Immunoinformatic Design and Evaluation of a Multi-Epitope mRNA Vaccine RP14914P Targeting Latent Tuberculosis Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Antigen Selection and Sequence Retrieval
2.2. Prediction and Screening of Immunodominant Epitopes
2.2.1. CTL Epitope Prediction
2.2.2. HTL Epitope Prediction
2.2.3. B-Cell Epitope Prediction
2.2.4. Epitope Homology Prediction
2.2.5. Epitope Safety Validation
2.3. mRNA Vaccine Construct Assembly
2.4. Analysis of Physicochemical Properties, Antigenicity, and Safety of the Candidate Vaccine
2.5. Selection of the Optimal Vaccine Construct
2.6. Secondary and Tertiary Structure Prediction and Validation
2.7. Immunoinformatics and Interaction Analysis
2.7.1. Population Coverage Analysis
2.7.2. Discontinuous B-Cell Epitope Prediction
2.7.3. Molecular Docking
2.7.4. Molecular Dynamics (MD) Simulation
2.7.5. Immune Simulation
2.8. Codon Optimization and mRNA Structure Prediction
3. Results
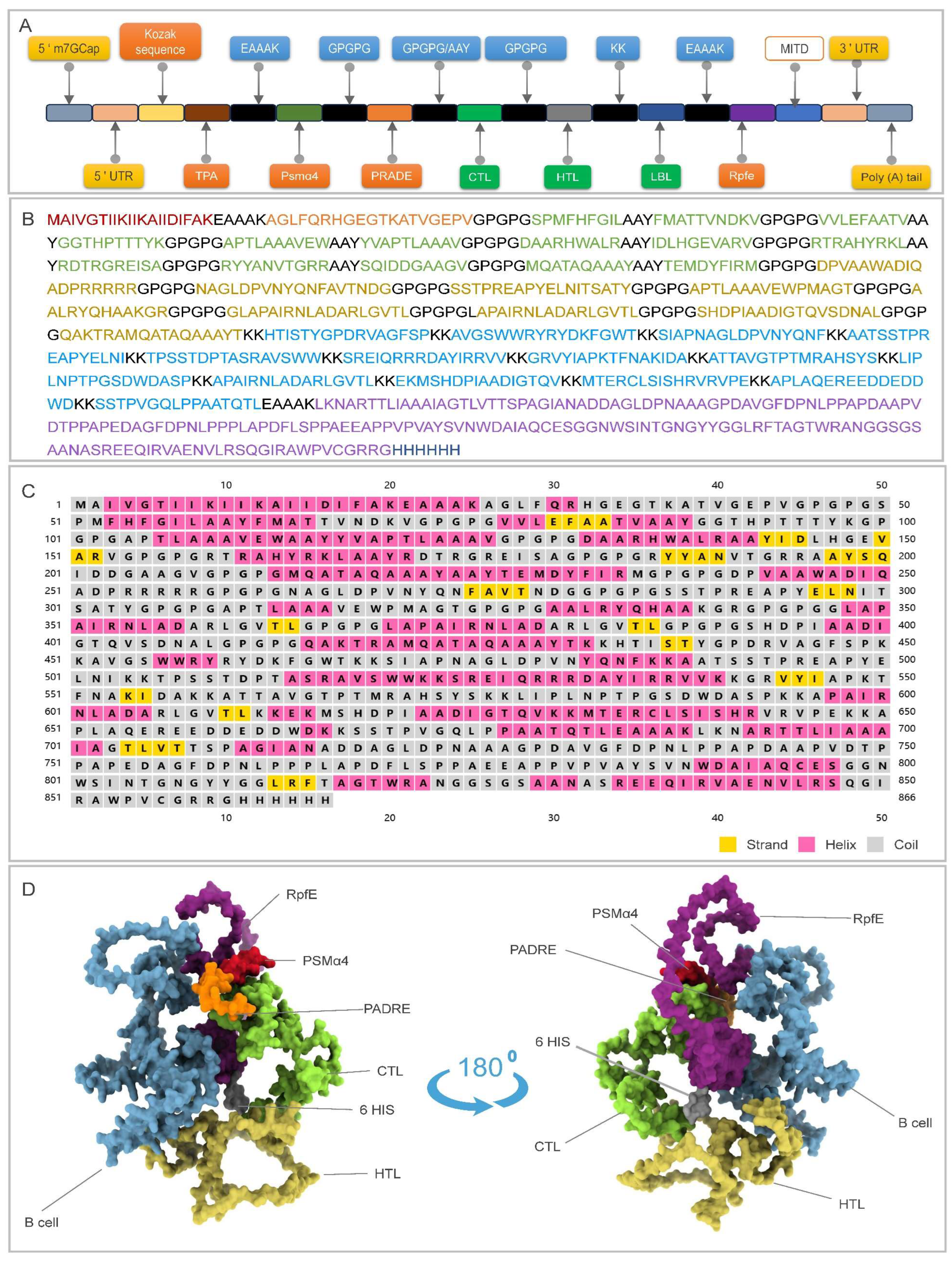
3.1. Rational Design and Assembly of the Multi-Epitope mRNA Vaccine RP14914P
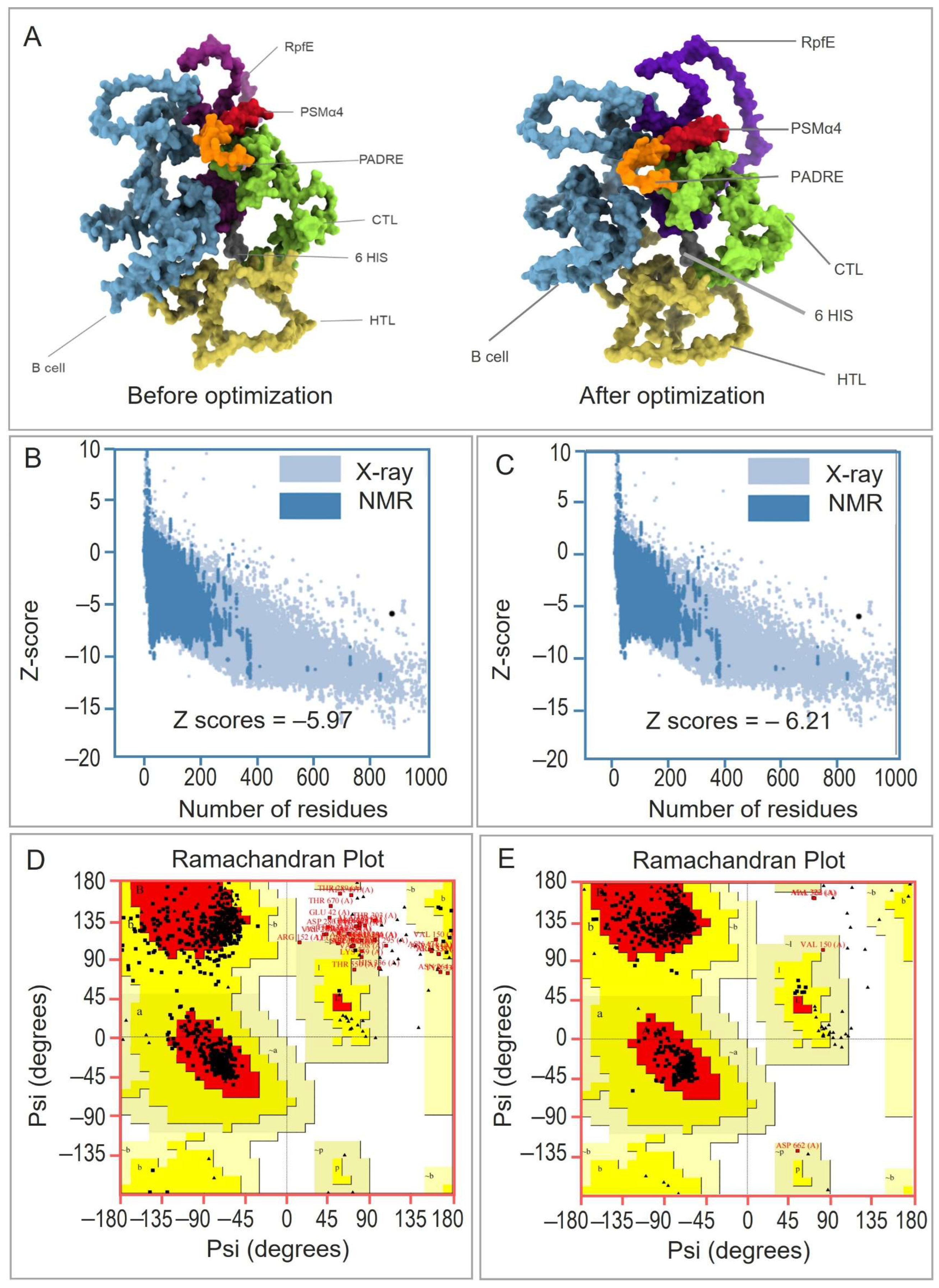
3.2. Prediction and Optimization of Secondary and Tertiary Structures
3.3. Favorable Physicochemical Profile, Safety, and Global Population Coverage of RP14914P
3.4. Conservation Analysis of Vaccine Epitopes Across Mycobacterium Strains
3.5. Prediction of Discontinuous B-Cell Epitopes
3.6. High-Affinity Docking to TLR2/4 and Robust MD Trajectories
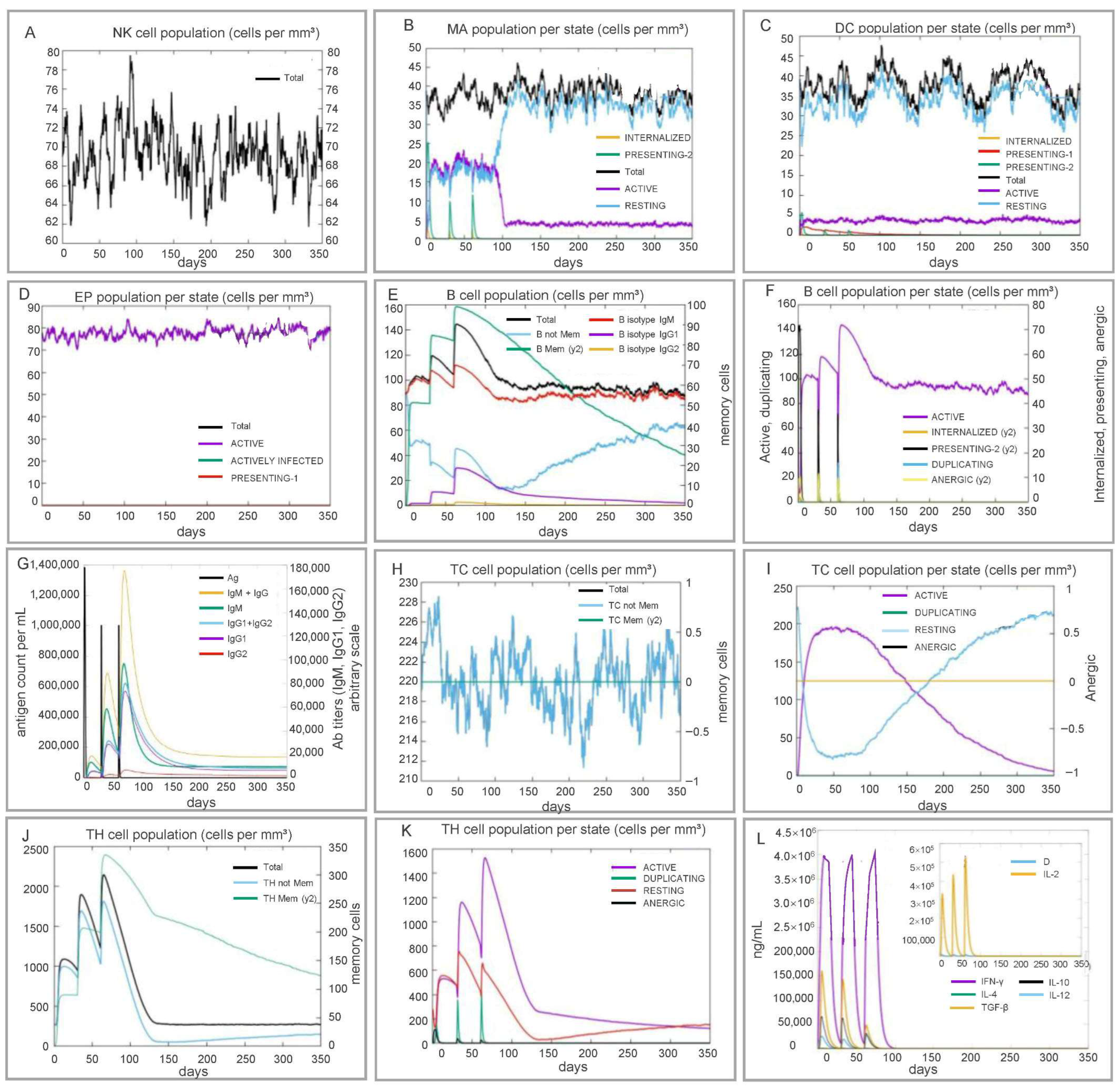
3.7. In Silico Immune Simulation Predicts Potent Multi-Layer Immunity
3.8. Codon Optimization and mRNA Secondary-Structure Stability
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ATB | Active Tuberculosis |
| BCG | Bacillus Calmette-Guérin |
| CTL | Cytotoxic T Lymphocyte |
| DCCM | Dynamic Cross-Correlation Matrix |
| HLA | Human Leukocyte Antigen |
| HTL | Helper T Lymphocyte |
| IFN-γ | Interferon-Gamma |
| IL-2 | Interleukin-2 |
| IL-4 | Interleukin-4 |
| IL-10 | Interleukin-10 |
| LNP | Lipid Nanoparticle |
| LTBI | Latent Tuberculosis Infection |
| MHC | Major Histocompatibility Complex |
| MTB | Mycobacterium Tuberculosis |
| MTBC | Mycobacterium Tuberculosis Complex |
| Rg | Radius of Gyration |
| RMSD | Root Mean Square Deviation |
| RMSF | Root Mean Square Fluctuation |
| TC | Cytotoxic T Cell |
| TH | Helper T Cell |
| TGF-β | Transforming Growth Factor-Beta |
| TLR | Toll-Like Receptor |
| WHO | World Health Organization |
| NK Cell | Natural Killer Cell |
References
- Liu, Y.; Yang, L.; Meskini, M.; Goel, A.; Opperman, M.; Shyamal, S.S.; Manaithiya, A.; Xiao, M.; Ni, R.; An, Y.; et al. Gut microbiota and tuberculosis. IMeta 2025, 4, e70054. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Wang, T.; Du, J.; Sun, L.; Wang, G.; Ni, R.; An, Y.; Fan, X.; Li, Y.; Guo, R.; et al. Decoding the WHO Global Tuberculosis Report 2024: A Critical Analysis of Global and Chinese Key Data. Zoonoses 2025, 5, 1. [Google Scholar] [CrossRef]
- An, Y.; Ni, R.; Zhuang, L.; Yang, L.; Ye, Z.; Li, L.; Parkkila, S.; Aspatwar, A.; Gong, W. Tuberculosis vaccines and therapeutic drug: Challenges and future directions. Mol. Biomed. 2025, 6, 4. [Google Scholar] [CrossRef]
- Hatzenbuehler, L.A.; Starke, J.R. Current Diagnosis and Treatment of Pediatric Latent Tuberculosis Infection. Curr. Pediatr. Rep. 2014, 2, 145–155. [Google Scholar] [CrossRef]
- Doshi, J.; Deokar, K.; Gaikwad, P. Novel tuberculosis skin tests for detecting latent tuberculosis infection. Monaldi Arch. Chest Dis. 2024, 95, 3156. [Google Scholar] [CrossRef] [PubMed]
- Chee, C.B.E.; Reves, R.; Zhang, Y.; Belknap, R. Latent tuberculosis infection: Opportunities and challenges. Respirology 2018, 23, 893–900. [Google Scholar] [CrossRef]
- Carranza, C.; Pedraza-Sanchez, S.; de Oyarzabal-Mendez, E.; Torres, M. Diagnosis for Latent Tuberculosis Infection: New Alternatives. Front. Immunol. 2020, 11, 2006. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Tuberculosis Report 2025; World Health Organization: Geneva, Switzerland, 2025; pp. 1–76. [Google Scholar]
- Gong, W.; Wu, X. Differential Diagnosis of Latent Tuberculosis Infection and Active Tuberculosis: A Key to a Successful Tuberculosis Control Strategy. Front. Microbiol. 2021, 12, 745592. [Google Scholar] [CrossRef]
- Mangtani, P.; Abubakar, I.; Ariti, C.; Beynon, R.; Pimpin, L.; Fine, P.E.; Rodrigues, L.C.; Smith, P.G.; Lipman, M.; Whiting, P.F.; et al. Protection by BCG vaccine against tuberculosis: A systematic review of randomized controlled trials. Clin. Infect. Dis. 2014, 58, 470–480. [Google Scholar] [CrossRef]
- Zhuang, L.; Ye, Z.; Li, L.; Yang, L.; Gong, W. Next-Generation TB Vaccines: Progress, Challenges, and Prospects. Vaccines 2023, 11, 1304. [Google Scholar] [CrossRef]
- Tait, D.R.; Hatherill, M.; Van Der Meeren, O.; Ginsberg, A.M.; Van Brakel, E.; Salaun, B.; Scriba, T.J.; Akite, E.J.; Ayles, H.M.; Bollaerts, A.; et al. Final Analysis of a Trial of M72/AS01(E) Vaccine to Prevent Tuberculosis. N. Engl. J. Med. 2019, 381, 2429–2439. [Google Scholar] [CrossRef]
- Luabeya, A.K.; Kagina, B.M.; Tameris, M.D.; Geldenhuys, H.; Hoff, S.T.; Shi, Z.; Kromann, I.; Hatherill, M.; Mahomed, H.; Hanekom, W.A.; et al. First-in-human trial of the post-exposure tuberculosis vaccine H56:IC31 in Mycobacterium tuberculosis infected and non-infected healthy adults. Vaccine 2015, 33, 4130–4140. [Google Scholar] [CrossRef]
- Aagaard, C.; Hoang, T.; Dietrich, J.; Cardona, P.J.; Izzo, A.; Dolganov, G.; Schoolnik, G.K.; Cassidy, J.P.; Billeskov, R.; Andersen, P. A multistage tuberculosis vaccine that confers efficient protection before and after exposure. Nat. Med. 2011, 17, 189–194. [Google Scholar] [CrossRef]
- O’Garra, A.; Redford, P.S.; McNab, F.W.; Bloom, C.I.; Wilkinson, R.J.; Berry, M.P. The immune response in tuberculosis. Annu. Rev. Immunol. 2013, 31, 475–527. [Google Scholar] [CrossRef] [PubMed]
- Rezk, N.; McClean, S. Harnessing the Potential of mRNA Vaccines Against Infectious Diseases. Microb. Biotechnol. 2025, 18, e70212. [Google Scholar] [CrossRef] [PubMed]
- Sethi, G.; Kim, Y.K.; Han, S.C.; Hwang, J.H. Designing a broad-spectrum multi-epitope subunit vaccine against leptospirosis using immunoinformatics and structural approaches. Front. Immunol. 2024, 15, 1503853. [Google Scholar] [CrossRef] [PubMed]
- Yazdani, Z.; Rafiei, A.; Irannejad, H.; Yazdani, M.; Valadan, R. Designing a novel multiepitope peptide vaccine against melanoma using immunoinformatics approach. J. Biomol. Struct. Dyn. 2022, 40, 3312–3324. [Google Scholar] [CrossRef]
- Tan, C.; Xiao, Y.; Liu, T.; Chen, S.; Zhou, J.; Zhang, S.; Hu, Y.; Wu, A.; Li, C. Development of multi-epitope mRNA vaccine against Clostridioides difficile using reverse vaccinology and immunoinformatics approaches. Synth. Syst. Biotechnol. 2024, 9, 667–683. [Google Scholar] [CrossRef]
- Ramprasadh, S.V.; Rajakumar, S.; Srinivasan, S.; Susha, D.; Sharma, S.; Chourasiya, R. Computer-Aided Multi-Epitope Based Vaccine Design Against Monkeypox Virus Surface Protein A30L: An Immunoinformatics Approach. Protein J. 2023, 42, 645–663. [Google Scholar] [CrossRef]
- Fathollahi, M.; Motamedi, H.; Hossainpour, H.; Abiri, R.; Shahlaei, M.; Moradi, S.; Dashtbin, S.; Moradi, J.; Alvandi, A. Designing a novel multi-epitopes pan-vaccine against SARS-CoV-2 and seasonal influenza: In silico and immunoinformatics approach. J. Biomol. Struct. Dyn. 2023, 42, 10761–10784. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Ali, S.L.; Tian, Y.; Abduldayeva, A.; Zhou, S.; An, Y.; Li, Y.; Ni, R.; Zhang, L.; Liu, Y.; et al. EP9158H: An Immunoinformatics-Designed mRNA Vaccine Encoding Multi-Epitope Antigens and Dual TLR Agonists for Tuberculosis Prevention. Bioengineering 2025, 12, 1378. [Google Scholar] [CrossRef]
- Shahrear, S.; Islam, A.B.M.M.K. Modeling of MT. P495, an mRNA-based vaccine against the phosphate-binding protein PstS1 of Mycobacterium tuberculosis. Mol. Divers. 2022, 27, 1613–1632. [Google Scholar] [CrossRef]
- Xu, S.; Yang, K.; Li, R.; Zhang, L. mRNA Vaccine Era-Mechanisms, Drug Platform and Clinical Prospection. Int. J. Mol. Sci. 2020, 21, 6582. [Google Scholar] [CrossRef]
- Honaker, R.W.; Stewart, A.; Schittone, S.; Izzo, A.; Klein, M.R.; Voskuil, M.I. Mycobacterium bovis BCG vaccine strains lack narK2 and narX induction and exhibit altered phenotypes during dormancy. Infect. Immun. 2008, 76, 2587–2593. [Google Scholar] [CrossRef]
- Cao, X.J.; Li, Y.P.; Wang, J.Y.; Zhou, J.; Guo, X.G. MPT64 assays for the rapid detection of Mycobacterium tuberculosis. BMC Infect. Dis. 2021, 21, 336. [Google Scholar] [CrossRef]
- Bai, X. Preparation of Four Tuberculosis Latent Proteins and the Evaluation of Their Immunological Characteristic. Doctoral Dissertation, Medical School of Chinese PLA, Beijing, China, 2014. [Google Scholar]
- Hinks, T.S.; Dosanjh, D.P.; Innes, J.A.; Pasvol, G.; Hackforth, S.; Varia, H.; Millington, K.A.; Liu, X.Q.; Bakir, M.; Soysal, A.; et al. Frequencies of region of difference 1 antigen-specific but not purified protein derivative-specific gamma interferon-secreting T cells correlate with the presence of tuberculosis disease but do not distinguish recent from remote latent infections. Infect. Immun. 2009, 77, 5486–5495. [Google Scholar] [CrossRef][Green Version]
- Mukherjee, P.; Dutta, M.; Datta, P.; Dasgupta, A.; Pradhan, R.; Pradhan, M.; Kundu, M.; Basu, J.; Chakrabarti, P. The RD1-encoded antigen Rv3872 of Mycobacterium tuberculosis as a potential candidate for serodiagnosis of tuberculosis. Clin. Microbiol. Infect. 2007, 13, 146–152. [Google Scholar] [CrossRef]
- Mahmoudi, S.; Pourakbari, B.; Mamishi, S. Interferon Gamma Release Assay in response to PE35/PPE68 proteins: A promising diagnostic method for diagnosis of latent tuberculosis. Eur. Cytokine Netw. 2017, 28, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Reynisson, B.; Alvarez, B.; Paul, S.; Peters, B.; Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: Improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 2020, 48, W449–W454. [Google Scholar] [CrossRef] [PubMed]
- Kaabinejadian, S.; Barra, C.; Alvarez, B.; Yari, H.; Hildebrand, W.H.; Nielsen, M. Accurate MHC Motif Deconvolution of Immunopeptidomics Data Reveals a Significant Contribution of DRB3, 4 and 5 to the Total DR Immunopeptidome. Front. Immunol. 2022, 13, 835454. [Google Scholar] [CrossRef] [PubMed]
- Arif, S.; Aslam, F. Immunoinformatics-driven construction of a next-generation epitope-based vaccine from conserved hypothetical proteins of M. tuberculosis for enhanced TB control. Comput. Biol. Med. 2025, 198, 111144. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Gupta, S.; Vir, P.; Raghava, G.P. Prediction of IL4 inducing peptides. Clin. Dev. Immunol. 2013, 2013, 263952. [Google Scholar] [CrossRef]
- Nagpal, G.; Usmani, S.S.; Dhanda, S.K.; Kaur, H.; Singh, S.; Sharma, M.; Raghava, G.P. Computer-aided designing of immunosuppressive peptides based on IL-10 inducing potential. Sci. Rep. 2017, 7, 42851. [Google Scholar] [CrossRef]
- Saha, S.; Raghava, G.P. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 2006, 65, 40–48. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007, 8, 4. [Google Scholar] [CrossRef]
- Dimitrov, I.; Bangov, I.; Flower, D.R.; Doytchinova, I. AllerTOP v.2—A server for in silico prediction of allergens. J. Mol. Model. 2014, 20, 2278. [Google Scholar] [CrossRef]
- Sharma, N.; Naorem, L.D.; Jain, S.; Raghava, G.P.S. ToxinPred2: An improved method for predicting toxicity of proteins. Brief. Bioinform. 2022, 23, bbac174. [Google Scholar] [CrossRef]
- Shi, H.; Zhu, Y.; Shang, K.; Tian, T.; Yin, Z.; Shi, J.; He, Y.; Ding, J.; Wang, Q.; Zhang, F. Development of innovative multi-epitope mRNA vaccine against central nervous system tuberculosis using in silico approaches. PLoS ONE 2024, 19, e0307877. [Google Scholar] [CrossRef] [PubMed]
- Reshetnikov, V.; Terenin, I.; Shepelkova, G.; Yeremeev, V.; Kolmykov, S.; Nagornykh, M.; Kolosova, E.; Sokolova, T.; Zaborova, O.; Kukushkin, I.; et al. Untranslated Region Sequences and the Efficacy of mRNA Vaccines against Tuberculosis. Int. J. Mol. Sci. 2024, 25, 888. [Google Scholar] [CrossRef]
- Saadh, M.J.; Muhammad, F.A.; Albadr, R.J.; Ballal, S.; Singh, A.; Devi, A.; Joshi, K.K.; Saidkhodjaeva, S.; Taher, W.M.; Alwan, M.; et al. Structure-based design of a multi-epitope vaccine candidate against marburg virus using immunoinformatics and dynamics simulations. J. Mol. Graph. Model. 2025, 141, 109130. [Google Scholar] [CrossRef]
- Hebditch, M.; Carballo-Amador, M.A.; Charonis, S.; Curtis, R.; Warwicker, J. Protein-Sol: A web tool for predicting protein solubility from sequence. Bioinformatics 2017, 33, 3098–3100. [Google Scholar] [CrossRef] [PubMed]
- Barman, A.; Deb, B.; Chakraborty, S. Prediction of potential epitopes for peptide vaccine formulation against teschovirus a using immunoinformatics. Int. J. Pept. Res. Ther. 2020, 26, 1137–1146. [Google Scholar] [CrossRef] [PubMed]
- Long, S.; Tian, P. Protein secondary structure prediction with context convolutional neural network. RSC Adv. 2019, 9, 38391–38396. [Google Scholar] [CrossRef]
- McGuffin, L.J.; Bryson, K.; Jones, D.T. The PSIPRED protein structure prediction server. Bioinformatics 2000, 16, 404–405. [Google Scholar] [CrossRef] [PubMed]
- Haron, F.N.; Azazi, A.; Chua, K.H.; Lim, Y.A.L.; Lee, P.C.; Chew, C.H. In silico structural modeling and quality assessment of Plasmodium knowlesi apical membrane antigen 1 using comparative protein models. Trop. Biomed. 2022, 39, 394–401. [Google Scholar] [CrossRef]
- Muccee, F.; Ghazanfar, S.; Ajmal, W.; Al-Zahrani, M. In-silico characterization of estrogen reactivating β-glucuronidase enzyme in git associated microbiota of normal human and breast cancer patients. Genes 2022, 13, 1545. [Google Scholar] [CrossRef]
- Gong, W.; Liang, Y.; Mi, J.; Jia, Z.; Xue, Y.; Wang, J.; Wang, L.; Zhou, Y.; Sun, S.; Wu, X. Peptides-Based Vaccine MP3RT Induced Protective Immunity Against Mycobacterium Tuberculosis Infection in a Humanized Mouse Model. Front. Immunol. 2021, 12, 666290. [Google Scholar] [CrossRef]
- Gong, W.; Liang, Y.; Mi, J.; Xue, Y.; Wang, J.; Wang, L.; Zhou, Y.; Sun, S.; Wu, X. A peptide-based vaccine ACP derived from antigens of Mycobacterium tuberculosis induced Th1 response but failed to enhance the protective efficacy of BCG in mice. Indian J. Tuberc. 2022, 69, 482–495. [Google Scholar] [CrossRef]
- Setlur, A.S.; K, C.; Pandey, S.; Sarkar, M.; Niranjan, V. Comprehensive Molecular Interaction Studies to Construe the Repellent/Kill Activity of Geraniol During Binding Event Against Aedes aegypti Proteins. Mol. Biotechnol. 2023, 65, 726–740. [Google Scholar] [CrossRef]
- Bui, H.H.; Sidney, J.; Dinh, K.; Southwood, S.; Newman, M.J.; Sette, A. Predicting population coverage of T-cell epitope-based diagnostics and vaccines. BMC Bioinform. 2006, 7, 153. [Google Scholar] [CrossRef]
- Ponomarenko, J.; Bui, H.H.; Li, W.; Fusseder, N.; Bourne, P.E.; Sette, A.; Peters, B. ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 2008, 9, 514. [Google Scholar] [CrossRef]
- Jones, G.; Jindal, A.; Ghani, U.; Kotelnikov, S.; Egbert, M.; Hashemi, N.; Vajda, S.; Padhorny, D.; Kozakov, D. Elucidation of protein function using computational docking and hotspot analysis by ClusPro and FTMap. Acta Crystallogr. D Struct. Biol. 2022, 78, 690–697. [Google Scholar] [CrossRef]
- Ullah, A.; Rehman, B.; Khan, S.; Almanaa, T.N.; Waheed, Y.; Hassan, M.; Naz, T.; Ul Haq, M.; Muhammad, R.; Sanami, S.; et al. An In Silico Multi-epitopes Vaccine Ensemble and Characterization Against Nosocomial Proteus penneri. Mol. Biotechnol. 2024, 66, 3498–3513. [Google Scholar] [CrossRef]
- Zhang, M.; Wei, N.; Lin, R.; Xu, Y.; Zhang, Q.; Jia, L.; Zhang, X.; Yang, X. Deferoxamine addresses metabolic dysregulation and urinary tract infections in weight-associated gestational diabetes mellitus. Eur. J. Med. Res. 2025, 30, 897. [Google Scholar] [CrossRef] [PubMed]
- Lopes, C.M.R.; de Araújo, L.P.; Mariano, C.P.; Falleiros, L.; de Azevedo Junior, W.F.; Coelho, L.F.L.; da Silveira, N.J.F. Bioisosteric and Virtual Screening Approach to Identify Natural Inhibitors of Chikunguya alphavirus nsP3. Cell Biochem. Biophys. 2025, 83, 5297–5305. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Pandit, S.; Singh, T.P.; Sharma, S.; Sharma, P. Disruption of histidine biosynthesis in Acinetobacter baumannii by Tubuloside B from Cistanche tubulosa. Comput. Biol. Med. 2025, 198, 111174. [Google Scholar] [CrossRef]
- Rapin, N.; Lund, O.; Bernaschi, M.; Castiglione, F. Computational immunology meets bioinformatics: The use of prediction tools for molecular binding in the simulation of the immune system. PLoS ONE 2010, 5, e9862. [Google Scholar] [CrossRef]
- Zaib, S.; Rana, N.; Areeba; Hussain, N.; Alrbyawi, H.; Dera, A.A.; Khan, I.; Khalid, M.; Khan, A.; Al-Harrasi, A. Designing multi-epitope monkeypox virus-specific vaccine using immunoinformatics approach. J. Infect. Public Health 2023, 16, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M.; Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981, 9, 133–148. [Google Scholar] [CrossRef]
- Salaikumaran, M.R.; Kasamuthu, P.S.; Aathmanathan, V.S.; Burra, V. An in silico approach to study the role of epitope order in the multi-epitope-based peptide (MEBP) vaccine design. Sci. Rep. 2022, 12, 12584. [Google Scholar] [CrossRef]
- Gonzalez, M.C.; Kostrzak, A.; Guetard, D.; Pniewski, T.; Sala, M. HIV-1 derived peptides fused to HBsAg affect its immunogenicity. Virus Res. 2009, 146, 107–114. [Google Scholar] [CrossRef]
- Yang, Y.; Xue, Y.; Wang, X.; Wang, L.; Wang, J.; Zhang, J.; Liu, Y.; Liang, Y.; Wu, X. Bioinformatics Analysis and Immunogenicity Assessment of the Novel Multi-Stage DNA Vaccine W541 Against Mycobacterium Tuberculosis. Immun. Inflamm. Dis. 2024, 12, e70074. [Google Scholar] [CrossRef] [PubMed]
- Tien, N.T.N.; Yen, N.T.H.; Phat, N.K.; Anh, N.K.; Thu, N.Q.; Eunsu, C.; Kim, H.S.; Hoa, V.D.; Nguyen, D.N.; Kim, D.H.; et al. Multiomics and Machine Learning Identify Immunometabolic Biomarkers for Active Tuberculosis Diagnosis Against Nontuberculous Mycobacteria and Latent Tuberculosis Infection. J. Proteome Res. 2025, 24, 3783–3797. [Google Scholar] [CrossRef]
- Bongomin, F.; Pitua, I.; Ssekamatte, P.; Sitenda, D.; Andia-Biraro, I.; Jonani, B. Agreement and systematic bias between QuantiFERON–chemiluminescent immunoassay and QuantiFERON–enzyme-linked immunosorbent assay in the detection of latent tuberculosis infection: A systematic review and meta-analysis. IJID Reg. 2025, 100824. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Wang, Y.; Zhang, Y.; Geng, S.; Wu, H.; Shao, Y.; Kang, G. Construction of Immune-Related Diagnostic Model for Latent Tuberculosis Infection and Active Tuberculosis. J. Inflamm. Res. 2024, 17, 2499–2511. [Google Scholar] [CrossRef]
- Li, F.; Dang, W.; Du, Y.; Xu, X.; He, P.; Zhou, Y.; Zhu, B. Tuberculosis Vaccines and T Cell Immune Memory. Vaccines 2024, 12, 483. [Google Scholar] [CrossRef] [PubMed]
- Moradi, M.; Vahedi, F.; Abbassioun, A.; Ramezanpour Shahi, A.; Sholeh, M.; Taheri-Anganeh, M.; Dargahi, Z.; Ghanavati, R.; Khatami, S.H.; Movahedpour, A. Liposomal delivery system/adjuvant for tuberculosis vaccine. Immun. Inflamm. Dis. 2023, 11, e867. [Google Scholar] [CrossRef]
- Schrager, L.K.; Vekemens, J.; Drager, N.; Lewinsohn, D.M.; Olesen, O.F. The status of tuberculosis vaccine development. Lancet Infect. Dis. 2020, 20, e28–e37. [Google Scholar] [CrossRef]
- Kazakova, A.; Zhelnov, P.; Sidorov, R.; Rogova, A.; Vasileva, O.; Ivanov, R.; Reshetnikov, V.; Muslimov, A. DNA and RNA vaccines against tuberculosis: A scoping review of human and animal studies. Front. Immunol. 2024, 15, 1457327. [Google Scholar] [CrossRef]
- Kozlova, A.; Pateev, I.; Shepelkova, G.; Vasileva, O.; Zakharova, N.; Yeremeev, V.; Ivanov, R.; Reshetnikov, V. A Cap-Optimized mRNA Encoding Multiepitope Antigen ESAT6 Induces Robust Cellular and Humoral Immune Responses Against Mycobacterium tuberculosis. Vaccines 2024, 12, 1267. [Google Scholar] [CrossRef] [PubMed]
- Xue, T.; Stavropoulos, E.; Yang, M.; Ragno, S.; Vordermeier, M.; Chambers, M.; Hewinson, G.; Lowrie, D.B.; Colston, M.J.; Tascon, R.E. RNA encoding the MPT83 antigen induces protective immune responses against Mycobacterium tuberculosis infection. Infect. Immun. 2004, 72, 6324–6329. [Google Scholar] [CrossRef] [PubMed]
- Lorenzi, J.C.; Trombone, A.P.; Rocha, C.D.; Almeida, L.P.; Lousada, R.L.; Malardo, T.; Fontoura, I.C.; Rossetti, R.A.; Gembre, A.F.; Silva, A.M.; et al. Intranasal vaccination with messenger RNA as a new approach in gene therapy: Use against tuberculosis. BMC Biotechnol. 2010, 10, 77. [Google Scholar] [CrossRef]
- Chugh, S.; Bahal, R.K.; Dhiman, R.; Singh, R. Antigen identification strategies and preclinical evaluation models for advancing tuberculosis vaccine development. npj Vaccines 2024, 9, 57. [Google Scholar] [CrossRef]
- Agrawal, N.; Ates, L.S.; Schille, S.; Chaturvedi, A.; Vogt, J.; Vukovic, N.; Vogel, A.B.; Diekmann, J.; Diken, M.; Şahin, U. mRNA-based tuberculosis vaccines BNT164a1 and BNT164b1 are immunogenic, well-tolerated and efficacious in rodent models. bioRxiv 2025. [Google Scholar] [CrossRef]
- Jiang, F.; Peng, C.; Cheng, P.; Wang, J.; Lian, J.; Gong, W. PP19128R, a Multiepitope Vaccine Designed to Prevent Latent Tuberculosis Infection, Induced Immune Responses In Silico and In Vitro Assays. Vaccines 2023, 11, 856. [Google Scholar] [CrossRef]
- Al Tbeishat, H. Novel In Silico mRNA vaccine design exploiting proteins of M. tuberculosis that modulates host immune responses by inducing epigenetic modifications. Sci. Rep. 2022, 12, 4645. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, L.; Ali, A.; Yang, L.; Ye, Z.; Li, L.; Ni, R.; An, Y.; Ali, S.L.; Gong, W. Leveraging computer-aided design and artificial intelligence to develop a next-generation multi-epitope tuberculosis vaccine candidate. Infect. Med. 2024, 3, 100148. [Google Scholar] [CrossRef]
- Kazakova, A.A.; Shepelkova, G.S.; Kukushkin, I.S.; Yeremeev, V.V.; Ivanov, R.A.; Reshetnikov, V.V. Multiepitope mRNA Vaccine mRNA-mEp21-FL-IDT Provides Efficient Protection against M. tuberculosis. Biochemistry 2025, 90, 754–772. [Google Scholar] [CrossRef]
- Nell, A.S.; D’Lom, E.; Bouic, P.; Sabaté, M.; Bosser, R.; Picas, J.; Amat, M.; Churchyard, G.; Cardona, P.J. Safety, tolerability, and immunogenicity of the novel antituberculous vaccine RUTI: Randomized, placebo-controlled phase II clinical trial in patients with latent tuberculosis infection. PLoS ONE 2014, 9, e89612. [Google Scholar] [CrossRef] [PubMed]
- Correia, B.E.; Bates, J.T.; Loomis, R.J.; Baneyx, G.; Carrico, C.; Jardine, J.G.; Rupert, P.; Correnti, C.; Kalyuzhniy, O.; Vittal, V.; et al. Proof of principle for epitope-focused vaccine design. Nature 2014, 507, 201–206. [Google Scholar] [CrossRef]
- Tahir Ul Qamar, M.; Saleem, S.; Ashfaq, U.A.; Bari, A.; Anwar, F.; Alqahtani, S. Epitope-based peptide vaccine design and target site depiction against Middle East Respiratory Syndrome Coronavirus: An immune-informatics study. J. Transl. Med. 2019, 17, 362. [Google Scholar] [CrossRef]
- Sela-Culang, I.; Kunik, V.; Ofran, Y. The structural basis of antibody-antigen recognition. Front. Immunol. 2013, 4, 302. [Google Scholar] [CrossRef] [PubMed]
- Cheng, P.; Jiang, F.; Wang, G.; Wang, J.; Xue, Y.; Wang, L.; Gong, W. Bioinformatics analysis and consistency verification of a novel tuberculosis vaccine candidate HP13138PB. Front. Immunol. 2023, 14, 1102578. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Kagan, J.C. A cross-disciplinary perspective on the innate immune responses to bacterial lipopolysaccharide. Mol. Cell 2014, 54, 212–223. [Google Scholar] [CrossRef]
- Nussinov, R.; Tsai, C.J.; Jang, H. Anticancer drug resistance: An update and perspective. Drug Resist. Updat. 2021, 59, 100796. [Google Scholar] [CrossRef]
- Ghisletti, S.; Barozzi, I.; Mietton, F.; Polletti, S.; De Santa, F.; Venturini, E.; Gregory, L.; Lonie, L.; Chew, A.; Wei, C.L.; et al. Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity 2010, 32, 317–328. [Google Scholar] [CrossRef]
- Blum, J.S.; Wearsch, P.A.; Cresswell, P. Pathways of antigen processing. Annu. Rev. Immunol. 2013, 31, 443–473. [Google Scholar] [CrossRef]
- Zhu, J.; Yamane, H.; Paul, W.E. Differentiation of effector CD4 T cell populations (*). Annu. Rev. Immunol. 2010, 28, 445–489. [Google Scholar] [CrossRef] [PubMed]
- Schoenborn, J.R.; Wilson, C.B. Regulation of interferon-gamma during innate and adaptive immune responses. Adv. Immunol. 2007, 96, 41–101. [Google Scholar] [CrossRef] [PubMed]
- Halle, S.; Keyser, K.A.; Stahl, F.R.; Busche, A.; Marquardt, A.; Zheng, X.; Galla, M.; Heissmeyer, V.; Heller, K.; Boelter, J.; et al. In Vivo Killing Capacity of Cytotoxic T Cells Is Limited and Involves Dynamic Interactions and T Cell Cooperativity. Immunity 2016, 44, 233–245. [Google Scholar] [CrossRef]
- Kaech, S.M.; Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 2012, 12, 749–761. [Google Scholar] [CrossRef] [PubMed]
- Badovinac, V.P.; Harty, J.T. Programming, demarcating, and manipulating CD8+ T-cell memory. Immunol. Rev. 2006, 211, 67–80. [Google Scholar] [CrossRef]
- Pape, K.A.; Taylor, J.J.; Maul, R.W.; Gearhart, P.J.; Jenkins, M.K. Different B cell populations mediate early and late memory during an endogenous immune response. Science 2011, 331, 1203–1207. [Google Scholar] [CrossRef]
- Plotkin, S.A. Correlates of protection induced by vaccination. Clin. Vaccine Immunol. 2010, 17, 1055–1065. [Google Scholar] [CrossRef]
- Larsen, S.E.; Erasmus, J.H.; Reese, V.A.; Pecor, T.; Archer, J.; Kandahar, A.; Hsu, F.C.; Nicholes, K.; Reed, S.G.; Baldwin, S.L.; et al. An RNA-Based Vaccine Platform for Use against Mycobacterium tuberculosis. Vaccines 2023, 11, 130. [Google Scholar] [CrossRef]
- Rais, M.; Abdelaal, H.; Reese, V.A.; Ferede, D.; Larsen, S.E.; Pecor, T.; Erasmus, J.H.; Archer, J.; Khandhar, A.P.; Cooper, S.K.; et al. Immunogenicity and protection against Mycobacterium avium with a heterologous RNA prime and protein boost vaccine regimen. Tuberculosis 2023, 138, 102302. [Google Scholar] [CrossRef]
- Jiang, F.; Han, Y.; Liu, Y.; Xue, Y.; Cheng, P.; Xiao, L.; Gong, W. A comprehensive approach to developing a multi-epitope vaccine against Mycobacterium tuberculosis: From in silico design to in vitro immunization evaluation. Front. Immunol. 2023, 14, 1280299. [Google Scholar] [CrossRef]
- Hu, Z.; Xia, J.; Wu, J.; Zhao, H.; Ji, P.; Gu, L.; Gu, W.; Chen, Z.; Xu, J.; Huang, X.; et al. A multistage Sendai virus vaccine incorporating latency-associated antigens induces protection against acute and latent tuberculosis. Emerg. Microbes Infect. 2024, 13, 2300463. [Google Scholar] [CrossRef] [PubMed]
- Weng, S.; Zhang, J.; Ma, H.; Zhou, J.; Jia, L.; Wan, Y.; Cui, P.; Ruan, Q.; Shao, L.; Wu, J.; et al. B21 DNA vaccine expressing ag85b, rv2029c, and rv1738 confers a robust therapeutic effect against latent Mycobacterium tuberculosis infection. Front. Immunol. 2022, 13, 1025931. [Google Scholar] [CrossRef] [PubMed]








| Antigen | Type a | Phase | Antigenicity b | Allergenicity c | Toxicity d | Function | Functional Category | Vaccine Development Status | LTBI Differential Diagnosis | References |
|---|---|---|---|---|---|---|---|---|---|---|
| Rv1736c | DosR | Latent | 0.4753 | N | N | Involved in nitrate reduction, and in the persistence in the host | Intermediary metabolism and respiration | It performs well in preclinical trials and may be a good candidate vaccine | Induced by MTB rather than 13 BCG strains or M. bovis | [25] |
| Rv1980c | Other | Latent | 0.6007 | N | N | Detoxification of reactive oxygen species, contributes to drug resistance | Cell wall and cell processes | NA | Sensitivity and specificity were 0.92 and 0.95, respectively, sensitivity of the MPT64 test was significantly higher in TB-infected children than in adults. | [26] |
| Rv2656c | NS | Latent | 0.5239 | N | N | NA | Insertion seqs and phages | NA | NA | NA |
| Rv2659c | NS | Latent | 0.5055 | N | N | Sequence integration. Integrase is necessary for integration of a phage into the host genome by site-specific recombination | Insertion seqs and phages | rBCG Aure C::hly | Higher IFN-γ-producing T cells in LTBI vs. aTB and HC | [27] |
| Rv3879c | Other | Latent | 0.6104 | N | N | NA | Cell wall and cell processes | NA | The immunodominance of Rv3879c is higher than that of Rv3878 and Rv3873 in aTB and LTBI subjects. | [28] |
| Rv3872 | Other | Latent | 0.4894 | N | N | NA | PE/PPE | NA | (1) Sensitivity and specificity of PE35 for detecting LTBI in children were 76% and80%. (2) Elicited stronger immunoreactivity and could discriminate TB from HC vaccinated with BCG (better than Rv3878). | [29,30] |
| Rv3873 | NS | Latent | 0.6381 | N | N | lmmune modulation, interacts with humanTLR2, stimulates IL-10 and MCP-1 secretion | PE/PPE | H107e/CAF®10b | Sensitivity and specificity of PE68 for detecting LTBI in children were 73% and 75%. | [30] |
| Protein | Peptide Sequence | Length | Alleles | Percentile Rank a | IC50 b | Antigenicity Score c | IFN-γ Score d | Immunogenicity Score e | ABC Pred Score f | IL4 g | IL10 h | AllerTOP V 2.0 i | Toxin Pred j |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTL epitopes | |||||||||||||
| Rv1736c | SPMFHFGIL | 9 | HLA-B*07:02/08:01 | 0.04 | 3.89 | 3.8900 | NA | 0.3089 | NA | NA | NA | Non | Non |
| FMATTVNDKV | 10 | HLA-A*02:03/02:01 | 0.28 | 4.27 | 4.2700 | NA | 0.0164 | NA | NA | NA | Non | Non | |
| VVLEFAATV | 9 | HLA-A*02:06/02:01/02:03 | 0.06 | 7.85 | 7.8500 | NA | 0.3037 | NA | NA | NA | Non | Non | |
| Rv1980c | GGTHPTTTYK | 10 | HLA-A*11:01 | 0.02 | 21.79 | 1.6270 | NA | 0.1393 | NA | NA | NA | Non | Non |
| Rv2656c | APTLAAAVEW | 10 | HLA-B*58:01 | 0.09 | 18.05 | 0.6194 | NA | 0.2078 | NA | NA | NA | Non | Non |
| YVAPTLAAAV | 10 | HLA-A*02:06 | 0.20 | 5.43 | 0.5868 | NA | 0.1213 | NA | NA | NA | Non | Non | |
| DAARHWALR | 9 | HLA-A*68:01 | 0.15 | 9.92 | 0.5705 | NA | 0.3313 | NA | NA | NA | Non | Non | |
| Rv2659 | IDLHGEVARV | 10 | HLA-A*02:03 | 0.24 | 20.23 | 1.4721 | NA | 0.2616 | NA | NA | NA | Non | Non |
| RTRAHYRKL | 9 | HLA-A*30:01 | 0.04 | 6.31 | 1.0337 | NA | 0.0019 | NA | NA | NA | Non | Non | |
| Rv3872 | SQIDDGAAGV | 10 | HLA-A*02:06 | 0.21 | 21.71 | 1.0865 | NA | 0.2098 | NA | NA | NA | Non | Non |
| Rv3873 | MQATAQAAAY | 10 | HLA-B*15:01/35:01 HLA-A*30:02 | 0.04 | 4.88 | 0.5750 | NA | 0.0698 | NA | NA | NA | Non | Non |
| TEMDYFIRM | 9 | HLA-B*44:03 | 0.04 | 32.71 | 0.5697 | NA | 0.2145 | NA | NA | NA | Non | Non | |
| Rv3879 | RDTRGREISA | 10 | HLA-A*30:01 | 0.36 | 28.91 | 2.5449 | NA | 0.2580 | NA | NA | NA | Non | Non |
| RYYANVTGRR | 10 | HLA-A*31:01 | 0.13 | 15.61 | 1.1056 | NA | 0.1675 | NA | NA | NA | Non | Non | |
| HTL epitopes | |||||||||||||
| Rv1736c | DPVAAWADIQADPRRRRR | 18 | HLA-DRB1*03:01/03:02/05:01 HLA-DQA1*01:01/03:01 | 0.47 | NA | 0.5475 | 1.6921 | NA | NA | Non | Non | Non | Non |
| Rv1980c | NAGLDPVNYQNFAVTNDG | 18 | HLA-DPA1*03:01/DPB1*04:02 HLA-DPA1*01:03/DPB1*04:0 HLA-DPA1*02:01/DPB1*01:01 HLA-DPA1*01:03/DPB1*02:01 HLA-DRB1*04:05/15:01 HLA-DPA1*02:01/DPB1*05:01 HLA-DQA1*01:01/DQB1*05:01 | 0.57 | NA | 0.5137 | 0.4837 | NA | NA | Non | Non | Non | Non |
| SSTPREAPYELNITSATY | 18 | HLA-DRB3*02:02 | 0.44 | NA | 0.7701 | 0.5404 | NA | NA | Non | Non | Non | Non | |
| Rv2656c | APTLAAAVEWPMAGT | 15 | HLA-DRB1*09:01 | 0.45 | NA | 0.5853 | 0.0141 | NA | NA | Non | Non | Non | Non |
| Rv2659c | AALRYQHAAKGR | 12 | HLA-DRB4*01:01 | 0.06 | NA | 1.0668 | 0.4385 | NA | NA | Non | Non | Non | Non |
| Rv3872 | SHDPIAADIGTQVSDNAL | 18 | HLA-DRB3*01:01 | 0.21 | NA | 0.5918 | 0.0183 | NA | NA | Non | Non | Non | Non |
| Rv3873 | QAKTRAMQATAQAAAYT | 17 | HLA-DQA1*01:02, HLA-DQB1*06:02 | 0.43 | NA | 0.7682 | 0.4548 | NA | NA | Non | Non | Non | Non |
| Rv3879c | GLAPAIRNLADARLGVTL | 18 | HLA-DQA1*01:01, HLA-DQB1*05:01 | 0.47 | NA | 0.6025 | 0.4770 | NA | NA | Non | Non | Non | Non |
| LAPAIRNLADARLGVTL | 17 | HLA-DQA1*01:01, HLA-DQB1*05:01 | 0.48 | NA | 0.5405 | 0.1502 | NA | NA | Non | Non | Non | Non | |
| B cellular epitopes | |||||||||||||
| Rv1736c | HTISTYGPDRVAGFSP | 16 | NA | NA | NA | NA | NA | NA | 0.94 | NA | NA | Non | Non |
| AVGSWWRYRYDKFGWT | 16 | NA | NA | NA | NA | NA | NA | 0.93 | NA | NA | Non | Non | |
| Rv1980c | SIAPNAGLDPVNYQNF | 16 | NA | NA | NA | NA | NA | NA | 0.92 | NA | NA | Non | Non |
| AATSSTPREAPYELNI | 16 | NA | NA | NA | NA | NA | NA | 0.92 | NA | NA | Non | Non | |
| Rv2656c | TPSSTDPTASRAVSWW | 16 | NA | NA | NA | NA | NA | NA | 0.93 | NA | NA | Non | Non |
| SREIQRRRDAYIRRVV | 16 | NA | NA | NA | NA | NA | NA | 0.81 | NA | NA | Non | Non | |
| Rv2659c | GRVYIAPKTFNAKIDA | 16 | NA | NA | NA | NA | NA | NA | 0.97 | NA | NA | Non | Non |
| ATTAVGTPTMRAHSYS | 16 | NA | NA | NA | NA | NA | NA | 0.9 | NA | NA | Non | Non | |
| Rv3872 | EKMSHDPIAADIGTQV | 16 | NA | NA | NA | NA | NA | NA | 0.87 | NA | NA | Non | Non |
| MTERCLSISHRVRVPE | 16 | NA | NA | NA | NA | NA | NA | 0.84 | NA | NA | Non | Non | |
| Rv3873 | APLAQEREEDDEDDWD | 16 | NA | NA | NA | NA | NA | NA | 0.93 | NA | NA | Non | Non |
| SSTPVGQLPPAATQTL | 16 | NA | NA | NA | NA | NA | NA | 0.85 | NA | NA | Non | Non | |
| Rv3879c | LIPLNPTPGSDWDASP | 16 | NA | NA | NA | NA | NA | NA | 0.86 | NA | NA | Non | Non |
| APAIRNLADARLGVTL | 16 | NA | NA | NA | NA | NA | NA | 0.84 | NA | NA | Non | Non | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Tian, Y.; Zhang, M.; Ali, S.L.; Abduldayeva, A.; Zhou, S.; An, Y.; Li, Y.; Ni, R.; Zhang, L.; Liu, Y.; et al. Immunoinformatic Design and Evaluation of a Multi-Epitope mRNA Vaccine RP14914P Targeting Latent Tuberculosis Infection. Pathogens 2026, 15, 297. https://doi.org/10.3390/pathogens15030297
Tian Y, Zhang M, Ali SL, Abduldayeva A, Zhou S, An Y, Li Y, Ni R, Zhang L, Liu Y, et al. Immunoinformatic Design and Evaluation of a Multi-Epitope mRNA Vaccine RP14914P Targeting Latent Tuberculosis Infection. Pathogens. 2026; 15(3):297. https://doi.org/10.3390/pathogens15030297
Chicago/Turabian StyleTian, Yuan, Mingming Zhang, Syed Luqman Ali, Aigul Abduldayeva, Shuang Zhou, Yajing An, Yufeng Li, Ruizi Ni, Lingxia Zhang, Yanhua Liu, and et al. 2026. "Immunoinformatic Design and Evaluation of a Multi-Epitope mRNA Vaccine RP14914P Targeting Latent Tuberculosis Infection" Pathogens 15, no. 3: 297. https://doi.org/10.3390/pathogens15030297
APA StyleTian, Y., Zhang, M., Ali, S. L., Abduldayeva, A., Zhou, S., An, Y., Li, Y., Ni, R., Zhang, L., Liu, Y., Sun, W., & Gong, W. (2026). Immunoinformatic Design and Evaluation of a Multi-Epitope mRNA Vaccine RP14914P Targeting Latent Tuberculosis Infection. Pathogens, 15(3), 297. https://doi.org/10.3390/pathogens15030297







