Molecular and Culture-Based Surveillance of Free-Living Amoebae in Human Related Sources in an Outermost Region
Abstract
1. Introduction
2. Materials and Methods
2.1. Geographical Location
2.2. Sample Collection and Processing
2.3. Amoebae Molecular Characterization
2.4. qPCR Assay
2.5. FLA Sequencing and Genotyping by Standard PCR
2.6. Phylogenetic Analysis
2.7. Tolerance Assays on Potentially Pathogenic Amoeba
3. Results
3.1. Pathogenic FLA Detection by qPCR
3.2. FLA Isolation by NNA Culture
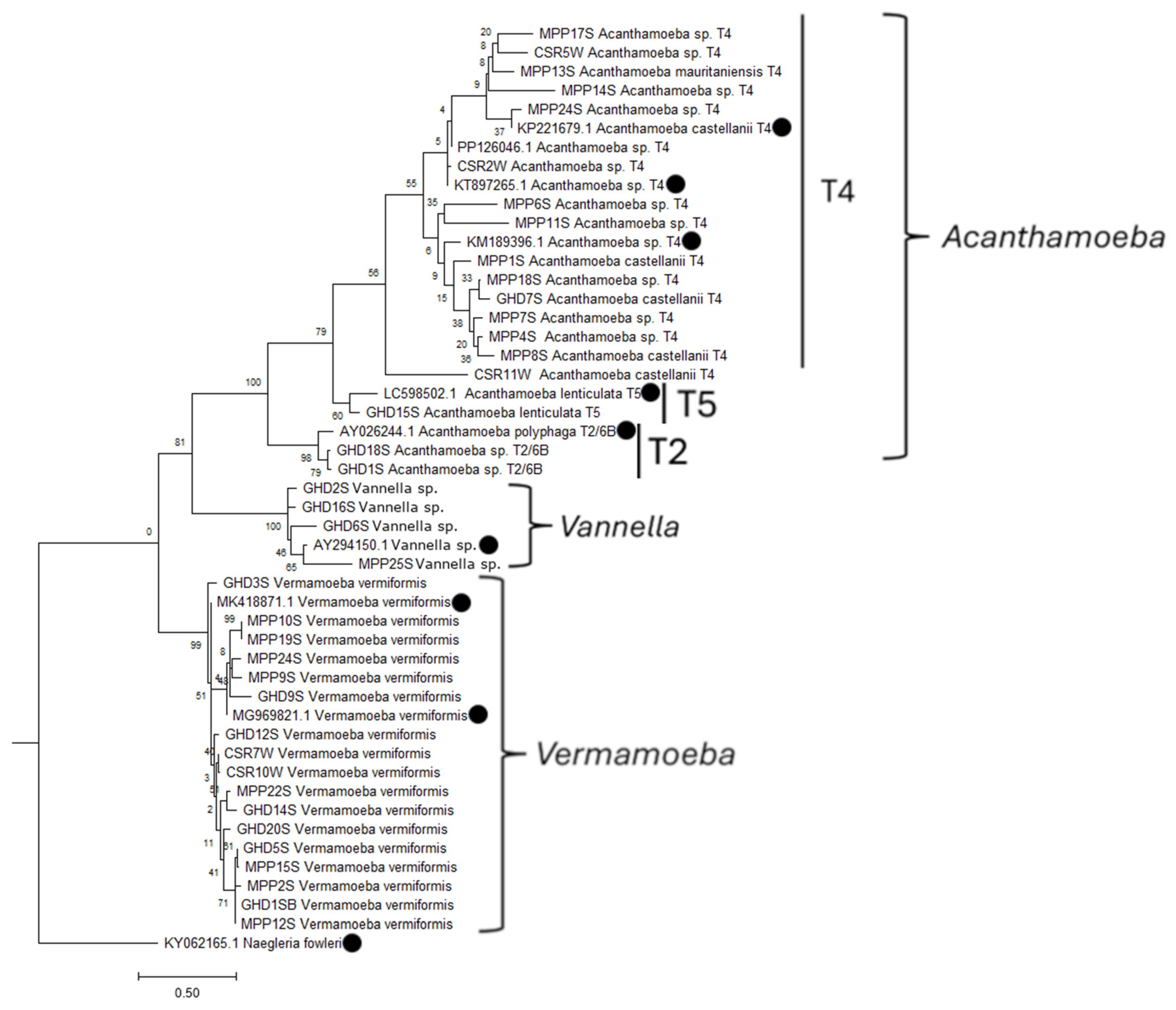
3.3. Phylogenetic Relationship
3.4. Osmotolerance and Thermotolerance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| FLA | Free living amoebae |
| qPCR | Quantitative Polymerase Chain Reaction |
| NNA | Non nutrient agar |
| AS | Agricultural Soil |
| PS | Playground Soil |
| RW | Refrigerated Water |
| PAS | Page’s Amoeba Solution or Page’s Saline |
References
- Martinez, A.J.; Visvesvara, G.S. Free-living, amphizoic and opportunistic amebas. Brain Pathol. 1997, 7, 583–598. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dupuy, M.; Berne, F.; Herbelin, P.; Binet, M.; Berthelot, N.; Rodier, M.H.; Soreau, S.; Héchard, Y. Sensitivity of free-living amoeba trophozoites and cysts to water disinfectants. Int. J. Hyg. Environ. Health 2014, 217, 335–339. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Zaragoza, S. Ecology of free-living amoebae. Crit. Rev. Microbiol. 1994, 20, 225–241. [Google Scholar] [CrossRef] [PubMed]
- Curds, C.R. The ecology and role of protozoa in aerobic sewage treatment processes. Annu. Rev. Microbiol. 1982, 36, 27–46. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, R.; Makhlouf, Z.; Khan, N.A. The increasing importance of Vermamoeba vermiformis. J. Eukaryot. Microbiol. 2021, 68, e12857. [Google Scholar] [CrossRef]
- Chaúque, B.J.M.; da Silva, T.C.B.; dos Santos, D.L.; Benitez, G.B.; Chaúque, L.G.H.; Benetti, A.D.; Zanette, R.A.; Rott, M.B. Global prevalence of free-living amoebae in solid matrices–A systematic review with meta-analysis. Acta Trop. 2023, 247, 107006. [Google Scholar] [CrossRef]
- Geisen, S.; Fiore-Donno, A.M.; Walochnik, J.; Bonkowski, M. Acanthamoeba everywhere: High diversity of Acanthamoeba in soils. Parasitol. Res. 2014, 113, 3151–3158. [Google Scholar] [CrossRef]
- Lorenzo-Morales, J.; Khan, N.A.; Walochnik, J. An update on Acanthamoeba keratitis: Diagnosis, pathogenesis and treatment. Parasite 2015, 22, 10. [Google Scholar] [CrossRef]
- Diehl, M.L.N.; Paes, J.; Rott, M.B. Genotype distribution of Acanthamoeba in keratitis: A systematic review. Parasitol. Res. 2021, 120, 3051–3063. [Google Scholar] [CrossRef]
- Delafont, V.; Rodier, M.-H.; Maisonneuve, E.; Cateau, E. Vermamoeba vermiformis: A Free-Living Amoeba of Interest. Microb. Ecol. 2018, 76, 991–1001. [Google Scholar] [CrossRef]
- Scheid, P.L.; Lâm, T.-T.; Sinsch, U.; Balczun, C. Vermamoeba vermiformis as etiological agent of a painful ulcer close to the eye. Parasitol. Res. 2019, 118, 1999–2004. [Google Scholar] [CrossRef] [PubMed]
- Moran, S.; Mooney, R.; Henriquez, F.L. Diagnostic Considerations for Non-Acanthamoeba Amoebic Keratitis and Clinical Outcomes. Pathogens 2022, 11, 219. [Google Scholar] [CrossRef] [PubMed]
- Leal Dos Santos, D.; Chaúque, B.J.M.; Virginio, V.G.; Cossa, V.C.; Pettan-Brewer, C.; Schrekker, H.S.; Rott, M.B. Occurrence of Naegleria fowleri and their implication for health-a look under the One Health approaches. Int. J. Hyg. Environ. Health 2022, 246, 114053. [Google Scholar] [CrossRef] [PubMed]
- Jahangeer, M.; Mahmood, Z.; Munir, N.; Waraich, U.E.A.; Tahir, I.M.; Akram, M.; Ali Shah, S.M.; Zulfqar, A.; Zainab, R. Naegleria fowleri: Sources of infection, pathophysiology, diagnosis, and management; a review. Clin. Exp. Pharmacol. Physiol. 2020, 47, 199–212. [Google Scholar] [CrossRef]
- Phung, N.T.N.; Pham, H.T.; Tran, T.T.; Dinh, V.H.; Tran, N.M.; Tran, N.A.N.; Ngo, M.Q.N.; Nguyen, H.T.T.; Tran, D.K.; Le, T.K.T.; et al. Naegleria fowleri: Portrait of a Cerebral Killer. Diagnostics 2025, 15, 89. [Google Scholar] [CrossRef]
- Cabello-Vílchez, A.M.; Reyes-Batlle, M.; Montalbán-Sandoval, E.; Martín-Navarro, C.M.; López-Arencibia, A.; Elias-Letts, R.; Guerra, H.; Gotuzzo, E.; Martínez-Carretero, E.; Piñero, J.E.; et al. The isolation of Balamuthia mandrillaris from environmental sources from Peru. Parasitol. Res. 2014, 113, 2509–2513. [Google Scholar] [CrossRef]
- Diaz, J.H. Behavioral and recreational risk factors for free-living amebic infections. J. Travel Med. 2011, 18, 130–137. [Google Scholar] [CrossRef][Green Version]
- Greub, G.; Raoult, D. Microorganisms resistant to free-living amoebae. Clin. Microbiol. Rev. 2004, 17, 413–433. [Google Scholar] [CrossRef]
- LeChevallier, M.W.; Prosser, T.; Stevens, M. Opportunistic Pathogens in Drinking Water Distribution Systems—A Review. Microorganisms 2024, 12, 916. [Google Scholar] [CrossRef]
- Pérez-Pérez, P.; Rodríguez-Escolar, I.; Córdoba-Lanús, E.; Domínguez-de-Barros, A.; García-Pérez, O.; Piñero, J.E.; Morchón, R.; Lorenzo-Morales, J. Assessment of the Presence of Free-Living Amoebae in Soil Samples from the Northwest Region of Spain Using Culture and Molecular Assays. Microorganisms 2025, 13, 1065. [Google Scholar] [CrossRef]
- Reyes-Batlle, M.; Díaz, F.J.; Sifaoui, I.; Rodríguez-Expósito, R.; Rizo-Liendo, A.; Piñero, J.E.; Lorenzo-Morales, J. Free living amoebae isolation in irrigation waters and soils of an insular arid agroecosystem. Sci. Total Environ. 2021, 753, 141833. [Google Scholar] [CrossRef] [PubMed]
- Page, F.C. A New Key to Freshwater and Soil Gymnamoebae: With Instructions for Culture; Freshwater Biological Association: Newby Bridge, UK, 1988. [Google Scholar]
- Córdoba-Lanús, E.; Reyes-Batlle, M.; Domínguez-de-Barros, A.; Pérez-Pérez, P.; Rodríguez-Expósito, R.L.; García-Ramos, A.; Sifaoui, I.; García-Pérez, O.; Aneiros-Giraldez, G.; Piñero, J.E.; et al. Multiplex Real-Time PCR assay to detect Acanthamoeba spp., Vermamoeba vermiformis, Naegleria fowleri, and Balamuthia mandrillaris in different water sources. Am. J. Trop. Med. Hyg. 2024, 111, 785–790. [Google Scholar] [CrossRef] [PubMed]
- Kuiper, M.W.; Valster, R.M.; Wullings, B.A.; Boonstra, H.; Smidt, H.; van der Kooij, D. Quantitative detection of the free-living amoeba Hartmannella vermiformis in surface water by using real-time PCR. Appl. Environ. Microbiol. 2006, 72, 3430–3435. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, J.M.; Booton, G.C.; Hay, J.; Niszl, I.A.; Seal, D.V.; Markus, M.B.; Fuerst, P.A.; Byers, T.J. Use of subgenic 18S rDNA PCR and sequencing for genus and genotype identification of Acanthamoebae from humans with keratitis and from sewage sludge. J. Clin. Microbiol. 2001, 39, 1903–1911. [Google Scholar] [CrossRef]
- De Jonckheere, J.F.; Brown, S. The identification of vahlkampfiid amoebae by ITS sequencing. Protist 2005, 156, 89–96. [Google Scholar] [CrossRef]
- Tsvetkova, N.; Schild, M.; Panaiotov, S.; Kurdova-Mintcheva, R.; Gottstein, B.; Walochnik, J.; Aspöck, H.; Lucas, M.S.; Müller, N. The identification of free-living environmental isolates of amoebae from Bulgaria. Parasitol. Res. 2004, 92, 405–413. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Latifi, A.; Salami, M.; Kazemirad, E.; Soleimani, M. Isolation and identification of free-living amoeba from the hot springs and beaches of the Caspian Sea. Parasite Epidemiol. Control 2020, 10, e00151. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.R-project.org/ (accessed on 1 September 2025).
- Smirnov, A.V.; Chao, E.; Nassonova, E.S.; Cavalier-Smith, T. A Revised Classification of Naked Lobose Amoebae (Amoebozoa: Lobosa). Protist 2011, 162, 545–570. [Google Scholar] [CrossRef]
- Page, F.C.; Blanton, R.L. The Heterolobosea (Sarcodina: Rhizopoda), a new class uniting the Schizopyrenida and the Acrasidae (Acrasida). Protistologica 1985, 21, 121. [Google Scholar]
- Johan, F. A Century of Research on the Amoeboflagellate Genus Naegleria. Acta Protozool. 2002, 41, 309–342. [Google Scholar]
- Öztoprak, H.; Walden, S.; Heger, T.; Bonkowski, M.; Dumack, K. What Drives the Diversity of the Most Abundant Terrestrial Cercozoan Family (Rhogostomidae, Cercozoa, Rhizaria)? Microorganisms 2020, 8, 1123. [Google Scholar] [CrossRef]
- Reyes-Batlle, M.; Hernández-Piñero, I.; Rizo-Liendo, A.; López-Arencibia, A.; Sifaoui, I.; Bethencourt-Estrella, C.J.; Chiboub, O.; Valladares, B.; Piñero, J.E.; Lorenzo-Morales, J. Isolation and molecular identification of free-living amoebae from dishcloths in Tenerife, Canary Islands, Spain. Parasitol. Res. 2019, 118, 927–933. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, X.; Wei, Z.; Cao, K.; Zhang, Z.; Liang, Q. The global epidemiology and clinical diagnosis of Acanthamoeba keratitis. J. Infect. Public Health 2023, 16, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, P.; Degorge, S.; Saint-Jean, C.; Yera, H.; Zekhnini, F.; Batellier, L.; Laroche, L.; Chaumeil, C. Resistance of Acanthamoeba to classic DNA extraction methods used for the diagnosis of corneal infections. Br. J. Ophthalmol. 2008, 92, 112–115. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Lim, H.J.; Son, A. Discrepancies in qPCR-based gene quantification and their dependencies on soil properties, inhibitor presence, and DNA extraction kit types. RSC Adv. 2025, 15, 19656–19664. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Thomas, V.; Loret, J.F.; Jousset, M.; Greub, G. Free-living amoebae and their intracellular pathogenic microorganisms: Risks for water quality. FEMS Microbiol. Rev. 2010, 34, 231–259. [Google Scholar] [CrossRef]
- Anwar, A.; Khan, N.A.; Siddiqui, R. Combating Acanthamoeba spp. cysts: What are the options? Parasites Vectors 2018, 11, 26. [Google Scholar] [CrossRef]
- König, S.; Priglinger, S.; Schaumberger, M.; Messmer, E.M. Tear Film Osmolarity in Normal Individuals: Comparison of Two Osmometers. Klin. Monatsblatter Fur Augenheilkd. 2020, 237, 649–654. [Google Scholar] [CrossRef]
- Arnalich-Montiel, F.; Lorenzo-Morales, J.; Irigoyen, C.; Morcillo-Laiz, R.; López-Vélez, R.; Muñoz-Negrete, F.; Piñero, J.E.; Valladares, B. Co-isolation of Vahlkampfia and Acanthamoeba in Acanthamoeba-like keratitis in a Spanish population. Cornea 2013, 32, 608–614. [Google Scholar] [CrossRef]
- Delumeau, A.; Quétel, I.; Harnais, F.; Sellin, A.; Gros, O.; Talarmin, A.; Marcelino, I. Bacterial microbiota management in free-living amoebae (Heterolobosea lineage) isolated from water: The impact of amoebae identity, grazing conditions, and passage number. Sci. Total Environ. 2023, 900, 165816. [Google Scholar] [CrossRef] [PubMed]
- Dumack, K.; Flues, S.; Hermanns, K.; Bonkowski, M. Rhogostomidae (Cercozoa) from soils, roots and plant leaves (Arabidopsis thaliana): Description of Rhogostoma epiphylla sp. nov. and R. cylindrica sp. nov. Eur. J. Protistol. 2017, 60, 76–86. [Google Scholar] [CrossRef] [PubMed]
- Pohl, N.; Solbach, M.D.; Dumack, K. The wastewater protist Rhogostoma minus (Thecofilosea, Rhizaria) is abundant, widespread, and hosts Legionellales. Water Res. 2021, 203, 117566. [Google Scholar] [CrossRef]
- Smirnov, A.V.; Nassonova, E.S.; Chao, E.; Cavalier-Smith, T. Phylogeny, Evolution, and Taxonomy of Vannellid Amoebae. Protist 2007, 158, 295–324. [Google Scholar] [CrossRef]
- Jensen, H.M.; Karami, A.M.; Mathiessen, H.; Al-Jubury, A.; Kania, P.W.; Buchmann, K. Gill amoebae from freshwater rainbow trout (Oncorhynchus mykiss): In vitro evaluation of antiparasitic compounds against Vannella sp. J. Fish Dis. 2020, 43, 665–672. [Google Scholar] [CrossRef] [PubMed]
- Robino, E.; Poirier, A.C.; Amraoui, H.; Le Bissonnais, S.; Perret, A.; Lopez-Joven, C.; Auguet, J.; Rubio, T.P.; Cazevieille, C.; Rolland, J.; et al. Resistance of the oyster pathogen Vibrio tasmaniensis LGP32 against grazing by Vannella sp. marine amoeba involves Vsm and CopA virulence factors. Environ. Microbiol. 2020, 22, 4183–4197. [Google Scholar] [CrossRef]
- Free Living Amebic Infections; Centers for Disease Control and Prevention (CDC) DPDx: Atlanta, GA, USA, 2024. Available online: https://www.cdc.gov/dpdx/freelivingamebic/index.html (accessed on 1 September 2025).









| FLA | Forward | Reverse |
|---|---|---|
| V. vermiformis HV1227F/VERM-RV [23,24] | 5′-TTACGAGGTCAGGACACTGT-3′ | 5′-TGCCTCAAACTTCCATTCGC-3 |
| Acanthamoeba spp. JDP1/JDP2 [25] | 5′-GGCCCAGATCGTTTACCGTGAA-3′ | 5′-TCTCACAAGCTGCTAGGGGAGTCA-3′ |
| Vahlkampfiidae VAHL1/VAHL2 [26] | 5′-CTCTTCGTAGGTGAACCTGC-3′ | 5′-CCGCTTACTGATATGCTTAA-3 |
| Universal for FLA FLA-F/FLA-R [27] | 5′-CGCGGTAATTCCAGCTCCAATAGC-3′ | 5′-CAGGTTAAGGTCTCGTTCGTTAAC-3′ |
| Primers | PCR Conditions | ||||
|---|---|---|---|---|---|
| Initial Denaturation | Denaturation | Annealing | Extension | Final Elongation | |
| V. vermiformis Hv1227F [24]/VermR [23] | 95 °C—3′ | 95 °C—20″ | 58 °C—30″ | 72 °C—40″ | 72 °C—10′ |
| 40 Cycles | |||||
| Acanthamoeba JDP1/JDP2 [25] | 95 °C—5′ | 95 °C—30″ | 50 °C—30″ | 72 °C—30″ | 72 °C—7′ |
| 35 Cycles | |||||
| Vahlkampfiidae VAHL1/VAHL2 [26] | 95 °C—2′ | 95 °C—1′ | 55 °C—1′30″ | 72 °C—2′ | 72 °C—7′ |
| 35 Cycles | |||||
| Universal for FLA FLAf/FLAr [27] | 95 °C—2′ | 95 °C—30″ | 58 °C—30″ | 72 °C—30″ | 72 °C—7′ |
| 40 cycles | |||||
| Score | Tolerance Levels |
|---|---|
| 0 | All amoebae are in the cyst stage, and none are outside the initial inoculation zone. |
| 1 | Some amoebae are in trophozoite stage and there are amoebae in any stage surrounding the initial inoculation area. |
| 2 | A considerable increase in the number of trophozoites, with slight movement across the agar plate. |
| 3 | A massive increase in the number of amoebae and widespread extension across the agar plate. |
| Sample Type | AS | PS | RW | Total | |
|---|---|---|---|---|---|
| Total samples (N) | 25 | 20 | 17 | 62 | |
| qPCR | Acanthamoeba (n/%) | 14/56% | 12/60% | 8/47.1% | 34/54.8% |
| V. vermiformis (n/%) | 12/48% | 17/85% | 8/47.1% | 37/59.7% | |
| B. mandrillaris (n/%) | 2/8% | 4/20% | 0 | 6/9.7% | |
| CT Mean value | Acanthamoeba | 33.32 (IQR: 2.3 Ct) | 32.02 (IQR: 1.7 Ct) | 33.36 (IQR: 2.9 Ct) | 32.74 (IQR: 3.1 Ct) |
| V. vermiformis | 33.52 (IQR: 2.5 Ct) | 30.88 (IQR: 2.2 Ct) | 33.69 (IQR: 3.4 Ct) | 32.43 (IQR: 3.9 Ct) | |
| B. mandrillaris | 33 (IQR: 4 Ct) | 33.43 (IQR: 3.8 Ct) | - | 33.21 (IQR: 2.9 Ct) | |
| Isolation/Culture | Monoxenic cultures | 27 | 17 | 7 | 51 |
| Acanthamoeba spp. (n/%) | 11/40.7% | 4/23.5% | 3/42.9% | 18/35.3% | |
| V. vermiformis (n/%) | 8/29.6% | 7/41.2% | 2/28.6% | 17/33.3% | |
| Vahlkampfiidae (n/%) | 7/25.9% | 2/11.8% | 2/28.6% | 11/21.6% | |
| Vannellidae (n/%) | 1/3.7% | 3/17.6% | 0 | 4/7.8% | |
| Rhogostoma sp. (n/%) | 0 | 1/5.9% | 0 | 1/2% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Peña-Prunell, M.D.; Reyes-Batlle, M.; Pérez-Pérez, P.; Rodríguez-Expósito, R.L.; Sifaoui, I.; García-Pérez, O.; Domínguez-de Barros, A.T.; Córdoba-Lanús, E.; Piñero, J.E.; Lorenzo-Morales, J. Molecular and Culture-Based Surveillance of Free-Living Amoebae in Human Related Sources in an Outermost Region. Pathogens 2026, 15, 73. https://doi.org/10.3390/pathogens15010073
Peña-Prunell MD, Reyes-Batlle M, Pérez-Pérez P, Rodríguez-Expósito RL, Sifaoui I, García-Pérez O, Domínguez-de Barros AT, Córdoba-Lanús E, Piñero JE, Lorenzo-Morales J. Molecular and Culture-Based Surveillance of Free-Living Amoebae in Human Related Sources in an Outermost Region. Pathogens. 2026; 15(1):73. https://doi.org/10.3390/pathogens15010073
Chicago/Turabian StylePeña-Prunell, Marco D., María Reyes-Batlle, Patricia Pérez-Pérez, Rubén L. Rodríguez-Expósito, Ines Sifaoui, Omar García-Pérez, Angélica T. Domínguez-de Barros, Elizabeth Córdoba-Lanús, José E. Piñero, and Jacob Lorenzo-Morales. 2026. "Molecular and Culture-Based Surveillance of Free-Living Amoebae in Human Related Sources in an Outermost Region" Pathogens 15, no. 1: 73. https://doi.org/10.3390/pathogens15010073
APA StylePeña-Prunell, M. D., Reyes-Batlle, M., Pérez-Pérez, P., Rodríguez-Expósito, R. L., Sifaoui, I., García-Pérez, O., Domínguez-de Barros, A. T., Córdoba-Lanús, E., Piñero, J. E., & Lorenzo-Morales, J. (2026). Molecular and Culture-Based Surveillance of Free-Living Amoebae in Human Related Sources in an Outermost Region. Pathogens, 15(1), 73. https://doi.org/10.3390/pathogens15010073










