Glnk Mediates Carbapenem Resistance Through the NtrB/NtrC-OprD Regulatory Pathway in Pseudomonas aeruginosa
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasmids, Primers, and Bacterial Strains
2.2. Construction of the glnK Mutant
2.3. Minimum Inhibitory Concentration (MIC) Assay
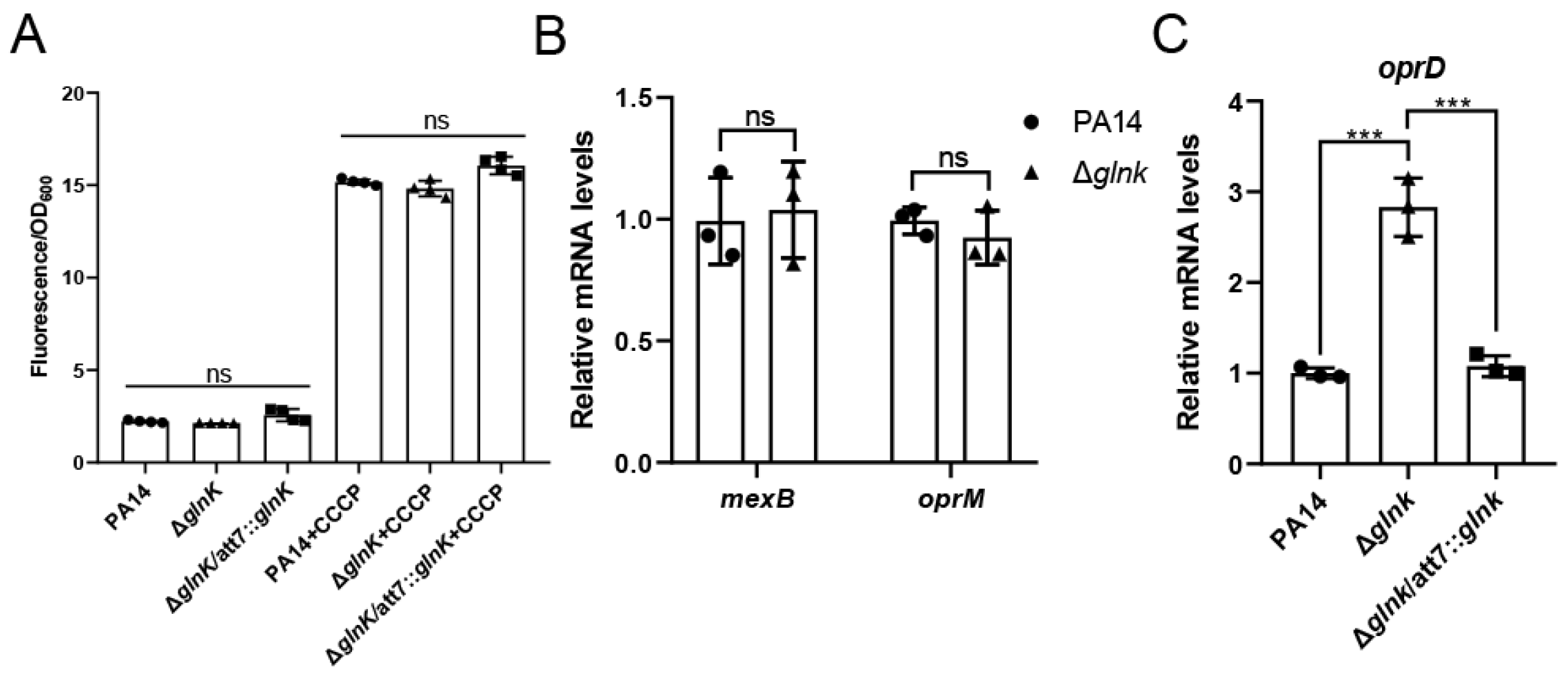
2.4. Ethidium Bromide Influx Assay
2.5. RNA Extraction and Real-Time Quantitative PCR
2.6. β-Galactosidase Activity Assay
2.7. Electrophoretic Mobility Shift Assay (EMSA)
2.8. Statistical Analysis
3. Results
3.1. Susceptibility of the glnK Mutant to Carbapenem Antibiotics
3.2. Upregulation of OprD in the glnK Mutant Contributes to the Increased Carbapenem Susceptibility
3.3. GlnK Regulates oprD Through the NtrB/NtrC Two-Component System
3.4. NtrC Directly Regulates oprD
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| MIC | minimum inhibitory concentration |
| MEM | meropenem |
| IMP | imipenem |
| DRPM | doripenem |
| FEP | cefepime |
| CAZ | ceftazidime |
| ATM | aztreonam |
| CAR | carbenicillin |
| FOX | cefoxitin |
| CCCP | carbonyl cyanide m-chlorophenylhydrazone |
References
- Carmeli, Y.; Cisneros, J.M.; Paul, M.; Daikos, G.L.; Wang, M.; Torre-Cisneros, J.; Singer, G.; Titov, I.; Gumenchuk, I.; Zhao, Y. Aztreonam-avibactam versus meropenem for the treatment of serious infections caused by Gram-negative bacteria (REVISIT): A descriptive, multinational, open-label, phase 3, randomised trial. Lancet Infect. Dis. 2025, 25, 218–230. [Google Scholar] [CrossRef]
- Jones, K.E.; Patel, N.G.; Levy, M.A.; Storeygard, A.; Balk, D.; Gittleman, J.L.; Daszak, P. Global trends in emerging infectious diseases. Nature 2008, 451, 990–993. [Google Scholar] [CrossRef]
- GBD 2021 Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance 1990–2021: A systematic analysis with forecasts to 2050. Lancet 2024, 404, 1199–1226. [Google Scholar] [CrossRef]
- Hotor, P.; Kotey, F.C.N.; Donkor, E.S. Antibiotic resistance in hospital wastewater in West Africa: A systematic review and meta-analysis. BMC Public Health 2025, 25, 1364. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Qi, G.; Feng, Y.; Du, C. Application of nematicide avermectin enriched antibiotic-resistant bacteria and antibiotic resistance genes in farmland soil. Environ. Res. 2023, 227, 115802. [Google Scholar] [CrossRef]
- Macesic, N.; Uhlemann, A.C.; Peleg, A.Y. Multidrug-resistant Gram-negative bacterial infections. Lancet 2025, 405, 257–272. [Google Scholar] [CrossRef]
- Letizia, M.A.; Diggle, S.A.; Whiteley, M.A. Pseudomonas aeruginosa: Ecology, evolution, pathogenesis and antimicrobial susceptibility. Nat. Rev. Microbiol. 2025, 23, 701–717. [Google Scholar] [CrossRef] [PubMed]
- Stover, C.K.; Pham, X.Q.; Erwin, A.L.; Mizoguchi, S.D.; Warrener, P.; Hickey, M.J.; Brinkman, F.S.; Hufnagle, W.O.; Kowalik, D.J.; Lagrou, M.; et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 2000, 406, 959–964. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Feng, J.; Wu, Y.; Ding, W.; Xie, L.; Han, L.; Wang, C. Whole Genome Features and Analysis of Antibiotic Resistance Determinants in Pseudomonas aeruginosa Strain CYZ. Microb. Drug Resist. 2025, 31, 403–412. [Google Scholar] [CrossRef]
- Dang, J.X.; Shu, J.; Wang, R.; Yu, H.; Chen, Z.; Yan, W.; Zhao, B.; Ding, L.; Wang, Y.; Hu, H. The glycopatterns of Pseudomonas aeruginosa as a potential biomarker for its carbapenem resistance. Microbiol. Spectr. 2023, 11, e0200123. [Google Scholar] [CrossRef] [PubMed]
- Pont, S.; Nilly, F.; Berry, L.; Bonhoure, A.; Alford, M.A.; Louis, M.; Nogaret, P.; Bains, M.; Lesouhaitier, O.; Hancock, R.E.W. Intracellular Pseudomonas aeruginosa persist and evade antibiotic treatment in a wound infection model. PLoS Pathog. 2025, 21, e1012922. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Liu, M.; Mo, C.; Tan, R.; Li, S.; Liang, H.; Li, M. Molecular characteristics and antibiotic resistance mechanisms of multidrug-resistant Pseudomonas aeruginosa in Nanning, China. BMC Microbiol. 2024, 24, 478. [Google Scholar] [CrossRef]
- Hernández-García, M.; Barbero-Herranz, R.; Bastón-Paz, N.; Díez-Aguilar, M.; López-Collazo, E.; Márquez-Garrido, F.J.; Hernández-Pérez, J.M.; Baquero, F.; Ekkelenkamp, M.B.; Fluit, A.C. Unravelling the mechanisms causing murepavadin resistance in Pseudomonas aeruginosa: Lipopolysaccharide alterations and its consequences. Front. Cell Infect. Microbiol. 2024, 14, 1446626. [Google Scholar] [CrossRef] [PubMed]
- Mei, L.; Song, Y.; Liu, X.; Li, K.; Guo, X.; Liu, L.; Liu, Y.; Kozlakidis, Z.A.; Cheong, I.H.; Wang, D. Characterization and Implications of IncP-2A Plasmid pMAS152 Harboring Multidrug Resistance Genes in Extensively Drug-Resistant Pseudomonas aeruginosa. Microorganisms 2024, 12, 562. [Google Scholar] [CrossRef] [PubMed]
- Wüllner, D.A.; Gesper, M.; Haupt, A.; Liang, X.X.; Zhou, P.A.; Dietze, P.; Narberhaus, F.A.; Bandow, J.A. Adaptive Responses of Pseudomonas aeruginosa to Treatment with Antibiotics. Antimicrob. Agents Chemother. 2022, 66, e0087821. [Google Scholar] [CrossRef]
- Glen, K.A.; Lamont, I.A. Penicillin-binding protein 3 sequence variations reduce susceptibility of Pseudomonas aeruginosa to β-lactams but inhibit cell division. J. Antimicrob. Chemother. 2024, 79, 2170–2178. [Google Scholar] [CrossRef]
- Monti, G.; Bradic, N.; Marzaroli, M.; Konkayev, A.; Fominskiy, E.; Kotani, Y.; Likhvantsev, V.V.; Momesso, E.; Nogtev, P.; Lobreglio, R. Continuous vs Intermittent Meropenem Administration in Critically Ill Patients with Sepsis: The MERCY Randomized Clinical Trial. JAMA 2023, 330, 141–151. [Google Scholar] [CrossRef]
- Reyes, J.; Komarow, L.; Chen, L.; Ge, L.; Hanson, B.M.; Cober, E.; Herc, E.; Alenazi, T.; Kaye, K.S.; Garcia-Diaz, J. Global epidemiology and clinical outcomes of carbapenem-resistant Pseudomonas aeruginosa and associated carbapenemases (POP): A prospective cohort study. Lancet Microbe 2023, 4, e159–e170. [Google Scholar] [CrossRef]
- Studemeister, A.E.; Quinn, J.P. Selective imipenem resistance in Pseudomonas aeruginosa associated with diminished outer membrane permeability. Antimicrob. Agents Chemother. 1988, 32, 1267–1268. [Google Scholar] [CrossRef]
- Yano, H.A.; Nakata, N.A.; Yahara, K.A.; Sugai, M.A.; Sato, Y.A. Specific variants in ftsI reduce carbapenem susceptibility in Pseudomonas aeruginosa. Microbiol. Spectr. 2025, 13, e0102725. [Google Scholar] [CrossRef]
- Yousefi, S.; Nazari, M.; Ramazanzadeh, R.; Sahebkar, A.; Safarzadeh, E.; Khademi, F.A. Association of carbapenem and multidrug resistance with the expression of efflux pump-encoding genes in Pseudomonas aeruginosa clinical isolates. Acta Microbiol. Immunol. Hung. 2023, 70, 161–166. [Google Scholar] [CrossRef]
- Kim, C.H.; Kang, H.Y.; Kim, B.R.; Jeon, H.; Lee, Y.C.; Lee, S.H.; Lee, J.C. Mutational inactivation of OprD in carbapenem-resistant Pseudomonas aeruginosa isolates from Korean hospitals. J. Microbiol. 2016, 54, 44–49. [Google Scholar] [CrossRef]
- Yano, H.A.; Hayashi, W.A.; Kawakami, S.; Aoki, S.; Anzai, E.; Zuo, H.; Kitamura, N.A.; Hirabayashi, A.A.; Kajihara, T.A.; Kayama, S.A. Nationwide genome surveillance of carbapenem-resistant Pseudomonas aeruginosa in Japan. Antimicrob. Agents Chemother. 2024, 68, e0166923. [Google Scholar] [CrossRef]
- Xie, X.; Liu, Z.; Huang, J.; Wang, X.; Tian, Y.; Xu, P.; Zheng, G. Molecular epidemiology and carbapenem resistance mechanisms of Pseudomonas aeruginosa isolated from a hospital in Fujian, China. Front. Microbiol. 2024, 15, 1431154. [Google Scholar] [CrossRef]
- Gerhardt, E.C.M.; Parize, E.; Gravina, F.; Pontes, F.L.D.; Santos, A.R.S.; Araujo, G.A.T.; Goedert, A.C.; Urbanski, A.H.; Steffens, M.B.R.; Chubatsu, L.S. The Protein-Protein Interaction Network Reveals a Novel Role of the Signal Transduction Protein PII in the Control of c-di-GMP Homeostasis in Azospirillum brasilense. mSystems 2020, 5, e00817-20. [Google Scholar] [CrossRef]
- Conroy, M.J.; Durand, A.; Lupo, D.; Li, X.D.; Bullough, P.A.; Winkler, F.K.; Merrick, M. The crystal structure of the Escherichia coli AmtB-GlnK complex reveals how GlnK regulates the ammonia channel. Proc. Natl. Acad. Sci. USA 2007, 104, 1213–1218. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, J.; Behrends, V.; Pan, Z.; Brown, D.R.; Heydenreich, F.; Lewis, M.R.; Bennett, M.H.; Razzaghi, B.; Komorowski, M.; Barahona, M. Nitrogen and carbon status are integrated at the transcriptional level by the nitrogen regulator NtrC in vivo. mBio 2013, 4, e00881-13. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, M.R.; Kamberov, E.S.; Weiss, R.L.; Ninfa, A.J. Reversible uridylylation of the Escherichia coli PII signal transduction protein regulates its ability to stimulate the dephosphorylation of the transcription factor nitrogen regulator I (NRI or NtrC). J. Biol. Chem. 1994, 269, 28288–28293. [Google Scholar] [CrossRef] [PubMed]
- Naren, N.; Zhang, X.X. Role of a local transcription factor in governing cellular carbon/nitrogen homeostasis in Pseudomonas fluorescens. Nucleic Acids Res. 2021, 49, 3204–3216. [Google Scholar] [CrossRef]
- Nguyen, D.; Joshi-Datar, A.; Lepine, F.; Bauerle, E.; Olakanmi, O.; Beer, K.; McKay, G.; Siehnel, R.; Schafhauser, J.; Wang, Y. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 2011, 334, 982–986. [Google Scholar] [CrossRef]
- Meylan, S.; Porter, C.B.M.; Yang, J.H.; Belenky, P.; Gutierrez, A.; Lobritz, M.A.; Park, J.; Kim, S.H.; Moskowitz, S.M.; Collins, J.J. Carbon Sources Tune Antibiotic Susceptibility in Pseudomonas aeruginosa via Tricarboxylic Acid Cycle Control. Cell Chem. Biol. 2017, 24, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Kok, M.; Zwep, L.B.; Jansen, R.S.; Hankemeier, T.; van Hasselt, J.G.C. Nutrients drive the antibiotic-specific evolution of resistance in Pseudomonas aeruginosa. Res. Microbiol. 2025. Epub ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Littlejohn, P.A.; Metcalfe-Roach, A.A.; Cardenas Poire, E.X.; Holani, R.A.; Bar-Yoseph, H.; Fan, Y.X.; Woodward, S.A.; Finlay, B.A. Multiple micronutrient deficiencies in early life cause multi-kingdom alterations in the gut microbiome and intrinsic antibiotic resistance genes in mice. Nat. Microbiol. 2023, 8, 2392–2405. [Google Scholar] [CrossRef]
- Liberati, N.T.; Urbach, J.M.; Miyata, S.; Lee, D.G.; Drenkard, E.; Wu, G.; Villanueva, J.; Wei, T.; Ausubel, F.M. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl. Acad. Sci. USA 2006, 103, 2833–2838. [Google Scholar] [CrossRef]
- Sun, X.; Du, Q.; Li, Y.; Gong, X.; Zhang, Y.; Jin, Y.; Jin, S.; Wu, W. GlnK Regulates the Type III Secretion System by Modulating NtrB-NtrC Homeostasis in Pseudomonas aeruginosa. Microorganisms 2026, 14, 339. [Google Scholar] [CrossRef]
- Ren, H.; Liu, Y.; Zhou, J.; Long, Y.; Liu, C.; Xia, B.; Shi, J.; Fan, Z.; Liang, Y.; Chen, S. Combination of Azithromycin and Gentamicin for Efficient Treatment of Pseudomonas aeruginosa Infections. J. Infect. Dis. 2019, 220, 1667–1678. [Google Scholar] [CrossRef]
- Choi, K.H.; Schweizer, H.P. mini-Tn7 insertion in bacteria with single attTn7 sites: Example Pseudomonas aeruginosa. Nat. Protoc. 2006, 1, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Morita, Y.; Sobel, M.L.; Poole, K. Antibiotic inducibility of the MexXY multidrug efflux system of Pseudomonas aeruginosa: Involvement of the antibiotic-inducible PA5471 gene product. J. Bacteriol. 2006, 188, 1847–1855. [Google Scholar] [CrossRef]
- Viveiros, M.; Rodrigues, L.; Martins, M.; Couto, I.; Spengler, G.; Martins, A.; Amaral, L. Evaluation of efflux activity of bacteria by a semi-automated fluorometric system. Methods Mol. Biol. 2010, 642, 159–172. [Google Scholar] [CrossRef]
- Yang, S.K.; Yusoff, K.; Ajat, M.; Yap, W.S.; Lim, S.E.; Lai, K.S. Antimicrobial activity and mode of action of terpene linalyl anthranilate against carbapenemase-producing Klebsiella pneumoniae. J. Pharm. Anal. 2021, 11, 210–219. [Google Scholar] [CrossRef]
- Dumas, J.L.; van Delden, C.; Perron, K.; Köhler, T. Analysis of antibiotic resistance gene expression in Pseudomonas aeruginosa by quantitative real-time-PCR. FEMS Microbiol. Lett. 2006, 254, 217–225. [Google Scholar] [CrossRef]
- Li, S.; Weng, Y.; Li, X.; Yue, Z.; Chai, Z.; Zhang, X.; Gong, X.; Pan, X.A.; Jin, Y.; Bai, F. Acetylation of the CspA family protein CspC controls the type III secretion system through translational regulation of exsA in Pseudomonas aeruginosa. Nucleic Acids Res. 2021, 49, 6756–6770. [Google Scholar] [CrossRef]
- Qu, J.; Yin, L.; Qin, S.; Sun, X.; Gong, X.; Li, S.; Pan, X.; Jin, Y.; Cheng, Z.; Jin, S. Identification of the Pseudomonas aeruginosa AgtR-CspC-RsaL pathway that controls Las quorum sensing in response to metabolic perturbation and Staphylococcus aureus. PLoS Pathog. 2025, 21, e1013054. [Google Scholar] [CrossRef]
- Pan, X.; Fan, Z.; Chen, L.; Liu, C.; Bai, F.; Wei, Y.; Tian, Z.; Dong, Y.; Shi, J.; Chen, H. PvrA is a novel regulator that contributes to Pseudomonas aeruginosa pathogenesis by controlling bacterial utilization of long chain fatty acids. Nucleic Acids Res. 2020, 48, 5967–5985. [Google Scholar] [CrossRef] [PubMed]
- Blair, J.M.; Piddock, L.J. How to Measure Export via Bacterial Multidrug Resistance Efflux Pumps. mBio 2016, 7, e00840-16. [Google Scholar] [CrossRef]
- Piddock, L.J.; Johnson, M.M. Accumulation of 10 fluoroquinolones by wild-type or efflux mutant Streptococcus pneumoniae. Antimicrob. Agents Chemother. 2002, 46, 813–820. [Google Scholar] [CrossRef]
- Morita, Y.; Komori, Y.; Mima, T.; Kuroda, T.; Mizushima, T.; Tsuchiya, T. Construction of a series of mutants lacking all of the four major mex operons for multidrug efflux pumps or possessing each one of the operons from Pseudomonas aeruginosa PAO1: MexCD-OprJ is an inducible pump. FEMS Microbiol. Lett. 2001, 202, 139–143. [Google Scholar] [CrossRef]
- Masuda, N.; Sakagawa, E.; Ohya, S.; Gotoh, N.; Tsujimoto, H.; Nishino, T. Substrate specificities of MexAB-OprM, MexCD-OprJ, and MexXY-oprM efflux pumps in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2000, 44, 3322–3327. [Google Scholar] [CrossRef] [PubMed]
- Richardot, C.; Plésiat, P.; Fournier, D.; Monlezun, L.; Broutin, I.; Llanes, C. Carbapenem resistance in cystic fibrosis strains of Pseudomonas aeruginosa as a result of amino acid substitutions in porin OprD. Int. J. Antimicrob. Agents 2015, 45, 529–532. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Hancock, R.E. Genetic definition of the substrate selectivity of outer membrane porin protein OprD of Pseudomonas aeruginosa. J. Bacteriol. 1993, 175, 7793–7800. [Google Scholar] [CrossRef]
- Ochs, M.M.; Lu, C.D.; Hancock, R.E.; Abdelal, A.T. Amino acid-mediated induction of the basic amino acid-specific outer membrane porin OprD from Pseudomonas aeruginosa. J. Bacteriol. 1999, 181, 5426–5432. [Google Scholar] [CrossRef]
- van Heeswijk, W.C.; Hoving, S.; Molenaar, D.; Stegeman, B.; Kahn, D.; Westerhoff, H.V. An alternative PII protein in the regulation of glutamine synthetase in Escherichia coli. Mol. Microbiol. 1996, 21, 133–146. [Google Scholar] [CrossRef]
- Atkinson, M.R.; Ninfa, A.J. Characterization of the GlnK protein of Escherichia coli. Mol. Microbiol. 1999, 32, 301–313. [Google Scholar] [CrossRef] [PubMed]
- Maier, S.; Schleberger, P.; Lü, W.; Wacker, T.; Pflüger, T.; Litz, C.; Andrade, S.L.A. Mechanism of disruption of the Amt-GlnK complex by P(II)-mediated sensing of 2-oxoglutarate. PLoS ONE 2011, 6, e26327. [Google Scholar] [CrossRef]
- Aparecida Gonçalves, A.C.; de Mello Damasco Nunes, T.; Parize, E.; Marques Gerhardt, E.C.; Antônio de Souza, G.; Scholl, J.; Forchhammer, K.; Huergo, L.F. The activity of the ribonucleotide monophosphatase UmpH is controlled by interaction with the GlnK signaling protein in Escherichia coli. J. Biol. Chem. 2024, 300, 107931. [Google Scholar] [CrossRef]
- Alford, M.A.; Baghela, A.; Yeung, A.T.Y.; Pletzer, D.; Hancock, R.E.W. NtrBC Regulates Invasiveness and Virulence of Pseudomonas aeruginosa During High-Density Infection. Front. Microbiol. 2020, 11, 773. [Google Scholar] [CrossRef] [PubMed]
- Turner, K.H.; Everett, J.; Trivedi, U.; Rumbaugh, K.P.; Whiteley, M. Requirements for Pseudomonas aeruginosa acute burn and chronic surgical wound infection. PLoS Genet. 2014, 10, e1004518. [Google Scholar] [CrossRef]
- Mima, T.; Kohira, N.; Li, Y.; Sekiya, H.; Ogawa, W.; Kuroda, T.; Tsuchiya, T. Gene cloning and characteristics of the RND-type multidrug efflux pump MuxABC-OpmB possessing two RND components in Pseudomonas aeruginosa. Microbiology 2009, 155, 3509–3517. [Google Scholar] [CrossRef] [PubMed]
- Alford, M.A.; Baquir, B.; An, A.; Choi, K.G.; Hancock, R.E.W. NtrBC Selectively Regulates Host-Pathogen Interactions, Virulence, and Ciprofloxacin Susceptibility of Pseudomonas aeruginosa. Front. Cell Infect. Microbiol. 2021, 11, 694789. [Google Scholar] [CrossRef]
- Rodríguez-Martínez, J.M.; Poirel, L.; Nordmann, P. Molecular epidemiology and mechanisms of carbapenem resistance in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2009, 53, 4783–4788. [Google Scholar] [CrossRef]
- Wang, M.; Zhang, Y.; Pei, F.; Liu, Y.; Zheng, Y. Loss of OprD function is sufficient for carbapenem-resistance-only but insufficient for multidrug resistance in Pseudomonas aeruginosa. BMC Microbiol. 2025, 25, 218. [Google Scholar] [CrossRef]
- amochi, T.; Ugajin, K.; On, R.; Inoue, S.; Ikeda, H.; Yamochi, T.; Takimoto, M.; Tokimatsu, I. Impact of meropenem exposure on fluoroquinolone and carbapenem resistance in Pseudomonas aeruginosa infection in inpatients in a Japanese university hospital: Insights into oprD mutations and efflux pump overexpression. J. Glob. Antimicrob. Resist. 2025, 41, 163–168. [Google Scholar] [CrossRef]
- Itani, R.; Khojah, H.M.J.; Mukattash, T.L.; Shuhaiber, P.; Raychouni, H.; Dib, C.; Hassan, M.; El-Lakany, A. Difficult-to-treat resistant Pseudomonas aeruginosa infections in Lebanese hospitals: Impact on mortality and the role of initial antibiotic therapy. PLoS ONE 2025, 20, e0321935. [Google Scholar] [CrossRef]
- Judd, W.R.; Ratliff, P.D.; Hickson, R.P.; Stephens, D.M.; Kennedy, C.A. Clinical and economic impact of meropenem resistance in Pseudomonas aeruginosa-infected patients. Am. J. Infect. Control 2016, 44, 1275–1279. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.R.; Ray, A.; Hodson, M.E.; Pitt, T.L. Increased sputum amino acid concentrations and auxotrophy of Pseudomonas aeruginosa in severe cystic fibrosis lung disease. Thorax 2000, 55, 795–797. [Google Scholar] [CrossRef] [PubMed]
- Wolter, D.J.; Acquazzino, D.; Goering, R.V.; Sammut, P.; Khalaf, N.; Hanson, N.D. Emergence of carbapenem resistance in Pseudomonas aeruginosa isolates from a patient with cystic fibrosis in the absence of carbapenem therapy. Clin. Infect. Dis. 2008, 46, e137–e141. [Google Scholar] [CrossRef] [PubMed]


| Strain/Plasmids/Primers | Description | Source/Purpose |
|---|---|---|
| PA14 | Wild-type strain | [34] |
| ΔglnK | PA14 glnK gene deletion mutant | [35] |
| ΔglnK/att7::glnK | ΔglnK complementation with glnK inserted on chromosome, Tcr | [35] |
| ΔoprD | PA14 oprD gene deletion mutant | [36] |
| ΔglnKΔoprD | PA14 glnK and oprD genes double deletion mutant | This study |
| ΔntrB | PA14 ntrB gene deletion mutant | [35] |
| ΔntrC | PA14 ntrC gene deletion mutant | [35] |
| ΔglnKΔntrB | PA14 glnK and ntrB genes double deletion mutant | [35] |
| ΔglnKΔntrC | PA14 glnK and ntrC genes double deletion mutant | [35] |
| ΔglnKΔntrBΔoprD | PA14 glnK, ntrB, and oprD genes triple deletion mutant | This study |
| Plasmids | ||
| pEx18Tc-ΔglnK | glnK gene deletion suicide plasmid; Tcr | [35] |
| pEx18Tc-ΔntrB | ntrB gene deletion suicide plasmid; Tcr | [35] |
| pEx18Tc-ΔntrC | ntrC gene deletion suicide plasmid; Tcr | [35] |
| pEx18Tc-ΔoprD | oprD gene deletion suicide plasmid; Tcr | [36] |
| pUC18T-mini-Tn7-Tc-glnK-his | chromosomal integration of glnK-his | [35] |
| pTNS3 | helper plasmid encoding the TnsABCD transposase subunits; Apr | [37] |
| pUCP20-PoprD-lacZ | lacZ with oprD promoter (500 bp); Apr | This study |
| pMMB67EH-ntrC-his | expression driven by an inducible tac promoter; Gmr | [35] |
| Primers | ||
| glnK-LF | CGAGCTCGAGTGCGATGGCCAGGT | glnK deletion |
| glnK-LR | TCGGTTGGGCGAAACTCTCTCCCGTGTT | glnK deletion |
| glnK-RF | GAGAGAGTTTCGCCCAACCGAACCCCCAAA | glnK deletion |
| glnK-RR | CGGGATCCCCATCGGACCGGCGGTG | glnK deletion |
| mini-glnK-F | CGAGCTCGCTGACGCAGGGGGCTTC | GlnK cloning |
| mini-glnK-R | CGGGATCCTTAGTGATGGTGATGGTGATGGATCGCGTCGGTATCGGTTTC | GlnK cloning |
| ntrB-LF ntrB-LR ntrB-RF ntrB-RR ntrC-LF ntrC-LR ntrC-RF ntrC-RR NtrC-F NtrC-R PoprD-F PoprD-R | CGAGCTCAGTTGCAACTGGTGCTGGACGGCAAACCCTACGGCACGCCC CTCTGATCGGCTCATGGGGCGGGCAGCTGTTCCAAGGTTGGGCAGG GGAACAGCTGCCCGCCCCATGAGCCGATCAGAGACCGTCTGGATCGTC CCAAGCTTGCTCCTGGGCGGCGCGGCTGAGGAAGTGCCGGGCCAG CGGGATCCTGGAGTACATGAACCCGGCAG GTGGATCAACGGGTCAATGCACTCCTTGTTCCAGGGGCA GTGCATTGACCGAATACCTGCCCAAGCCGTTCGAC CCAAGCTTCGAGCTGGTGATGAATGCCTCTGGAG CGGAATTCATGAGCCGATCAGAGACCGTCTG CGGGATCCTTAGTGATGGTGATGGTGATGGTCGTCGCCTTCGTCGTC CGGAATTCCAAACGCATTCGCCACAGACAAC CGGGATCCTGTGATTGCTCCTTTGGTTTTG | ntrB deletion ntrB deletion ntrB deletion ntrB deletion ntrC deletion ntrC deletion ntrC deletion ntrC deletion NtrC cloning NtrC cloning PoprD-lacZ cloning PoprD-lacZ cloning |
| Probe-oprD-F | CAAACGCATTCGCCACAG | EMSA |
| Probe-oprD-R Negative-oprD-F Negative-oprD-R | TGTGATTGCTCCTTTGGTTTTG GTGGAGCGCCATTGCACT CTTCGAGTTCGCTGCTCTG | EMSA EMSA EMSA |
| RT-mexB-F | GTGGGTGATCGCCTTGGTG | RT-PCR |
| RT-mexB-R | GCTCACCTGCACGGCGATG | RT-PCR |
| RT-oprM-F | CCGCCTACCTGACGCTGA | RT-PCR |
| RT-oprM-R | ACGCCGACGTCGTAGCTG | RT-PCR |
| RT-oprD-F | CACTCAGTTCGCCGTGGC | RT-PCR |
| RT-oprD-R | GCCGCTCTTGCCGTCACG | RT-PCR |
| RT-mexE-F | CACTTCTCCTGGCGCTAC | RT-PCR |
| RT-mexE-R | CTTCGGCGACGCTGACCT | RT-PCR |
| RT-mexC-F | GTTGGCAGGTTGTGGGCC | RT-PCR |
| RT-mexC-R | ACTCAGCGCCAGGGACTCG | RT-PCR |
| RT-mexY-F | CTGGCGATCCGCTTCCTGC | RT-PCR |
| RT-mexY-R | CCTCGACCACCTTGGCCGAGGC | RT-PCR |
| Strain | MIC a (μg/mL) | |||||||
|---|---|---|---|---|---|---|---|---|
| MEM b | IMP c | DRPM d | FEP e | CAZ f | ATM g | CB h | FOX i | |
| PA14 | 0.25 (0.29) | 1 (0.00) | 0.125 (0.00) | 0.5 (0.00) | 2 (0.00) | 2 (0.00) | 32 (0.00) | 256 (0.00) |
| ΔglnK | 0.0625 (0.02) * | 0.5 (0.00) * | 0.0625 (0.00) * | 0.5 (0.00) | 2 (0.00) | 2 (0.00) | 32 (0.00) | 256 (0.00) |
| ΔglnK/att7::glnK | 0.25 (0.29) | 1 (0.00) | 0.125 (0.00) | 0.5 (0.00) | 2 (0.00) | 2 (0.00) | 32 (0.00) | 256 (0.00) |
| Antibiotics | MIC a (μg/mL) | |||
|---|---|---|---|---|
| PA14 | ΔglnK | ΔoprD | ΔglnKΔoprD | |
| MEM | 0.25 (0.00) * | 0.0625 (0.02) | 16 (0.00) * | 16 (0.00) * |
| IMP | 1 (0.00) * | 0.5 (0.00) | 8 (0.05) * | 8 (0.05) * |
| DRPM | 0.125 (0.00) * | 0.0625 (0.00) | 4 (0.00) * | 4 (0.00) * |
| Antibiotics | MIC (μg/mL) | |||||
|---|---|---|---|---|---|---|
| PA14 | ΔglnK | ΔntrB | ΔglnKΔntrB | ΔntrC | ΔglnKΔntrC | |
| MEM | 0.25 (0.00) * | 0.0625 (0.02) | 0.25 (0.00) * | 0.25 (0.00) * | 0.25 (0.00) * | 0.25 (0.00) * |
| IMP | 1 (0.00) * | 0.5 (0.00) | 2 (0.29) * | 2 (0.29) * | 1 (0.00) * | 1 (0.00) * |
| DRPM | 0.125 (0.00) * | 0.0625 (0.00) | 0.125 (0.00) * | 0.125 (0.00) * | 0.125 (0.00) * | 0.125 (0.00) * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Sun, X.; Li, Y.; Gong, X.; Du, Q.; Jin, Y.; Cheng, Z.; Jin, S.; Wu, W. Glnk Mediates Carbapenem Resistance Through the NtrB/NtrC-OprD Regulatory Pathway in Pseudomonas aeruginosa. Pathogens 2026, 15, 289. https://doi.org/10.3390/pathogens15030289
Sun X, Li Y, Gong X, Du Q, Jin Y, Cheng Z, Jin S, Wu W. Glnk Mediates Carbapenem Resistance Through the NtrB/NtrC-OprD Regulatory Pathway in Pseudomonas aeruginosa. Pathogens. 2026; 15(3):289. https://doi.org/10.3390/pathogens15030289
Chicago/Turabian StyleSun, Xiaomeng, Yiming Li, Xuetao Gong, Qitong Du, Yongxin Jin, Zhihui Cheng, Shouguang Jin, and Weihui Wu. 2026. "Glnk Mediates Carbapenem Resistance Through the NtrB/NtrC-OprD Regulatory Pathway in Pseudomonas aeruginosa" Pathogens 15, no. 3: 289. https://doi.org/10.3390/pathogens15030289
APA StyleSun, X., Li, Y., Gong, X., Du, Q., Jin, Y., Cheng, Z., Jin, S., & Wu, W. (2026). Glnk Mediates Carbapenem Resistance Through the NtrB/NtrC-OprD Regulatory Pathway in Pseudomonas aeruginosa. Pathogens, 15(3), 289. https://doi.org/10.3390/pathogens15030289





