Burkholderia pseudomallei in Sarawak, Malaysian Borneo, Remains Highly Susceptible to Trimethoprim-Sulfamethoxazole Despite Resistance to Its Individual Components
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Bacterial Strains
2.3. Antibiotic Susceptibility Testing (AST)
2.3.1. Kirby–Bauer Disk Diffusion Testing
2.3.2. E-Test
2.3.3. Broth Microdilution
2.4. Statistical Data Analysis
- κ = 0.00–0.20 indicates slight agreement
- κ = 0.21–0.40 indicates fair agreement
- κ = 0.41–0.60 indicates moderate agreement
- κ = 0.61–0.80 indicates substantial agreement
- κ = 0.81–1.00 indicates almost perfect agreement
2.5. Genomic Analysis
2.5.1. Primer Design
2.5.2. Polymerase Chain Reaction (PCR)
3. Results
3.1. Disk Diffusion Testing Revealed a High Prevalence of Trimethoprim-Sulfamethoxazole Susceptibility Among the Sarawak B. pseudomallei
3.2. E-Test Indicates Evidence of a Trimethoprim-Sulfamethoxazole False-Resistance Profile Among the Sarawak B. pseudomallei Clinical Isolates
3.3. Broth Microdilution Demonstrated the Resistance of Sarawak B. pseudomallei Clinical Isolates to Trimethoprim and Sulfamethoxazole Despite Trimethoprim-Sulfamethoxazole Susceptibility
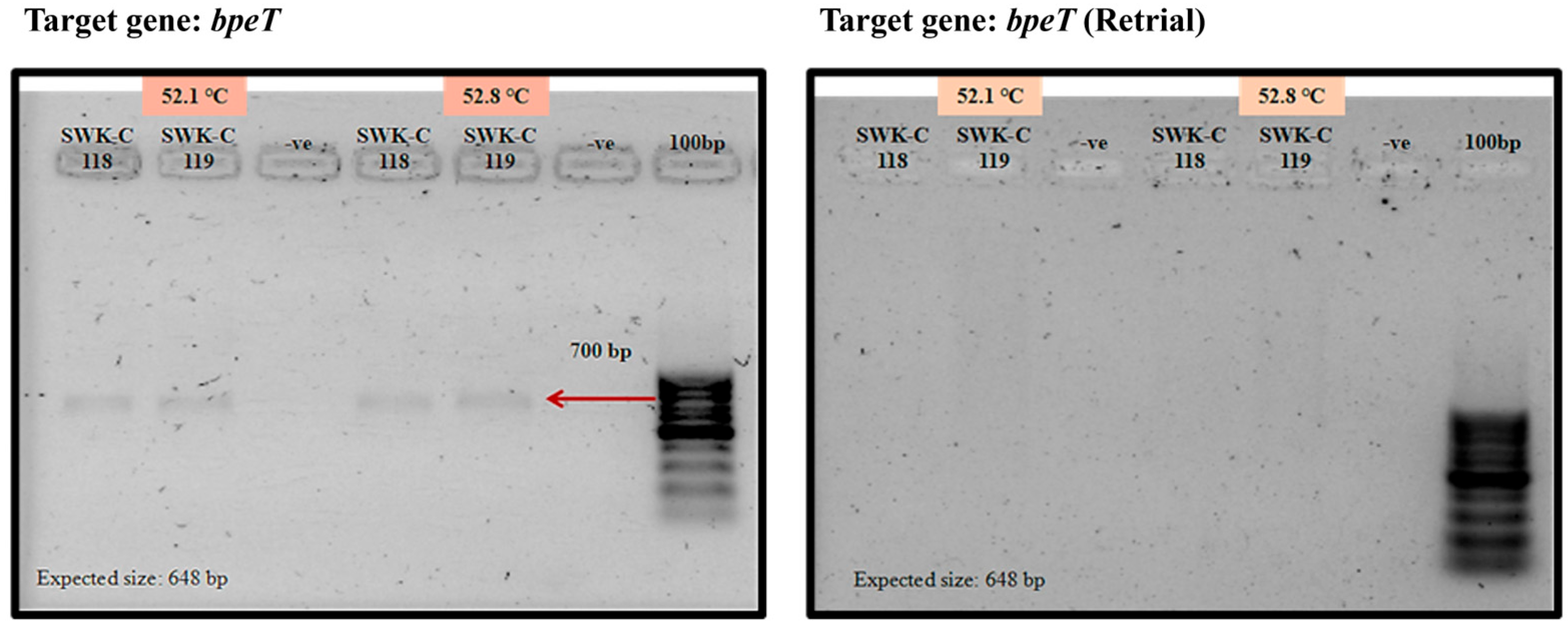
3.4. PCR Analysis Suggests Divergence in the bpeEF-oprC Efflux Pump Gene Cluster in Sarawak B. pseudomallei Clinical Isolates
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MIC | Minimal inhibitory concentration |
| CLSI | Clinical Laboratory Standards Institute |
| EUCAST | European Committee on Antimicrobial Susceptibility Testing |
| QC | Quality Control |
| IZ | Inhibition zone |
| AST | Antimicrobial Susceptibility Testing |
| MIC50 | Minimal inhibitory concentration at 50% inhibited growth |
| MIC80 | Minimal inhibitory concentration at 80% inhibited growth |
| OD | Optical density |
| GenS | Gentamicin-susceptible |
| TMP | Trimethoprim |
| SMX | Sulfamethoxazole |
| PCR | Polymerase Chain Reaction |
| ST | Sequence type |
Appendix A
| Isolates | Origin | MIC50, mm (for Type C IZ Only) | MIC80, mm (CLSI/EUCAST Interpretation) | Type of Inhibition Zone |
|---|---|---|---|---|
| SWK-C 101 | Bintulu | NA | 19 (S/S) | |
| SWK-C 103 1,3 | Bintulu | NA | 10 (R/R) | Type B 1 |
| SWK-C 105 | Bintulu | NA | 23 (S/S) | |
| SWK-C 106 1,3 | Bintulu | NA | 12 (I/R) | Type B 1 |
| SWK-C 107 | Bintulu | NA | 25 (S/S) | |
| SWK-C 108 1,3 | Bintulu | NA | 11 (I/R) | Type B 1 |
| SWK-C 109 1,3 | Bintulu | NA | 18 (S/R) | Type B 1 |
| SWK-C 111 | Bintulu | NA | 18 (S/S) | |
| SWK-C 114 | Bintulu | NA | 26 (S/S) | |
| SWK-C 115 | Bintulu | NA | 21 (S/S) | |
| SWK-C 118 1 | Bintulu | NA | 12 (I/R) | Type B 1 |
| SWK-C 119 1 | Bintulu | NA | 15 (I/R) | Type B 1 |
| SWK-C 120 | Bintulu | NA | 34 (S/S) | |
| SWK-C 123 1,3 | Bintulu | NA | 14 (I/R) | Type B 1 |
| SWK-C 143 | Bintulu | NA | 24 (S/S) | |
| SWK-C 145 1 | Bintulu | NA | 20 (S/S) | Type B 1 |
| SWK-C 146 (a) | Bintulu | NA | 19 (S/S) | |
| SWK-C 146 (b) | Bintulu | NA | 21 (S/S) | |
| SWK-C 146 (c) 1,3 | Bintulu | NA | 15 (I/R) | Type B 1 |
| SWK-C 147 | Bintulu | NA | 19 (S/S) | |
| SWK-C 148 | Bintulu | NA | 19 (S/S) | |
| SWK-C 149 (a) | Bintulu | NA | 18 (S/S) | |
| SWK-C 149 (b) 1 | Bintulu | NA | 16 (S/R) | Type B 1 |
| SWK-C 150 1,3 | Bintulu | NA | 15 (I/R) | Type B 1 |
| SWK-C 151 | Bintulu | NA | 20 (S/S) | |
| SWK-C 154 (b) 1,3 | Bintulu | NA | 12 (I/I) | Type B 1 |
| SWK-C 154 (c) 1 | Bintulu | NA | 14 (I/R) | Type B 1 |
| SWK-C 155 (a) 1 | Bintulu | NA | 12 (I/R) | Type B 1 |
| SWK-C 155 (b) 1 | Bintulu | NA | 18 (S/R) | Type B 1 |
| SWK-C 156 | Bintulu | NA | 20 (S/S) | |
| SWK-C 158 1,3 | Bintulu | NA | 12 (I/R) | Type B 1 |
| SWK-C 159 | Bintulu | NA | 17 (S/S) | |
| SWK-C 160 | Bintulu | NA | 37 (S/S) | |
| SWK-C 161 1 | Bintulu | NA | 37 (S/R) | Type B 1 |
| SWK-C 162 | Bintulu | NA | 25 (S/S) | |
| SWK-C 163 (a) 1,3 | Bintulu | NA | 15 (I/R) | Type B 1 |
| SWK-C 163 (b) 1,3 | Bintulu | NA | 10 (R/R) | Type B 1 |
| SWK-C 164 | Bintulu | NA | 20 (S/S) | |
| SWK-C 167 | Bintulu | NA | 21 (S/S) | |
| SWK-C 172 | Bintulu | NA | 21 (S/S) | |
| SWK-C 175 | Bintulu | NA | 21 (S/S) | |
| SWK-C 177 3 | Bintulu | NA | 15 (I/R) | |
| SWK-C 180 | Bintulu | NA | 25 (S/S) | |
| SWK-C 182 | Bintulu | NA | 23 (S/S) | |
| SWK-C 184 | Bintulu | NA | 24 (S/S) | |
| SWK-C 185 | Bintulu | NA | 24 (S/S) | |
| MSHR 7905 | Bintulu | NA | 22 (S/S) | |
| MSHR 7895 | Bintulu | NA | 28 (S/S) | |
| MSHR 7903 | Bintulu | NA | 21 (S/S) | |
| MSHR 7881 | Bintulu | NA | 17 (S/S) | |
| MSHR 7882 | Bintulu | NA | 20 (S/S) | |
| MSHR 7906 | Bintulu | NA | 20 (S/S) | |
| MSHR 7887 3 | Bintulu | NA | 22 (S/S) | |
| MSHR 7888 | Bintulu | NA | 25 (S/S) | |
| SWK-C 192 | Bintulu | NA | 20 (S/S) | |
| SWK-C 193 | Bintulu | NA | 30 (S/S) | |
| SWK-C 194 1 | Bintulu | NA | 15 (I/R) | Type B 1 |
| SWK-C 195 | Bintulu | NA | 21 (S/S) | |
| SWK-C 196 | Bintulu | NA | 20 (S/S) | |
| SWK-C 197 | Bintulu | NA | 28 (S/S) | |
| SWK-C 199 1 | Bintulu | NA | 16 (S/R) | Type B 1 |
| SWK-C 243 | Bintulu | NA | 16 (S/S) | |
| SWK-C 267 | Bintulu | NA | 12 (I/I) | |
| SWK-C 269 | Bintulu | NA | 13 (I/I) | |
| SWK-C 270 | Bintulu | NA | 20 (S/S) | |
| SWK-C 271 | Bintulu | NA | 16 (S/S) | |
| SWK-C 290 | Bintulu | NA | 15 (I/S) | |
| SWK-C 124 | Kapit | NA | 19 (S/S) | |
| SWK-C 125 | Kapit | NA | 24 (S/S) | |
| SWK-C 126 | Kapit | NA | 22 (S/S) | |
| SWK-C 127 | Kapit | NA | 19 (S/S) | |
| SWK-C 130 | Kapit | NA | 22 (S/S) | |
| SWK-C 131 1 | Kapit | NA | 19 (S/S) | Type A 1 |
| SWK-C 132 1 | Kapit | NA | 20 (S/S) | Type A 1 |
| SWK-C 133 1 | Kapit | NA | 21 (S/S) | Type A 1 |
| SWK-C 134 1 | Kapit | NA | 19 (S/S) | Type A 1 |
| SWK-C 136 1 | Kapit | NA | 21 (S/S) | Type A 1 |
| SWK-C 138 1 | Kapit | NA | 16 (S/S) | Type A 1 |
| SWK-C 139 1 | Kapit | NA | 15 (R/S) | Type B 1 |
| SWK-C 140 1 | Kapit | NA | 15 (I/R) | Type B 1 |
| SWK-C 141 | Kapit | NA | 24 (S/S) | |
| SWK-C 214 | Kapit | NA | 22 (S/S) | |
| SWK-C 215 | Kapit | NA | 24 (S/S) | |
| SWK-C 216 | Kapit | NA | 22 (S/S) | |
| SWK-C 218 | Kapit | NA | 19 (S/S) | |
| SWK-C 220 | Kapit | NA | 20 (S/S) | |
| SWK-C 221 1 | Kapit | NA | 20 (S/R) | Type B 1 |
| SWK-C 222 | Kapit | NA | 19 (S/S) | |
| SWK-C 223 1 | Kapit | NA | 12 (I/I) | Type B 1 |
| SWK-C 226 | Kapit | NA | 16 (S/S) | |
| SWK-C 227 | Kapit | NA | 26 (S/S) | |
| SWK-C 229 | Kapit | NA | 30 (S/S) | |
| SWK-C 230 | Kapit | NA | 26 (S/S) | |
| SWK-C 231 | Kapit | NA | 19 (S/S) | |
| SWK-C 233 1 | Kapit | NA | 15 (I/R) 1 | Type B 1 |
| SWK-C 234 | Kapit | NA | 20 (S/S) | |
| SWK-C 235 | Kapit | NA | 30 (S/S) | |
| SWK-C 236 | Kapit | NA | 19 (S/S) | |
| SWK-C 237 | Kapit | NA | 29 (S/S) | |
| SWK-C 238 | Kapit | NA | 21 (S/S) | |
| SWK-C 241 | Kapit | NA | 21 (S/S) | |
| SWK-C 242 | Kapit | NA | 33 (S/S) | |
| SWK-C 089 | Kapit | NA | 21 (S/S) | |
| SWK-C 097 | Kuching | NA | 20 (S/S) | |
| SWK-C 187 | Kuching | NA | 20 (S/S) | |
| MSHR 7891 | Miri | NA | 21 (S/S) | |
| MSHR 7894 | Miri | NA | 25 (S/S) | |
| SWK-C 063 3 | Sibu | NA | 30 (S/S) | |
| SWK-C 064 3 | Sibu | NA | 20.7 (S/S) | |
| MSHR 6392 2 | Sibu | NA | 31 (S/S) | |
| MSHR 6802 2 | Sibu | NA | 29 (S/S) | |
| MSHR 6401 2 | Sibu | NA | 25 (S/S) | |
| MSHR 6404 2 | Sibu | NA | 18 (S/S) | |
| MSHR 6816 2 | Sibu | NA | 31 (S/S) | |
| SWK-C 084 | Sibu | NA | 22 (S/S) | |
| SWK-C 085 | Sibu | NA | 26 (S/S) | |
| SWK-C 087 | Sibu | NA | 21 (S/S) | |
| SWK-C 102 | Bintulu | 35 | 21 (S/S) | |
| SWK-C 104 | Bintulu | 32 | 22 (S/S) | |
| SWK-C 112 1 | Bintulu | 23 | 15 (I/S) 1 | Type C 1 |
| SWK-C 113 1,3 | Bintulu | 22 | 12 (I/I) 1 | Type C 1 |
| SWK-C 116 | Bintulu | 35 | 21 (S/S) | |
| SWK-C 117 | Bintulu | 28 | 20 (S/S) | |
| SWK-C 152 | Bintulu | 30 | 16 (S/S) | |
| SWK-C 153 | Bintulu | 33 | 24 (S/S) | |
| SWK-C 154 (a) 1 | Bintulu | 24 | 16 (S/S) 1 | Type C 1 |
| SWK-C 157 | Bintulu | 34 | 20 (S/S) | |
| SWK-C 165 | Bintulu | 29 | 20 (S/S) | |
| SWK-C 168 | Bintulu | 28 | 18(S/S) | |
| SWK-C 169 | Bintulu | 29 | 20 (S/S) | |
| SWK-C 170 | Bintulu | 25 | 19 (S/S) | |
| SWK-C 173 | Bintulu | 27 | 17 (S/S) | |
| SWK-C 176 | Bintulu | 30 | 18 (S/S) | |
| MSHR 7883 | Bintulu | 29 | 24 (S/S) | |
| MSHR 7884 | Bintulu | 27 | 22 (S/S) | |
| MSHR 7896 | Bintulu | 35 | 20 (S/S) | |
| MSHR 7904 | Bintulu | 32 | 17 (S/S) | |
| MSHR 7885 | Bintulu | 35 | 26 (S/S) | |
| MSHR 7897 1 | Bintulu | 32 | 15 (I/S) 1 | Type C 1 |
| MSHR 7886 | Bintulu | 29 | 18 (S/S) | |
| SWK-C 200 | Bintulu | 30 | 20 (S/S) | |
| SWK-C 201 | Bintulu | 24 | 17 (S/S) | |
| SWK-C 204 | Bintulu | 27 | 20 (S/S) | |
| SWK-C 207 | Bintulu | 31 | 21 (S/S) | |
| SWK-C 209 | Bintulu | 25 | 20 (S/S) | |
| SWK-C 210 | Bintulu | 32 | 25 (S/S) | |
| SWK-C 212 | Bintulu | 33 | 22 (S/S) | |
| SWK-C 262 3 | Bintulu | 26 | 16 (S/S) | |
| SWK-C 129 1 | Kapit | 26 | 15 (I/S) 1 | Type C 1 |
| SWK-C 213 | Kapit | 33 | 22 (S/S) | |
| SWK-C 219 1 | Kapit | 28 | 9 (R/R) 1 | Type C 1 |
| SWK-C 224 | Kapit | 28 | 22 (S/S) | |
| SWK-C 225 | Kapit | 35 | 24 (S/S) | |
| SWK-C 228 | Kapit | 25 | 23 (S/S) | |
| SWK-C 232 1 | Kapit | 33 | 12 (I/I) 1 | Type C 1 |
| SWK-C 239 1 | Kapit | 33 | 15 (I/S) 1 | Type C 1 |
| SWK-C 240 | Kapit | 32 | 24 (S/S) | |
| SWK-C 096 1 | Kuching | 28 | 18 (S/S) 1 | Type C 1 |
| SWK-C 100 | Kuching | 30 | 20 (S/S) | |
| MSHR 7898/M2 | Miri | 27 | 21 (S/S) | |
| MSHR 7899/M3 | Miri | 31 | 22 (S/S) | |
| MSHR 7889/M4 | Miri | 26 | 19 (S/S) | |
| MSHR 7890/M5 | Miri | 28 | 19 (S/S) | |
| MSHR 7892/M7 | Miri | 30 | 20 (S/S) |
| Isolate(s) | Origin | Minimal Inhibitory Concentration ((µg/mL) | MIC Interpretation (CLSI and EUCAST) |
|---|---|---|---|
| SWK-C 103 2 | Bintulu | 1 | S |
| SWK-C 106 2 | Bintulu | 2 | S |
| SWK-C 108 2 | Bintulu | 0.75 | S |
| SWK-C 109 2 | Bintulu | 1 | S |
| SWK-C 112 | Bintulu | 0.5 | S |
| SWK-C 113 2 | Bintulu | 0.75 | S |
| SWK-C 118 | Bintulu | 3 1 | I |
| SWK-C 119 | Bintulu | 1.5 | S |
| SWK-C 123 2 | Bintulu | 1 | S |
| SWK-C 145 | Bintulu | 0.38 | S |
| SWK-C 146 (c) 2 | Bintulu | 1.5 | S |
| SWK-C 149 (b) | Bintulu | 2 | S |
| SWK-C 150 2 | Bintulu | 1.5 | S |
| SWK-C 154 (a) | Bintulu | 2 | S |
| SWK-C 154 (b) 2 | Bintulu | 2 | S |
| SWK-C 154 (c) | Bintulu | 2 | S |
| SWK-C 155 (a) | Bintulu | 1.5 | S |
| SWK-C 155 (b) | Bintulu | 1.5 | S |
| SWK-C 158 | Bintulu | 1 | S |
| SWK-C 161 | Bintulu | 0.5 | S |
| SWK-C 163 (a) 2 | Bintulu | 1.5 | S |
| SWK-C 163 (b) 2 | Bintulu | 2 | S |
| MSHR 7897 | Bintulu | 2 | S |
| SWK-C 194 | Bintulu | 0.38 | S |
| SWK-C 199 | Bintulu | 0.5 | S |
| SWK-C 129 | Kapit | 0.75 | S |
| SWK-C 131 | Kapit | 1 | S |
| SWK-C 132 | Kapit | 1 | S |
| SWK-C 133 | Kapit | 0.75 | S |
| SWK-C 134 | Kapit | 0.5 | S |
| SWK-C 136 | Kapit | 1.5 | S |
| SWK-C 138 | Kapit | 0.5 | S |
| SWK-C 139 | Kapit | 0.75 | S |
| SWK-C 140 | Kapit | 1 | S |
| SWK-C 219 | Kapit | 2 | S |
| SWK-C 221 | Kapit | 0.5 | S |
| SWK-C 223 | Kapit | 0.75 | S |
| SWK-C 226 | Kapit | 1.5 | S |
| SWK-C 232 | Kapit | 1 | S |
| SWK-C 233 | Kapit | 0.75 | S |
| SWK-C 239 | Kapit | 0.75 | S |
| SWK-C 096 | Kuching | 0.75 | S |
References
- Currie, B.J.; Kaestli, M. A global picture of melioidosis. Nature 2016, 529, 290–291. [Google Scholar] [CrossRef]
- Limmathurotsakul, D.; Golding, N.; Dance, D.A.B.; Messina, J.P.; Pigott, D.M.; Moyes, C.L.; Rolim, D.B.; Bertherat, E.; Day, N.P.J.; Peacock, S.J.; et al. Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat. Microbiol. 2016, 1, 15008. [Google Scholar] [CrossRef]
- Wiersinga, W.J.; Virk, H.S.; Torres, A.G.; Currie, B.J.; Peacock, S.J.; Dance, D.A.B.; Limmathurotsakul, D. Melioidosis. Nat. Rev. Dis. Primers 2018, 4, 17107. [Google Scholar] [CrossRef]
- Sanchez-Villamil, J.I.; Torres, A.G. Melioidosis in Mexico, Central America, and the Caribbean. Trop. Med. Infect. Dis. 2018, 3, 24. [Google Scholar] [CrossRef] [PubMed]
- Nathan, S.; Chieng, S.; Kingsley, P.V.; Mohan, A.; Podin, Y.; Ooi, M.-H.; Mariappan, V.; Vellasamy, K.M.; Vadivelu, J.; Daim, S.; et al. Melioidosis in Malaysia: Incidence, Clinical Challenges, and Advances in Understanding Pathogenesis. Trop. Med. Infect. Dis. 2018, 3, 25. [Google Scholar] [CrossRef]
- Sia, T.L.L.; Mohan, A.; Ooi, M.-H.; Chien, S.-L.; Tan, L.-S.; Goh, C.; Pang, D.C.L.; Currie, B.J.; Wong, J.-S.; Podin, Y. Epidemiological and Clinical Characteristics of Melioidosis Caused by Gentamicin-Susceptible Burkholderia pseudomallei in Sarawak, Malaysia. Open Forum Infect. Dis. 2021, 8, ofab460. [Google Scholar] [CrossRef] [PubMed]
- Mohan, A.; Podin, Y.; Tai, N.; Chieng, C.-H.; Rigas, V.; Machunter, B.; Mayo, M.; Wong, D.; Chien, S.-L.; Tan, L.-S.; et al. Pediatric melioidosis in Sarawak, Malaysia: Epidemiological, clinical and microbiological characteristics. PLoS Neglected Trop. Dis. 2017, 11, e0005650. [Google Scholar] [CrossRef] [PubMed]
- White, N. Melioidosis. Lancet 2003, 361, 1715–1722. [Google Scholar] [CrossRef]
- Dance, D. Treatment and prophylaxis of melioidosis. Int. J. Antimicrob. Agents 2014, 43, 310–318. [Google Scholar] [CrossRef]
- Vima, V.; Ling, H.-W.; Cho, W.-M.; Norhuzaimah, J. Incidence, Risk Factors and Clinical Epidemiology of Melioidosis in Miri Hospital, Sarawak, Malaysia. Sarawak Health J. 2016, 3, 22–23. [Google Scholar]
- Yong, K.-Y.; Tang, A.S.-O.; Teh, Y.-C.; Fam, T.-L.; Chua, H.-H. Prevalence of Antibiotic Resistance in Burkholderia pseudomallei Cases Presented to Miri General Hospital. Sarawak Health J. 2016, 3, 26. [Google Scholar]
- Podin, Y.; Kaestli, M.; McMahon, N.; Hennessy, J.; Ngian, H.U.; Wong, J.S.; Mohana, A.; Wong, S.C.; William, T.; Mayo, M.; et al. Reliability of Automated Biochemical Identification of Burkholderia pseudomallei Is Regionally Dependent. J. Clin. Microbiol. 2013, 51, 3076–3078. [Google Scholar] [CrossRef]
- Podin, Y.; Sarovich, D.S.; Price, E.P.; Kaestli, M.; Mayo, M.; Hii, K.; Ngian, H.; Wong, S.; Wong, I.; Wong, J.; et al. Burkholderia pseudomallei Isolates from Sarawak, Malaysian Borneo, Are Predominantly Susceptible to Aminoglycosides and Macrolides. Antimicrob. Agents Chemother. 2013, 58, 162–166. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing 27th ed. CLSI Supplement M100; Clinical Laboratory Standards Institute: Wayne, PA, USA; Available online: https://www.google.com/url?sa=t&source=web&rct=j&opi=89978449&url=https://www.researchgate.net/profile/Nikolaos_Andritsos/post/Where_to_find_MICs_of_antimicrobial_agents/attachment/59d659e279197b80779af319/AS%253A544238144323584%25401506767893696/download/2017_CLSI_M100_Performance%2BStandards%2Bfor%2BAntimicrobial%2BSusceptibility%2BTesting_27th%2Bed..pdf&ved=2ahUKEwjwu9rQ-paSAxVp2TQHHZHCOzcQFnoECBoQAQ&usg=AOvVaw1tGIX4d98p2DVfRY_2WrOh (accessed on 12 November 2018).
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Version 8.1, Valid from 2018-05-15. 2018. Available online: https://www.eucast.org/ast_of_bacteria/previous_versions_of_documents/ (accessed on 12 November 2018).
- CLSI. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically 11th ed CLSI Standard M07; Clinical Laboratory Standards Institute: Wayne, PA, USA, 2018; Available online: https://clsi.org/shop/standards/m07/ (accessed on 20 January 2019).
- Warrens, M.J. Five Ways to Look at Cohen’s Kappa. J. Psychol. Psychother. 2015, 5, 1000197. [Google Scholar] [CrossRef]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed]
- Holden, M.T.G.; Titball, R.W.; Peacock, S.J.; Cerdeño-Tárraga, A.M.; Atkins, T.; Crossman, L.C.; Pitt, T.; Churcher, C.; Mungall, K.; Bentley, S.D.; et al. Genomic plasticity of the causative agent of melioidosis, Burkholderia pseudomallei. Proc. Natl. Acad. Sci. USA 2004, 101, 14240–14245. [Google Scholar] [CrossRef]
- Biot, F.V.; Valade, E.; Garnotel, E.; Chevalier, J.; Villard, C.; Thibault, F.M.; Vidal, D.R.; Pagès, J.-M. Involvement of the Efflux Pumps in Chloramphenicol Selected Strains of Burkholderia thailandensis: Proteomic and Mechanistic Evidence. PLoS ONE 2011, 6, e16892. [Google Scholar] [CrossRef]
- Podnecky, N.L.; Wuthiekanun, V.; Peacock, S.J.; Schweizer, H.P. The BpeEF-OprC Efflux Pump Is Responsible for Widespread Trimethoprim Resistance in Clinical and Environmental Burkholderia pseudomallei Isolates. Antimicrob. Agents Chemother. 2013, 57, 4381–4386. [Google Scholar] [CrossRef]
- Payne, G.W.; Vandamme, P.; Morgan, S.H.; LiPuma, J.J.; Coenye, T.; Weightman, A.J.; Jones, T.H.; Mahenthiralingam, E. Development of a recA Gene-Based Identification Approach for the Entire Burkholderia Genus. Appl. Environ. Microbiol. 2005, 71, 3917–3927. [Google Scholar] [CrossRef]
- Sia, T.L.-L.; Lai, C.D.; Manan, K.; Khiu, F.-L.; Bakhtiar, S.Z.; Chor, Y.-K.; Chien, S.-L.; Tan, L.-S.; Ooi, M.-H.; Mohan, A. Ceftazidime-resistance in pediatric melioidosis: A case report and literature review. IDCases 2025, 39, e02149. [Google Scholar] [CrossRef] [PubMed]
- Dance, D.; Davong, V.; Soeng, S.; Phetsouvanh, R.; Newton, P.; Turner, P. Trimethoprim/sulfamethoxazole resistance in Burkholderia pseudomallei. Int. J. Antimicrob. Agents 2014, 44, 368–369. [Google Scholar] [CrossRef] [PubMed]
- Saiprom, N.; Amornchai, P.; Wuthiekanun, V.; Day, N.P.; Limmathurotsakul, D.; Peacock, S.J.; Chantratita, N. Trimethoprim/sulfamethoxazole resistance in clinical isolates of Burkholderia pseudomallei from Thailand. Int. J. Antimicrob. Agents 2015, 45, 557–559. [Google Scholar] [CrossRef]
- Dutta, S.; Haq, S.; Hasan, M.R.; Haq, J.A. Antimicrobial susceptibility pattern of clinical isolates of Burkholderia pseudomallei in Bangladesh. BMC Res. Notes 2017, 10, 299. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, N.; Hashim, R.; Noor, A.M. The In Vitro Antibiotic Susceptibility of Malaysian Isolates of Burkholderia pseudomallei. Int. J. Microbiol. 2013, 2013, 121845. [Google Scholar] [CrossRef] [PubMed]
- Karatuna, O.; Dance, D.; Matuschek, E.; Åhman, J.; Turner, P.; Hopkins, J.; Amornchai, P.; Wuthiekanun, V.; Cusack, T.-P.; Baird, R.; et al. Burkholderia pseudomallei multi-centre study to establish EUCAST MIC and zone diameter distributions and epidemiological cut-off values. Clin. Microbiol. Infect. 2021, 27, 736–741. [Google Scholar] [CrossRef]
- Dance, D.A.; Wuthiekanun, V.; Baird, R.W.; Norton, R.; Limmathurotsakul, D.; Currie, B.J. Interpreting Burkholderia pseudomallei disc diffusion susceptibility test results by the EUCAST method. Clin. Microbiol. Infect. 2021, 27, 827–829. [Google Scholar] [CrossRef]
- Kahlmeter, G.; Giske, C.G.; Kirn, T.J.; Sharp, S.E. Point-Counterpoint: Differences between the European Committee on Antimicrobial Susceptibility Testing and Clinical and Laboratory Standards Institute Recommendations for Reporting Antimicrobial Susceptibility Results. J. Clin. Microbiol. 2019, 57. [Google Scholar] [CrossRef]
- Podnecky, N.L.; Rhodes, K.A.; Mima, T.; Drew, H.R.; Chirakul, S.; Wuthiekanun, V.; Schupp, J.M.; Sarovich, D.S.; Currie, B.J.; Keim, P.; et al. Mechanisms of Resistance to Folate Pathway Inhibitors in Burkholderia pseudomallei: Deviation from the Norm. mBio 2017, 8, e01357-17. [Google Scholar] [CrossRef]





| Hospital | Location | No. of Isolates Collected |
|---|---|---|
| Bintulu Hospital | Bintulu, Sarawak | 98 |
| Sibu Hospital | Sibu, Sarawak | 10 |
| Kapit Hospital | Kapit, Sarawak | 45 |
| Miri Hospital | Miri, Sarawak | 7 |
| Sarawak General Hospital | Kuching, Sarawak | 2 |
| Borneo Medical Centre | Kuching, Sarawak | 2 |
| Antibiotic | MIC Interpretation According to Both CLSI and EUCAST Standards (µg/mL) | ||
|---|---|---|---|
| S≤ | I | R≥ | |
| Trimethoprim-sulfamethoxazole (SXT) | 2 | - | 4 |
| Amoxicillin-clavulanate (AMC) | 8/4 | 16/8 | 32/16 |
| Azithromycin (AZ) | 2 | 4 | 8 |
| Ceftazidime (TZ) | 8 | 16 | 32 |
| Doxycycline (DC) | 4 | 8 | 16 |
| Gentamicin (GEN) | 4 | 8 | 16 |
| Meropenem (MEM) | 2 | 8 | 2 |
| Isolates | Description | Zone Diameter, mm | MIC, µg/mL | ||||
|---|---|---|---|---|---|---|---|
| Trial 1 | Trial 2 | Trial 3 | Trial 1 | Trial 2 | Trial 3 | ||
| SWK-C 106 | SXTI in this study 1 | 12 | 16 | 14 | 2 | 3 | 2 |
| SWK-C 118 | SXTI in this study 1 | 12 | 16 | 12 | 3 | 2 | 2 |
| SWK-C 145 | SXTS in this study 2 | 20 | 22 | 22 | 0.75 | 0.75 | 0.5 |
| E. coli ATCC 25922 | QC recommended by CLSI and EUCAST | NZ 3 | NZ 3 | NZ 3 | NZ 3 | NZ 3 | NZ 3 |
| E. coli ATCC 11775 | Alternative QC proposed in this study | 32 | 32 | 33 | 0.125 | 0.125 | 0.125 |
| Isolates | Zone Diameter (mm) | MIC (µg/mL) | ||||
|---|---|---|---|---|---|---|
| 18 h | 24 h | 36 h | 18 h | 24 h | 36 h | |
| SWK-C 106 | 16 | 11 | NZ | 3 | 4 | NZ |
| SWK-C 118 | 16 | 16 | 9 | 2 | 4 | NZ |
| SWK-C 145 | 22 | 16 | 13 | 0.75 | 1.5 | 3 |
| E. coli ATCC 25922 | NZ | NZ | NZ | NZ | NZ | NZ |
| E. coli ATCC 11775 | 32 | 32 | 32 | 0.125 | 0.125 | 0.125 |
| Target Gene | Primer | Primer Sequence (5′->3′) [Expected Base Pair, bp] |
|---|---|---|
| bpeT | BpeT_F1 BpeT_R1 BpeT_F2 BpeT_R2 | 5′- TGC GCA AAC ATA TGA CGA AC -3′ 5′- CGA ATT CCA CTC ACG CTA CC -3′ [768 bp] 5′- GCG GCT CGA AAA GTA GTT GA -3′ 5′- ACA ATT CAC GTC CCC TGA AC -3′ [684 bp] |
| llpe | llpE_F1 llpE_R1 llpE_F2 llpE_R2 | 5′- GAT TGT TCA GGG GAC GTG A -3′ 5′- GAG CGA ATA ATC GAC CGA CA -3′ [392 bp] 5′- CGG TGG TGC TTT ATT TCC AC -3’ 5′- CGG GAA GTA CGC AAG ATA GC -3′ [774 bp] |
| bpeE | BpeE_F1 BpeE_R1 BpeE_F2 BpeE_R2 | 5′- CGA CAA CCT GAG GGG TTT T -3′ 5′- GCC GAT GTA TTG CAG GTA GG -3′ [730 bp] 5′- TTA CGA CGA GAA GCA GAA CG -3′ 5′- TGA AAG GCT CTG TCT GAT TGG -3′ [846 bp] |
| bpeF | BpeF_F1 BpeF_R1 BpeF_F2 BpeF_R2 BpeF_F3 BpeF_R3 BpeF_F4 BpeF_R4 BpeF_F5 BpeF_R5 | 5′- CCC AAT CAG ACA GAG CCT TT -3′ 5′- CGA ACT CGT CCT CGT TCT G -3′ [769 bp] 5′- ATT CGC GAG CAG AAC GTG -3′ 5′- GTC ATC GCG AAC TGC TTG TA -3′ [797 bp] 5′- CTA TTC GAT CAA CGC GCT CT -3′ 5′- CCG CGT ACT TCT GGT TCA G -3′ [824 bp] 5′- GTG AAC GGC TTC ACG AAC A -3′ 5′- TGA TCG GAA ACA CCC AGA AC -3′ [796 bp] 5′- GAC ATC CTG CAA CTG AAG ACG -3’ 5′- GCG TTC GTT GAT GTT GGT CT -3′ [850 bp] |
| oprC | OprC_F1 OprC_R1 OprC_F2 OprC_R2 | 5′- CGG ACG CTT GAG GAT AGA AA -3′ 5′- CTC GCT GAA CGA GAA ATC C -3′ [882 bp] 5′- CGC GGA TTT CTC GTT CAG -3′ 5′- CGA CAT TCG CAT TTC GTC -3′ [785 bp] |
| IZD 1 (mm) | ≤10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | 26 | 27 | 28 | 29 | ≥30 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MIC80, 1 IZ 2 | 2 | 1 | 7 | 1 | 2 | 10 | 6 | 2 | 5 | 11 | 15 5 | 15 5 | 7 | 2 | 6 | 6 | 4 | 0 | 2 | 2 | 11 |
| MIC50, 2 IZs 3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 2 | 3 | 3 | 4 | 6 5 | 4 | 23 |
| MIC80, 2 IZs 4 | 1 | 0 | 2 | 0 | 0 | 4 | 3 | 3 | 4 | 3 | 10 5 | 4 | 6 | 1 | 4 | 1 | 1 | 0 | 0 | 0 | 0 |
| Total, N = 164 (MIC80) | 3 | 1 | 9 | 1 | 2 | 14 | 9 | 5 | 9 | 14 | 25 5 | 19 | 13 | 3 | 10 | 7 | 5 | 0 | 2 | 2 | 11 |
| MIC’s Interpretive Standard | CLSI Interpretive Standard | EUCAST Interpretive Standard | ||
|---|---|---|---|---|
| Frequency, % (n/N) | 95% CI | Frequency, % (n/N) | 95% CI | |
| Susceptible Isolates | 81.7% (134/164) | 0.749 ~ 0.873 | 79.9% (131/164) | 0.729 ~ 0.857 |
| Intermediate Isolates | 15.9% (26/164) | 0.106 ~ 0.224 | 3.7% (6/164) | 0.014 ~ 0.078 |
| Resistant Isolates | 2.4% (4/164) | 0.007 ~ 0.061 | 16.5% (27/164) | 0.111 ~ 0.230 |
| Cohen’s Kappa value, κ (95% CI) | κ = 0.458 (0.340 ~ 0.570) | |||
| Frequency (%) | CLSI’s Standard | EUCAST’s Standard | ||
|---|---|---|---|---|
| Disk Diffusion | E-Test | Disk Diffusion | E-Test | |
| Susceptibility, % (n/N) | 81.7% (134/164) | 96.3% (158/164) | 79.9% (131/164) | 97.6% (160/164) |
| Intermediate, % (n/N) | 15.9% (26/164) | 3.7% (6/164) | 3.7% (6/164) | 1.2% (2/164) |
| Resistance, % (n/N) | 2.4% (4/164) | - | 16.5% (27/164) | 1.2% (2/164) |
| SXT Phenotype (Disk Diffusion) | Isolate(s) | SXT | GEN | AZ | MEM | TZ | DC | AMC |
|---|---|---|---|---|---|---|---|---|
| Susceptible | SWK-C 145 | 0.38 (S) | NZ (R) | NZ (R) | 1.5 (S) | 1.5 (S) | 1 (S) | 3 (S) |
| SWK-C 133 | 0.5 (S) | 1 (S) | 3 (S) | 1 (S) | 2 (S) | 0.75 (S) | 4 (S) | |
| SWK-C 136 | 0.5 (S) | 2 (S) | 4 (S) | 0.75 (S) | 1.5 (S) | 0.75 (S) | 3 (S) | |
| SWK-C 161 | 0.5 (S) | 2 (S) | 4 (S) | 2 (S) | 2 (S) | 0.75 (S) | 3 (S) | |
| Resistance | SWK-C 219 | 0.5 (S) | 96 (R) | NZ (R) | 2 (S) | 3 (S) | 1.5 (S) | 4 (S) |
| SWK-C 109 | 1 (S) | 1.5 (S) | 6(S) | 1 (S) | 1.5 (S) | 0.75 (S) | 6 (S) | |
| SWK-C 118 | 3 (I) | 6 (S) | 6(S) | 1.5 (S) | 2 (S) | 1 (S) | 6 (S) | |
| Intermediate | SWK-C 106 | 2 (S) | 1.5 (S) | 4 (S) | 1 (S) | 2 (S) | 0.75 (S) | 4 (S) |
| SWK-C 140 | 2 (S) | 6 (S) | 4 (S) | 1.5 (S) | 2 (S) | 0.75 (S) | 4 (S) | |
| SWK-C 149 (b) | 2 (S) | 1.5 (S) | 4 (S) | 1.5 (S) | 3 (S) | 0.75 (S) | 6 (S) | |
| SWK-C 154 (a) | 2 (S) | 1.5 (S) | 3 (S) | 1.5 (S) | 2 (S) | 0.75 (S) | 4 (S) | |
| SWK-C 154 (b) | 2 (S) | 1.5 (S) | 4 (S) | 2 (S) | 2 (S) | 0.75 (S) | 6 (S) | |
| SWK-C 154 (c) | 2 (S) | 4 (S) | 4 (S) | 2 (S) | 1.5 (S) | 0.75 (S) | 4 (S) | |
| SWK-C 163 (b) | 2 (S) | 2 (S) | 4 (S) | 1.5 (S) | 3 (S) | 1 (S) | 4 (S) | |
| E-test Susceptibility frequency (%) | 92.9% | 85.7% | 85.7% | 100% | 100% | 100% | 100% | |
| Isolate(s) | Disk Diffusion IZD (mm) | E-Test MIC (µg/mL) | SMX, Broth Microdilution MIC (µg/mL) | TMP, Broth Microdilution MIC (µg/mL) | ||
|---|---|---|---|---|---|---|
| MIC50 | MIC80 | MIC50 | MIC80 | |||
| SWK-C 219 GenR | 9 (R) | 2 (S) | 8 (S) | 128 (S) | 2 (S) | 16 (R) |
| SWK-C 163b GenS | 10 (R) | 2 (S) | 32 (S) | 1024 (R) 1 | 4 (S) | 32 (R) 1 |
| SWK-C 106 GenS | 12 (I) | 2 (S) | 32 (S) | 512 (R) 1 | 8 (S) | 64 (R) 1 |
| SWK-C 118 GenS | 12 (I) | 3 (I) | 8 (S) | 256 (S) | 4 (S) | 64 (R) |
| SWK-C 154b GenS | 12 (I) | 2 (S) | 8 (S) | 128 (S) | 2 (S) | 32 (R) |
| SWK-C 140 GenS | 15 (I) | 2 (S) | 16 (S) | 1024 (R) 1 | 4 (S) | 32 (R) 1 |
| SWK-C 154c GenS | 14 (I) | 2 (S) | 16 (S) | 512 (R) 1 | 4 (S) | 32 (R) 1 |
| SWK-C 149b GenS | 16 (S) | 2 (S) | 16 (S) | 256 (S) | 4 (S) | 64 (R) |
| SWK-C 154a GenS | 16 (S) | 2 (S) | 32 (S) | ETR 1,2 | 16 (R) | 64 (R) 1 |
| SWK-C 109 GenS | 18 (S) | 1 (S) | 32 (S) | ETR 1,2 | 2 (S) | 16 (R) 1 |
| SWK-C 145 GenR | 20 (S) | 0.75 (S) | 4 (S) | 32 (S) | 1 (S) | 8 (S) |
| SWK-C 133 GenS | 21 (S) | 0.75 (S) | 4 (S) | 128 (S) | 2 (S) | 16 (R) |
| SWK-C 136 GenS | 21 (S) | 0.25 (S) | 4 (S) | 32 (S) | 1 (S) | 16 (R) |
| SWK-C 161 GenS | 37 (S) | 0.5 (S) | 32 (S) | ETR 1,2 | 4 (S) | 32 (R) 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Sumbu, L.L.; Sia, T.L.-L.; Ooi, M.-H.; Mohan, A.; Wong, J.-S.; Podin, Y. Burkholderia pseudomallei in Sarawak, Malaysian Borneo, Remains Highly Susceptible to Trimethoprim-Sulfamethoxazole Despite Resistance to Its Individual Components. Pathogens 2026, 15, 110. https://doi.org/10.3390/pathogens15010110
Sumbu LL, Sia TL-L, Ooi M-H, Mohan A, Wong J-S, Podin Y. Burkholderia pseudomallei in Sarawak, Malaysian Borneo, Remains Highly Susceptible to Trimethoprim-Sulfamethoxazole Despite Resistance to Its Individual Components. Pathogens. 2026; 15(1):110. https://doi.org/10.3390/pathogens15010110
Chicago/Turabian StyleSumbu, Liana Lantong, Tonnii Loong-Loong Sia, Mong-How Ooi, Anand Mohan, Jin-Shyan Wong, and Yuwana Podin. 2026. "Burkholderia pseudomallei in Sarawak, Malaysian Borneo, Remains Highly Susceptible to Trimethoprim-Sulfamethoxazole Despite Resistance to Its Individual Components" Pathogens 15, no. 1: 110. https://doi.org/10.3390/pathogens15010110
APA StyleSumbu, L. L., Sia, T. L.-L., Ooi, M.-H., Mohan, A., Wong, J.-S., & Podin, Y. (2026). Burkholderia pseudomallei in Sarawak, Malaysian Borneo, Remains Highly Susceptible to Trimethoprim-Sulfamethoxazole Despite Resistance to Its Individual Components. Pathogens, 15(1), 110. https://doi.org/10.3390/pathogens15010110






