Physalin F Promotes AFG3L2-Mediated Degradation of VISA/MAVS to Suppress Innate Immune Response to RNA Virus
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents, Antibodies, Viruses, and Cells
2.2. Constructs
2.3. Transfection and Reporter Assay
2.4. Screening of Compounds
2.5. RT-qPCR
2.6. CRISPR/Cas9 Gene Editing
2.7. Immunoblotting Analysis
2.8. Mice
2.9. ELISA
2.10. Preparation of BMDMs and BMDCs
2.11. Limited Proteolysis Coupled with Mass Spectrometry (LiP-MS)
2.12. Cellular Thermal Shift Assay (CETSA)
2.13. Statistical Analysis
3. Results
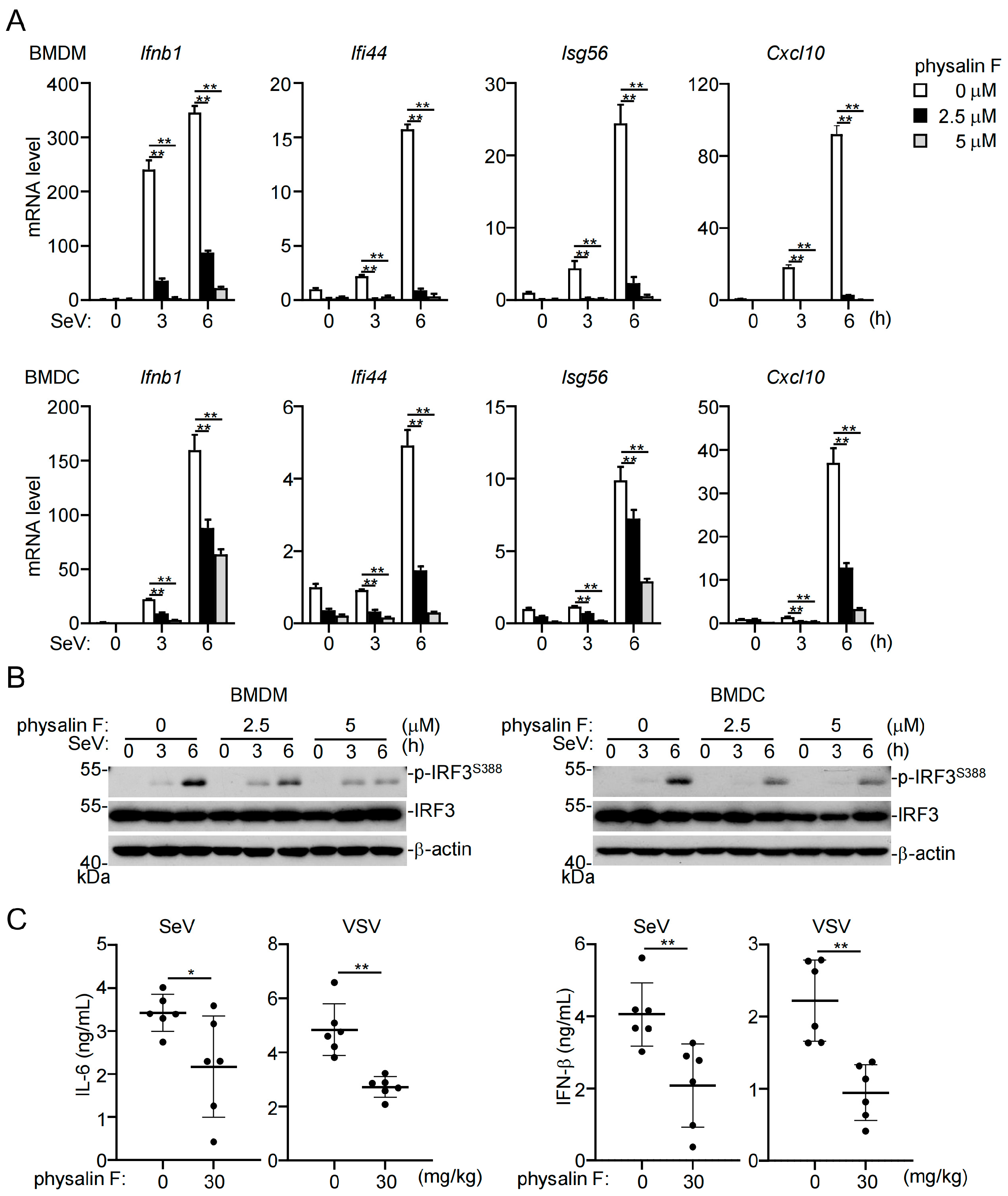
3.1. Identification of Physalin F as an Inhibitor of RNA Virus-Induced Innate Antiviral Response
3.2. Physalin F Directly Interacts with the Mitochondrial m-AAA Protease AFG3L2
3.3. Inhibition of Innate Antiviral Signaling by Physalin F Is Mediated by AFG3L2
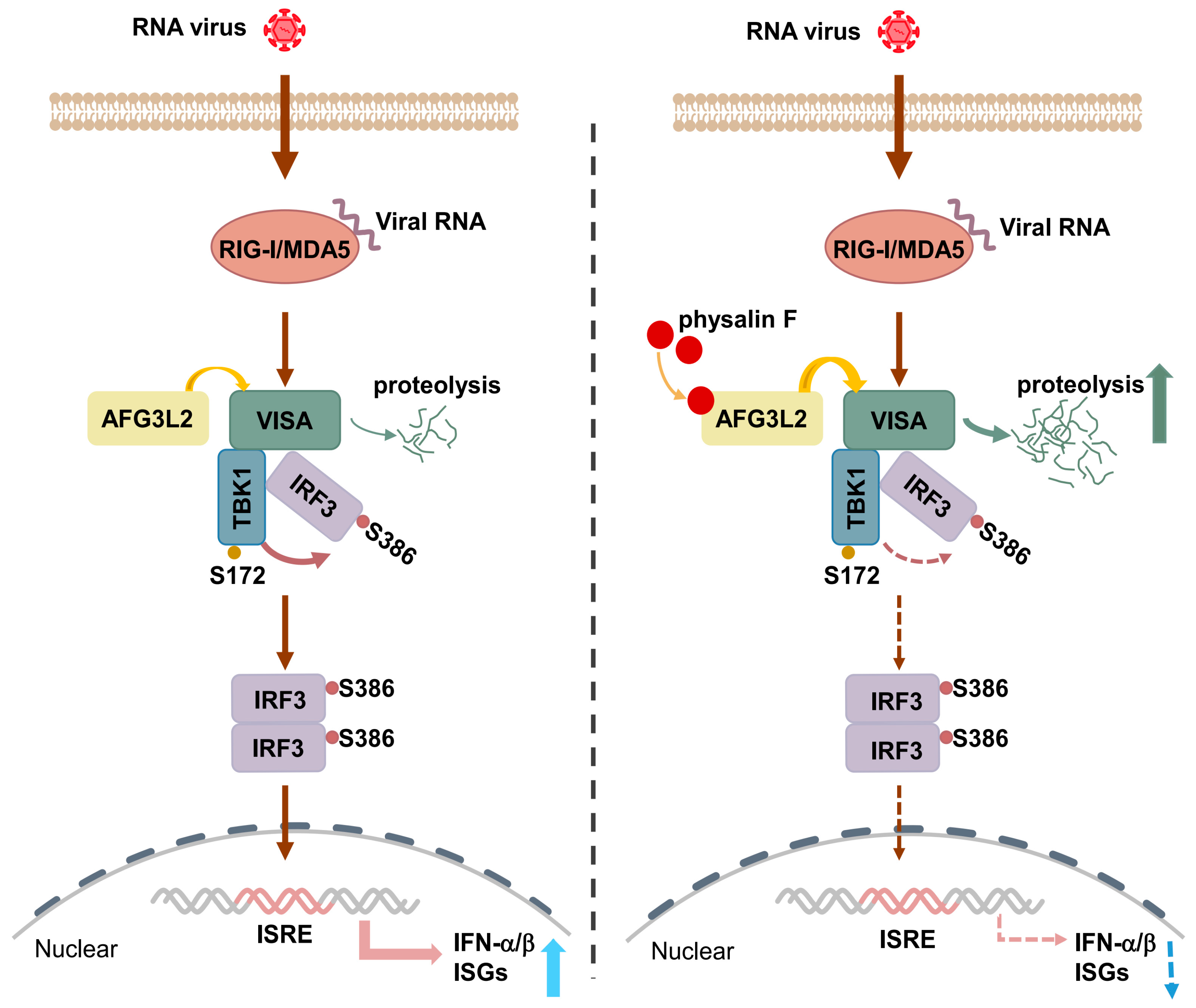
3.4. AFG3L2 Mediates Degradation of VISA
3.5. Physalin F Interacts with AFG3L2 to Enhance Its Proteolytic Activity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AFG | ATPase family gene |
| AFG3L2 | AFG3-like matrix AAA peptidase subunit 2 |
| AGS | Aicardi Goutieres syndrome |
| BMDCs | bone marrow-derived dendritic cells |
| BMDMs | bone marrow-derived macrophages |
| CARD | caspase activation and recruitment domain |
| CETSA | cellular thermal shift assay |
| CRISPR | clustered regularly interspaced short palindromic repeats |
| DMSO | dimethyl sulfoxide |
| EGCG | epigallocatechin gallate |
| EMRE | essential MCU regulator element |
| GPATCH3 | G patch domain containing protein 3 |
| HEK293T | human embryonic kidney |
| HTLV-1 | human T lymphotropic virus type 1 |
| IC50 | half-maximal inhibitory concentration |
| IFNs | type I interferons |
| IRF3 | interferon regulatory transcription factor 3 |
| IMM | inner mitochondrial membrane |
| ISRE | IFN-stimulated response element |
| LiP-MS | Limited Proteolysis coupled with Mass Spectrometry |
| LPS | Lipopolysaccharide |
| m-AAA | matrix ATPase associated with diverse cellular activities |
| MDA5 | melanoma differentiation-associated gene 5 |
| MICOS | mitochondrial contact site and cristae organizing system |
| mPTP | mitochondrial permeability transition pore |
| MUC | mitochondrial calcium uniporter |
| OMM | outer mitochondrial membrane |
| OXPHOS | oxidative phosphorylation |
| RIG-I | retinoic acid-inducible gene I |
| RLRs | RIG I like receptors |
| ROS | reactive oxygen species |
| PRRs | pattern recognition receptors |
| SeV | Sendai virus |
| SLE | Systemic lupus erythematosus |
| SNX8 | Several proteins, including sorting nexin 8 |
| Tagg | aggregation temperature |
| TBK1 | TANK-binding kinase 1 |
| THP1 | human monocytes |
| TNFα | tumor necrosis factor alpha |
| TRX2 | Thioredoxin 2 |
| VISA | virus-induced signaling adaptor |
| VSV | Vesicular stomatitis virus |
References
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen Recognition and Innate Immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef]
- Barbalat, R.; Ewald, S.E.; Mouchess, M.L.; Barton, G.M. Nucleic Acid Recognition by the Innate Immune System. Annu. Rev. Immunol. 2011, 29, 185–214. [Google Scholar] [CrossRef]
- Hu, M.-M.; Shu, H.-B. Cytoplasmic Mechanisms of Recognition and Defense of Microbial Nucleic Acids. Annu. Rev. Cell Dev. Biol. 2018, 34, 357–379. [Google Scholar] [CrossRef]
- Heim, M.H.; Thimme, R. Innate and Adaptive Immune Responses in HCV Infections. J. Hepatol. 2014, 61, S14–S25. [Google Scholar] [CrossRef]
- Kato, H.; Takeuchi, O.; Sato, S.; Yoneyama, M.; Yamamoto, M.; Matsui, K.; Uematsu, S.; Jung, A.; Kawai, T.; Ishii, K.J.; et al. Differential Roles of MDA5 and RIG-I Helicases in the Recognition of RNA Viruses. Nature 2006, 441, 101–105. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, M.; Kikuchi, M.; Natsukawa, T.; Shinobu, N.; Imaizumi, T.; Miyagishi, M.; Taira, K.; Akira, S.; Fujita, T. The RNA Helicase RIG-I Has an Essential Function in Double-Stranded RNA-Induced Innate Antiviral Responses. Nat. Immunol. 2004, 5, 730–737. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.-G.; Wang, Y.-Y.; Han, K.-J.; Li, L.-Y.; Zhai, Z.; Shu, H.-B. VISA Is an Adapter Protein Required for Virus-Triggered IFN-β Signaling. Mol. Cell 2005, 19, 727–740. [Google Scholar] [CrossRef] [PubMed]
- Seth, R.B.; Sun, L.; Ea, C.-K.; Chen, Z.J. Identification and Characterization of MAVS, a Mitochondrial Antiviral Signaling Protein That Activates NF-κB and IRF3. Cell 2005, 122, 669–682. [Google Scholar] [CrossRef]
- Kawai, T.; Takahashi, K.; Sato, S.; Coban, C.; Kumar, H.; Kato, H.; Ishii, K.J.; Takeuchi, O.; Akira, S. IPS-1, an Adaptor Triggering RIG-I- and Mda5-Mediated Type I Interferon Induction. Nat. Immunol. 2005, 6, 981–988. [Google Scholar] [CrossRef]
- Hou, F.; Sun, L.; Zheng, H.; Skaug, B.; Jiang, Q.-X.; Chen, Z.J. MAVS Forms Functional Prion-like Aggregates to Activate and Propagate Antiviral Innate Immune Response. Cell 2011, 146, 448–461. [Google Scholar] [CrossRef]
- Saha, S.K.; Pietras, E.M.; He, J.Q.; Kang, J.R.; Liu, S.; Oganesyan, G.; Shahangian, A.; Zarnegar, B.; Shiba, T.L.; Wang, Y.; et al. Regulation of Antiviral Responses by a Direct and Specific Interaction between TRAF3 and Cardif. EMBO J. 2006, 25, 3257–3263. [Google Scholar] [CrossRef]
- Liu, S.; Chen, J.; Cai, X.; Wu, J.; Chen, X.; Wu, Y.-T.; Sun, L.; Chen, Z.J. MAVS Recruits Multiple Ubiquitin E3 Ligases to Activate Antiviral Signaling Cascades. eLife 2013, 2, e00785. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Cai, X.; Wu, J.; Cong, Q.; Chen, X.; Li, T.; Du, F.; Ren, J.; Wu, Y.-T.; Grishin, N.V.; et al. Phosphorylation of Innate Immune Adaptor Proteins MAVS, STING, and TRIF Induces IRF3 Activation. Science 2015, 347, aaa2630. [Google Scholar] [CrossRef]
- Guo, W.; Wei, J.; Zhong, X.; Zang, R.; Lian, H.; Hu, M.-M.; Li, S.; Shu, H.-B.; Yang, Q. SNX8 Modulates the Innate Immune Response to RNA Viruses by Regulating the Aggregation of VISA. Cell. Mol. Immunol. 2020, 17, 1126–1135. [Google Scholar] [CrossRef]
- Liu, B.; Zhang, M.; Chu, H.; Zhang, H.; Wu, H.; Song, G.; Wang, P.; Zhao, K.; Hou, J.; Wang, X.; et al. The Ubiquitin E3 Ligase TRIM31 Promotes Aggregation and Activation of the Signaling Adaptor MAVS through Lys63-Linked Polyubiquitination. Nat. Immunol. 2017, 18, 214–224. [Google Scholar] [CrossRef]
- Nie, Y.; Ran, Y.; Zhang, H.-Y.; Huang, Z.-F.; Pan, Z.-Y.; Wang, S.-Y.; Wang, Y.-Y. GPATCH3 Negatively Regulates RLR-Mediated Innate Antiviral Responses by Disrupting the Assembly of VISA Signalosome. PLoS Pathog. 2017, 13, e1006328. [Google Scholar] [CrossRef]
- Li, D.; Yang, W.; Ru, Y.; Ren, J.; Liu, X.; Zheng, H. Thioredoxin 2 Negatively Regulates Innate Immunity to RNA Viruses by Disrupting the Assembly of the Virus-Induced Signaling Adaptor Complex. J. Virol. 2020, 94, e01756-19. [Google Scholar] [CrossRef]
- Zhong, B.; Zhang, Y.; Tan, B.; Liu, T.-T.; Wang, Y.-Y.; Shu, H.-B. The E3 Ubiquitin Ligase RNF5 Targets Virus-Induced Signaling Adaptor for Ubiquitination and Degradation. J. Immunol. 2010, 184, 6249–6255. [Google Scholar] [CrossRef]
- Yan, B.-R.; Zhou, L.; Hu, M.-M.; Li, M.; Lin, H.; Yang, Y.; Wang, Y.-Y.; Shu, H.-B. PKACs Attenuate Innate Antiviral Response by Phosphorylating VISA and Priming It for MARCH5-Mediated Degradation. PLoS Pathog. 2017, 13, e1006648. [Google Scholar] [CrossRef] [PubMed]
- Yoo, Y.-S.; Park, Y.-Y.; Kim, J.-H.; Cho, H.; Kim, S.-H.; Lee, H.-S.; Kim, T.-H.; Sun Kim, Y.; Lee, Y.; Kim, C.-J.; et al. The Mitochondrial Ubiquitin Ligase MARCH5 Resolves MAVS Aggregates during Antiviral Signalling. Nat. Commun. 2015, 6, 7910. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Li, R.; Meng, J.-L.; Mao, H.-T.; Zhang, Y.; Zhang, J. Smurf2 Negatively Modulates RIG-I–Dependent Antiviral Response by Targeting VISA/MAVS for Ubiquitination and Degradation. J. Immunol. 2014, 192, 4758–4764. [Google Scholar] [CrossRef] [PubMed]
- Kretschmer, S.; Lee-Kirsch, M.A. Type I Interferon-Mediated Autoinflammation and Autoimmunity. Curr. Opin. Immunol. 2017, 49, 96–102. [Google Scholar] [CrossRef]
- Rönnblom, L.; Eloranta, M.-L. The Interferon Signature in Autoimmune Diseases. Curr. Opin. Rheumatol. 2013, 25, 248. [Google Scholar] [CrossRef]
- Varga, Z.; Rafay Ali Sabzwari, S.; Vargova, V. Cardiovascular Risk of Nonsteroidal Anti-Inflammatory Drugs: An Under-Recognized Public Health Issue. Cureus 2017, 9, e1144. [Google Scholar] [CrossRef]
- Schneider, B.J.; Naidoo, J.; Santomasso, B.D.; Lacchetti, C.; Adkins, S.; Anadkat, M.; Atkins, M.B.; Brassil, K.J.; Caterino, J.M.; Chau, I.; et al. Management of Immune-Related Adverse Events in Patients Treated With Immune Checkpoint Inhibitor Therapy: ASCO Guideline Update. J. Clin. Oncol. 2021, 39, 4073–4126. [Google Scholar] [CrossRef]
- Baker, D.D.; Alvi, K.A. Small-Molecule Natural Products: New Structures, New Activities. Curr. Opin. Biotechnol. 2004, 15, 576–583. [Google Scholar] [CrossRef]
- Gasmi, A.; Shanaida, M.; Oleshchuk, O.; Semenova, Y.; Mujawdiya, P.K.; Ivankiv, Y.; Pokryshko, O.; Noor, S.; Piscopo, S.; Adamiv, S.; et al. Natural Ingredients to Improve Immunity. Pharmaceuticals 2023, 16, 528. [Google Scholar] [CrossRef]
- Dai, J.; Huang, Y.-J.; He, X.; Zhao, M.; Wang, X.; Liu, Z.-S.; Xue, W.; Cai, H.; Zhan, X.-Y.; Huang, S.-Y.; et al. Acetylation Blocks cGAS Activity and Inhibits Self-DNA-Induced Autoimmunity. Cell 2019, 176, 1447–1460.e14. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.-S.; Cai, H.; Xue, W.; Wang, M.; Xia, T.; Li, W.-J.; Xing, J.-Q.; Zhao, M.; Huang, Y.-J.; Chen, S.; et al. G3BP1 Promotes DNA Binding and Activation of cGAS. Nat. Immunol. 2019, 20, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-S.; Chiang, H.-C.; Kan, W.-S.; Hone, E.; Shih, S.-J.; Won, M.-H. Immunomodulatory Activity of Various Fractions Derived from Physalis angulata L Extract. Am. J. Chin. Med. 1992, 20, 233–243. [Google Scholar] [CrossRef]
- Silva, D.K.C.; Novo, L.B.d.C.; Ribeiro, I.M.; Barreto, B.C.; Opretzka, L.C.F.; Meira, C.S.; Soares, M.B.P. Physalin F, a Potent Inhibitor of Lymphocyte Function, Is a Calcineurin Inhibitor and Has Synergistic Effect with Dexamethasone. Molecules 2025, 30, 916. [Google Scholar] [CrossRef]
- Meira, C.S.; Soares, J.W.C.; dos Reis, B.P.Z.C.; Pacheco, L.V.; Santos, I.P.; Silva, D.K.C.; de Lacerda, J.C.; Daltro, S.R.T.; Guimarães, E.T.; Soares, M.B.P. Therapeutic Applications of Physalins: Powerful Natural Weapons. Front. Pharmacol. 2022, 13, 864714. [Google Scholar] [CrossRef]
- Pinto, L.A.; Meira, C.S.; Villarreal, C.F.; Vannier-Santos, M.A.; de Souza, C.V.C.; Ribeiro, I.M.; Tomassini, T.C.B.; Galvão-Castro, B.; Soares, M.B.P.; Grassi, M.F.R. Physalin F, a Seco-Steroid from Physalis angulata L., Has Immunosuppressive Activity in Peripheral Blood Mononuclear Cells from Patients with HTLV1-Associated Myelopathy. Biomed. Pharmacother. 2016, 79, 129–134. [Google Scholar] [CrossRef]
- Ooi, K.L.; Tengku Muhammad, T.S.; Sulaiman, S.F. Physalin F from Physalis minima L. Triggers Apoptosis-Based Cytotoxic Mechanism in T-47D Cells through the Activation Caspase-3- and c-Myc-Dependent Pathways. J. Ethnopharmacol. 2013, 150, 382–388. [Google Scholar] [CrossRef]
- Wu, S.-Y.; Leu, Y.-L.; Chang, Y.-L.; Wu, T.-S.; Kuo, P.-C.; Liao, Y.-R.; Teng, C.-M.; Pan, S.-L. Physalin F Induces Cell Apoptosis in Human Renal Carcinoma Cells by Targeting NF-kappaB and Generating Reactive Oxygen Species. PLoS ONE 2012, 7, e40727. [Google Scholar] [CrossRef]
- Patron, M.; Sprenger, H.-G.; Langer, T. M-AAA Proteases, Mitochondrial Calcium Homeostasis and Neurodegeneration. Cell Res. 2018, 28, 296–306. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, R.; Zhou, Q.; Xu, Z.; Li, C.; Wang, S.; Mao, A.; Zhang, X.; He, W.; Shu, H.-B. LSm14A Is a Processing Body-Associated Sensor of Viral Nucleic Acids That Initiates Cellular Antiviral Response in the Early Phase of Viral Infection. Proc. Natl. Acad. Sci. USA 2012, 109, 11770–11775. [Google Scholar] [CrossRef]
- Luo, W.-W.; Li, S.; Li, C.; Zheng, Z.-Q.; Cao, P.; Tong, Z.; Lian, H.; Wang, S.-Y.; Shu, H.-B.; Wang, Y.-Y. iRhom2 Is Essential for Innate Immunity to RNA Virus by Antagonizing ER- and Mitochondria-Associated Degradation of VISA. PLoS Pathog. 2017, 13, e1006693. [Google Scholar] [CrossRef] [PubMed]
- Cao, L.-B.; Ruan, Z.-L.; Yang, Y.-L.; Zhang, N.-C.; Gao, C.; Cai, C.; Zhang, J.; Hu, M.-M.; Shu, H.-B. Estrogen Receptor α-Mediated Signaling Inhibits Type I Interferon Response to Promote Breast Carcinogenesis. J. Mol. Cell Biol. 2023, 15, mjad047. [Google Scholar] [CrossRef]
- Sanjana, N.E.; Shalem, O.; Zhang, F. Improved Vectors and Genome-Wide Libraries for CRISPR Screening. Nat. Methods 2014, 11, 783–784. [Google Scholar] [CrossRef] [PubMed]
- Shalem, O.; Sanjana, N.E.; Hartenian, E.; Shi, X.; Scott, D.A.; Mikkelsen, T.S.; Heckl, D.; Ebert, B.L.; Root, D.E.; Doench, J.G.; et al. Genome-Scale CRISPR-Cas9 Knockout Screening in Human Cells. Science 2014, 343, 84–87. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Shu, H.-B. Dephosphorylation of cGAS by PPP6C Impairs Its Substrate Binding Activity and Innate Antiviral Response. Protein Cell 2020, 11, 584–599. [Google Scholar] [CrossRef]
- Piazza, I.; Kochanowski, K.; Cappelletti, V.; Fuhrer, T.; Noor, E.; Sauer, U.; Picotti, P. A Map of Protein-Metabolite Interactions Reveals Principles of Chemical Communication. Cell 2018, 172, 358–372.e23. [Google Scholar] [CrossRef] [PubMed]
- Grentzmann, G.; Ingram, J.A.; Kelly, P.J.; Gesteland, R.F.; Atkins, J.F. A Dual-Luciferase Reporter System for Studying Recoding Signals. RNA 1998, 4, 479–486. [Google Scholar] [PubMed]
- Zhang, R.; Zheng, H. Luciferase Reporter Assay for Determining the Signaling Activity of Interferons. In Antiviral Innate Immunity; Zheng, C., Ed.; Springer: New York, NY, USA, 2025; pp. 19–28. ISBN 978-1-0716-4108-8. [Google Scholar]
- Jafari, R.; Almqvist, H.; Axelsson, H.; Ignatushchenko, M.; Lundbäck, T.; Nordlund, P.; Molina, D.M. The Cellular Thermal Shift Assay for Evaluating Drug Target Interactions in Cells. Nat. Protoc. 2014, 9, 2100–2122. [Google Scholar] [CrossRef]
- Augustin, S.; Gerdes, F.; Lee, S.; Tsai, F.T.F.; Langer, T.; Tatsuta, T. An Intersubunit Signaling Network Coordinates ATP Hydrolysis by M-AAA Proteases. Mol. Cell 2009, 35, 574–585. [Google Scholar] [CrossRef]
- Ding, B.; Martin, D.W.; Rampello, A.J.; Glynn, S.E. Dissecting Substrate Specificities of the Mitochondrial AFG3L2 Protease. Biochemistry 2018, 57, 4225–4235. [Google Scholar] [CrossRef]
- Puchades, C.; Ding, B.; Song, A.; Wiseman, R.L.; Lander, G.C.; Glynn, S.E. Unique Structural Features of the Mitochondrial AAA+ Protease AFG3L2 Reveal the Molecular Basis for Activity in Health and Disease. Mol. Cell 2019, 75, 1073–1085.e6. [Google Scholar] [CrossRef]
- Ablasser, A.; Hertrich, C.; Waßermann, R.; Hornung, V. Nucleic Acid Driven Sterile Inflammation. Clin. Immunol. 2013, 147, 207–215. [Google Scholar] [CrossRef]
- Roers, A.; Hiller, B.; Hornung, V. Recognition of Endogenous Nucleic Acids by the Innate Immune System. Immunity 2016, 44, 739–754. [Google Scholar] [CrossRef]
- Jiang, J.; Zhao, M.; Chang, C.; Wu, H.; Lu, Q. Type I Interferons in the Pathogenesis and Treatment of Autoimmune Diseases. Clin. Rev. Allergy Immunol. 2020, 59, 248–272. [Google Scholar] [CrossRef]
- Liang, Y.; Liang, L.; Shi, R.; Luo, R.; Yue, Y.; Yu, J.; Wang, X.; Lin, J.; Zhou, T.; Yang, M.; et al. Genus Physalis L.: A Review of Resources and Cultivation, Chemical Composition, Pharmacological Effects and Applications. J. Ethnopharmacol. 2024, 324, 117736. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Li, C.; Wang, Y.; Yue, Y.; Zhang, H.; Yang, M.; Cao, X.; Zhao, M.; Du, J.; Peng, M.; et al. Physalis alkekengi L. Var. Franchetii (Mast.) Makino: A Review of the Pharmacognosy, Chemical Constituents, Pharmacological Effects, Quality Control, and Applications. Phytomedicine 2022, 105, 154328. [Google Scholar] [CrossRef]
- Chen, C.; Zhu, D.; Zhang, H.; Han, C.; Xue, G.; Zhu, T.; Luo, J.; Kong, L. YAP-Dependent Ubiquitination and Degradation of β-Catenin Mediates Inhibition of Wnt Signalling Induced by Physalin F in Colorectal Cancer. Cell Death Dis. 2018, 9, 591. [Google Scholar] [CrossRef]
- Soares, M.B.P.; Brustolim, D.; Santos, L.A.; Bellintani, M.C.; Paiva, F.P.; Ribeiro, Y.M.; Tomassini, T.C.B.; Ribeiro dos Santos, R. Physalins B, F and G, Seco-Steroids Purified from Physalis angulata L., Inhibit Lymphocyte Function and Allogeneic Transplant Rejection. Int. Immunopharmacol. 2006, 6, 408–414. [Google Scholar] [CrossRef]
- Soares, M.B.P.; Bellintani, M.C.; Ribeiro, I.M.; Tomassini, T.C.B.; Ribeiro dos Santos, R. Inhibition of Macrophage Activation and Lipopolysaccaride-Induced Death by Seco-Steroids Purified from Physalis angulata L. Eur. J. Pharmacol. 2003, 459, 107–112. [Google Scholar] [CrossRef] [PubMed]
- Atorino, L.; Silvestri, L.; Koppen, M.; Cassina, L.; Ballabio, A.; Marconi, R.; Langer, T.; Casari, G. Loss of M-AAA Protease in Mitochondria Causes Complex I Deficiency and Increased Sensitivity to Oxidative Stress in Hereditary Spastic Paraplegia. J. Cell Biol. 2003, 163, 777–787. [Google Scholar] [CrossRef]
- Maltecca, F.; Magnoni, R.; Cerri, F.; Cox, G.A.; Quattrini, A.; Casari, G. Haploinsufficiency of AFG3L2, the Gene Responsible for Spinocerebellar Ataxia Type 28, Causes Mitochondria-Mediated Purkinje Cell Dark Degeneration. J. Neurosci. 2009, 29, 9244–9254. [Google Scholar] [CrossRef]
- Almajan, E.R.; Richter, R.; Paeger, L.; Martinelli, P.; Barth, E.; Decker, T.; Larsson, N.-G.; Kloppenburg, P.; Langer, T.; Rugarli, E.I. AFG3L2 Supports Mitochondrial Protein Synthesis and Purkinje Cell Survival. J. Clin. Investig. 2012, 122, 4048–4058. [Google Scholar] [CrossRef] [PubMed]
- Mancini, C.; Hoxha, E.; Iommarini, L.; Brussino, A.; Richter, U.; Montarolo, F.; Cagnoli, C.; Parolisi, R.; Gondor Morosini, D.I.; Nicolò, V.; et al. Mice Harbouring a SCA28 Patient Mutation in AFG3L2 Develop Late-Onset Ataxia Associated with Enhanced Mitochondrial Proteotoxicity. Neurobiol. Dis. 2019, 124, 14–28. [Google Scholar] [CrossRef]
- Pareek, G.; Pallanck, L.J. Inactivation of the Mitochondrial Protease Afg3l2 Results in Severely Diminished Respiratory Chain Activity and Widespread Defects in Mitochondrial Gene Expression. PLoS Genet. 2020, 16, e1009118. [Google Scholar] [CrossRef]
- Nolden, M.; Ehses, S.; Koppen, M.; Bernacchia, A.; Rugarli, E.I.; Langer, T. The M-AAA Protease Defective in Hereditary Spastic Paraplegia Controls Ribosome Assembly in Mitochondria. Cell 2005, 123, 277–289. [Google Scholar] [CrossRef]
- Ehses, S.; Raschke, I.; Mancuso, G.; Bernacchia, A.; Geimer, S.; Tondera, D.; Martinou, J.-C.; Westermann, B.; Rugarli, E.I.; Langer, T. Regulation of OPA1 Processing and Mitochondrial Fusion by M-AAA Protease Isoenzymes and OMA1. J. Cell Biol. 2009, 187, 1023–1036. [Google Scholar] [CrossRef]
- König, T.; Tröder, S.E.; Bakka, K.; Korwitz, A.; Richter-Dennerlein, R.; Lampe, P.A.; Patron, M.; Mühlmeister, M.; Guerrero-Castillo, S.; Brandt, U.; et al. The m-AAA Protease Associated with Neurodegeneration Limits MCU Activity in Mitochondria. Mol. Cell 2016, 64, 148–162. [Google Scholar] [CrossRef] [PubMed]
- Wei, K.; Chen, T.; Fang, H.; Shen, X.; Tang, Z.; Zhao, J. Mitochondrial DNA Release via the Mitochondrial Permeability Transition Pore Activates the cGAS-STING Pathway, Exacerbating Inflammation in Acute Kawasaki Disease. Cell Commun. Signal. CCS 2024, 22, 328. [Google Scholar] [CrossRef]
- Shi, X.; DeCiucis, M.; Grabinska, K.A.; Kanyo, J.; Liu, A.; Lam, T.T.; Shen, H. Dual Regulation of SLC25A39 by AFG3L2 and Iron Controls Mitochondrial Glutathione Homeostasis. Mol. Cell 2024, 84, 802–810.e6. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Yang, J.; Chen, Z.; Li, Y.; Luo, G.; Hu, K.; Lu, Z.; Lv, X.; Cao, Y.; Yuan, H.; et al. SLC25A39 Overexpression Exacerbates Lung Adenocarcinoma Progression and Is Negatively Regulated by AFG3L2. npj Precis. Oncol. 2025, 9, 317. [Google Scholar] [CrossRef] [PubMed]
- Van Der Laan, M.; Horvath, S.E.; Pfanner, N. Mitochondrial Contact Site and Cristae Organizing System. Curr. Opin. Cell Biol. 2016, 41, 33–42. [Google Scholar] [CrossRef]
- Wu, X.; Li, L.; Jiang, H. Mitochondrial Inner-Membrane Protease Yme1 Degrades Outer-Membrane Proteins Tom22 and Om45. J. Cell Biol. 2017, 217, 139–149. [Google Scholar] [CrossRef]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Gao, X.-N.; Shu, H.-B.; Li, M. Physalin F Promotes AFG3L2-Mediated Degradation of VISA/MAVS to Suppress Innate Immune Response to RNA Virus. Pathogens 2026, 15, 74. https://doi.org/10.3390/pathogens15010074
Gao X-N, Shu H-B, Li M. Physalin F Promotes AFG3L2-Mediated Degradation of VISA/MAVS to Suppress Innate Immune Response to RNA Virus. Pathogens. 2026; 15(1):74. https://doi.org/10.3390/pathogens15010074
Chicago/Turabian StyleGao, Xiao-Nan, Hong-Bing Shu, and Mi Li. 2026. "Physalin F Promotes AFG3L2-Mediated Degradation of VISA/MAVS to Suppress Innate Immune Response to RNA Virus" Pathogens 15, no. 1: 74. https://doi.org/10.3390/pathogens15010074
APA StyleGao, X.-N., Shu, H.-B., & Li, M. (2026). Physalin F Promotes AFG3L2-Mediated Degradation of VISA/MAVS to Suppress Innate Immune Response to RNA Virus. Pathogens, 15(1), 74. https://doi.org/10.3390/pathogens15010074





