Multiplex Real-Time RT-PCR Assays for Detection and Differentiation of Porcine Enteric Coronaviruses
Abstract
1. Introduction
2. Materials and Methods
2.1. Nucleic Acids from Field Samples and Isolates
2.2. RNA Standards for Testing and Validation
2.3. Selection of Primers and Probes
2.4. RT-qPCR Protocol
2.5. Optimization of Primer and Probe Concentrations
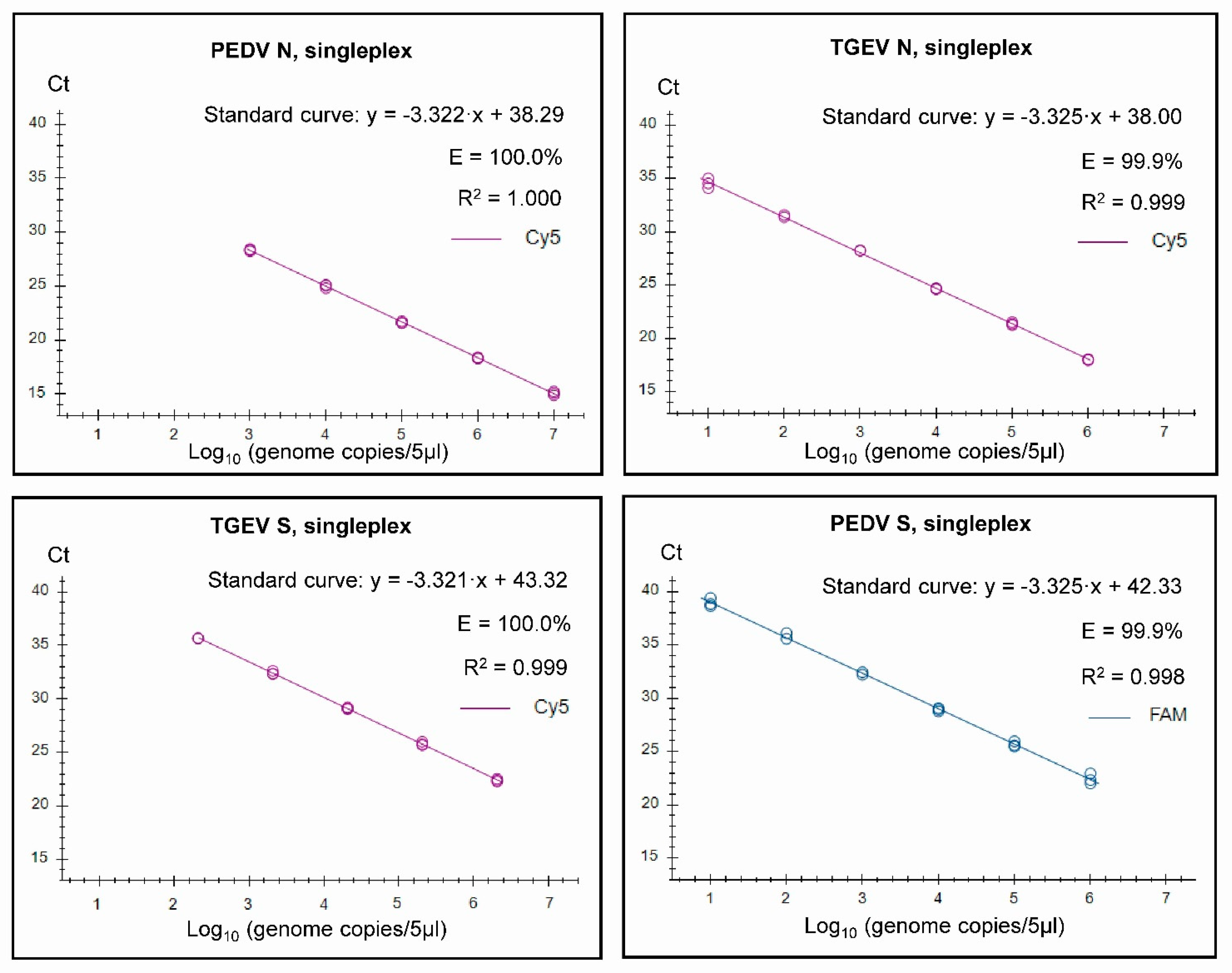
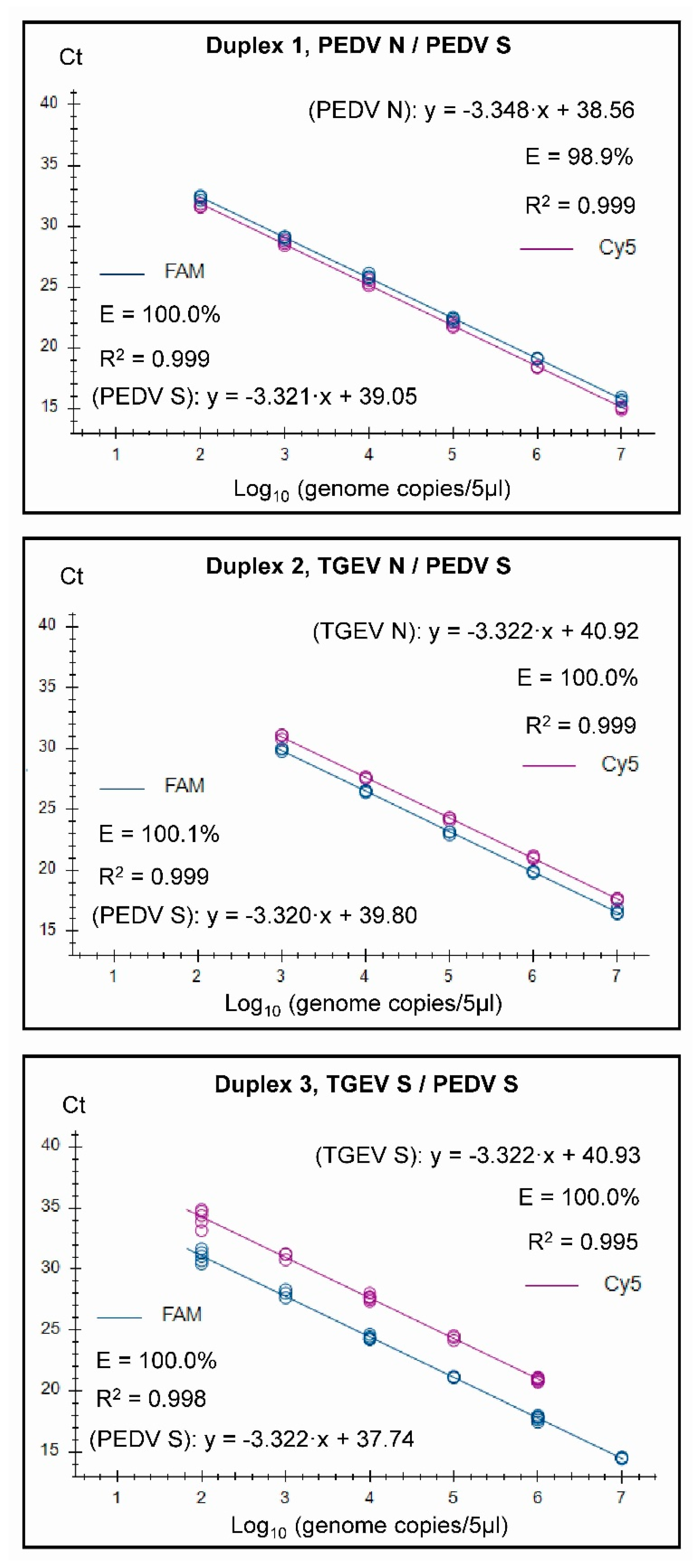
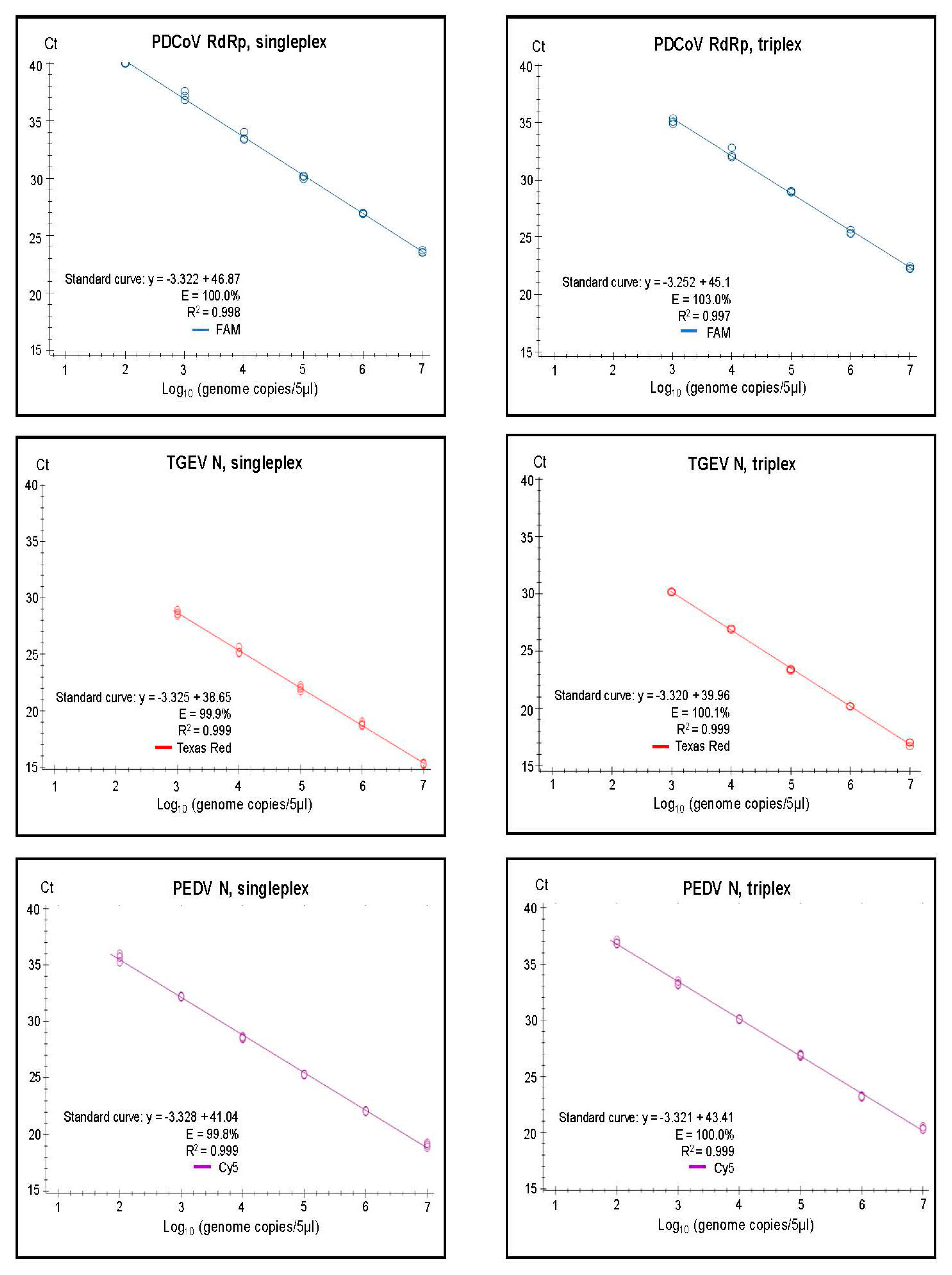
2.6. Standard Curves
2.7. Validation Steps
3. Results
3.1. Assay Design
3.2. Assay Characteristics
3.3. Specificity and Senstivity Testing of the Assays
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jung, K.; Saif, L.J.; Wang, Q. Porcine epidemic diarrhea virus (PEDV): An update on etiology, transmission, pathogenesis, and prevention and control. Virus Res. 2020, 286, 198045. [Google Scholar] [CrossRef] [PubMed]
- Lee, C. Porcine epidemic diarrhea virus: An emerging and re-emerging epizootic swine virus. Virol. J. 2015, 12, 193. [Google Scholar] [CrossRef]
- Chen, F.; Knutson, T.P.; Rossow, S.; Saif, L.J.; Marthaler, D.G. Decline of transmissible gastroenteritis virus and its complex evolutionary relationship with porcine respiratory coronavirus in the United States. Sci. Rep. 2019, 9, 3953. [Google Scholar] [CrossRef]
- Zhou, P.; Fan, H.; Lan, T.; Yang, X.-L.; Shi, W.-F.; Zhang, W.; Zhu, Y.; Zhang, Y.-W.; Xie, Q.-M.; Mani, S.; et al. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 2018, 556, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lau, C.C.Y.; Tsang, A.K.L.; Lau, J.H.N.; Bai, R.; Teng, J.L.L.; Tsang, C.C.C.; Wang, M.; et al. Discovery of Seven Novel Mammalian and Avian Coronaviruses in the Genus Deltacoronavirus Supports Bat Coronaviruses as the Gene Source of Alphacoronavirus and Betacoronavirus and Avian Coronaviruses as the Gene Source of Gammacoronavirus and Deltacoronavi. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Byrum, B.; Zhang, Y. Detection and genetic characterization of deltacoronavirus in pigs, Ohio, USA, 2014. Emerg. Infect. Dis. 2014, 20, 1227–1230. [Google Scholar] [CrossRef]
- Akimkin, V.; Beer, M.; Blome, S.; Hanke, D.; Höper, D.; Jenckel, M.; Pohlmann, A. New chimeric porcine coronavirus in Swine Feces, Germany, 2012. Emerg. Infect. Dis. 2016, 22, 1314–1315. [Google Scholar] [CrossRef]
- Belsham, G.J.; Rasmussen, T.B.; Normann, P.; Vaclavek, P.; Strandbygaard, B.; Bøtner, A. Characterization of a Novel Chimeric Swine Enteric Coronavirus from Diseased Pigs in Central Eastern Europe in 2016. Transbound. Emerg. Dis. 2016, 63, 595–601. [Google Scholar] [CrossRef]
- Boniotti, M.B.; Papetti, A.; Lavazza, A.; Alborali, G.; Sozzi, E.; Chiapponi, C.; Faccini, S.; Bonilauri, P.; Cordioli, P.; Marthaler, D. Porcine epidemic diarrhea virus and discovery of a recombinant swine enteric coronavirus, Italy. Emerg. Infect. Dis. 2016, 22, 83–87. [Google Scholar] [CrossRef]
- de Nova, P.J.G.; Cortey, M.; Díaz, I.; Puente, H.; Rubio, P.; Martín, M.; Carvajal, A. A retrospective study of porcine epidemic diarrhoea virus (PEDV) reveals the presence of swine enteric coronavirus (SeCoV) since 1993 and the recent introduction of a recombinant PEDV-SeCoV in Spain. Transbound. Emerg. Dis. 2020, 67, 2911–2922. [Google Scholar] [CrossRef]
- Papetti, A.; Bonilauri, P.; Chiapponi, C.; Baioni, L.; Boniotti, M.B. Complete Genome Sequence of an Italian Swine Enteric Coronavirus Strain 77590/2019. Microbiol. Resour. Announc. 2022, 11, e00386-22. [Google Scholar] [CrossRef] [PubMed]
- Boniotti, M.B.; Papetti, A.; Bertasio, C.; Giacomini, E.; Lazzaro, M.; Cerioli, M.; Faccini, S.; Bonilauri, P.; Vezzoli, F.; Lavazza, A.; et al. Porcine Epidemic Diarrhoea Virus in Italy: Disease spread and the role of transportation. Transbound. Emerg. Dis. 2018, 65, 1935–1942. [Google Scholar] [CrossRef]
- Valkó, A.; Biksi, I.; Cságola, A.; Tuboly, T.; Kiss, K.; Ursu, K.; Dán, Á. Porcine epidemic diarrhoea virus with a recombinant S gene detected in Hungary, 2016. Acta Vet. Hung. 2017, 65, 253–261. [Google Scholar] [CrossRef]
- Valkó, A.; Albert, E.; Cságola, A.; Varga, T.; Kiss, K.; Farkas, R.; Rónai, Z.; Biksi, I.; Dán, Á. Isolation and characterisation of porcine epidemic diarrhoea virus in Hungary. Acta Vet. Hung. 2019, 67, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Mackay, I.M.; Arden, K.E.; Nitsche, A. Real-time PCR in virology. Nucleic Acids Res. 2002, 30, 1292–1305. [Google Scholar] [CrossRef]
- Bertasio, C.; Giacomini, E.; Lazzaro, M.; Perulli, S.; Papetti, A.; Lavazza, A.; Lelli, D.; Alborali, G.; Boniotti, M.B. Porcine epidemic diarrhea virus shedding and antibody response in swine farms: A longitudinal study. Front. Microbiol. 2016, 7, 2009. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Kim, I.J.; Pyo, H.M.; Tark, D.S.; Song, J.Y.; Hyun, B.H. Multiplex real-time RT-PCR for the simultaneous detection and quantification of transmissible gastroenteritis virus and porcine epidemic diarrhea virus. J. Virol. Methods 2007, 146, 172–177. [Google Scholar] [CrossRef]
- Bigault, L.; Brown, P.; Bernard, C.; Blanchard, Y.; Grasland, B. Porcine epidemic diarrhea virus: Viral RNA detection and quantification using a validated one-step real time RT-PCR. J. Virol. Methods 2020, 283, 113906. [Google Scholar] [CrossRef]
- Alonso, C.; Goede, D.P.; Morrison, R.B.; Davies, P.R.; Rovira, A.; Marthaler, D.G.; Torremorell, M. Evidence of infectivity of airborne porcine epidemic diarrhea virus and detection of airborne viral RNA at long distances from infected herds. Vet. Res. 2014, 45, 73. [Google Scholar] [CrossRef]
- Chen, Q.; Li, G.; Stasko, J.; Thomas, J.T.; Stensland, W.R.; Pillatzki, A.E.; Gauger, P.C.; Schwartz, K.J.; Madson, D.; Yoon, K.J.; et al. Isolation and characterization of porcine epidemic diarrhea viruses associated with the 2013 disease outbreak among swine in the united states. J. Clin. Microbiol. 2014, 52, 234–243. [Google Scholar] [CrossRef]
- Vemulapalli, R.; Gulani, J.; Santrich, C. A real-time TaqMan® RT-PCR assay with an internal amplification control for rapid detection of transmissible gastroenteritis virus in swine fecal samples. J. Virol. Methods 2009, 162, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Liu, Y.; Chen, Y.; Xing, G.; Hao, H.; Wei, Q.; Liang, Y.; Xie, W.; Li, D.; Huang, H.; et al. A novel duplex TaqMan probe-based real-time RT-qPCR for detecting and differentiating classical and variant porcine epidemic diarrhea viruses. Mol. Cell. Probes 2018, 37, 6–11. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Broeders, S.; Huber, I.; Grohmann, L.; Berben, G.; Taverniers, I.; Mazzara, M.; Roosens, N.; Morisset, D. Guidelines for validation of qualitative real-time PCR methods. Trends Food Sci. Technol. 2014, 37, 115–126. [Google Scholar] [CrossRef]
- Costantini, V.; Lewis, P.; Alsop, J.; Templeton, C.; Saif, L.J. Respiratory and fecal shedding of Porcine respiratory coronavirus (PRCV) in sentinel weaned pigs and sequence of the partial S-gene of the PRCV isolates. Arch. Virol. 2004, 149, 957–974. [Google Scholar] [CrossRef]
- Ding, G.; Fu, Y.; Li, B.; Chen, J.; Wang, J.; Yin, B.; Sha, W.; Liu, G. Development of a multiplex RT-PCR for the detection of major diarrhoeal viruses in pig herds in China. Transbound. Emerg. Dis. 2020, 67, 678–685. [Google Scholar] [CrossRef]
- Masuda, T.; Tsuchiaka, S.; Ashiba, T.; Yamasato, H.; Fukunari, K.; Omatsu, T.; Furuya, T.; Shirai, J.; Mizutani, T.; Nagai, M. Development of one-step real-time reverse transcriptase-PCR-based assays for the rapid and simultaneous detection of four viruses causing porcine diarrhea. Jpn. J. Vet. Res. 2016, 64, 5–14. [Google Scholar] [CrossRef]
- Si, G.; Niu, J.; Zhou, X.; Xie, Y.; Chen, Z.; Li, G.; Chen, R.; He, D. Use of dual priming oligonucleotide system-based multiplex RT-PCR assay to detect five diarrhea viruses in pig herds in South China. AMB Express 2021, 11, 99. [Google Scholar] [CrossRef]
- Goecke, N.B.; Hjulsager, C.K.; Krog, J.S.; Skovgaard, K.; Larsen, L.E. Development of a high-throughput real-time PCR system for detection of enzootic pathogens in pigs. J. Vet. Diagn. Investig. 2020, 32, 51–64. [Google Scholar] [CrossRef]
- Puente, H.; Argüello, H.; Mencía-Ares, Ó.; Gómez-García, M.; Rubio, P.; Carvajal, A. Detection and Genetic Diversity of Porcine Coronavirus Involved in Diarrhea Outbreaks in Spain. Front. Vet. Sci. 2021, 8, 651999. [Google Scholar] [CrossRef]
- de Souza Luna, L.K.; Heiser, V.; Regamey, N.; Panning, M.; Drexler, J.F.; Mulangu, S.; Poon, L.; Baumgarte, S.; Haijema, B.J.; Kaiser, L.; et al. Generic Detection of Coronaviruses and Differentiation at the Prototype Strain Level by Reverse Transcription-PCR and Nonfluorescent Low-Density Microarray. J. Clin. Microbiol. 2007, 45, 1049–1052. [Google Scholar] [CrossRef] [PubMed]
- Escutenaire, S.; Mohamed, N.; Isaksson, M.; Thorén, P.; Klingeborn, B.; Belák, S.; Berg, M.; Blomberg, J. SYBR Green Real-Time Reverse Transcription-Polymerase Chain Reaction Assay for the Generic Detection of Coronaviruses. Arch. Virol. 2007, 152, 41–58. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Jung, K.; Wang, Q.; Saif, L.J.; Vlasova, A.N. Development of a one-step RT-PCR assay for detection of pancoronaviruses (α-, β-, γ-, and δ-coronaviruses) using newly designed degenerate primers for porcine and avian fecal samples. J. Virol. Methods 2018, 256, 116–122. [Google Scholar] [CrossRef]
- Vijgen, L.; Moës, E.; Keyaerts, E.; Li, S.; Van Ranst, M. A Pancoronavirus RT-PCR Assay for Detection of All Known Coronaviruses. Methods Mol. Biol. 2008, 454, 3–12. [Google Scholar] [CrossRef]
- Lazov, C.M.; Chriél, M.; Baagøe, H.J.; Fjederholt, E.; Deng, Y.; Kooi, E.A.; Belsham, G.J.; Bøtner, A.; Rasmussen, T.B. Detection and Characterization of Distinct Alphacoronaviruses in Five Different Bat Species in Denmark. Viruses 2018, 10, 486. [Google Scholar] [CrossRef]
- Kim, S.Y.; Song, D.S.; Park, B.K. Differential detection of transmissible gastroenteritis virus and porcine epidemic diarrhea virus by duplex RT-PCR. J. Vet. Diagn. Investig. 2001, 13, 516–520. [Google Scholar] [CrossRef] [PubMed]
| Oligo | Sequence 5′–3′ | Location ** | References | |
|---|---|---|---|---|
| Duplex assays | PEDV-N-R | TTGCCTCTGTTGTTACTTGGAGAT | 26,853–26,876 | [17] |
| PEDV-N-F | CGCAAAGACTGAACCCACTAATTT | 26,679–26,702 | ||
| PEDV-N-US-F | CGCAAAGACTGAACCCACTAACCT | 26,679–26,702 | [17,20] | |
| PEDV-N-P | TGTTGCCATTRCCACGACTCCTGC | 26,819–26,842 | ||
| TGEV-N-F | GCAGGTARAGGTGATGTGACAA | 27,637–27,658 | [17] | |
| TGEV-N-R | ACATTCAGCCARTTGTGGGTAA | 27,735–27,756 | ||
| TGEV-N-P | TGGCACTGCTCCCATTGGCAACGA | 27,707–27,730 | ||
| PEDV-S-F | ACGTCCCTTTACTTTCAATTCACA | 22,474–22,497 | [19] | |
| PEDV-S-R | TATACTTGGTACACACATCCAGAGTCA | 22,559–22,585 | ||
| PEDV-S-P | TGAGTTGATTACTGGCACGCCTAAACCAC | 22,503–22,531 | ||
| TGEV-S-F | TCTGCTGAAGGTGCTATTATATGC | 20,734–20,757 | [21] | |
| TGEV-S-R | CCACAATTTGCCTCTGAATTAGAA * | 20,856–20,879 | ||
| TGEV-S-P | * AAGGGCTCACCACCTACTACCACCA | 20,764–20,788 | ||
| Triplex assay | PDCoV-RdRp-F | AACTGACATGAATGTTGGCCCT | 13,777–13,798 | This study |
| PDCoV-RdRp-R | CATGCACCCAGAATGCGAGA | 13,874–13,893 | ||
| PDCoV-RdRp-P | AGCATACTGTGTTAGCAGAGCATGATGGT | 13,815–13,843 | ||
| TGEV-N-F | GCAGGTARAGGTGATGTGACAA | 27,637–27,658 | [17] | |
| Triplex-TGEV-N-R | TGCTRGACACAGATGGAACACA | 27,754–27,775 | ||
| Triplex-TGEV-N-P | GGAGCAGTGCCAAGCATTACCCACAA | 27,719–27,744 | ||
| Triplex-PEDV-N-F | CGCAAAGACTGAACCCACTAAC | 26,679–26,699 | [22] | |
| Triplex-PEDV-N-R | TGGTTRTTGCCTCTGTTGTTACT | 26,860–26,882 | ||
| Triplex-PEDV-N-P | TGTTGCCATTGCCACGACTCCTGC | 26,819–26,842 |
| Expected Assay Characteristics | Duplex 1 | Duplex 2 | Duplex 3 | Triplex |
|---|---|---|---|---|
| Target genes | PDCoV | |||
| PEDV N | TGEV N | TGEV S | + | |
| + | + | + | TGEV N | |
| PEDV S | PEDV S | PEDV S | + | |
| PEDV N | ||||
| Detects PEDV | Yes | Yes | Yes | Yes |
| Detects SeCoV | Yes | Yes | Yes | Yes |
| Detects TGEV | No | Yes | Yes | Yes |
| Detects PRCV | No | Yes | No | Yes |
| Detects PDCoV | No | No | No | Yes |
| Distinguishes PEDV from SeCoV | Yes | Yes | No | Yes |
| Distinguishes SeCoV from TGEV/PRCV | N/A | Yes | Yes | No |
| Distinguishes TGEV from PRCV | N/A | No | N/A | No |
| Assay | Gene Target (Virus Detected) | Forward Primer | Conc. | Reverse Primer | Conc. | Probe | Conc. |
|---|---|---|---|---|---|---|---|
| Duplex 1 | PEDV N (PEDV) | PEDV-N-F | 400 nM | PEDV-N-R | 800 nM | PEDV-N-probe (Cy5) | 240 nM |
| PEDV-N-US-F | 400 nM | ||||||
| PEDV S (PEDV + SeCoV) | PEDV-S-F | 900 nM | PEDV-S-R | 900 nM | PEDV-S-probe (FAM) | 200 nM | |
| Duplex 2 | TGEV N (TGEV/PRCV + SeCoV) | TGEV-N-F | 700 nM | TGEV-N-R | 700 nM | TGEV-N-probe (Cy5) | 200 nM |
| PEDV S (PEDV + SeCoV) | PEDV-S-F | 900 nM | PEDV-S-R | 900 nM | PEDV-S-probe (FAM) | 200 nM | |
| Duplex 3 | TGEV S (TGEV) | TGEV-S-F | 900 nM | TGEV-S-R | 900 nM | TGEV-S-probe (Cy5) | 200 nM |
| PEDV S (PEDV + SeCoV) | PEDV-S-F | 900 nM | PEDV-S-R | 900 nM | PEDV-S-probe (FAM) | 200 nM | |
| Triplex | PDCoV RdRp (PDCoV) | PDCoV-RdRp-F | 900 nM | PDCoV-RdRp-R | 900 nM | PDCoV-RdRp-P (FAM) | 200 nM |
| TGEV N (TGEV/PRCV + SeCoV) | TGEV-N-F | 900 nM | TGEV-N-R | 900 nM | TGEV-N-P (Texas Red) | 200 nM | |
| PEDV N (PEDV) | PEDV-N-F | 700 nM | PEDV-N-R | 700 nM | PEDV-N-P (Cy5) | 200 nM |
| LOD of Duplex Assays | LOD of Triplex Assay | |||||
|---|---|---|---|---|---|---|
| Assay | Singleplex | Duplex 1 | Duplex 2 | Duplex 3 | Singleplex | Triplex |
| PEDV N | 1000 * | 50 | - | - | ||
| PEDV S | 10 | 25 | 25 | 25 | ||
| TGEV N | 25 | - | 100 | - | ||
| TGEV S ** | 50 | - | - | 100 | ||
| PDCoV RdRp | 100 | 1000 | ||||
| TGEV N | 25 | 25 | ||||
| PEDV N | 10 | 25 | ||||
| Tested Sample for Repeatability | Tested Sample for Reproducibility | ||||||
|---|---|---|---|---|---|---|---|
| Assay | High Conc. | Intermediate Conc. | Low Conc. | High Conc. | Intermediate Conc. | Low Conc. | |
| Duplex 1 | PEDV N | 4% | 6% | 17% | 27% | 27% | 16% |
| PEDV S | 7% | 8% | 31% | 29% | 34% | 22% | |
| Duplex 2 | TGEV N | 4% | 6% | 43% | 27% | 27% | 24% |
| PEDV S | 6% | 7% | 19% | 29% | 33% | 43% | |
| Duplex 3 | TGEV S | 7% | 11% | 110% | 19% | 21% | * 37% |
| PEDV S | 4% | 3% | 11% | 32% | 34% | 35% | |
| Triplex | PDCoV RdRp | 10% | 16% | 9% | 14% | 16% | 4% |
| TGEV N | 5% | 7% | 13% | 10% | 6% | 32% | |
| PEDV N | 5% | 8% | 22% | 5% | 7% | 16% | |
| Duplex 1 | Duplex 2 | Duplex 3 | Triplex | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Pathogen Samples | No. Tested | PEDV N Positive (%) | PEDV S Positive (%) | TGEV N Positive (%) | PEDV S Positive (%) | TGEV S Positive (%) | PEDV S Positive (%) | PDCoV RdRp Positive (%) | TGEV N Positive (%) | PEDV N Positive (%) |
| PEDV | 24 | 23 (96) | 24 (100) | 0 (0) | 24 (100) | 0 (0) | 24 (100) | 0 (0) | 0 (0) | 23 (96) |
| Rec. SeCoV/PEDV | 22 | 22 (100) | 22 (100) | 0 (0) | 22 (100) | 0 (0) | 22 (100) | 0 (0) | 0 (0) | 20 (91) |
| SeCoV | 9 | 0 (0) | 9 (100) | 9 (100) | 9 (100) | 0 (0) | 9 (100) | 0 (0) | 9 (100) | 0 (0) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lazov, C.M.; Papetti, A.; Belsham, G.J.; Bøtner, A.; Rasmussen, T.B.; Boniotti, M.B. Multiplex Real-Time RT-PCR Assays for Detection and Differentiation of Porcine Enteric Coronaviruses. Pathogens 2023, 12, 1040. https://doi.org/10.3390/pathogens12081040
Lazov CM, Papetti A, Belsham GJ, Bøtner A, Rasmussen TB, Boniotti MB. Multiplex Real-Time RT-PCR Assays for Detection and Differentiation of Porcine Enteric Coronaviruses. Pathogens. 2023; 12(8):1040. https://doi.org/10.3390/pathogens12081040
Chicago/Turabian StyleLazov, Christina M., Alice Papetti, Graham J. Belsham, Anette Bøtner, Thomas Bruun Rasmussen, and Maria Beatrice Boniotti. 2023. "Multiplex Real-Time RT-PCR Assays for Detection and Differentiation of Porcine Enteric Coronaviruses" Pathogens 12, no. 8: 1040. https://doi.org/10.3390/pathogens12081040
APA StyleLazov, C. M., Papetti, A., Belsham, G. J., Bøtner, A., Rasmussen, T. B., & Boniotti, M. B. (2023). Multiplex Real-Time RT-PCR Assays for Detection and Differentiation of Porcine Enteric Coronaviruses. Pathogens, 12(8), 1040. https://doi.org/10.3390/pathogens12081040






