Evaluating the Performance of Loop-Mediated Isothermal Amplification for the Detection of Listeria monocytogenes Biofilms on Stainless Steel Surfaces
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Inoculum Preparation
2.2. Biofilm Formation
2.3. Bacterial Recovery
2.4. Sample Processing and DNA Extraction
2.5. Molecular Detection
2.5.1. Real-Time LAMP
2.5.2. Colorimetric LAMP
2.5.3. qPCR
2.6. Confocal Microscopy
2.7. Data Representation and Statistical Analyses
3. Results
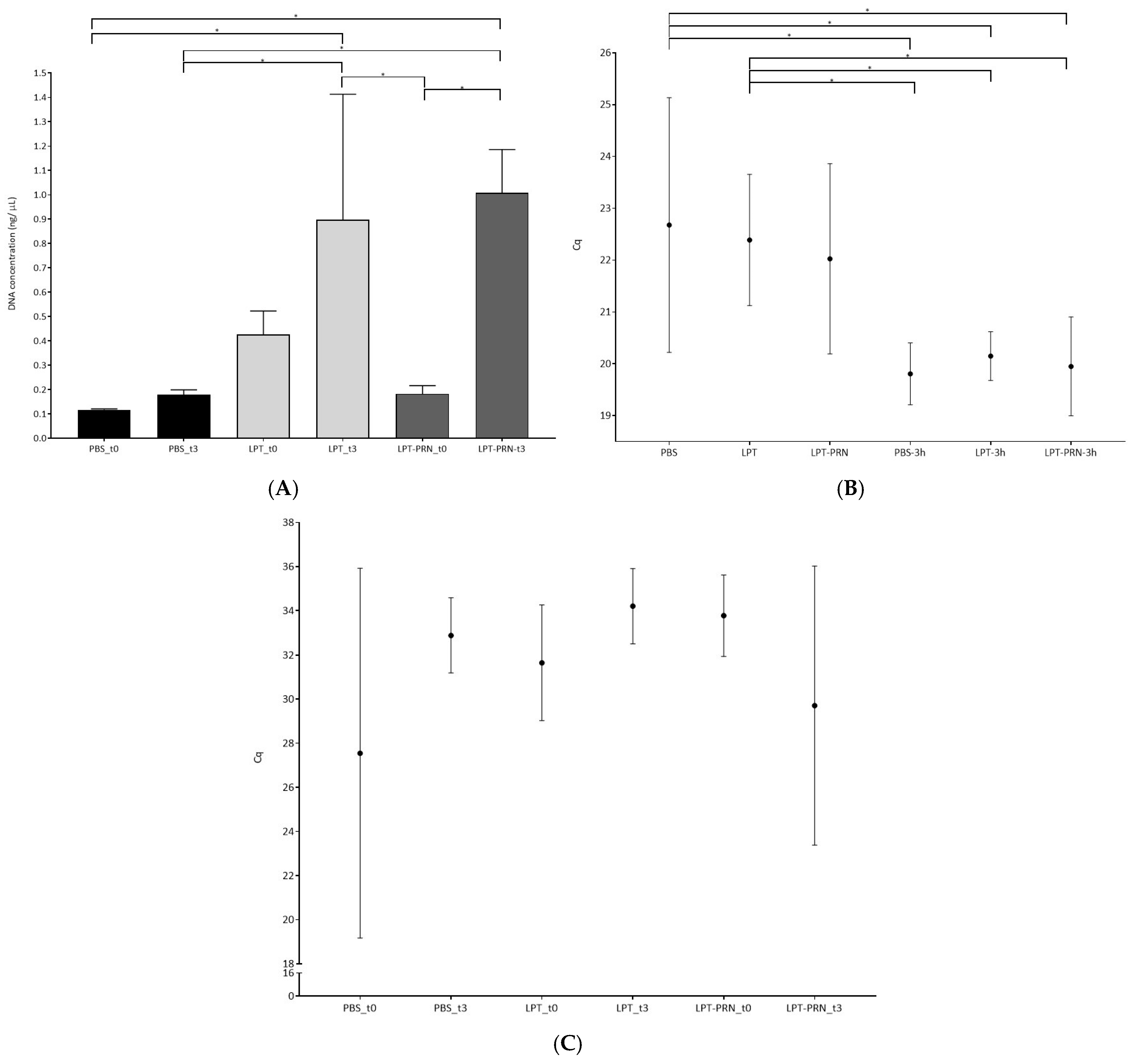
3.1. Screening of L. monocytogenes Recovery Buffers on Monospecies Biofilms
3.2. Screening of L. monocytogenes Recovery Buffers on Multispecies Biofilms
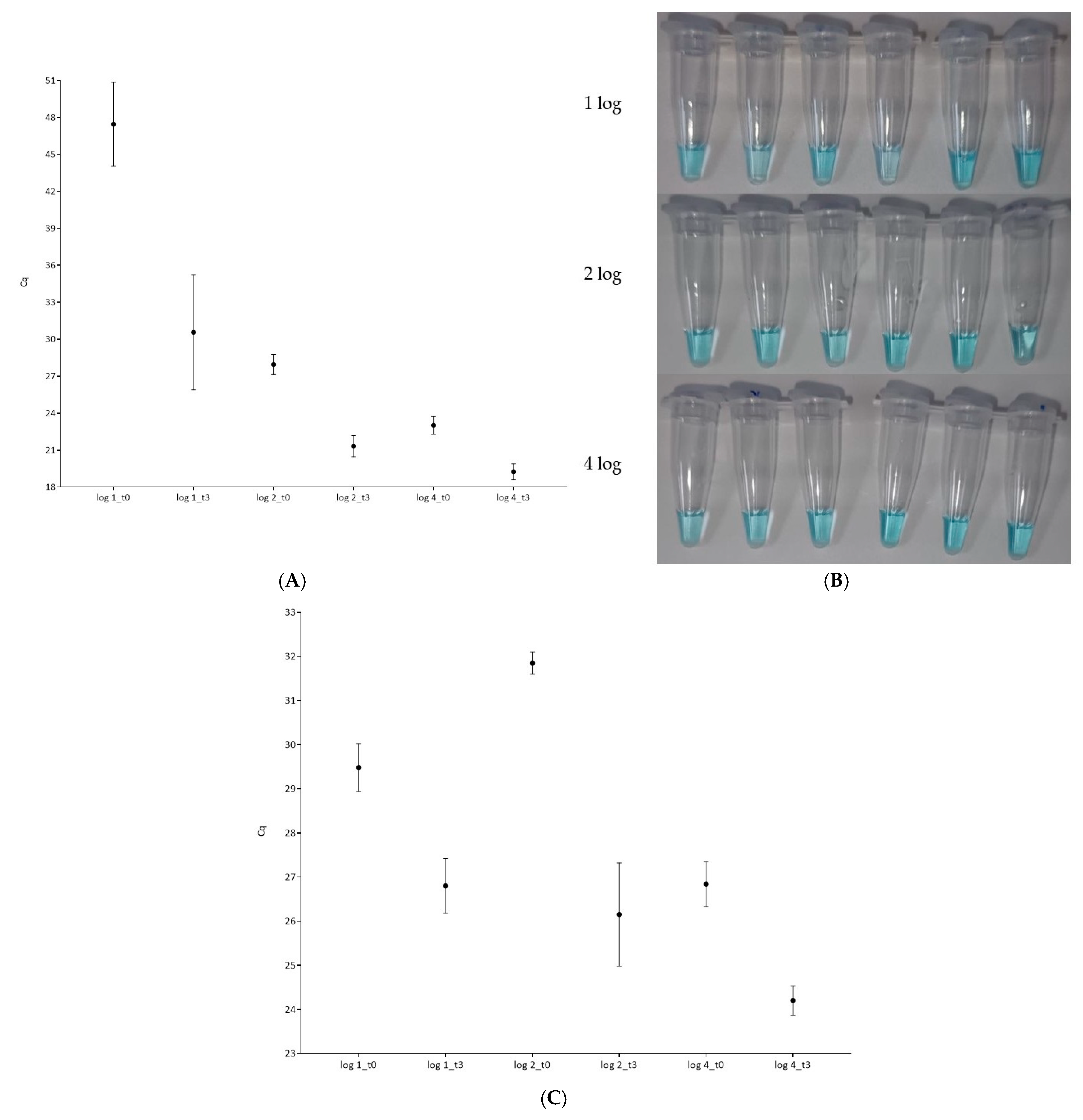
3.3. Effect of Decreasing L. monocytogenes Concentration in Mixed Biofilms
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lomonaco, S.; Nucera, D.; Filipello, V. The Evolution and Epidemiology of Listeria monocytogenes in Europe and the United States. Infect. Genet. Evol. 2015, 35, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Qiao, J.; Lu, Z.; Fen, Z.; Tao, Y.; Lv, F.; Zhao, H.; Zhang, C.; Bie, X. Influence of Different Factors on Biofilm Formation of Listeria monocytogenes and the Regulation of cheY Gene. Food Res. Int. 2020, 137, 109405. [Google Scholar] [CrossRef] [PubMed]
- Warriner, K.; Namvar, A. What Is the Hysteria with Listeria? Trends Food Sci. Technol. 2009, 20, 245–254. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA). European Centre for Disease Prevention and Control (ECDC). In The European Union One Health 2023 Zoonoses Report; John Wiley and Sons Inc.: Hoboken, NJ, USA, 2024; Volume 22. [Google Scholar]
- Brehm-Stecher, B.; Young, C.; Jaykus, L.A.; Mary Lou, T. Sample Preparation: The Forgotten Beginning. J. Food Prot. 2009, 72, 1774–1789. [Google Scholar] [CrossRef]
- Spanu, C.; Jordan, K. Listeria monocytogenes Environmental Sampling Program in Ready-to-Eat Processing Facilities: A Practical Approach. Compr. Rev. Food Sci. Food Saf. 2020, 19, 2843–2861. [Google Scholar] [CrossRef]
- Rawlinson, S.; Ciric, L.; Cloutman-Green, E. How to Carry out Microbiological Sampling of Healthcare Environment Surfaces? A Review of Current Evidence. J. Hosp. Infect. 2019, 103, 363–374. [Google Scholar] [CrossRef]
- Manso, B.; Melero, B.; Stessl, B.; Fernández-Natal, I.; Jaime, I.; Hernández, M.; Wagner, M.; Rovira, J.; Rodríguez-Lázaro, D. Characterization of Virulence and Persistence Abilities of Listeria monocytogenes Strains Isolated from Food Processing Premises. J. Food Prot. 2019, 82, 1922–1930. [Google Scholar] [CrossRef] [PubMed]
- Melero, B.; Stessl, B.; Manso, B.; Wagner, M.; Esteban-Carbonero, Ó.J.; Hernández, M.; Rovira, J.; Rodriguez-Lázaro, D. Listeria monocytogenes Colonization in a Newly Established Dairy Processing Facility. Int. J. Food Microbiol. 2019, 289, 64–71. [Google Scholar] [CrossRef]
- Melero, B.; Manso, B.; Stessl, B.; Hernández, M.; Wagner, M.; Rovira, J.; Rodríguez-Lázaro, D. Distribution and Persistence of Listeria monocytogenes in a Heavily Contaminated Poultry Processing Facility. J. Food Prot. 2019, 82, 1524–1531. [Google Scholar] [CrossRef]
- Ceuppens, S.; Li, D.; Uyttendaele, M.; Renault, P.; Ross, P.; Ranst, M.V.; Cocolin, L.; Donaghy, J. Molecular Methods in Food Safety Microbiology: Interpretation and Implications of Nucleic Acid Detection. Compr. Rev. Food Sci. Food Saf. 2014, 13, 551–577. [Google Scholar] [CrossRef]
- Umesha, S.; Manukumar, H.M. Advanced Molecular Diagnostic Techniques for Detection of Food-Borne Pathogens: Current Applications and Future Challenges. Crit. Rev. Food Sci. Nutr. 2018, 58, 84–104. [Google Scholar] [CrossRef]
- Hameed, S.; Xie, L.; Ying, Y. Conventional and Emerging Detection Techniques for Pathogenic Bacteria in Food Science: A Review. Trends Food Sci. Technol. 2018, 81, 61–73. [Google Scholar] [CrossRef]
- Mangal, M.; Bansal, S.; Sharma, S.K.; Gupta, R.K. Molecular Detection of Foodborne Pathogens: A Rapid and Accurate Answer to Food Safety. Crit. Rev. Food Sci. Nutr. 2016, 56, 1568–1584. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, X.; Shi, X.; Shi, H.; Wang, Z.; Peng, C. Review in Isothermal Amplification Technology in Food Microbiological Detection. Food Sci. Biotechnol. 2022, 31, 1501–1511. [Google Scholar] [CrossRef] [PubMed]
- Garrido-Maestu, A.; Prado, M. Naked-Eye Detection Strategies Coupled with Isothermal Nucleic Acid Amplification Techniques for the Detection of Human Pathogens. Compr. Rev. Food Sci. Food Saf. 2022, 21, 1913–1939. [Google Scholar] [CrossRef] [PubMed]
- Abalo, M.; Lamas, A.; Teixeira, C.; Prado, M.; Garrido-Maestu, A. Surface Monitoring of L. monocytogenes by Real-Time Fluorescence and Colorimetric LAMP. Appl. Microbiol. Biotechnol. 2024, 108, 510. [Google Scholar] [CrossRef]
- Kim, U.; Kim, J.H.; Oh, S.W. Review of Multi-Species Biofilm Formation from Foodborne Pathogens: Multi-Species Biofilms and Removal Methodology. Crit. Rev. Food Sci. Nutr. 2022, 62, 5783–5793. [Google Scholar] [CrossRef]
- Norwood, D.; Gilmour, A. The Differential Adherence Capabilities of Two Listeria monocytogenes Strains in Monoculture and Multispecies Biofilms as a Function of Temperature. Lett. Appl. Microbiol. 2001, 33, 320–324. [Google Scholar] [CrossRef]
- Rodríguez-López, P.; Puga, C.H.; Orgaz, B.; Cabo, M.L. Quantifying the Combined Effects of Pronase and Benzalkonium Chloride in Removing Late-Stage Listeria monocytogenes–Escherichia coli Dual-Species Biofilms. Biofouling 2017, 33, 690–702. [Google Scholar] [CrossRef]
- Puga, C.H.; Rodríguez-López, P.; Cabo, M.L.; SanJose, C.; Orgaz, B. Enzymatic Dispersal of Dual-Species Biofilms Carrying Listeria monocytogenes and Other Associated Food Industry Bacteria. Food Control 2018, 94, 222–228. [Google Scholar] [CrossRef]
- Sweeney, P.J.; Walker, J.M. Pronase (EC 3.4.24.4). In Enzymes of Molecular Biology; Burrell, M.M., Ed.; Humana Press: Totowa, NJ, USA, 1993; pp. 271–276. ISBN 978-1-59259-503-7. [Google Scholar]
- Kim, J.S.; Lim, M.C.; Kim, S.M.; Lee, J.Y. Extracellular Matrix-Degrading Enzymes as a Biofilm Control Strategy for Food-Related Microorganisms. Food Sci. Biotechnol. 2023, 32, 1745–1761. [Google Scholar] [CrossRef]
- Nogueira, R.; Cabo, M.L.; García-Sanmartín, L.; Sánchez-Ruiloba, L.; Rodríguez-Herrera, J.J. Risk Factor-Based Clustering of Listeria monocytogenes in Food Processing Environments Using Principal Component Analysis. Food Res. Int. 2023, 170, 112989. [Google Scholar] [CrossRef]
- Rodríguez-López, P.; Cabo, M.L. Tolerance Development in Listeria monocytogenes-Escherichia coli Dual-Species Biofilms after Sublethal Exposures to Pronase-Benzalkonium Chloride Combined Treatments. Food Microbiol. 2017, 67, 58–66. [Google Scholar] [CrossRef]
- Roumani, F.; Azinheiro, S.; Carvalho, J.; Prado, M.; Garrido-Maestu, A. Loop-Mediated Isothermal Amplification Combined with Immunomagnetic Separation and Propidium Monoazide for the Specific Detection of Viable Listeria monocytogenes in Milk Products, with an Internal Amplification Control. Food Control 2021, 125, 107975. [Google Scholar] [CrossRef]
- Garrido-Maestu, A.; Azinheiro, S.; Carvalho, J.; Fuciños, P.; Prado, M. Development and Evaluation of Loop-Mediated Isothermal Amplification, and Recombinase Polymerase Amplification Methodologies, for the Detection of Listeria monocytogenes in Ready-to-Eat Food Samples. Food Control 2018, 86, 27–34. [Google Scholar] [CrossRef]
- Garrido-Maestu, A.; Fuciños, P.; Azinheiro, S.; Carvalho, J.; Prado, M. Systematic Loop-Mediated Isothermal Amplification Assays for Rapid Detection and Characterization of Salmonella spp., Enteritidis and Typhimurium in Food Samples. Food Control 2017, 80, 297–306. [Google Scholar] [CrossRef]
- Azinheiro, S.; Rodríguez-López, P.; Lozano-León, A.; Guedes, H.; Regal, P.; Franco, C.M.; Cepeda, A.; Teixeira, P.; Melo, L.D.R.R.; Silva, D.; et al. Interlaboratory Validation of a Multiplex QPCR Method for the Detection of Listeria monocytogenes in a Ready-to-Eat Seafood Product. Food Control 2023, 150, 109769. [Google Scholar] [CrossRef]
- Manyi-Loh, C.E.; Lues, R. Listeria monocytogenes and Listeriosis: The Global Enigma. Foods 2025, 14, 1266. [Google Scholar] [CrossRef]
- Dygico, L.K.; Gahan, C.G.M.; Grogan, H.; Burgess, C.M. The Ability of Listeria monocytogenes to Form Biofilm on Surfaces Relevant to the Mushroom Production Environment. Int. J. Food Microbiol. 2020, 317, 108385. [Google Scholar] [CrossRef]
- Dey, B.P.; Engley, F.B. Comparison of Dey and Engley (D/E) Neutralizing Medium to Letheen Medium and Standard Methods Medium for Recovery of Staphylococcus Aureus from Sanitized Surfaces. J. Ind. Microbiol. 1995, 14, 21–25. [Google Scholar] [CrossRef]
- Dey, B.P.; Engley, F.B. Neutralization of Antimicrobial Chemicals by Recovery Media. J. Microbiol. Methods 1994, 19, 51–58. [Google Scholar] [CrossRef]
- Colagiorgi, A.; Di Ciccio, P.; Zanardi, E.; Ghidini, S.; Ianieri, A. A Look inside the Listeria monocytogenes Biofilms Extracellular Matrix. Microorganisms 2016, 4, 22. [Google Scholar] [CrossRef]
- Burmølle, M.; Ren, D.; Bjarnsholt, T.; Sørensen, S.J. Interactions in Multispecies Biofilms: Do They Actually Matter? Trends Microbiol. 2014, 22, 84–91. [Google Scholar] [CrossRef]
- Yang, L.; Liu, Y.; Wu, H.; Høiby, N.; Molin, S.; Song, Z.J. Current Understanding of Multispecies Biofilms. Int. J. Oral Sci. 2011, 3, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Pant, K.; Palmer, J.; Flint, S. Evaluation of Single and Multispecies Biofilm Formed in the Static and Continuous Systems. Int. J. Food Microbiol. 2025, 429, 111030. [Google Scholar] [CrossRef] [PubMed]
- Broten, C.J.; Wydallis, J.B.; Iii, T.H.R.; Bisha, B. Development and Evaluation of a Paper-Based Microfluidic Device for Detection of Listeria monocytogenes on Food Contact and Non-Food Contact Surfaces. Foods 2022, 11, 947. [Google Scholar] [CrossRef]
- Ríos-Castillo, A.G.; Ripolles-Avila, C.; Rodríguez-Jerez, J. Detection by Real-Time PCR and Conventional Culture of Salmonella Typhimurium and Listeria monocytogenes Adhered to Stainless Steel Surfaces under Dry Conditions. Food Control 2022, 137, 108971. [Google Scholar] [CrossRef]
- Fachmann, M.S.R.; Löfström, C.; Hoorfar, J.; Hansen, F.; Christensen, J.; Mansdal, S.; Josefsen, M.H.M.H.; Mandal, S.; Josefsen, M.H.M.H. Detection of Salmonella Enterica in Meat in Less than 5 Hours by a Low-Cost and Noncomplex Sample Preparation Method. Appl. Environ. Microbiol. 2017, 83, e03151-16. [Google Scholar] [CrossRef]
- Wang, Y.; Kang, Z.; Gao, H.; Gao, Y.; Qin, L.; Lin, H.; Yu, F.; Qi, X.; Wang, X. A One-Step Reverse Transcription Loop-Mediated Isothermal Amplification for Detection and Discrimination of Infectious Bursal Disease Virus. Virol. J. 2011, 8, 108. [Google Scholar] [CrossRef][Green Version]
- Nguyen, H.Q.; Nguyen, V.D.; Van Nguyen, H.; Seo, T.S. Quantification of Colorimetric Isothermal Amplification on the Smartphone and Its Open-Source App for Point-of-Care Pathogen Detection. Sci. Rep. 2020, 10, 15123. [Google Scholar] [CrossRef]
- Panich, W.; Puttharugsa, C.; Tejangkura, T.; Chontananarth, T. A Simple Color Absorption Analysis of Colorimetric Loop-Mediated Isothermal Amplification for Detection of Raillietina spp. in Clinical Samples Using a 3D-Printed Tube Holder Coupled with a Smartphone Camera and Notebook Screen. Microchim. Acta 2024, 191, 603. [Google Scholar] [CrossRef] [PubMed]


| Technique | Primer | Sequence (5′ → 3′) | Concentration (nM) | Reference |
|---|---|---|---|---|
| LAMP | F3 | TTC AAA AGC TTA TAC AGA TGG AA | 200 | [17] |
| B3 | AAG CTA AAC CAG TGC ATT C | 200 | ||
| FIP | TGA ACA ATT TCG TTA CCT TCA GGA T tttt TCG ATC ACT CTG GAG GAT AC | 1000 | ||
| BIP | GGA GCG AAA ACA ATA AAA GCA AGC T tttt GCG TAA ACA TTA ATA TTT CTC GC | 1000 | ||
| LF | CAT CCC AAG AAA TGT TGA ATT GAG C | 800 | ||
| LB | TCG TCC ATC TAT TTG CCA GGT A | 800 | ||
| qPCR * | hly-P3F | CGC AAC AAA CTG AAG CAA AGG A | 200 | [26] |
| hly-P3R | CGA TTG GCG TCT TAG GAC TTG C | 200 | ||
| hly-P3P | FAM—CAT GGC ACC/ZEN/ACC AGC ATC TCC G—IABkFQ | 150 | ||
| IAC-F | AGT TGC ACA CAG TTA GTT CGA G | 100 | [27] | |
| IAC-R | TGG AGT GCT GGA CGA TTT GAA G | 100 | ||
| IAC-P | YY—AGT GGC GGT/ZEN/GAC ACT GTT GAC CT—IABkFQ | 100 | [28] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Garrido-Pérez, C.P.; López-Cabo, M.; Garrido-Maestu, A. Evaluating the Performance of Loop-Mediated Isothermal Amplification for the Detection of Listeria monocytogenes Biofilms on Stainless Steel Surfaces. Pathogens 2026, 15, 49. https://doi.org/10.3390/pathogens15010049
Garrido-Pérez CP, López-Cabo M, Garrido-Maestu A. Evaluating the Performance of Loop-Mediated Isothermal Amplification for the Detection of Listeria monocytogenes Biofilms on Stainless Steel Surfaces. Pathogens. 2026; 15(1):49. https://doi.org/10.3390/pathogens15010049
Chicago/Turabian StyleGarrido-Pérez, Carmen Pilar, Marta López-Cabo, and Alejandro Garrido-Maestu. 2026. "Evaluating the Performance of Loop-Mediated Isothermal Amplification for the Detection of Listeria monocytogenes Biofilms on Stainless Steel Surfaces" Pathogens 15, no. 1: 49. https://doi.org/10.3390/pathogens15010049
APA StyleGarrido-Pérez, C. P., López-Cabo, M., & Garrido-Maestu, A. (2026). Evaluating the Performance of Loop-Mediated Isothermal Amplification for the Detection of Listeria monocytogenes Biofilms on Stainless Steel Surfaces. Pathogens, 15(1), 49. https://doi.org/10.3390/pathogens15010049







