks, M.P.; Bennedsgaard, T.W.; Biggs, A.; De Vliegher, S.; Mateo, D.H.; et al. Genomic Analysis of European Bovine Staphylococcus Aureus from Clinical versus Subclinical Mastitis. Sci. Rep. 2020, 10, 18172. [Google Scholar] [CrossRef]
- Sakwinska, O.; Morisset, D.; Madec, J.-Y.; Waldvogel, A.; Moreillon, P.; Haenni, M. Link between Genotype and Antimicrobial Resistance in Bovine Mastitis-Related Staphylococcus Aureus Strains, Determined by Comparing Swiss and French Isolates from the Rhône Valley. Appl. Environ. Microbiol. 2011, 77, 3428–3432. [Google Scholar] [CrossRef] [PubMed]
- Fitzgerald, J.R. Livestock-Associated Staphylococcus Aureus: Origin, Evolution and Public Health Threat. Trends Microbiol. 2012, 20, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Moser, A.; Stephan, R.; Corti, S.; Lehner, A. Resistance Profiles and Genetic Diversity of Escherichia Coli Strains Isolated from Acute Bovine Mastitis. Schweiz. Arch. Für Tierheilkd. 2013, 155, 351–357. [Google Scholar] [CrossRef][Green Version]
- Boss, R.; Cosandey, A.; Luini, M.; Artursson, K.; Bardiau, M.; Breitenwieser, F.; Hehenberger, E.; Lam, T.; Mansfeld, M.; Michel, A.; et al. Bovine Staphylococcus Aureus: Subtyping, Evolution, and Zoonotic Transfer. J. Dairy Sci. 2016, 99, 515–528. [Google Scholar] [CrossRef]
- Ramezanigardaloud, N.; Loncaric, I.; Mikuni-Mester, P.; Alinaghi, M.; Ehling-Schulz, M.; Khol, J.L.; Grunert, T. Prevalence and Diversity of Staphylococcus Aureus in Bulk Tank Milk from Community-Based Alpine Dairy Pastures in Tyrol, Austria. Animals 2025, 15, 2153. [Google Scholar] [CrossRef] [PubMed]
- Graber, H.U.; Naskova, J.; Studer, E.; Kaufmann, T.; Kirchhofer, M.; Brechbühl, M.; Schaeren, W.; Steiner, A.; Fournier, C. Mastitis-Related Subtypes of Bovine Staphylococcus Aureus Are Characterized by Different Clinical Properties. J. Dairy Sci. 2009, 92, 1442–1451. [Google Scholar] [CrossRef]
- Hummerjohann, J.; Naskova, J.; Baumgartner, A.; Graber, H.U. Enterotoxin-Producing Staphylococcus Aureus Genotype B as a Major Contaminant in Swiss Raw Milk Cheese. J. Dairy Sci. 2014, 97, 1305–1312. [Google Scholar] [CrossRef]
- Kaakoush, N.O.; Castaño-Rodríguez, N.; Mitchell, H.M.; Man, S.M. Global Epidemiology of Campylobacter Infection. Clin. Microbiol. Rev. 2015, 28, 687–720. [Google Scholar] [CrossRef]
- Humphrey, T.; O’Brien, S.; Madsen, M. Campylobacters as Zoonotic Pathogens: A Food Production Perspective. Int. J. Food Microbiol. 2007, 117, 237–257. [Google Scholar] [CrossRef]
- Bronowski, C.; James, C.E.; Winstanley, C. Role of Environmental Survival in Transmission of Campylobacter jejuni. FEMS Microbiol. Lett. 2014, 356, 8–19. [Google Scholar] [CrossRef]
- Gras, L.M.; Smid, J.H.; Wagenaar, J.A.; Koene, M.G.J.; Havelaar, A.H.; Friesema, I.H.M.; French, N.P.; Flemming, C.; Galson, J.D.; Graziani, C.; et al. Increased Risk for Campylobacter jejuni and C. coli Infection of Pet Origin in Dog Owners and Evidence for Genetic Association between Strains Causing Infection in Humans and Their Pets. Epidemiol. Infect. 2013, 141, 2526–2535. [Google Scholar] [CrossRef] [PubMed]
- Al-Saigh, H.; Zweifel, C.; Blanco, J.; Blanco, J.E.; Blanco, M.; Usera, M.A.; Stephan, R. Fecal Shedding of Escherichia Coli O157, Salmonella, and Campylobacter in Swiss Cattle at Slaughter. J. Food Prot. 2004, 67, 679–684. [Google Scholar] [CrossRef]
- Taghizadeh, M.; Nematollahi, A.; Bashiry, M.; Javanmardi, F.; Mousavi, M.; Hosseini, H. The Global Prevalence of Campylobacter Spp. in Milk: A Systematic Review and Meta-Analysis. Int. Dairy J. 2022, 133, 105423. [Google Scholar] [CrossRef]
- Wysok, B.; Rudowska, M.; Wiszniewska-Łaszczych, A. The Transmission of Campylobacter Strains in Dairy Herds in Different Housing Systems. Pathogens 2024, 13, 317. [Google Scholar] [CrossRef] [PubMed]
- Oliver, S.P.; Jayarao, B.M.; Almeida, R.A. Foodborne Pathogens in Milk and the Dairy Farm Environment: Food Safety and Public Health Implications. Foodborne Pathog. Dis. 2005, 2, 115–129. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA); European Centre for Disease Prevention and Control (ECDC). The European Union One Health 2024 Zoonoses Report. EFSA J. 2025, 23, e9759. [Google Scholar] [CrossRef]
- Jonas, R.; Kittl, S.; Overesch, G.; Kuhnert, P. Genotypes and Antibiotic Resistance of Bovine Campylobacter and Their Contribution to Human Campylobacteriosis. Epidemiol. Infect. 2015, 143, 2373–2380. [Google Scholar] [CrossRef]
- French, N.; Barrigas, M.; Brown, P.; Ribiero, P.; Williams, N.; Leatherbarrow, H.; Birtles, R.; Bolton, E.; Fearnhead, P.; Fox, A. Spatial Epidemiology and Natural Population Structure of Campylobacter jejuni Colonizing a Farmland Ecosystem. Environ. Microbiol. 2005, 7, 1116–1126. [Google Scholar] [CrossRef]
- Ghielmetti, G.; Seth-Smith, H.M.B.; Roloff, T.; Cernela, N.; Biggel, M.; Stephan, R.; Egli, A. Whole-Genome-Based Characterization of Campylobacter jejuni from Human Patients with Gastroenteritis Collected over an 18 Year Period Reveals Increasing Prevalence of Antimicrobial Resistance. Microb. Genom. 2023, 9, 000941. [Google Scholar] [CrossRef]
- Terentjeva, M.; Ķibilds, J.; Gradovska, S.; Alksne, L.; Streikiša, M.; Meistere, I.; Valciņa, O. Prevalence, Virulence Determinants, and Genetic Diversity in Yersinia Enterocolitica Isolated from Slaughtered Pigs and Pig Carcasses. Int. J. Food Microbiol. 2022, 376, 109756. [Google Scholar] [CrossRef]
- McNally, A.; Cheasty, T.; Fearnley, C.; Dalziel, R.W.; Paiba, G.A.; Manning, G.; Newell, D.G. Comparison of the Biotypes of Yersinia Enterocolitica Isolated from Pigs, Cattle and Sheep at Slaughter and from Humans with Yersiniosis in Great Britain during 1999–2000. Lett. Appl. Microbiol. 2004, 39, 103–108. [Google Scholar] [CrossRef]
- Grahek-Ogden, D.; Schimmer, B.; Cudjoe, K.S.; Nygård, K.; Kapperud, G. Outbreak of Yersinia Enterocolitica Serogroup O:9 Infection and Processed Pork, Norway. Emerg. Infect. Dis. 2007, 13, 754–756. [Google Scholar] [CrossRef]
- Gruber, J.F.; Morris, S.; Warren, K.A.; Kline, K.E.; Schroeder, B.; Dettinger, L.; Husband, B.; Pollard, K.; Davis, C.; Miller, J.; et al. Yersinia enterocolitica Outbreak Associated with Pasteurized Milk. Foodborne Pathog. Dis. 2021, 18, 448–454. [Google Scholar] [CrossRef] [PubMed]
- Savin, C.; Fredriksen, N.; Calba, C.; Puzin, L.; Felten, A.; Larréché, S.; Roche, A.; Denis, M.; Le Guern, A.-S.; Chereau, F.; et al. Cross-Border Outbreak of Yersinia Enterocolitica Bioserotype 2/O:9 Infections Associated with Consumption of French Unpasteurised Soft Goat’s Milk Cheese, 2024. Eurosurveillance 2025, 30, 2500002. [Google Scholar] [CrossRef]
- Espenhain, L.; Riess, M.; Müller, L.; Colombe, S.; Ethelberg, S.; Litrup, E.; Jernberg, C.; Kühlmann-Berenzon, S.; Lindblad, M.; Hove, N.K.; et al. Cross-Border Outbreak of Yersinia Enterocolitica O3 Associated with Imported Fresh Spinach, Sweden and Denmark, March 2019. Eurosurveillance 2019, 24, 1900368. [Google Scholar] [CrossRef]
- MacDonald, E.; Heier, B.T.; Nygård, K.; Stalheim, T.; Cudjoe, K.S.; Skjerdal, T.; Wester, A.L.; Lindstedt, B.-A.; Stavnes, T.-L.; Vold, L. Yersinia enterocolitica Outbreak Associated with Ready-to-Eat Salad Mix, Norway, 2011. Emerg. Infect. Dis. 2012, 18, 1496–1499. [Google Scholar] [CrossRef] [PubMed]
- Wauters, G.; Kandolo, K.; Janssens, M. Revised Biogrouping Scheme of Yersinia Enterocolitica. Contrib. Microbiol. Immunol. 1987, 9, 14–21. [Google Scholar] [PubMed]
- Bottone, E.J. Yersinia Enterocolitica: Overview and Epidemiologic Correlates. Microbes Infect. 1999, 1, 323–333. [Google Scholar] [CrossRef]
- Rakin, A.; Noelting, C.; Schubert, S.; Heesemann, J. Common and Specific Characteristics of the High-Pathogenicity Island of Yersinia Enterocolitica. Infect. Immun. 1999, 67, 5265–5274. [Google Scholar] [CrossRef]
- Bancerz-Kisiel, A.; Pieczywek, M.; Łada, P.; Szweda, W. The Most Important Virulence Markers of Yersinia Enterocolitica and Their Role during Infection. Genes 2018, 9, 235. [Google Scholar] [CrossRef]
- Wren, B.W. The Yersiniae—A Model Genus to Study the Rapid Evolution of Bacterial Pathogens. Nat. Rev. Microbiol. 2003, 1, 55–64. [Google Scholar] [CrossRef]
- Grant, T.; Bennett-Wood, V.; Robins-Browne, R.M. Identification of Virulence-Associated Characteristics in Clinical Isolates of Yersinia enterocolitica Lacking Classical Virulence Markers. Infect. Immun. 1998, 66, 1113–1120. [Google Scholar] [CrossRef]
- Seabaugh, J.A.; Anderson, D.M. Pathogenicity and Virulence of Yersinia. Virulence 2024, 15, 2316439. [Google Scholar] [CrossRef] [PubMed]
- Nieckarz, M.; Jaworska, K.; Raczkowska, A.; Brzostek, K. The Regulatory Circuit Underlying Downregulation of a Type III Secretion System in Yersinia Enterocolitica by Transcription Factor OmpR. Int. J. Mol. Sci. 2022, 23, 4758. [Google Scholar] [CrossRef]
- Sihvonen, L.M.; Hallanvuo, S.; Haukka, K.; Skurnik, M.; Siitonen, A. The Ail Gene Is Present in Some Yersinia enterocolitica Biotype 1A Strains. Foodborne Pathog. Dis. 2011, 8, 455–457. [Google Scholar] [CrossRef]
- Platt-Samoraj, A. Toxigenic Properties of Yersinia Enterocolitica Biotype 1A. Toxins 2022, 14, 118. [Google Scholar] [CrossRef]
- Ermoli, F.; Malengo, G.; Spahn, C.; Brianceau, C.; Glatter, T.; Diepold, A. Yersinia Actively Downregulates Type III Secretion and Adhesion at Higher Cell Densities. PLoS Pathog. 2025, 21, e1013423. [Google Scholar] [CrossRef]
- Ruusunen, M.; Salonen, M.; Pulkkinen, H.; Huuskonen, M.; Hellström, S.; Revez, J.; Hänninen, M.-L.; Fredriksson-Ahomaa, M.; Lindström, M. Pathogenic Bacteria in Finnish Bulk Tank Milk. Foodborne Pathog. Dis. 2013, 10, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Jayarao, B.M.; Henning, D.R. Prevalence of Foodborne Pathogens in Bulk Tank Milk. J. Dairy Sci. 2001, 84, 2157–2162. [Google Scholar] [CrossRef]
- Jayarao, B.M.; Donaldson, S.C.; Straley, B.A.; Sawant, A.A.; Hegde, N.V.; Brown, J.L. A Survey of Foodborne Pathogens in Bulk Tank Milk and Raw Milk Consumption Among Farm Families in Pennsylvania. J. Dairy Sci. 2006, 89, 2451–2458. [Google Scholar] [CrossRef]
- Łada, P.; Kończyk-Kmiecik, K.; Bancerz-Kisiel, A. Isolation, Characterization and Antimicrobial Resistance of Yersinia Enterocolitica from Polish Cattle and Their Carcasses. BMC Vet. Res. 2023, 19, 143. [Google Scholar] [CrossRef]
- Bottone, E.J. Yersinia Enterocolitica: Revisitation of an Enduring Human Pathogen. Clin. Microbiol. Newsl. 2015, 37, 1–8. [Google Scholar] [CrossRef]
- Bari, M.L.; Hossain, M.A.; Isshiki, K.; Ukuku, D. Behavior of Yersinia enterocolitica in Foods. J. Pathog. 2011, 2011, 420732. [Google Scholar] [CrossRef]
- Hunter, E.; Greig, D.R.; Schaefer, U.; Wright, M.J.; Dallman, T.J.; McNally, A.; Jenkins, C. Identification and Typing of Yersinia Enterocolitica and Yersinia Pseudotuberculosis Isolated from Human Clinical Specimens in England between 2004 and 2018. J. Med. Microbiol. 2019, 68, 538–548. [Google Scholar] [CrossRef] [PubMed]
- Piras, F.; Siddi, G.; Le Guern, A.-S.; Brémont, S.; Fredriksson-Ahomaa, M.; Sanna, R.; Meloni, M.P.; De Santis, E.P.L.; Scarano, C. Traceability, Virulence and Antimicrobial Resistance of Yersinia Enterocolitica in Two Industrial Cheese-Making Plants. Int. J. Food Microbiol. 2023, 398, 110225. [Google Scholar] [CrossRef]
- Yu, M.; Wang, Y.; Hao, X.; Jiang, L.; Liu, D.; Sun, Y.; Xu, J.; Peng, Z. Phylogenetic Relationship, Genomic Heterogeneity, and Population Structure of Yersinia Enterocolitica Biotype 1A Isolated from Pork and Poultry Meat. Food Res. Int. 2025, 212, 116405. [Google Scholar] [CrossRef]
- Stevens, M.J.A.; Barmettler, K.; Kelbert, L.; Stephan, R.; Nüesch-Inderbinen, M. Genome Based Characterization of Yersinia Enterocolitica from Different Food Matrices in Switzerland in 2024. Infect. Genet. Evol. 2025, 128, 105719. [Google Scholar] [CrossRef]
- Shoaib, M.; Shehzad, A.; Raza, H.; Niazi, S.; Khan, I.M.; Akhtar, W.; Safdar, W.; Wang, Z. A Comprehensive Review on the Prevalence, Pathogenesis and Detection of Yersinia Enterocolitica. RSC Adv. 2019, 9, 41010–41021. [Google Scholar] [CrossRef]
- Oellerich, M.F.; Jacobi, C.A.; Freund, S.; Niedung, K.; Bach, A.; Heesemann, J.; Trülzsch, K. Yersinia enterocolitica Infection of Mice Reveals Clonal Invasion and Abscess Formation. Infect. Immun. 2007, 75, 3802–3811. [Google Scholar] [CrossRef]
- Hunt, K.; Drummond, N.; Murphy, M.; Butler, F.; Buckley, J.; Jordan, K. A Case of Bovine Raw Milk Contamination with Listeria Monocytogenes. Ir. Vet. J. 2012, 65, 13. [Google Scholar] [CrossRef]
- Husu, J.R.; Seppänen, J.T.; Sivelä, S.K.; Rauramaa, A.L. Contamination of Raw Milk by Listeria monocytogenes on Dairy Farms. J. Vet. Med. Ser. B 1990, 37, 268–275. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, A.C.; Almeida, F.A.D.; Medeiros, M.M.; Miranda, B.R.; Pinto, U.M.; Alves, V.F. Listeria Monocytogenes: An Inconvenient Hurdle for the Dairy Industry. Dairy 2023, 4, 316–344. [Google Scholar] [CrossRef]
- Champagne, C.P.; Laing, R.R.; Roy, D.; Mafu, A.A.; Griffiths, M.W.; White, C. Psychrotrophs in Dairy Products: Their Effects and Their Control. Crit. Rev. Food Sci. Nutr. 1994, 34, 1–30. [Google Scholar] [CrossRef]
- EFSA Panel on Biological Hazards (BIOHAZ); Ricci, A.; Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Fernández Escámez, P.S.; Girones, R.; Herman, L.; Koutsoumanis, K.; et al. Listeria Monocytogenes Contamination of Ready-to-eat Foods and the Risk for Human Health in the EU. EFSA J. 2018, 16, e05134. [Google Scholar] [CrossRef] [PubMed]
- Fotopoulou, E.T.; Jenkins, C.; Painset, A.; Amar, C. Listeria Monocytogenes: The Silent Assassin. J. Med. Microbiol. 2024, 73, 001800. [Google Scholar] [CrossRef]
- Spoerry Serrano, N.; Zweifel, C.; Corti, S.; Stephan, R. Microbiological Quality and Presence of Foodborne Pathogens in Raw Milk Cheeses and Raw Meat Products Marketed at Farm Level in Switzerland. Ital. J. Food Saf. 2018, 7, 7337. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, D.M.; Barbaro, A.; Gallina, S.; Vitale, N.; Chiavacci, L.; Caramelli, M.; Decastelli, L. Monitoring of Foodborne Pathogenic Bacteria in Vending Machine Raw Milk in Piedmont, Italy. Food Control 2013, 32, 435–439. [Google Scholar] [CrossRef]
- Chiarlone, S.A.; Gori, A.; Ravetta, S.; Armani, A.; Guardone, L.; Pedonese, F.; Bavetta, S.; Fiannacca, C.; Pussini, N.; Maurella, C.; et al. Microbiological Analysis Conducted on Raw Milk Collected During Official Sampling in Liguria (North-West Italy) over a Ten-Year Period (2014–2023). Animals 2025, 15, 286. [Google Scholar] [CrossRef]
- Raschle, S.; Stephan, R.; Stevens, M.J.A.; Cernela, N.; Zurfluh, K.; Muchaamba, F.; Nüesch-Inderbinen, M. Environmental Dissemination of Pathogenic Listeria Monocytogenes in Flowing Surface Waters in Switzerland. Sci. Rep. 2021, 11, 9066. [Google Scholar] [CrossRef]
- Fagerlund, A.; Idland, L.; Heir, E.; Møretrø, T.; Aspholm, M.; Lindbäck, T.; Langsrud, S. Whole-Genome Sequencing Analysis of Listeria Monocytogenes from Rural, Urban, and Farm Environments in Norway: Genetic Diversity, Persistence, and Relation to Clinical and Food Isolates. Appl. Environ. Microbiol. 2022, 88, e02136-21. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control (ECDC). Listeriosis: Annual Epidemiological Report for 2022; ECDC: Stockholm, Sweden, 2024. [Google Scholar]
- European Centre for Disease Prevention and Control (ECDC). Listeriosis: Annual Epidemiological Report for 2023; ECDC: Stockholm, Sweden, 2025. [Google Scholar]
- Lassen, B.; Olsen, A.; Sandberg, M.; Müller, L.; Torpdahl, M.; Petersen, C.K. Annual Resport on Zoonoses in Denmark 2022; Technical University of Denmark: Kogens Lyngby, Denmark, 2023. [Google Scholar]
- Bundesamt für Lebensmittelsicherheit und Veterinärwesen (BLV) Fachinformation: Salmonellose. 2022. Available online: https://www.blv.admin.ch/blv/de/home/tiere/tierseuchen/uebersicht-seuchen/alle-tierseuchen/salmonellose.html (accessed on 27 October 2025).
- Spier, S.J.; Smith, B.P.; Cullor, J.S.; Olander, H.J.; Da Roden, L.; Dilling, G.W. Persistent Experimental Salmonella dublin Intramammary Infection in Dairy Cows. J. Vet. Intern. Med. 1991, 5, 341–350. [Google Scholar] [CrossRef]
- Dargatz, D.A.; Kopral, C.A.; Erdman, M.M.; Fedorka-Cray, P.J. Prevalence and Antimicrobial Resistance of Salmonella Isolated from Cattle Feces in United States Feedlots in 2011. Foodborne Pathog. Dis. 2016, 13, 483–489. [Google Scholar] [CrossRef]
- Bundesamt für Lebensmittelsicherheit und Veterinärwesen (BLV). Mikrobiologische Risikoevaluation Shigatoxin Produzierende E. coli (STEC) in Lebensmitteln; Bundesamt für Lebensmittelsicherheit und Veterinärwesen: Bern, Switzerland, 2019. [Google Scholar]
- EFSA BIOHAZ Panel; Koutsoumanis, K.; Allende, A.; Alvarez-Ordóñez, A.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Herman, L.; Hilbert, F.; et al. Pathogenicity Assessment of Shiga Toxin-producing Escherichia Coli (STEC) and the Public Health Risk Posed by Contamination of Food with STEC. EFSA J. 2020, 18, e05967. [Google Scholar] [CrossRef]
- Butcher, H.; Elson, R.; Chattaway, M.A.; Featherstone, C.A.; Willis, C.; Jorgensen, F.; Dallman, T.J.; Jenkins, C.; Mclauchlin, J.; Beck, C.R.; et al. Whole Genome Sequencing Improved Case Ascertainment in an Outbreak of Shiga Toxin-Producing Escherichia coli O157 Associated with Raw Drinking Milk. Epidemiol. Infect. 2016, 144, 2812–2823. [Google Scholar] [CrossRef]
- Treacy, J.; Jenkins, C.; Paranthaman, K.; Jorgensen, F.; Mueller-Doblies, D.; Anjum, M.; Kaindama, L.; Hartman, H.; Kirchner, M.; Carson, T.; et al. Outbreak of Shiga Toxin-Producing Escherichia Coli O157:H7 Linked to Raw Drinking Milk Resolved by Rapid Application of Advanced Pathogen Characterisation Methods, England, August to October 2017. Eurosurveillance 2019, 24, 1800191. [Google Scholar] [CrossRef] [PubMed]
- Kirchner, M.; Dildei, C.; Runge, M.; Brix, A.; Claussen, K.; Weiss, U.; Fruth, A.; Prager, R.; Mellmann, A.; Beutin, L.; et al. Outbreak of Non-Sorbitol-Fermenting Shiga Toxin-Producing E. Coli O157:H7 Infections among School Children Associated with Raw Milk Consumption in Germany. J. Food Saf. Food Qual. Arch. Für Leb. 2013, 64, 68–74. [Google Scholar] [CrossRef]
- Huber, H.; Koller, S.; Giezendanner, N.; Stephan, R.; Zweifel, C. Prevalence and Characteristics of Meticillin-Resistant Staphylococcus Aureus in Humans in Contact with Farm Animals, in Livestock, and in Food of Animal Origin, Switzerland, 2009. Euro Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull. 2010, 15, 19542. [Google Scholar] [CrossRef]
- Paterson, G.K.; Larsen, J.; Harrison, E.M.; Larsen, A.R.; Morgan, F.J.; Peacock, S.J.; Parkhill, J.; Zadoks, R.N.; Holmes, M.A. First Detection of Livestock-Associated Meticillin-Resistant Staphylococcus Aureus CC398 in Bulk Tank Milk in the United Kingdom, January to July 2012. Euro Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull. 2012, 17, 20337. [Google Scholar] [CrossRef]
- Schnitt, A.; Lienen, T.; Wichmann-Schauer, H.; Cuny, C.; Tenhagen, B.-A. The Occurrence and Distribution of Livestock-Associated Methicillin-Resistant Staphylococcus Aureus ST398 on German Dairy Farms. J. Dairy Sci. 2020, 103, 11806–11819. [Google Scholar] [CrossRef]
- Vanderhaeghen, W.; Cerpentier, T.; Adriaensen, C.; Vicca, J.; Hermans, K.; Butaye, P. Methicillin-Resistant Staphylococcus Aureus (MRSA) ST398 Associated with Clinical and Subclinical Mastitis in Belgian Cows. Vet. Microbiol. 2010, 144, 166–171. [Google Scholar] [CrossRef] [PubMed]
- Katayama, Y.; Ito, T.; Hiramatsu, K. A New Class of Genetic Element, Staphylococcus Cassette Chromosome Mec, Encodes Methicillin Resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 2000, 44, 1549–1555. [Google Scholar] [CrossRef]
- Nüesch-Inderbinen, M.; Stephan, R. Epidemiology of Extended-Spectrum β-Lactamase-Producing Escherichia Coli in the Human-Livestock Environment. Curr. Clin. Microbiol. Rep. 2016, 3, 1–9. [Google Scholar] [CrossRef]
- Bush, K. Proliferation and Significance of Clinically Relevant Β-lactamases. Ann. N. Y. Acad. Sci. 2013, 1277, 84–90. [Google Scholar] [CrossRef]
- Geser, N.; Stephan, R.; Hächler, H. Occurrence and Characteristics of Extended-Spectrum β-Lactamase (ESBL) Producing Enterobacteriaceae in Food Producing Animals, Minced Meat and Raw Milk. BMC Vet. Res. 2012, 8, 21. [Google Scholar] [CrossRef] [PubMed]
- Odenthal, S.; Akineden, Ö.; Usleber, E. Extended-Spectrum β-Lactamase Producing Enterobacteriaceae in Bulk Tank Milk from German Dairy Farms. Int. J. Food Microbiol. 2016, 238, 72–78. [Google Scholar] [CrossRef] [PubMed]


| Pseudomonas spp. | Acinetobacter spp. | Lactococcus spp. | Staphylococcus spp. | Yersinia spp. | Other Genera 1 |
|---|---|---|---|---|---|
| Pseudomonas fragi (6 *) | Acinetobacter johnsonii (3) | Lactococcus lactis (2) | Staphylococcus xylosus (1) | Yersinia enterocolitica (1) | Carnobacterium maltaromaticum (2) |
| Pseudomonas lundensis (3) | Acinetobacter guilloiae (1) | Lactococcus raffinolactis (1) | Staphylococcus hominis (1) | Enterobacter hormaechei (1) | |
| Pseudomonas brenneri (1) | Acinetobacter albensis (1) | Rahnella inusitata (1) | |||
| Pseudomonas corrugate (1) | Raoultella terrigena (1) | ||||
| Pseudomonas antarctica (1) | Erwinia persicina (1) | ||||
| Pseudomonas libanensis (1) | Hafnia alvei (1) | ||||
| Pseudomonas taetrolens (1) | Comamonas testosteroni (1) |
| Sample ID | Bacterial Count (CFU/mL) | ST | Enterotoxin Genes (SE) |
|---|---|---|---|
| TP 150 | 8 × 101 | 8 | |
| TP 151 | 1 × 102 | 8 | |
| TP 159 | 3 × 101 | 97 | |
| TP 95 | 1.3 × 102 | 151 | |
| TP 98 | 3 × 101 | 151 | |
| TP 2 | 3 × 101 | 352 | |
| TP 5 | 1 × 101 | 352 | |
| TP 28 | 3 × 101 | 352 | |
| TP 87 | 3 × 101 | 352 | |
| TP 89 | 4 × 101 | 352 | |
| TP 100 | 2 × 101 | 352 | |
| TP 56 | 2.2 × 102 | 389 | |
| TP 59 | 5 × 101 | 504 | |
| TP 106 | 1 × 101 | 504 | sec, sell |
| TP 137 | 9 × 101 | 582 |
| Sample ID | Bacterial Count (CFU/mL) (If Quantitative Detected) | BT | ST |
|---|---|---|---|
| TP 1 | - | 1A | n/d * |
| TP 3 | 1.0 × 103 | 1A | 3 |
| TP 5 | - | 1A | 118 |
| TP 7 | - | 1A | n/d |
| TP 8 | 1.3 × 103 | 1A | n/d |
| TP 11 | 1.0 × 101 | 1A | n/d |
| TP 12 | - | 1A | n/d |
| TP 14 | - | 1A | n/d |
| TP 22 | - | 1A | n/d |
| TP 27 | - | 1A | n/d |
| TP 29 | 4.7 × 102 | 1A | n/d |
| TP 45 | - | 1A | n/d |
| TP 57 | 1.1 × 105 | 1A | 118 |
| TP 59 | - | 1A | 118 |
| TP 63 | - | 1A | 8 |
| TP 69 | 2.2 × 102 | 1A | 8 |
| TP 73 | - | - | - |
| TP 77 | 9.2 × 104 | - | - |
| TP 93 | - | 1A | 3 |
| TP 101 | 1.4 × 102 | 1A | n/d |
| TP 104 | 2.3 × 104 | 1A | 118 |
| TP 106 | - | 1A | 118 |
| TP 111 | - | n/d | n/d |
| TP 113 | 6.4 × 104 | 1A | 157 |
| TP 116 | - | 1A | 118 |
| TP 118 | - | 1A | n/d |
| TP 123 | 3.2 × 102 | 1A | 118 |
| TP 125 | 1.9 × 106 | 1A | 3 |
| TP 126 | - | 1A | n/d |
| TP 127 | 2.0 × 105 | 1A | n/d |
| TP 138 | - | 1A | 3 |
| TP 142 | 2.2 × 103 | 1A | n/d |
| TP 149 | 4.0 × 102 | 1A | n/d |
| TP 152 | 1.0 × 103 | 1A | n/d |
| TP 154 | - | 1A | n/d |
| TP 156 | 9.0 × 103 | 1A | n/d |
| TP 161 | - | 1A | n/d |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Paravicini, T.; Stevens, M.J.A.; Barmettler, K.; Cernela, N.; Stephan, R. Assessment of the Microbiological Quality of Raw Milk Sold Through Vending Machines at the Farm Level in Switzerland. Pathogens 2026, 15, 322. https://doi.org/10.3390/pathogens15030322
Paravicini T, Stevens MJA, Barmettler K, Cernela N, Stephan R. Assessment of the Microbiological Quality of Raw Milk Sold Through Vending Machines at the Farm Level in Switzerland. Pathogens. 2026; 15(3):322. https://doi.org/10.3390/pathogens15030322
Chicago/Turabian StyleParavicini, Thomas, Marc J. A. Stevens, Karen Barmettler, Nicole Cernela, and Roger Stephan. 2026. "Assessment of the Microbiological Quality of Raw Milk Sold Through Vending Machines at the Farm Level in Switzerland" Pathogens 15, no. 3: 322. https://doi.org/10.3390/pathogens15030322
APA StyleParavicini, T., Stevens, M. J. A., Barmettler, K., Cernela, N., & Stephan, R. (2026). Assessment of the Microbiological Quality of Raw Milk Sold Through Vending Machines at the Farm Level in Switzerland. Pathogens, 15(3), 322. https://doi.org/10.3390/pathogens15030322





