Colonization of Group B Streptococcus in Pregnant Women and Their Neonates from a Sri Lankan Hospital
Abstract
:1. Introduction
2. Results
2.1. Serotype and Multilocus Sequence Typing (MLST) Distribution
2.2. Antimicrobial Susceptibility
2.3. Genome-Based Phylogenetic Analysis
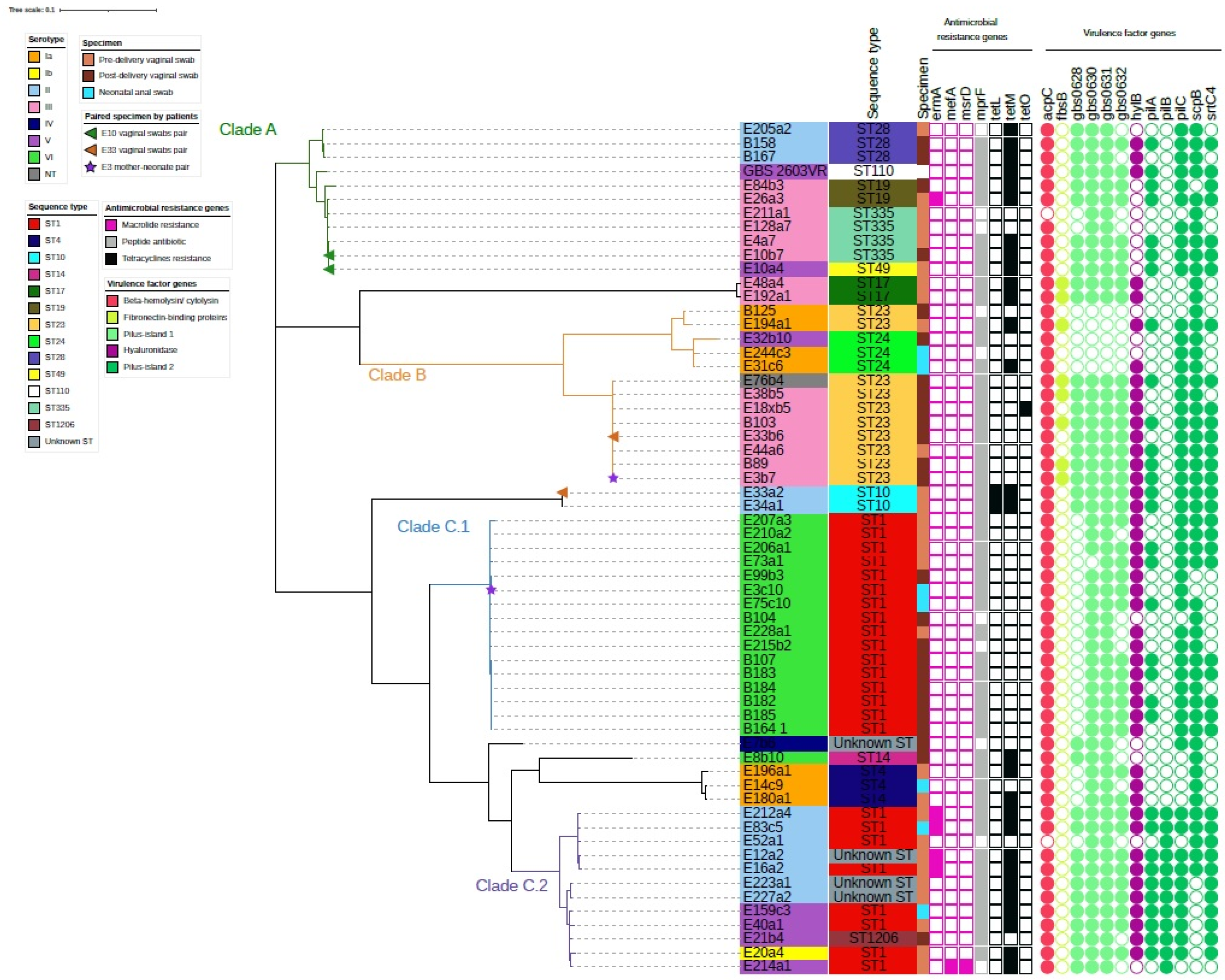
2.4. Virulence Genes and Antibiotic Resistance Genes among the Three GBS Clusters
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains
4.2. GBS Serotyping and Antibiotic Susceptibility Test
4.3. Whole-Genome Sequencing of GBS Strains
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gajdács, M.; Ábrók, M.; Lázár, A.; Burián, K. Beta-Haemolytic Group A, C and G Streptococcal Infections in Southern Hungary: A 10-Year Population-Based Retrospective Survey (2008–2017) and a Review of the Literature. Infect. Drug Resistance 2020, 13, 4739–4749. [Google Scholar] [CrossRef] [PubMed]
- Parks, T.; Barrett, L.; Jones, N. Invasive streptococcal disease: A review for clinicians. Br. Med. Bull. 2015, 115, 77–89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawn, J.; Cousens, S.; Zupan, J. 4 million neonatal deaths: When? Where? Why? Lancet 2005, 365, 891–900. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention, (CDC). Early-onset and late-onset neonatal group B streptococcal disease—United States, 1996–2004. MMWR Morb. Mortal. Wkly. Rep. 2005, 54, 1205–1208. [Google Scholar]
- Russell, N.; Seale, A.; O’Sullivan, C.; Le Doare, K.; Heath, P.; Lawn, J.; Bartlett, L.; Cutland, C.; Gravett, M.; Ip, M.; et al. Risk of Early-Onset Neonatal Group B Streptococcal Disease with Maternal Colonization Worldwide: Systematic Review and Meta-analyses. Clin. Infect. Dis. 2017, 65, S152–S159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bianchi-Jassir, F.; Seale, A.; Kohli-Lynch, M.; Lawn, J.; Baker, C.; Bartlett, L.; Cutland, C.; Gravett, M.; Heath, P.; Ip, M.; et al. Preterm Birth Associated with Group B Streptococcus Maternal Colonization Worldwide: Systematic Review and Meta-analyses. Clin. Infect. Dis. 2017, 65, S133–S142. [Google Scholar] [CrossRef] [Green Version]
- Le Doare, K.; O’Driscoll, M.; Turner, K.; Seedat, F.; Russell, N.; Seale, A.; Heath, P.; Lawn, J.; Baker, C.; Bartlett, L.; et al. Intrapartum Antibiotic Chemoprophylaxis Policies for the Prevention of Group B Streptococcal Disease Worldwide: Systematic Review. Clin. Infect. Dis. 2017, 65, S143–S151. [Google Scholar] [CrossRef] [Green Version]
- Baker, C.; Barrett, F. Transmission of group B streptococci among parturient women and their neonates. J. Pediatrics 1973, 83, 919–925. [Google Scholar] [CrossRef]
- Verani, J.R.; McGee, L.; Schrag, S.J. Prevention of Perinatal Group B Streptococcal Disease: Revised Guidelines from CDC, 2010; US Department of Health and Human Services, CDC: Atlanta, GA, USA, 2010; Volume 59, pp. 1–32. [Google Scholar]
- Russell, N.; Seale, A.; O’Driscoll, M.; O’Sullivan, C.; Bianchi-Jassir, F.; Gonzalez-Guarin, J.; Lawn, J.; Baker, C.; Bartlett, L.; Cutland, C.; et al. Maternal Colonization with Group B Streptococcus and Serotype Distribution Worldwide: Systematic Review and Meta-analyses. Clin. Infect. Dis. 2017, 65, S100–S111. [Google Scholar] [CrossRef]
- World Health Organization. Number of Infant Deaths (Child Mortality). 2022. Available online: https://www.who.int/data/gho/data/indicators/indicator-details/GHO/number-of-infant-deaths (accessed on 3 January 2022).
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 27th ed.; CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2017. [Google Scholar]
- Dilrukshi, G.; Kottahachchi, J.; Dissanayake, D.; Pathiraja, R.; Karunasingha, J.; Sampath, M.; Vidanage, U.; Fernando, S. Group B Streptococcus colonisation and their antimicrobial susceptibility among pregnant women attending antenatal clinics in tertiary care hospitals in the Western Province of Sri Lanka. J. Obstet. Gynaecol. 2020, 41, 1–6. [Google Scholar] [CrossRef]
- Kudagammana, H.; Rathnayaka, R.; Weerasooriya, B.; Karunathilaka, K.; Kumara, J. Group B Streptococcal colonisation among Sri Lankan mothers. SCIREA J. Clin. Med. 2019, 4, 209–215. [Google Scholar]
- Saha, S.; Ahmed, Z.; Modak, J.; Naziat, H.; Saha, S.; Uddin, M.; Islam, M.; Baqui, A.; Darmstadt, G.; Schrag, S. Group B Streptococcus among Pregnant Women and Newborns in Mirzapur, Bangladesh: Colonization, Vertical Transmission, and Serotype Distribution. J. Clin. Microbiol. 2017, 55, 2406–2412. [Google Scholar] [CrossRef] [Green Version]
- Shabayek, S.; Spellerberg, B. Group B Streptococcal Colonization, Molecular Characteristics, and Epidemiology. Front. Microbiol. 2018, 9, 437. [Google Scholar] [CrossRef] [PubMed]
- Madrid, L.; Seale, A.; Kohli-Lynch, M.; Edmond, K.; Lawn, J.; Heath, P.; Madhi, S.; Baker, C.; Bartlett, L.; Cutland, C.; et al. Infant Group B Streptococcal Disease Incidence and Serotypes Worldwide: Systematic Review and Meta-analyses. Clin. Infect. Dis. 2017, 65, S160–S172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teatero, S.; Ferrieri, P.; Martin, I.; Demczuk, W.; McGeer, A.; Fittipaldi, N. Serotype Distribution, Population Structure, and Antimicrobial Resistance of Group B Streptococcus Strains Recovered from Colonized Pregnant Women. J. Clin. Microbiol. 2017, 55, 412–422. [Google Scholar] [CrossRef] [Green Version]
- Shibata, M.; Morozumi, M.; Maeda, N.; Komiyama, O.; Shiro, H.; Iwata, S.; Ubukata, K. Relationship between intrapartum antibiotic prophylaxis and group B streptococcal colonization dynamics in Japanese mother–neonate pairs. J. Infect. Chemother. 2021, 27, 977–983. [Google Scholar] [CrossRef]
- Chaudhary, M.; Rench, M.; Baker, C.; Singh, P.; Hans, C.; Edwards, M. Group B Streptococcal Colonization Among Pregnant Women in Delhi, India. Pediatric Infect. Dis. J. 2017, 36, 665–669. [Google Scholar] [CrossRef]
- Shrestha, K.; Sah, A.; Singh, N.; Parajuli, P.; Adhikari, R. Molecular Characterization of Streptococcus agalactiae Isolates from Pregnant Women in Kathmandu City. J. Trop. Med. 2020, 2020, 4046703. [Google Scholar] [CrossRef]
- Critchfield, A.; Lievense, S.; Raker, C.; Matteson, K. Group B Streptococcus prophylaxis in patients who report a penicillin allergy: A follow-up study. Am. J. Obstet. Gynecol. 2011, 204, 150.e1–150.e8. [Google Scholar] [CrossRef] [Green Version]
- Hays, C.; Louis, M.; Plainvert, C.; Dmytruk, N.; Touak, G.; Trieu-Cuot, P.; Poyart, C.; Tazi, A. Changing Epidemiology of Group B Streptococcus Susceptibility to Fluoroquinolones and Aminoglycosides in France. Antimicrob. Agents Chemother. 2016, 60, 7424–7430. [Google Scholar] [CrossRef] [Green Version]
- Sharmila, V.; Babu, T. Antibiotic susceptibility pattern of group B streptococcal isolates from maternal genital tract. Int. J. Reprod. Contracept. Obstet. Gynecol. 2019, 8, 2738–2741. [Google Scholar] [CrossRef]
- Shafiq, S.; Ch, S.; Salee, S. Antibiotic susceptibility pattern of group B Streptococci isolated from pregnant women. J. Sheikh Zayed Med. Coll. 2016, 7, 974–976. [Google Scholar]
- Da Cunha, V.; Davies, M.; Douarre, P.; Rosinski-Chupin, I.; Margarit, I.; Spinali, S.; Perkins, T.; Lechat, P.; Dmytruk, N.; Sauvage, E.; et al. Streptococcus agalactiae clones infecting humans were selected and fixed through the extensive use of tetracycline. Nat. Commun. 2014, 5, 4544. [Google Scholar] [CrossRef] [PubMed]
- Poyart, C.; Jardy, L.; Quesne, G.; Berche, P.; Trieu-Cuot, P. Genetic Basis of Antibiotic Resistance in Streptococcus agalactiae Strains Isolated in a French Hospital. Antimicrob. Agents Chemother. 2003, 47, 794–797. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothen, J.; Pothier, J.; Foucault, F.; Blom, J.; Nanayakkara, D.; Li, C.; Ip, M.; Tanner, M.; Vogel, G.; Pflüger, V.; et al. Subspecies Typing of Streptococcus agalactiae Based on Ribosomal Subunit Protein Mass Variation by MALDI-TOF MS. Front. Microbiol. 2019, 10, 471. [Google Scholar] [CrossRef] [Green Version]
- Rothen, J.; Sapugahawatte, D.; Li, C.; Lo, N.; Vogel, G.; Foucault, F.; Pflüger, V.; Pothier, J.; Blom, J.; Daubenberger, C.; et al. A simple, rapid typing method for Streptococcus agalactiae based on ribosomal subunit proteins by MALDI-TOF MS. Sci. Rep. 2020, 10, 8788. [Google Scholar] [CrossRef]
- Ábrók, M.; Tigyi, P.; Kostrzewa, M.; Burián, K.; Deák, J. Evaluation of the Results of Group B Streptococcus Screening by MALDI-TOF MS among Pregnant Women in a Hungarian Hospital. Pathogens 2019, 9, 1. [Google Scholar] [CrossRef] [Green Version]
- Nanayakkara, D.; Liyanapathirana, V.; Kandauda, C.; Gihan, C.; Ekanayake, A.; Adasooriya, D. Maternal vaginal colonization with selected potential pathogens of neonatal sepsis in the era of antimicrobial resistance, a single center experience from Sri Lanka. BMC Infect. Dis. 2018, 18, 351. [Google Scholar] [CrossRef]
- Imperi, M.; Pataracchia, M.; Alfarone, G.; Baldassarri, L.; Orefici, G.; Creti, R. A multiplex PCR assay for the direct identification of the capsular type (Ia to IX) of Streptococcus agalactiae. J. Microbiol. Methods 2010, 80, 212–214. [Google Scholar] [CrossRef]
- Li, C.; Sapugahawatte, D.; Yang, Y.; Wong, K.; Lo, N.; Ip, M. Multidrug-Resistant Streptococcus agalactiae Strains Found in Human and Fish with High Penicillin and Cefotaxime Non-Susceptibilities. Microorganisms 2020, 8, 1055. [Google Scholar] [CrossRef]
- Sapugahawatte, D.N.; Li, C.; Dharmaratne, P.; Zhu, C.; Yeoh, Y.K.; Yang, J.; Lo, N.W.S.; Wong, K.T.; Ip, M. Prevalence and Char-acteristics of Streptococcus agalactiae from Freshwater Fish and Pork in Hong Kong Wet Markets. Antibiotics 2022, 11, 397. [Google Scholar] [CrossRef]
- Yang, Y.; Luo, M.; Zhou, H.; Li, C.; Luk, A.; Zhao, G.; Fung, K.; Ip, M. Role of Two-Component System Response Regulator bceR in the Antimicrobial Resistance, Virulence, Biofilm Formation, and Stress Response of Group B Streptococcus. Front. Microbiol. 2019, 10, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Serotypes | Total No. Swabs (n = 60) | Pre-Delivery Vaginal Swab (n = 28) | Post-Delivery Vaginal Swabs (n = 25) | Neonate Anal Swabs (n = 7) | p-Value (Pre- vs Post-Delivery Vaginal Swabs) ^ | p-Value (Pre-Delivery Vaginal Swabs vs Neonatal Anal Swabs) ^ | p-Value (Post-Delivery Vaginal Swabs vs Neonatal Anal Swabs) ^ |
|---|---|---|---|---|---|---|---|
| Ia | 7 (11.7%) | 3 (10.7%) | 1 (4%) | 3 (42.9%) | 0.6 | 0.07 | 0.03 |
| Ib | 1 (1.67%) | 1 (3.57%) | 0 (0%) | 0 (0%) | N/A | N/A | N/A |
| II | 12 (20%) | 9 (32.1%) | 2 (8%) | 1 (14.3%) | 0.04 | 0.6 | 1 |
| III | 15 (25%) | 7 (25%) | 8 (32%) | 0 (0%) | 0.76 | N/A | N/A |
| IV | 1 (1.67%) | 0 (0%) | 1 (4%) | 0 (0%) | N/A | N/A | N/A |
| V | 6 (10%) | 3 (10.7%) | 2 (8%) | 1 (14.3%) | 1 | 1 | 1 |
| VI | 17 (28.3%) | 5 (17.9%) | 10 (40%) | 2 (28.6%) | 0.12 | 0.6 | 0.7 |
| NT # | 1 (1.67%) | 0 (0%) | 1 (4%) | 0 (0%) | N/A | N/A | N/A |
| Sequence Type (ST) | Serotypes | |||||||
|---|---|---|---|---|---|---|---|---|
| Ia (n = 7) | Ib (n = 1) | II (n = 12) | III (n = 15) | IV (n = 1) | V (n = 16) | VI (n = 17) | NT (n = 1) # | |
| ST1 | - | 1 (100%) | 4 (33.3%) | - | - | 3 (50%) | 16 (94.1%) | - |
| ST23 | 2 (28.6%) | - | - | 7 (46.7%) | - | - | - | 1 (100%) |
| ST335 | - | - | - | 4 (26.7%) | - | - | - | - |
| ST28 | - | - | 3 (25%) | - | - | - | - | - |
| ST19 | - | - | - | 2 (13.3%) | - | - | - | - |
| ST4 | 3 (42.8%) | - | - | - | - | - | - | - |
| ST24 | 2 (28.6%) | - | - | - | - | 1 (16.7%) | - | - |
| ST10 | - | - | 2 (16.7%) | - | - | - | - | - |
| ST17 | - | - | - | 2 (13.3%) | - | - | - | - |
| ST14 | - | - | - | - | - | - | 1 (5.9%) | - |
| ST49 | - | - | - | - | - | 1 (16.7%) | - | - |
| ST1206 | - | - | - | - | 1 (16.7%) | - | - | |
| Unknown STs | - | - | 3 (25%) | - | 1 (100%) | - | - | |
| Class of Antibiotic | Antibiotic | MIC (μg/mL) a | No. of Resistance Strains (%) | ||
|---|---|---|---|---|---|
| Range | MIC50 | MIC90 | |||
| Penicillins | Penicillin | 2–0.0625 | 0.06 | 0.06 | 0 |
| Oxacillin | 32–0.03 | 0.25 | 0.5 | 0 | |
| Cephalosporins | Cefotaxime | 2–0.12 | 0.12 | 0.12 | 0 |
| Ceftibuten | 32–0.03 | 16 | 16 | 0 | |
| Glycopeptides | Vancomycin | 8–0.25 | ≤0.12 | ≤0.12 | 0 |
| Amino glycosides | Gentamicin | 4–0.25 | >4 | >4 | 60 (100%) |
| Tetracyclines | Doxycycline ^ | 32–1 | 32 | >32 | 34 (56.7%) |
| Minocycline ^ | 32–1 | 32 | >32 | 34 (56.7%) | |
| Tetracycline | 32–1 | 16 | 32 | 35 (58.3%) | |
| Oxazolidinones | Linezolid | 64–0.06 | 1 | 1 | 0 |
| Chloramphenicol | Chloramphenicol | 64–2 | ≤1 | 2 | 0 |
| Macrolides | Erythromycin | 4–0.12 | ≤0.06 | 1 | 9 (15%) |
| Lincosamides | Clindamycin | 4–0.12 | ≤0.06 | ≤0.06 | 0 |
| Lincomycin | 32–0.03 | 0.12 | 0.25 | 0 | |
| Fluoroquinolones | Ciprofloxacin | 64–0.06 | 0.25 | 0.5 | 0 |
| Levofloxacin | 32–1 | ≤0.5 | 1 | 0 | |
| Gatifloxacin | 4–0.25 | ≤0.5 | ≤0.5 | 0 | |
| Moxifloxacin * | 2–0.12 | ≤0.12 | ≤0.12 | 0 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sapugahawatte, D.N.; Li, C.; Liyanapathirana, V.; Kandauda, C.; Gihan, C.; Zhu, C.; Lo, N.W.S.; Wong, K.T.; Ip, M. Colonization of Group B Streptococcus in Pregnant Women and Their Neonates from a Sri Lankan Hospital. Pathogens 2022, 11, 386. https://doi.org/10.3390/pathogens11040386
Sapugahawatte DN, Li C, Liyanapathirana V, Kandauda C, Gihan C, Zhu C, Lo NWS, Wong KT, Ip M. Colonization of Group B Streptococcus in Pregnant Women and Their Neonates from a Sri Lankan Hospital. Pathogens. 2022; 11(4):386. https://doi.org/10.3390/pathogens11040386
Chicago/Turabian StyleSapugahawatte, Dulmini Nanayakkara, Carmen Li, Veranja Liyanapathirana, Chaminda Kandauda, Champika Gihan, Chendi Zhu, Norman Wai Sing Lo, Kam Tak Wong, and Margaret Ip. 2022. "Colonization of Group B Streptococcus in Pregnant Women and Their Neonates from a Sri Lankan Hospital" Pathogens 11, no. 4: 386. https://doi.org/10.3390/pathogens11040386
APA StyleSapugahawatte, D. N., Li, C., Liyanapathirana, V., Kandauda, C., Gihan, C., Zhu, C., Lo, N. W. S., Wong, K. T., & Ip, M. (2022). Colonization of Group B Streptococcus in Pregnant Women and Their Neonates from a Sri Lankan Hospital. Pathogens, 11(4), 386. https://doi.org/10.3390/pathogens11040386






